Abstract
Cryptic genetic diversity and erroneous morphological species determination represent frequent problems in biodiversity research. Here, examination of 138 specimens of Synodontis (Mochokidae, Siluriformes) from the Nile River and Lake Turkana revealed the presence of both S. schall-like and S. frontosus-like morphotypes, with a phenotypic gradient between them. We concluded phylogenetic and population genetic analyses based on two mitochondrial and one nuclear marker including 131 coxI (565 bp), 96 cytb (973 bp) and 19 RAG2 (896 bp) sequences from the Nile-Turkana population, plus additional GenBank data of Synodontis spp. Whilst nuclear data were inconclusive, mitochondrial sequences suggested that both morphotypes and intermediate forms are conspecific. The results imply probable synonymy of S. frontosus with S. schall. Conversely, a strong biogeographical signal was revealed among widely distributed and supposedly conspecific S. schall-like catfish of the Nilo-Sudanian ichthyological province. Synodontis schall sensu stricto (=Eastern clade), as defined by type locality in the Nile, is apparently restricted to the eastern part of the Nilo-Sudanian ichthyological province (e.g. Nile, Turkana, Chad). Synodontis schall Western clade (Senegambia, Niger, Chad) most probably represents a cryptic taxon, unrecognized thus far due to the absence of distinctive morphological differences.
Similar content being viewed by others
Introduction
Determining and identifying independent lineages and species is critical not only for taxonomy, but also for understanding the processes leading to diversification and the origin of population structure. Due to the extensive sampling, discovering cryptic species within the scope of population genetic studies is not a rare phenomenon1,2,3. Traditionally, cryptic species/diversity in fishes is mostly discovered in highly variable taxa, especially in the species rich lineages of ray-finned fishes, such as the neotropical Characidae4,5 or Cichlidae6,7. Cryptic diversity was also found - albeit less frequently - in other lineages, e.g. spiny eels8 and Neotropical9,10,11 and African catfish12,13, including the family Mochokidae14,15,16. Generally, the main contributing factor to the undiscovered cryptic diversity of some species is the lack of clear distinguishing morphological features in combination with the absence of relevant molecular data3. This not only underestimates the species richness but also limits our understanding of species geographical distribution.
Within the scope of research on fish parasites in the Nile River and Lake Turkana, we examined a total of 138 Synodontis specimens. Two of three Synodontis spp. in our sample, Synodontis nigrita Valenciennes, 1840 and Synodontis serratus Rüppell, 1829 (both absent in Lake Turkana) could be readily morphologically determined. The third species Synodontis schall (Bloch et Schneider, 1801) showed deviation from both, the data in the literature, as well as the differential diagnosis and neotype proposed recently by Musschoot & Lalèyè17. Among Synodontis spp. occurring in the Nile and Turkana basins, S. schall might be confused with the morphologically very similar Synodontis frontosus Vaillant, 1895. According to the literature, S. schall is common and abundant in both the Nile River and Lake Turkana18,19,20,21. Although the two species might co-occur in both basins, S. frontosus is considered absent in Lake Turkana itself, reportedly being restricted to its main source, the Omo River22. Diagnostic features of both species are similar and partly overlapping, therefore, distinguishing between the two in the field is difficult.
Resolving the species status of S. schall is important as the species has been used as a model in parasitological research23. Therefore, we decided to base the taxonomical assignment of our Nile-Turkana S. schall specimens on a multidisciplinary approach, particularly on molecular identification with an assessment of selected morphological traits. Herein, we assess the identity of S. schall-like catfish from the Nile and Turkana basins by means of phylogenetic and haplotype analyses of a representative set of Synodontis spp. coxI, cytb and RAG2 sequences, revealing a previously unrecognized biogeographical pattern. To the best of our knowledge, this is the first confrontation of morphological features traditionally used for determination of S. schall and similar Nilo-Sudanian catfishes with molecular data to aid proper distinction of these catfishes of high significance from both an ichthyological and regional fisheries perspective.
Material and Methods
Study group
The Mochokidae (Actinopterygii, Siluriformes) is an endemic African family of freshwater catfish containing 211 species in nine genera24. With 131 nominal species currently recognized, Synodontis Cuvier, 1817 is the most speciose mochokid genus widely distributed in all African ichthyological provinces except for Maghreb and South Africa18,21,25. Despite numerous studies based on genetic markers26,27,28 morphology remains the main classification tool for identification of Synodontis spp.17,18,29. Several molecular phylogenetic studies were published during the last 13 years which elucidated general biogeographic patterns and relationships within the genus26,27,28,30,31. Synodontis schall is traditionally considered widespread throughout the Nilo-Sudanian ichthyological province from the Atlantic to Indian Ocean17. Its widespread distribution, abundance and easy harvesting make it one of the most important subsistence fish species in Sub-Saharan Africa north of equator and along the whole Nile valley28,32,33,34. Importantly, this ecologically plastic species28 is less likely to be affected by climatic changes and anthropogenic drainage alterations, making it a prospective source for fisheries in the future35. Despite its significance for fisheries and a long taxonomical history, morphology-based distinction between S. schall and similar species such as S. frontosus, remains unclear.
Sampling and fish determination
Sampled fish were obtained in seven different localities in the Nile and Turkana basins (S. schall n = 120, S. nigrita n = 8, S. serratus n = 10, Total: 138; see Table 1 for details). The two Nile localities cover both the White (Kostí) and Blue Nile (Sennar). The five Turkana localities cover the salinity gradient present in the lake, i.e. the freshwater Omo River delta (Todonyang) and the saline (brackish) main part of the lake (central Turkana saline: Kalokol, Central Island; central Turkana freshwater refugium: Kerio River delta; south Turkana: El-Molo Bay). Fish examined were obtained in fish markets and/or from fishermen and processed under supervision of Kenya Marine and Fisheries Research Institute (KMFRI) authorities. Fish that were still alive when obtained were euthanized by dorsal pithing (spinal cord and blood vessels cut immediately behind the head), a method congruent with Kenyan, Sudanian, Czech and European legislation. Fish were destined for human consumption; therefore, no permission was needed to collect these fish. A small piece of liver tissue from each individual was cut off and preserved in 96% ethanol for molecular analyses. Fish were photo-documented and their standard length was measured. Their mouthparts including tooth plates and barbels were removed and preserved in 4% formaldehyde for further morphological examination.
