Abstract
Evolutionary theory predicts potential shifts between cooperative and uncooperative behaviour under fluctuating environmental conditions. This leads to unstable benefits to the partners and restricts the evolution of dependence. High dependence is usually found in those hosts in which vertically transmitted symbionts provide nutrients reliably. Here we study host dependence in the marine, giant colonial ciliate Zoothamnium niveum and its vertically transmitted, nutritional, thiotrophic symbiont from an unstable environment of degrading wood. Previously, we have shown that sulphidic conditions lead to high host fitness and oxic conditions to low fitness, but the fate of the symbiont has not been studied. We combine several experimental approaches to provide evidence for a sulphide-tolerant host with striking polyphenism involving two discrete morphs, a symbiotic and an aposymbiotic one. The two differ significantly in colony growth form and fitness. This polyphenism is triggered by chemical conditions and elicited by the symbiont’s presence on the dispersing swarmer. We provide evidence of a single aposymbiotic morph found in nature. We propose that despite a high fitness loss when aposymbiotic, the ciliate has retained a facultative life style and may use the option to live without its symbiont to overcome spatial and temporal shortage of sulphide in nature.
Similar content being viewed by others
Introduction
The notion that ‘all development is co-development’1 refers to the fact that hardly any animal or plant lives without microbial symbionts2,3. This entrenchment of environmental microbes4 has served innumerable times as an integral element for phenotypic construction and phenotypic novelty in eukaryote hosts5,6,7,8,9. Symbiont-induced developmental change in host traits has yielded complex phenotypes with adapted physiology to interact with the symbionts3,8,10,11,12,13.
The major transition from individuality of symbiotic partners in mutualistic relationships to a new integrated organism14 requires mutual dependence and alignment of partner interests15,16. Some symbionts are considered as being strictly required for normal host development and reproduction9,10, e.g. Buchnera aphidicola – aphids3, ‘Candidatus Endoriftia persephone’ – giant tubeworm17, ‘Candidatus Kentron’ – ciliate Kentrophoros18,19, Polynucleobacter – ciliate Euplotes20,21,22,23,24,25,26. Such obligate hosts bearing symbionts can display phenotypic plasticity – the ability of an organism to express different phenotypes in response to the varying environmental conditions4,27,28,29 - in various ways, but importantly, they do not retain the option to live aposymbiotically30.
In some other associations, however, not becoming irreversibly dependent has major evolutionary advantages. As environmental conditions strongly impact the outcome of interactions2,3, the elimination of aposymbiotic hosts is not selected for. Such facultative hosts have the option to interact with the partner as well as to live aposymbiotically1,8. For example, legumes develop root nodules in response to rhizobia infection only when fields are not fertilized with nitrogen31. The symbiotic and aposymbiotic anemone Anthopleura elegantissima occurs, according to light regime, either with Symbiodinum on sun-exposed or without these microalgae on shady substrates32. The endosymbiotic R-body producer Caedibacter transforms the host Paramecium into a killer of other paramecia including sometimes aposymbiotic individuals of the same species33,34. Most often Paramecium bursaria is found with photoautotrophic Chlorella variabilis35 but also natural aposymbiotic hosts exist36.
Some hosts, however, are never found in nature without their symbionts although they can be experimentally purged from them and still grow and sometimes even reproduce, e.g. Aliivibrio fischeri and bobtail squids37, methanogen bacteria and Metopus contortus ciliate38. To estimate host dependence over a range of animal and plant associations, the relative drop of host fitness between symbiotic and the experimentally purged or naturally occurring aposymbiotic hosts was calculated39. Overall hosts tend to be more dependent on vertically transmitted and nutritional symbionts than on horizontally transmitted and protective symbionts39. In ciliates fitness differences between symbiotic and aposymbiotic populations indicate the mutualistic advantage for the partners in nutritional38 and defensive associations34. Fitness, however, was found to be highly context dependent40. Under a variety of abiotic and biotic conditions ciliate mutualism might shift to a host exploiting the symbiont41 or a symbiont turning into a parasite42.