Upon field examination, all fish were determined based on the following combinations of qualitative diagnostic features: S. nigrita: body, head and fins with black spots, inner mandibular barbels bearing thick and tuberculate ramifications, maxillary barbels with a well visible broad black membrane, short and small adipose fin; S. serratus: maxillary barbels with a very large light-colored membrane, humeral process pointed, non-keeled, distinctive serrations on both edges of pectoral spine, prolonged rostrum20,21; for S. schall, see Table 2 and Results and Discussion sections. In congruence with taxonomic literature, the species identification was based on visual inspection of qualitative diagnostic features only as no relevant morphometric diagnostic traits are available for other Synodontis spp.
DNA extraction, PCR and sequencing
DNA was isolated using the phenol:chloroform:isoamyl alcohol (25:24:1) method described in Sambrook & Russell36. Two mitochondrial genes and one nuclear gene were used for phylogenetic analyses, cytochrome oxidase I (coxI), cytochrome b (cytb) and recombination activating gene 2 (RAG2), respectively (see Table S1 in Supplementary Data for a complete list of sequences and other details). Mitochondrial genes were chosen as they are known indicators of recently divided species and also because coxI and cytb are available in GenBank for a broad range of Synodontis spp. Due to the high abundance of RAG2 for genus Synodontis in GenBank, this nuclear gene was selected to supplement mtDNA data in selected Synodontis spp. specimens – 23 representatives from sampling localities in the Nile and Turkana. Amplification conditions and PCR primers FishF2 and VF2_t1 for partial sequence of coxI gene (~800 bp) were taken from Ivanova et al.37. PCR reactions contained 12.5 μl of PPP Master mix, 10.5 μl of ultra pure water, 5 pM of each PCR primer and 0.5 μl of isolated DNA (100 to 170 ng). Published primer sequences (5′-GAC-TTGAAGAACCACCGTTG-3′ forward and 5′-TTTAGAATTCTGG CTTTGGGAG-3′ reverse) to amplify part of cytb (~1000 bp) according to Pinton et al.27. For RAG2 (~1100 bp) amplification primers (forward 5′-TGY TAT CTC CCA CCT CTG CGY TAC C-3′ and reverse 5′-TCA TCC TCC TCA TCK TCC TCW TTG TA-3′) and reaction conditions followed those provided by Sullivan et al.38. Amplification reactions for cytb and RAG2 contained the same PCR reagents and amount of DNA as mentioned above. All products were verified on a 1% agarose gel and purified using enzymes exonuclease I (Exo I) and shrimp alkaline phosphatase (SAP)39. Purified fragments were directly sequenced by Macrogen Inc. (Amsterdam, Netherlands) using the same primers used for PCR amplification.
Phylogenetic, haplotype analyses and population genetic statistics
Obtained sequences were assembled and inspected for errors using the GENEIOUS Pro software package version 6.140. In addition to our original sequences (131 coxI, 96 cytb and 19 RAG2, Table S1), we used GenBank sequences of Synodontis spp. (n = 115 coxI sequences from 77 Synodontis spp., n = 80 cytb sequences from 52 taxa, and n = 102 RAG2 from 69 species) to interpret our results in a broader phylogenetic context (see Table S2 in Supplementary Data for complete list of sequences and additional information).
Alignments were constructed in MAFFT v. 641 and corrected manually using the GENEIOUS Pro software package version 6.1. Best-fit models of molecular evolution were selected using AIC criteria in jMODELTEST v2.1.442. Phylogenetic analyses were conducted for individual gene datasets plus a concatenated dataset of the two mtDNA genes (where data for both genes were available, see Tables S1 and S2). Phylogenetic analyses were performed using Bayesian inference (BI) in MrBayes v3.2.2.43,44 and maximum likelihood (ML) in PHYML v3.045. BI analyses were performed under the same conditions for all datasets - using 5 million MCMC replications, with four chains and three independent runs. The following nucleotide substitution models were used: GTR + I + G for coxI data, GTR + G for cytb, HKY + I + G for RAG2 and GTR + I + G for concatenated mt data. ML analyses were run using bootstrap analyses with 1000 replications and models mentioned above. Convergence for BI runs was estimated by TRACER v1.746, ESS values were assessed and found to exceed 200 for all parameters. Trees computed for cytb were rooted using Auchenoglanis occidentalis (HM880259, KY483731, EU781895) and Mochokus niloticus (KY483655) as an outgroup. The remaining trees (coxI, RAG2 and concatenated data) were rooted using GenBank sequences of Microsynodontis sp. (DQ886604 for cytb and HF565741 for RAG2). All trees were deposited in TreeBASE database (treebase.org) and are accessed under the following link http://purl.org/phylo/treebase/phylows/study/TB2:S23311.
Pairwise genetic distances were computed to assess the divergence of S. schall sequences and included sequences of a closely related taxon (S. serratus) and a distantly related congener (S. nigrita) were included for comparison. Due to the limited availability of the nuclear data, genetic distances were computed for mitochondrial data only.
To examine the level of population variability in the dataset, statistical parsimony networks based on pairwise differences were constructed for individual S. schall clades revealed by phylogenetic analyses using PopART v1.747 with settings for TCS networks48.
Statistics of Haplotype diversity (Hd), Fst values, genetic distances and differentiation were calculated using DNASP v549. Due to the limited number of GenBank sequences from the western part of S. schall geographic range, population statistics were performed only for S. schall samples obtained for this study (Nile and Turkana); data were split according to their resemblance to the S. schall-like and S. frontosus-like morphotypes (see Table S2).
Results
Morphology
Standard length of examined S. schall specimens ranged from 142 to 300 mm (mean ± SD; 209.8 ± 33.2 mm). Weight ranged from 75 to 783 g (mean ± SD, 227.1 ± 118.9 g). Synodontis schall from the Nile and Turkana showed remarkable variation in body shape, proportions, as well as coloration and other qualitative diagnostic features (see Fig. 1 and Table 2 for details). We observed a phenotypic gradient from the typical S. schall morphotype (Fig. 1a,d,i,k), which dominated the sample, to the typical S. frontosus morphotype (Fig. 1c,h,j,m), which was less frequent. These observations, together with the common occurrence of intermediate forms hampered reliable morphology-based determination of numerous specimens. In the majority of Nile-Turkana specimens, there was an obvious, blunt, low, relatively wide, longitudinally striated keel on the ventral side of the humeral process fitting the S. schall type (Fig. 1d,e,h). Only in some specimens the keel was absent, resembling that of the S. frontosus type (Fig. 1f,g). The humeral process shape showed gradual variation from S. schall-like (margins straight and/or dorsally concave and/or ventrally convex) (Fig. 1d,e) to S. frontosus-like (straight both dorsally and ventrally) (Fig. 1h). Gradual variation was observed also in anterior pectoral spine serrations, from S. schall-like (fine, sharp, distinct) (Fig. 1i) to S. frontosus-like (barely discernible, sometimes due to pigmentation in dark-colored specimens) (Fig. 1j). Remarkable gradual variation was present also in maxillary barbels from S. schall-like (devoid of a basal membrane or with a hardly visible slat-like rudiment) (Fig. 1k,l) to S. frontosus-like (distinct, well developed dark basal membrane) (Fig. 1m). Mandibular teeth numbered from 26 to 46 in total; these teeth numbers were outside ranges of both S. schall (18–39) and S. frontosus (33–48) morphotypes (Table 2). When only a single (main) row of mandibular teeth was considered, the count was 21 to 40. Mandibular teeth appeared to have formed from the outer margin of tooth plate towards the mouth interior and the old ones lost their sharp tips and eventually disappear, while new ones start to grow from the outer side (Fig. 2). The number of new mandibular teeth emerging in front of the main teeth row varied from 1 to 12.
Qualitative morphological diagnostic features in live specimens of Synodontis schall from Lake Turkana, Kenya. Corresponding humeral processes, pectoral spines and mouthparts are shown for two specimens representing typical S. schall morphotype (a,d,i,k) and typical S. frontosus morphotype (1c,h,j,m); remaining pictures originating from different individuals show additional variation. (a) Todonyang (Omo River delta); (b) El-Molo Bay (southern Lake Turkana); (c) Todonyang; (d–h) Humeral process variants (see Table 2 for details); (i) Pectoral spine with well visible fine sharp anterior serrations (starting from small arrow towards left side of picture); (j) Pectoral spine with anterior serrations barely discernible (partly due to pigmentation); (k) Maxillary barbel without membrane; (l) Rudimentary slat-like form (arrowheads); (m) Distinct dark membrane (arrows). Note: Depicted specimens show numerous dark specks in various body parts, but specimens almost lacking specks might occur too.
Synodontis schall mandibular dentition, Lake Turkana, Kenya. (a) High magnification image showing new teeth emerging at the outer margin of the tooth line (arrow, only one marked), main row of fully grown teeth with sharp tips (unmarked), and old teeth that already lost their sharp tips positioned at the inner margin of the tooth row (arrowhead, only one marked); (b) Composite image showing variability of teeth arrangement and number: 28 (upper left), 37 (upper right), 39 (bottom).
In three main Turkana sampling localities, the ratio of S. schall/S. frontosus/intermediate morphotypes was as follows: El-Molo Bay 35/16/7, Kalokol 19/11/0, Todonyang 32/31/14, implying somewhat higher relative incidence of S. frontosus-like morphotype in the freshwater part of Turkana (Chi-square test = 10.3; df = 4; P < 0.05) (Table S3; morphotype ratios for the Nile sample are not provided, because damage of fish from markets hampered assessment of all the focal features in some specimens).
In addition, several other features corresponding either to S. schall or S. frontosus have been observed, but not systematically recorded. For example, pointed snout combined with narrower interorbital region and eyes well within lateral margin of the head in dorsal view (as in S. schall) vs. snout more broadly rounded with wider interorbital region and orbits impinging on the lateral outline of the head in dorsal view (as in S. frontosus) (sensu Hopson22). The two states are well demonstrated by the two specimens showing S. schall and S. frontosus morphotype in Fig. 1a,c, respectively.
Phylogenetic analyses
In total, 131 sequences of coxI (565 bp), and two smaller sets comprising 96 cytb (973 bp) and 19 RAG2 (896 bp) sequences were generated. Data representing individual populations obtained from the Nile River and Lake Turkana were used for phylogenetic analyses together with Synodontis spp. sequences retrieved from GenBank. The newly generated sequence data were deposited in GenBank under accession numbers coxI: KY483652-KY483782, cytb: KY483783-KY483878, RAG2: MF136700-MF136718, MF150161- MF150164 (Table S1). As BI and ML phylogenetic analyses provided similar outcomes, only BI final trees are presented with posterior probabilities as branch supports, whereas ML trees are presented as Supplementary Data.
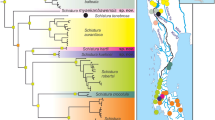
In coxI analysis, sequences of 131 Synodontis spp. obtained for this study and 115 sequences from GenBank were used to construct the final tree (Fig. 3). As expected, our samples of S. nigrita and S. serratus from the Nile clustered with conspecific sequences from GenBank (see Fig. S1 for composition of the collapsed clades). All Nile-Turkana S. schall samples, i.e. S. schall-like, S. frontosus-like and intermediate morphotypes, clustered together and created a well-supported group. Samples from different Turkana or Nile localities did not show any geographical pattern or affiliation to their sampling localities. According to the predominant morphological patterns observed in the Nile-Turkana sample and location of the type locality of S. schall in the Nile, we named this clade “S. schall Eastern clade”. Importantly, all GenBank sequences labeled S. frontosus (HF565883–89) and one sequence of S. caudovittatus (HF565865), all from the Nile, also clustered within the Eastern clade in all analyses. In contrast, GenBank sequences labeled S. schall (HF565950, HF565953-56), Synodontis ouemeensis HF565953, all from western part of Nilo-Sudanian and Eburneo-Ghanian provinces, together with sequences labeled Synodontis aff. bastiani and Synodontis aff. haugi HF565950, HF565954-56 (pet trade), created a separate group with high nodal support. Due to the clear separation of this group from the “S. schall Eastern clade” and taxonomic assignment of most its sequences, we named the clade “S. schall Western clade”.