The mutualism between the vertically transmitted and nutritional, sulphur-oxidizing, chemoautotrophic (thiotrophic) ectosymbiont ‘Candidatus Thiobios zoothamnicoli’ and its giant colonial ciliate host Zoothamnium niveum43,44 is most suitable to test the hypothesis that symbiont function and transmission influence host dependence39. According to phylogenetic analyses based on the 18S rRNA gene sequence, Z. niveum belongs to the monophyletic clade 245 (termed clade 146) within a non-monophyletic genus45,46,47,48,49,50,51. Morphologically clade 2 species exhibit a colony growth pattern of alternate branches45. They share with all Zoothamnium species a suite of morphological characters, e.g. a common stalk connecting all zooids and containing a continuous spasmoneme that facilitates contraction in a “zigzag“ pattern45,46,49. Besides Z. niveum, most other clade 2 species are associated with ectosymbiotic bacteria: Z. alternans52,53, Z. ignavum51, Z. pelagicum54,55,56. Only in Z. plumula symbionts are not mentioned57,58.
In contrast to other Zoothamnium species of clade 2, which occur in oxic environments51,53,54,55,57,59, the Z. niveum partnership thrives in unstable and highly disturbed decaying plants and whale bones60 in the presence of hydrogen sulphide (∑H2S, i.e. sum of all forms of dissolved sulphide61, hereafter termed sulphide). The gammaproteobacterial symbiont has genes for sulphur oxidation and carbon fixation indicating a thiotrophic metabolism62.
Individual cells (termed zooids) of the colony are differentiated with different functions for division (terminal zooids), feeding (microzooids), and asexual reproduction (macrozooids). The terminal zooid on the tip of the stalk produces the terminal zooids on each branch, which in turn produce feeding microzooids and the macrozooids on the branches63. Vertical transmission of the ectosymbiont is through macrozooids that leave the colony as swarmers43,44, recruit to sulphide-emitting surfaces and grow into new colonies64,65. To date, the symbiont has neither been detected free-living in the environment nor has it been cultivated (MB unpublished data).
Through colony contraction and expansion, the colony dips into the sulphidic layer close to the substrate and extends into the oxic layer above, thereby alternately providing access to sulphide and oxygen as a byproduct benefit to the symbiont60. The symbiont fixes substantial amounts of inorganic carbon66. Host nourishment involves translocation of organic carbon to the host through both passive release, considered a byproduct benefit, and through digestion of symbionts66. Trading of goods in the mutualism, however, is interrupted when sulphide ceases during the cold season in temperate waters67.
The Z. niveum mutualism is currently the only thiotrophic association that can be cultivated over several generations68. Experiments of the host under oxic, sulphide supplemented, flow-through conditions lead to fast growth, long life span, and high reproduction. The colonies are white because they are covered by symbionts that store elemental sulphur. Under oxic, flow-through conditions host reproduction was reduced and colonies were pale and short-lived68. At that time we did not investigate whether these small colonies still carried symbionts. We therefore hypothesized that swarmers lose their symbionts during dispersal, that settlement is preferentially on sulphide-emitting surfaces but may also be possible on oxic surfaces, and that aposymbiotic swarmers grow into aposymbiotic colonies.
To test these hypotheses, we designed laboratory experiments under oxic conditions that mimic cessation of sulphide flux in nature to explore the response of swarmers and colonies, and the fate of the symbiont on the host. Here, we report on experimental evidence for a striking polyphenism, i.e., the development of discrete alternative phenotypes30. We show that the environmental conditions encountered by the swarmer lead either to loss or maintenance of the symbiont. This triggers the developmental switch, expressed in two distinct colony growth forms. We developed a growth form index for statistical comparison. Our experiments show that each growth form performed better under the respective environmental conditions. The 18S rRNA gene sequences of two colonies collected in the field – one resembling the aposymbiotically grown morph, the other the symbiotic morph – were identical. This confirmed that both morphs occur in nature.
Results
Swarmers during dispersal
All swarmers kept under oxic, stagnant conditions for 4 h were fully covered by the symbiont, (n = 11, Fig. 1a,b and Supplementary Table S1). After 24 h, swarmers (n = 16) were either still fully covered (31%, Fig. 1c,d), partially covered (31%, Fig. 1e,f), or aposymbiotic (38%, Fig. 1g,h). All swarmers were aposymbiotic after 48 h (n = 14, Fig. 1i,j).