Bayesian phylogeny for coxI. Posterior probabilities are shown as branch supports. Tree is rooted with Microsynodontis sp. sequence. Samples of S. schall Eastern clade are collapsed; detailed view of Eastern clade structure is presented with sampling localities. Valid names of Synodontis spp. following25 are used for convenience. Original assignments retrieved from GenBank, together with sequence accession numbers are provided in Fig. S2 and Tables S1, S2.
The dataset for cytb included sequences representing 52 Synodontis spp. (Fig. 4). Our sequences clustered with all GenBank S. frontosus sequences (FM878850–54, HF566018–23), as well as S. schall sequences (EU781905–07, EU781915) and EU781915 from the Nile and/or Chad basin. This clade clearly corresponds with the S. schall Eastern clade, hence its name in the tree. The rest of S. schall sequences (EU781902–04, EU781908–14, EU781916) from Senegambia, Niger and Chad basin created a group corresponding to the S. schall Western clade, closely related to, but clearly distinct from the S. schall Eastern clade. Interestingly, cytb sequences of S. ouemeensis (HF566058), S. aff. bastiani (HF566004) and S. aff. haugi (HF566027) clustered with the S. schall Western clade data. This result is similar to coxI analysis, however; with a stronger branch support and placing sequences of S. aff. bastiani and S. aff. haugi within the Western clade. In conclusion, analyses of both mitochondrial markers revealed very similar patterns with strong biogeographic signals, indicating phylogenetic distinctiveness and geographic affinity of two clades, i.e. S. schall Eastern and S. schall Western clade (Figs 3–5).
Bayesian phylogeny for cytb. Posterior probabilities are shown as branch supports Tree is rooted to Microsynodontis sp. sequence. Samples of S. schall Eastern clade are collapsed; detailed view of Eastern clade structure is presented with sampling localities. Valid names of Synodontis spp. following25 are used for convenience. Original assignments retrieved from GenBank, together with sequence accession numbers are provided in Fig. S2 and Tables S1, S2.
Map of the Nilo-Sudanian ichthyological province (solid grey line) showing geographical origin of Synodontis schall Eastern and Western clade samples as revealed by phylogenetic analyses. Dashed line indicates western and eastern limits of Chad basin; asterisk indicates Eburneo-Ghanian province; only major lakes are shown outside the area of interest.
Similarly as in the tree based solely on cytb sequences, the S. schall Eastern and S. schall Western clades were clearly separated from each other in the concatenated mtDNA dataset too (Fig. S3).
The phylogenetic analysis of the nuclear gene (RAG2) was less informative compared to the clear separation of Eastern and Western clades in all analyses performed for both mitochondrial genes. A total of 19 RAG2 sequences of Synodontis spp. were analyzed (Fig. 6) together with 102 sequences from GenBank. Our S. schall Eastern clade samples from the Nile and Turkana comprised an internally unresolved cluster together with the samples of S. frontosus (HF565870-74), S. caudovittatus (HF565755), S. violacea (HF565838) and S. aff. bastiani (HF565751) from Eastern Nilo-Sudan province, as well as S. schall (HF565817-19) from Western Nilo-Sudan province. Although sequences of S. serratus from our sample and from GenBank clustered together with high support, the S. serratus branch was placed within the S. schall clade, probably as a result of low sequence variability in the whole dataset (100-94.5%).
Bayesian phylogeny for RAG2. Posterior probabilities are shown as branch supports Tree is rooted with Microsynodontis sp. Synodontis schall Eastern clade is displayed in and branches for other Synodontis species are collapsed for better visualization. Valid names of Synodontis spp. following Froese & Pauly (2017) are used for convenience. Original assignments retrieved from GenBank, together with sequence accession numbers are provided in Fig. S3 and Tables S1, S2.
Genetic distance
Genetic distances between S. schall Eastern clade and Western clade samples range from 5.5% to 6.4% and from 5.7% to 8.3% for coxI and cytb, respectively (Table 3). In addition, the difference between the two S. schall clades is very similar to the distance between morphologically and molecularly well distinguished species S. nigrita and S. serratus (from 9.6% to 10.4%, see Table 3). The coxI data showed smaller range of variability, which might be due to the shorter sequence of the gene or simply, coxI is more conservative than cytb. Nevertheless, the distinction between S. schall clades is still more than 5% for both coxI and cytb mitochondrial genes.
Haplotype networks
The S. schall Western clade produced a relatively compact network, exhibiting similar levels of diversity in both genes (Figs 7 and 8), with only a slight increase of distances between individual haplotypes in cytb. CoxI data for the Eastern clade also created a compact network, represented by one big central haplotype surrounded by derived haplotypes (Fig. 7), whereas cytb showed a more complicated network structure with some haplotypes divided by more than ten mutations (Fig. 8). Although the two Eastern clade networks might appear very different, similar phenomena can be seen in both datasets, most remarkably the sharing of haplotypes between the Nile and Turkana basins. The differences between structures of haplotype networks are probably due to a longer stretch of cytb sequence analyzed and a putatively higher mutation rate of cytb compared to coxI in vertebrates50.
Haplotype networks for coxI constructed in software PopART v1.7. Networks were constructed only for samples clustering in S. schall Eastern and Western clades. Sizes of haplo-nodes are relative to the sample size. The two networks were separated by 28 steps. Accession numbers for GenBank sequences are listed in the picture along with sampling locations.