Swarmer recruitment to sulphide-emitting surfaces
In oxic, stagnant seawater only about 1% of the released swarmers settled within 72 h (n ≈ 3,000). These swarmers were pale. Because of the low swarmer recruitment, we used a preference experiment (n = 6) to test whether reduced sulphur species (sulphide, thiosulphate) or low oxygen concentrations act as a settlement cue. Settlement checked after 24 h revealed an overall median of 22.5% recruited swarmers (interquartile range IQR 6.6%) exclusively to sulphide-emitting surfaces. Settlement was not significantly different between high and low sulphide concentrations (Wilcoxon-Mann-Whitney test, p-value 0.85, Supplementary Table S2). No settlement occurred at the thiosulphate, the reduced oxygen membranes, or the walls of the experimental chamber exposed to oxic seawater.
Because the swarmer recruitment experiment was carried out without chemical measurements, this experiment was repeated without swarmers to measure oxygen and sulphide concentration at the four membrane surfaces and revealed the presence of higher sulphide and lower oxygen at the high sulphide membrane and lower sulphide and higher oxygen at the low sulphide membrane until the end of the experiment. At the thiosulphate membrane, the oxygen concentration was similar to the concentration measured in the chamber, and sulphide was absent, whereas oxygen was very low and sulphide was absent at the membrane of the N2-bubbled vial (Supplementary Table S2).
To investigate whether this selective host behaviour was influenced by the presence of the symbiont on the host over time, a mix of aposymbiotic and symbiotic swarmers was exposed to sulphide under stagnant conditions to avoid removal of swarmers from the chamber under flow-through conditions (n = 17). A median of 47.7% (IQR 20.4%) swarmers recruited to surfaces within 23 h. Distance covariance tests showed that the time of sulphide exposure – between 2 and 23 h – had no influence on the number of settled symbiotic (p-value 0.28), aposymbiotic (p-value 0.27) and total swarmers (p-value 0.21) (Supplementary Table S3).
Effects of sulphide and food supply on aposymbiotic host traits
Settled pale swarmers grew into colonies both under oxic, flow-through and oxic, sulphide-supplemented, flow-through conditions. At the end of the experiment at day 7 scanning electron micrographs (SEM) confirmed that most colonies were without microbial overgrowth (Fig. 2). Because in a few colonies some patches of microbes were seen, we performed fluorescence in situ hybridisation (FISH) on semithin sections using the symbiont-specific and the EUBmix and Arch915 probes to identify the microbes. Also FISH micrographs revealed that most colonies were aposymbiotic (Fig. 3a–d). In some cases, however, other microbes colonized the host surfaces in small patches, which were labelled with the EUBmix and Arch915 probes but not with the symbiont-specific probe (Fig. 3e–h). In contrast, the symbiotic, white colonies were covered with a monolayer of symbionts with positive EUBmix and Arch915 and symbiont-specific labels (Fig. 3i–l). During the 7-day experiments no colony switched from pale to white under either chemical conditions. Under either condition, a few macrozooids were produced (Fig. 2b,h). Release of macrozooids and settlement of swarmers could not be followed in this experiment.
FISH micrographs of aposymbiotic morph grown under sulfidic conditions for seven days, stained with DAPI (a,e) (n macronuclei of host cells), labelled with EUBmix and Arch915 probes (b,f), and symbiont-specific probe (c,g), and overlay (d,h) arrows points to microbes other than symbionts (e,f). Symbiotic morph grown under sulfidic conditions for seven days stained with DAPI (i)(n macronuclei of host cells), labelled with EUBmix and Arch915 probes (j), and symbiont-specific probe (k) and overlay (l); arrows point to symbionts. Note that all micrographs are the same magnification.