Haplotype networks for cytb constructed in software PopART v1.7. Networks were constructed only for samples clustering in S. schall Eastern and Western clades. Sizes of haplo-nodes are relative to the sample size. The two networks were separated by 51 steps. Accession numbers for GenBank sequences are listed in the picture along with sampling locations.
Population genetic statistics of the Eastern clade
Population genetic statistics and genetic distances performed for our data did not show profound differences between the sequences of S. schall-like and S. frontosus-like specimens belonging to the Eastern clade (see Table S4). Genetic distances between the individuals of the same respective morphotype were similar to the distances between individuals belonging to different morphotypes (approximately 0 to 1% and 0 to 2% for coxI and cytb, respectively, see Table S4). The overall FST values between the two morphotypes were 0.08 and 0.04 for coxI and cytb, respectively. The summed chi-square values, for the test of genetic differentiation, were moderately significant for coxI sequences (0.001 < P < 0.01) and non-significant for cytb (P > 0.05). Similarly, a difference was seen in haplotype diversities for coxI, but not in cytb (Table S4). Moreover, the statistics of genetic differentiation between localities within Turkana (Snn test, Table S4) revealed a significant value for coxI sequences, but not in cytb. Results of all these tests provide evidence of low level of the genetic differentiation corresponding to an intraspecific level.
Discussion
Identity of S. schall-like catfish
In the Nile and Turkana basins, typical morphotypes of both S. schall and S. frontosus, as well as a phenotypic gradient between the two, have been identified. Regardless of morphological patterns, phylogenetic and haplotype analyses suggest that all the Nile-Turkana samples are conspecific, including both S. schall-like and S. frontosus-like specimens. It is to be noted, that the taxonomical incongruence between taxonomic assignment of GenBank sequences as identified by other authors e.g. Day et al.18, Pinton et al.19, Day et al.36 and supported by our phylogenetic analyses of coxI and cytb data clearly indicated both S. schall and S. frontosus morpho-species among samples revealed herein as conspecific (see original sequence assignments retained in Fig. 4 and Table S2). This result casts doubt on features presumably differentiating S. schall and S. frontosus. Especially, the relatively frequent presence of distinct dark membranes on maxillary barbels because its “absence, or a presence of only hardly visible rudiment” (sensu Paugy & Roberts21 and Musschoot & Lalèyè17) was considered a basic feature distinguishing S. schall from S. frontosus. The genetic uniformity of the Nile-Turkana sample on one hand, and its remarkable morphological variability lead us to assume that diagnostic features of S. frontosus merely describe part of the intraspecific variability of S. schall. In other words, explicit evidence of remarkable intraspecific variability provided herein renders the two nominal species from East Africa, S. schall and S. frontosus, most probably synonymous, hence requiring taxonomical reassessment. Additionally, from a strictly phylogenetic point of view, the clustering of Synodontis sequences assigned to several species (S. ouemeensis, S. aff. bastiani, S. aff. haugi and S. schall) implies that these data might represent a single species S. schall Western clade, especially considering uncertain status of many Synodontis nominal species.
Based on the following criteria, we assign the Nile-Turkana population, i.e. S. schall Eastern clade, as represented in our sample to S. schall sensu stricto: (i) the most prevalent morphotype matching S. schall, (ii) phylogenetic affinity (see below), (iii) geographic origin matching with the type locality in the Nile and (iv) with respect to probable synonymy, the specific name S. schall has priority over S. frontosus. The interpretation is further indirectly supported by the supposed absence of S. frontosus in Lake Turkana itself, being presumably limited to Omo River (as highlighted by Hopson 1982 and other authors). Although in our Turkana sample S. frontosus morphotype was not restricted to the freshwater Omo delta, it was most prevalent here, whereas the S. schall morphotype was the most prevalent in the Turkana saline and medium salinity parts. Differences in the prevalence of morphotypes between the localities may also coincide with the moderate (intraspecific) level of genetic differentiation found in the population genetic analysis of coxI (Snn test, Table S4). The coxI pattern is probably caused by an ongoing process of lineage sorting. No evidence for morphotype or genetic differentiation was found in the genetically more diverse cytb.
Mitochondrial data revealed no significant differentiation between specimens representing S. schall and S. frontosus morphotypes, indicating their possible conspecificity. However, it should be noted that an alternative evolutionary scenario could generate such a genetic pattern - a complete fixation (or replacement) of mitochondrial DNA introgressed from one species to another. Unfortunately, we were not able to corroborate the mitochondrial results by an independent marker. The phylogeny based on the nuclear RAG2 gene was not informative, due to the lack of resolution in the respective part of the tree. Lack of representative data from the Western Nilo-Sudanian province, together with the generally low variability of the gene did not allow clear distinction of mitochondrial lineages in the nuclear dataset. Although generally rare, complete fixation of introgressed mitochondrial DNA was found for example in African cichlids51,52,53 and coral reef fish54. Analyses of more variable nuclear loci (e.g. introns or whole genome analyses) would be required to completely exclude such a possibility in S. schall-like catfishes. However, considering the weak differentiation of morphological traits defining S. schall and S. frontosus, as opposed to the high level of cytb diversity required to be introgressed between the two species across all sampled localities, we consider the “introgression and fixation” alternative to be much less probable than conspecificity.
Concurrent morphological and molecular sampling throughout the Nile basin, including terra typica of both S. schall and S. frontosus, i.e. Nile River in Assouan and in Khartoum, respectively, is necessary for sound taxonomical measures (we presume synonymy of S. frontosus and S. schall). Unfortunately, studies of other authors do not aid further resolution, because even recent taxonomic studies17,20,37,38 on Nilo-Sudanian Synodontis spp. lack molecular phylogenetic analyses, and vice versa, all available phylogenetic studies lack clues for the determination of sampled specimens, and/or the exact origin of the samples17,28.
Hidden diversity of S. schall-like catfish of the Nilo-Sudanian ichtyological province
The combined Nile-Turkana and GenBank dataset comprises all major limnic systems of the Nilo-Sudanian province, i.e. rivers Gambia, Senegal, Volta, Niger, Nile (including Lake Albert), endorheic basins of Chad and Turkana, as well as Eburneo-Ghanian (sub) province. As such, the presented dataset allows us to draw conclusions regarding diversity of S. schall-like catfish throughout the Nilo-Sudanian province. The results of the phylogenetic reconstruction, haplotype analyses and genetic distance recognized different S. schall clades. Phylogenetic analyses of both mitochondrial markers unequivocally revealed separate S. schall Eastern and Western clades, each comprising a mixture of samples assigned by different authors to S. schall, S. frontosus or other taxa. In addition to that, the genetic similarities between S. schall Eastern and S. schall Western clades sequences showed more than 5% divergence for both mitochondrial genes, a phenomenon generally associated with taxa recognized as distinct species55. Regardless of GenBank taxonomic sequence assignment, the two well supported clades, S. schall Eastern and Western clade, exclusively comprise either samples from eastern part of the Nilo-Sudanian ichthyological province (Nile, Turkana, Chad), or those from its western part (e.g. Senegambia, Niger, Chad), respectively. The taxonomically incongruent, but geographically non-random phylogenetic pattern reveals a strong biogeographical signal among the supposedly homogenous, conspecific and widely distributed S. schall-like catfish.
Synodontis schall was long considered morphologically uniform and widely distributed throughout the Nilo-Sudanian ichtyological province from the Atlantic to Indian Ocean17,21. In contrast, our results imply the presence of two relatively distant phylogenetic lineages hidden under this name. Synodontis schall sensu stricto, as defined by location of type locality in the Nile, corresponds to the Eastern clade in our analyses, and is restricted to the province’s eastern part, i.e. Nile, Turkana and Chad basins, and possibly elsewhere in north-east Africa (Fig. 6). On the other hand, S. schall-like catfish from the western part of the Nilo-Sudanian province (S. schall Western clade in the present study) probably represent a cryptic species more closely related to congeners from the same region than to their eastern siblings. Interestingly, there seems to be an overlap in distribution ranges of the “Eastern clade” and the “Western clade” in the Chad basin, probably as a result of independent and temporally distinct colonization events.
The phenomenon of cryptic, geographically determined diversity in African fish described herein is not a new discovery. A similar pattern was described for African tigerfish genus Hydrocynus56. The cryptic diversity, i.e. presence of three lineages among Hydrocynus vittatus populations from three separate drainages (Zambezi, Congo and Okavango), was revealed based on a combination of morphological and molecular (cytb) data. Similarly, as in S. schall-like catfishes in this study, the diversity of H. vittatus has long been overlooked due to the absence of definitive morphological features and molecular evidence. Another possible example of overlooked diversity among African fish, but revealed morphologically in this case, might be two cyprinid species, Labeo niloticus and L. senegalensis. The two Labeo spp. show a distribution pattern almost identical to that of S. schall-like catfishes are also distinguished by subtle, sometimes overlapping morphological features. Morphological studies including samples from western (L. senegalensis) and eastern (L. niloticus) parts of Nilo-Sudan ichthyological province revealed apparent hidden diversity correlated with geographical origin of specimens19,57. However, molecular data are not yet available for L. senegalensis and L. niloticus and their species status remains to be confirmed. The multiple examples of overlooked diversity in H. vittatus and Labeo spp., as well as S. schall-like catfish herein, suggest that the phenomenon of hidden diversity among long known and geographically widely distributed African fishes might be more common than previously thought.
Conclusions
Synodontis schall (Bloch et Schneider, 1801), as it has been defined and recognized for more than two centuries, apparently includes two distinct phylogenetic units (Eastern and Western clades), i.e. two distinct species are hidden within this name. The Eastern clade represents S. schall sensu stricto sensu stricto as defined by the type locality in the Nile, while the Western clade from the western part of Nilo-Sudanian province probably represents a cryptic species, which remained unrecognized due to a lack of differentiating morphological traits. A candidate name Synodontis gambiensis Günther, 1864 (Terra typica: Gambia), currently considered a junior synonym of S. schall, is available for eventual taxonomical amendment of the Western clade. In contrast, another long-recognized nominal species, S. frontosus Vaillant, 1895, might be merely a junior synonym of S. schall sensu stricto. An interesting, rather unusual taxonomical case has emerged from this study, since concurrent taxonomical deflation (nominal species synonymisation) and taxonomical split (cryptic species formal recognition) might be inevitable in the future for the two long recognized congeners.
Change history
20 December 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Gill, B. A. et al. Cryptic species diversity reveals biogeographic support for the ‘mountain passes are higher in the tropics’ hypothesis. Proc. R. Soc. B Biol. Sci. 283, 20160553 (2016).
Hebert, P. D. N., Penton, E. H., Burns, J. M., Janzen, D. H. & Hallwachs, W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci. 101, 14812–14817 (2004).
Jörger, K. M. & Schrödl, M. How to describe a cryptic species? Practical challenges of molecular taxonomy. Front. Zool. 10, 59 (2013).
Cooke, G. M., Chao, N. L. & Beheregaray, L. B. Divergent natural selection with gene flow along major environmental gradients in Amazonia: insights from genome scans, population genetics and phylogeography of the characin fish Triportheus albus. Mol. Ecol. 21, 2410–27 (2012).
Piggott, M. P., Chao, N. L. & Beheregaray, L. B. Three fishes in one: Cryptic species in an Amazonian floodplain forest specialist. Biol. J. Linn. Soc. 102, 391–403 (2011).
Souza, C. R., de Mello Affonso, P. R. A., de Araújo Bitencourt, J., Sampaio, I. & Carneiro, P. L. S. Species validation and cryptic diversity in the Geophagus brasiliensis Quoy & Gaimard, 1824 complex (Teleostei, Cichlidae) from Brazilian coastal basins as revealed by DNA analyses. Hydrobiologia 809, 309–321 (2018).