At in situ microbial abundance (MA) aposymbiotic colonies had an estimated life span of 13.2 d (95% confidence interval 12.1, 15.3) under oxic, sulphide supplemented and 8.1 d (95% confidence interval 7.9, 8.2) under oxic conditions (Supplementary Table S4, Fig. 4a,d). The estimated maximal colony size reached a median of 7.2 branches (95% confidence interval 7.0, 7.4) in 4.0 d (95% confidence interval 4.0, 4.1) under oxic conditions and only 5.8 branches (95% confidence interval 5.2, 6.4) in 6.4 d (95% confidence interval 6.0, 7.3) under sulphidic conditions (Supplementary Table S4, Fig. 4a,d). Accordingly, colonies in oxic seawater grew faster with a relatively short, estimated life span, but to larger sizes, while oxic conditions supplemented with sulphide led to slow growth (hence longer life span) and smaller sizes.
Box plots of growth (number of branches) of aposymbiotic colonies with in situ (a), low (b) and high (c) MA in sulphidic and oxic seawater (Wilcoxon-Mann-Whitney comparisons for each day; *p-value < 0.05, **p-value < 0.01, ***p-value < 0.001); (d) Estimated life span and maximal colony size of the aposymbiotic phenotype under sulphidic (red lines) and oxic (blue lines) conditions with reduced (thin line), in situ (medium line), and enhanced (thick line) MA, are inferred from the parabolas and 95% confidence intervals (dotted lines); (e) Box plots of growth of symbiotic and aposymbiotic colonies grown in the same chamber for seven days (Wilcoxon-Mann-Whitney comparisons for each day; **p-value < 0.01, ***p-value < 0.001); (f) Counted branches and total number of zooids in symbiotic and aposymbiotic colonies of Z. niveum and Z. ignavum, in double logarithmic scale of base 10 with power law fits.
Food density had no direct influence on the growth of aposymbiotic ciliates over a MA range of 2.0 × 105 to 1.6 × 106 mL−1. Under oxic conditions, however, reduced MA restricted maximal colony sizes versus the larger sizes under in situ and enhanced MA (Fig. 4a–d). Survival of aposymbiotic colonies under oxic conditions ranged from 0% (enhanced MA), 25% (reduced MA) to 30% (in situ MA) (Supplementary Table S4). Under oxic, sulphide supplemented conditions, reduced and in situ MA led to similar estimated maximal colony sizes, whereas enhanced MA resulted in larger sizes (Fig. 4a–d). Survival of aposymbiotic colonies under oxic, sulphide supplemented conditions ranged from 28% (reduced MA), 64% (enhanced MA) to 100% (in situ MA).
Growth of symbiotic and aposymbiotic Z. niveum colonies
The morphology differed remarkably between colonies grown from aposymbiotic and symbiotic swarmers under oxic, sulphide supplemented conditions (Supplementary Table S5). Survival was 100% in aposymbiotic morphs and 66% in symbiotic morphs (Supplementary Table S5). After settlement, the small colonies were morphologically similar (Fig. 2c,d,f,g). During further development, however, the colonies with 10 (Fig. 2e,h) or 11 branches (Supplementary Fig. S2) differed between aposymbiotic and symbiotic phenotypes. This indicates that the proliferation activity of the terminal zooid of each branch was comparatively higher in the aposymbiotic than in the symbiotic phenotype. In contrast, the proliferation activity of the top terminal zooid was comparatively higher in the symbiotic compared to the aposymbiotic phenotype (Fig. 4e, Supplementary Table S5) leading to a long but narrow symbiotic morph and a short but wide, aposymbiotic morph. Aposymbiotic colonies grown from aposymbiotic swarmers under oxic conditions were identical in morphology to those grown under oxic, sulphide supplemented conditions (Fig. 2).
Comparing both morphs under oxic, sulphide supplemented, in situ MA conditions, growth was significantly higher in the symbiotic phenotype from day 2 to day 7 (Fig. 4e, Supplementary Fig. S1, Wilcoxon-Mann-Whitney tests, p < 0.001). At the end of the experiment on day 7, symbiotic colonies exhibited a median of 22 branches bearing an estimated 107 zooids, while aposymbiotic colonies had 6 branches with an estimated 24 zooids only (Supplementary Table S5). The aposymbiotic life span was limited to an estimated 13.2 d, but the symbiotic growth followed a monotonically increasing function (e.g. positive parabolic linear fit r2 0.70, p-value 2 × 10−44, n = 169). The symbiotic morph therefore grew faster and to larger sizes. The estimated growth curve showed that the expected life span is much longer in the symbiotic than the aposymbiotic morph (Fig. 4e) confirming previous results68.