Katongo, C., Koblmüller, S., Duftner, N., Makasa, L. & Sturmbauer, C. Phylogeography and speciation in the Pseudocrenilabrus philander species complex in Zambian Rivers. Hydrobiologia 542, 221–233 (2005).
Day, J. J. et al. Multiple independent colonizations into the Congo Basin during the continental radiation of African Mastacembelus spiny eels. J. Biogeogr. 44, 2308–2318 (2017).
Martin, A. P. & Bermingham, E. Regional endemism and cryptic species revealed by molecular and morphological analysis of a widespread species of Neotropical catfish. R. Soc. 267, 1135–1141 (2009).
Souza-Shibatta, L. et al. Cryptic species of the genus Pimelodella (siluriformes: Heptapteridae) from the Miranda river, Paraguay River basin, Pantanal of Mato Grosso do Sul, Central Brazil. Neotrop. Ichthyol. 11, 101–109 (2013).
Prizon, A. C. et al. Hidden diversity in the populations of the armored catfish Ancistrus Kner, 1854 (Loricariidae, Hypostominae) from the Paraná River basin revealed by molecular and cytogenetic data. Front. Genet. 8 (2017).
Wright, J. J. & Bailey, R. M. Systematic revision of the formerly monotypic genus Tanganikallabes (Siluriformes: Clariidae). Zool. J. Linn. Soc. 165, 121–142 (2012).
Barasa, J. E. et al. Genetic diversity and population structure of the African catfish, Clarias gariepinus (Burchell, 1822) in Kenya: implication for conservation and aquaculture. Belgian J. Zool. 147, 105–127 (2017).
Schmidt, R. C., Bart, H. L. & Pezold, F. High levels of endemism in suckermouth catfishes (Mochokidae: Chiloglanis) from the Upper Guinean forests of West Africa. Mol. Phylogenet. Evol. 100, 199–205 (2016).
Schmidt, R. C., Bart, H. L., Nyingi, D. W. & Gichuki, N. N. Phylogeny of suckermouth catfishes (Mochokidae: Chiloglanis) from Kenya: The utility of Growth Hormone introns in species level phylogenies. Mol. Phylogenet. Evol. 79, 415–421 (2014).
Morris, J. et al. High levels of genetic structure and striking phenotypic variability in a sexually dimorphic suckermouth catfish from the African Highveld. Biol. J. Linn. Soc. 117, 528–546 (2016).
Musschoot, T. & Lalèyè, P. Designation of a neotype for Synodontis schall (Bloch and Schneider, 1801) and description of two new species of Synodontis (Siluriformes: Mochokidae). J. Nat. Hist. 42, 1303–1331 (2008).
Poll, M. Révision des Synodontis africains (famille Mochokidae). Ann. – Musée R. l’Afrique Cent. Zool. 191, 1–497 (1971).
Lévêque, C., Paugy, D. & Teugels, G. G. Annotated check-list of the freshwater fisltes of the Nilo-sudan river basins in Africa. Changes 24, 131–154 (1991).
Bailey, R. G. Guide to the fishes of the River Nile in the Republic of the Sudan. J. Nat. Hist. https://doi.org/10.1080/00222939400770501(1994).
Paugy, D., Roberts, T. R. & Mochokidae, P. In C. Lévêque, D. Paugy and G.G. Teugels (eds.) Faune des poissons d’eaux douce et saumâtres de l’Afrique de l’Ouest, Tome 2. Coll. 195–268 (Faune et Flore tropicales 40. Musée Royal de l’Afrique Centrale, Tervuren, Belgique, Museum National d’Histoire Naturalle, Paris, France and Institut de Recherche pour le Développement, Paris, France. 815 p., 2003).
Hopson, A. Lake Turkana; A Report on the Finding of the Lake Turkana Project 1972–1975, Vols. 1–6. (Overseas Development Administration, London, 1982).
Jirsová, D., Štefka, J. & Jirků, M. Discordant population histories of host and its parasite: A role for ecological permeability of extreme environment? PLoS One 12, 1–25 (2017).
Eschmeyer, W. N., Fricke, R. & van der Laan, R. Catalog of fishes: genera, species, references, (http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp). Electronic version accessed 22/07/2017 (2017).
Froese, R. & Pauly, D. Fishbase. World Wide Web electronic publication, www.fishbase.org, version (02/2017). (2017). Available at, www.fishbase.org version (02/2017).
Koblmüller, S., Sturmbauer, C., Verheyen, E., Meyer, A. & Salzburger, W. Mitochondrial phylogeny and phylogeography of East African squeaker catfishes (Siluriformes: Synodontis). BMC Evol. Biol. 6, 49 (2006).
Pinton, A., Agnèse, J. F., Paugy, D. & Otero, O. A large-scale phylogeny of Synodontis (Mochokidae, Siluriformes) reveals the influence of geological events on continental diversity during the Cenozoic. Mol. Phylogenet. Evol. 66, 1027–40 (2013).
Day, J. J. et al. Continental diversification of an African catfish radiation (Mochokidae: Synodontis). Syst. Biol. 62, 351–365 (2013).
Sands, D. D. Catfishes of the World. Vol. 2: Mochokidae. (Dunure Enterprises, Dunure, Scotland, 1983).
Day, J. J. & Wilkinson, M. On the origin of the Synodontis catfish species flock from Lake Tanganyika. Biol. Lett. 2, 548–552 (2006).
Day, J. J., Bills, R. & Friel, J. P. Lacustrine radiations in African Synodontis catfish. J. Evol. Biol. 22, 805–17 (2009).
Mohamed, E. H. A. Characterization of two Synodontis (Siluriformes: Mochokidae) catfish species in the White Nile and Lake Nubia. Environ. Biol. Fishes 88, 17–23 (2010).
Luff, R. M. & Bailey, G. The Aquatic Basis of Ancient Civilisaticlns:the case of Synodontis schal2 and the Nile Valley. in Human Ecodynamics. Symposia of the Association for Environmental Archaeology (eds. Charles, R. & Winder, N.) 100–113 (Oxbow Books, 2000).
El-Kasheif, M. A., Authman, M. M. N. & Ibrahim, S. A. Environmental studies on Synodontis schall (Bloch and Schneider, 1801) (Pisces: Siluriformes: Mochokidae) in The River Nile at Gizza Sector, Egypt: biological aspects and population dynamics. J. Fish. Aquat. Sci. 7, 104–133 (2012).
Gownaris, N. J., Pikitch, E. K., Ojwang, W. O., Michener, R. & Kaufman, L. Predicting species’ vulnerability in a massively perturbed system: The fishes of Lake Turkana, Kenya. PLoS One 10, 1–24 (2015).
Sambrook, J. & Russell, D. W. Molecular Cloning: A Laboratory Manual. Cold Spring Harb. Lab. Press. Cold Spring Harb. NY 999 (2001).
Ivanova, N. V., Zemlak, T. S., Hanner, R. H. & Hebert, P. D. N. Universal primer cocktails for fish DNA barcoding. Mol. Ecol. Notes 7, 544–548 (2007).
Sullivan, J. P., Lundberg, J. G. & Hardman, M. A phylogenetic analysis of the major groups of catfishes (Teleostei: Siluriformes) using rag1 and rag2 nuclear gene sequences. Mol. Phylogenet. Evol. 41, 636–62 (2006).
Werle, E., Schneider, C., Renner, M., Völker, M. & Fiehn, W. Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res. 22, 4354–4355 (1994).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Posada, D. jModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 25, 1253–1256 (2008).
Huelsenbeck, J. P., Ronquist, F., Nielsen, R. & Bollback, J. P. Bayesian Inference of Phylogeny and Its Impact on Evolutionary Biology. Science (80-.). 294, 2310–2314 (2001).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Rambaut, A., Drummond, A., Xie, D., Baele, G. & Suchard, M. Tracer v1.7, Available from, http://beast.community/tracer (2018).
Leigh, J. W. & Bryant, D. Popart: Full-Feature Software for Haplotype Network Construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
Clement, M., Posada, D. & Crandall, Ka TCS: a computer program to estimate gene genealogies. Mol. Ecol. 9, 1657–9 (2000).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–2 (2009).
Tobe, S. S., Kitchener, A. & Linacre, A. Cytochrome b or cytochrome c oxidase subunit I for mammalian species identification-An answer to the debate. Forensic Sci. Int. Genet. Suppl. Ser. 2, 306–307 (2009).
Meyer, B. S., Matschiner, M. & Salzburger, W. Disentangling incomplete lineage sorting and introgression to refine species-tree estimates for Lake Tanganyika Cichlid fishes. Syst. Biol. 66, 531–550 (2017).
Koblmüller, S. et al. Phylogeny and phylogeography of Altolamprologus: ancient introgression and recent divergence in a rock-dwelling Lake Tanganyika cichlid genus. Hydrobiologia 791, 35–50 (2017).
Koblmüller, S. et al. Reticulate phylogeny of gastropod-shell-breeding cichlids from Lake Tanganyika - The result of repeated introgressive hybridization. BMC Evol. Biol. 7, 1–13 (2007).
Harrison, H. B. et al. Widespread hybridization and bidirectional introgression in sympatric species of coral reef fish. Mol. Ecol. 26, 5692–5704 (2017).
Bradley, R. D. & Baker, R. J. A test of the Genetic Species Concept: cytochrome-b sequences and mammals. J. Mammal. 82, 960–973 (2001).
Goodier, S. A. M., Cotterill, F. P. D., O’Ryan, C., Skelton, P. H. & de Wit, M. J. Cryptic diversity of African tigerfish (genus Hydrocynus) reveals palaeogeographic signatures of linked Neogene geotectonic events. PLoS One 6 (2011).
Paugy, D., Zaiss, R. & Troubat, J. J. ‘Faunafri’. World Wide Web electronic publication, http://www.poissons-afrique.ird.fr/faunafri/, version (09/2017) (2008).
Acknowledgements
Our thanks are due to Turkana fishermen, field assistants and chiefs, as well as Father S.J. Ochieng and Father Fernando Aguirre, Our Lady Queen of Peace Todonyang Catholic Mission. Nevertheless, we also thank members of KMFRI for the supervision and help during the fieldwork. Sampling in the Nile was possible thanks to White Nile Fisheries Research Station, Kostí. We thank Thomas Vigliotta, Cornell University, for comments on the first draft and Ashlie Hartigan (Natural History Museum London) for correction of English. Our thanks are also due to reviewers and editor, who significantly aided the final version of manuscript. Authors greatly acknowledge Czech Science Foundation (P505/12/G112, 18-13458S), Institute of Parasitology (RVO: 60077344) and Grant Agency of University of South Bohemia (037/2011/P, 04-135/2010/P). Computational resources were provided by the CESNET LM2015042 and the CERIT Scientific Cloud LM2015085, provided under the programme “Projects of Large Research, Development, and Innovations Infrastructures”.
Author information
Authors and Affiliations
Contributions
D.J., M.J. and J.S. designed the study, planned the fieldwork, contributed to data analyses and interpretation and revised the manuscript. J.O.M., D.E.L. and Z.N.M. collected, imaged and measured the specimens. D.J. and M.J. performed fieldwork and sample collection, D.J. performed molecular work and analyzed sequences; R.B. and M.J. obtained morphometric data. D.J., J.S. and M.J. and contributed to data analyses and interpretation and wrote the manuscript. M.J. provided financial support and necessary experience of African geography. J.S. contributed to sequence analysis and manuscript writing. R.B. and M.J. morphologically identified the samples. All authors contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jirsová, D., Štefka, J., Blažek, R. et al. From taxonomic deflation to newly detected cryptic species: Hidden diversity in a widespread African squeaker catfish. Sci Rep 9, 15748 (2019). https://doi.org/10.1038/s41598-019-52306-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52306-2
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.