Under oxic, in situ MA conditions, none of the symbiotic morphs were found after seven days. Survival of aposymbiotic morphs was 30% (Supplementary Table S4). Recruited white swarmers turned pale within the first 24 h and grew into aposymbiotic morphs, which was confirmed with SEM at the end of the experiment at day 7.
Growth form index (GFI)
A colony GFI (exponent of the power law relating zooids and branches) was obtained from aposymbiotic and symbiotic Z. niveum. Because the aposymbiotic morph highly resembled the symbiotic Z. ignavum in morphology we also obtained a GFI for comparison from this closely related species collected in the field51. The aposymbiotic morph had a GFI of 0.47 (n = 45, 95% confidence interval 0.40, 0.54), significantly different (p-value 5 × 10−9, analysis of covariance) from the symbiotic morph (GFI of 0.73, n = 51, 95% confidence interval 0.67, 0.78) (Fig. 4f, Supplementary Fig. S2). Symbiotic Z. ignavum showed a GFI of 0.49 (n = 30, 95% confidence interval 0.46, 0.52), non-significantly different to the aposymbiotic Z. niveum phenotype (p-value 0.73) but significantly different to the symbiotic Z. niveum phenotype (p-value 4 × 10−12) (Fig. 4f, Supplementary Fig. S2).
Host 18S rRNA and symbiont 16S rRNA genes sequencing and phylogenetic analysis
Most of the colonies collected from minimally degraded wood and resembling the pale aposymbiotic phenotype were too low in DNA content to be sequenced (#4697/2-4,6,7). From one of these colonies (#4697/5, DNA content 3.6 ng/µL) 1,466 bp of the 18S rRNA gene could be retrieved. From a symbiotic, white colony collected from highly degraded wood (#4577, DNA content 32.4 ng/µL) 1,537 bp of the 18S rRNA gene were obtained. The sequence was identical to the pale colony (Supplementary Figs S3, S4) but differed to Z. niveum colony from USA49 (Sequence similarity 99.5%) (Supplementary Fig. S4). All three Z. niveum colonies build a monophyletic subclade in clade 2 supported by 100% posterior probability (Bayesian inference, BI) and 100% bootstrap support (Maximum Likelihood, ML; Maximum Parsimony, MP; Fig. 5). The presence of the symbiont 16S rRNA gene using general primers could only be obtained from the symbiotic colony indicating that the aposymbiotic phenotype was containing microbes (including the symbiont) too low in abundance to be sequenced (Supplementary Fig. S3).
Bayesian tree inferred from the nucleotide sequences of the small subunit 18S rRNA gene of the monophyletic clade 2 of Zoothamnium and combined with colony drawings and life style. Support metrics are provided (BI/ML/MP). Scale bar corresponds to 1 substitution per 200 nt positions; numbers in parentheses are the NCBI accession numbers for each species. All colony drawings, including Z. alternans53, Z. pelagicum54, Z. plumula57, Z. ignavum51 and Z. niveum symbiotic43 and aposymbiotic Z. niveum (drawing from a colony grown in a flow-through chamber) show macrozooids in grey. Colony drawing of Z. plumula57 is under copyright and its use was granted by Magnolia Press www.mapress.com/j/zt. Colony drawings reproduction of Z. alternans53 was granted by the Instytut Biologii Doświadczalnej im. M. Nenckiego, and of Z. niveum43 by Elsevier. Colony drawing of Z. pelagicum54 is under copyright and its use was granted by CNRS Éditions (M. Laval; Zoothamnium pelagicum du Plessis. Cilié Péritriche planctonique: morphologie, croissance et comportement in Protistologica n°4 ©CNRS Éditions, 1968).
Discussion
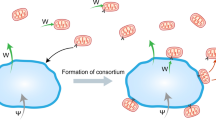
The thiotrophic mutualism between Zoothamnium niveum and its symbiont faces the challenge of maintaining long-term stability in a notoriously unstable environment. Sulphide and oxygen are not always available, and unless the host provides access to these chemicals to the symbiont, the symbiont cannot provide organic carbon to the host41. This study reports the discovery of one aposymbiotic ciliate in nature and shows experimentally, how this ciliate loses its symbiont when trading of goods between partners is interrupted under oxic conditions (Fig. 6). Most interestingly this ciliate exhibits a polyphenism in colony growth form. Despite high dependency, indicated by a fitness drop of 73% growth reduction in the aposymbiotic phenotype (Supplementary Table S5) and consistent with other hosts of vertically transmitted and nutritional symbionts39, this points to a facultative host. We propose that the aposymbiotic life style may ensure survival of the host at a temporal or spatial lack of sulphide in the environment.
Proposed development of symbiotic morphs from symbiotic swarmers released from colony and short migration through oxic water prior to settlement on sulphide-emitting wood surface. In contrast, long migration of swarmers in oxic seawater leads to loss of symbionts and development of aposymbiotic morphs under both oxic and sulphidic conditions.
New colonies can arise from swarmers under prolonged oxic conditions68. Here, we showed that swarmers easily lose the symbiont. Swarmers preferentially seek sulphidic surfaces for settlement to optimize symbiosis survival, but can also settle and grow aposymbiotically under unfavorable, oxic conditions, albeit to a very low percentage (Fig. 6). The time window for the swarmer to maintain its symbiont under oxic conditions is between 24 and 48 h. At a swarmer swim speed69 of 5 mm s−1, between 400 m and 800 m can be covered in one and two days, respectively. Accordingly, failing to find a sulphidic settlement near its release site leads to symbiont loss. The lack of a fully developed cytopharynx in swarmers suggests that symbionts cannot be digested43. It remains to be determined whether the host eliminates the symbiont because no benefits are provided and/or additional costs arise for the host to carry the symbiont during dispersal70, or whether the symbiont dies or leaves the host due to adverse conditions.
Our experiments indicate that sulphide is the settlement cue. This is consistent with previous experiments in which swarmers settled at the edge of sulphide point sources on cut-out blocks of mangrove peat placed in aquaria65. Only 1% of swarmers settled within 72 h during dispersal in oxic seawater. Although the percentage of recruited aposymbiotic colonies is low, in nature the high density of symbiotic populations producing large numbers of swarmers might nevertheless allow survival of aposymbiotic host populations. Based on an average number of 15.5 ± 2.8 (mean ± standard error) macrozooids per colony63 and 1,200 colonies m−2 on mangrove peat64) we estimate that well over 150 aposymbiotic colonies would be produced under oxic conditions from populations inhabiting a single square meter. Equally densely colonized wood (Supplementary Fig. S5), whale bones71, and seagrass debris68 have been documented elsewhere as well.
Most remarkably, this colonial ciliate exhibits a polyphenism in colony growth form, discernible as a long and narrow form in the symbiotic morph and a short and wide form in the aposymbiotic morph. The two morphs differ significantly in their growth form index. Polyphenism is well known from predator – prey, parasitic, and competitive interactions72. Other microbe – ciliate mutualisms are well known for their facultative host life style73,74. Often aposymbiotic and symbiotic morphs are quite similar in these unicellular hosts. Metopus contortus can be grown with its methanogen endosymbionts as well as aposymbiotically, revealing little morphological changes38. In the ciliate Euplotidium itoi, however, the presence of epixenosomes, unique ectosymbionts related to Verrucomicrobia75 defending their host against predators76, symbiotic morphs differ from artificially purged aposymbiotic morphs in the presence of a widened cortical region that the ectosymbionts are attached to77.
In the presence of sulphide in our experimental chamber, symbiotic and aposymbiotic swarmers gave rise to symbiotic and aposymbiotic morphs, respectively. In the absence of sulphide, only the aposymbiotic morph developed. In both cases the alternative phenotype exhibited significantly reduced growth indicating that each phenotype may be adaptive for the respective environment. The growth form expressed in oxic environments without symbionts experimentally would no doubt help the population to survive when swarmers fail to find a sulphidic substrate when such substrates are scarce or temporarily lacking at winter temperatures67. The finding of a single aposymbiotic morphotype of Z. niveum in nature may point to a much larger and more diverse habitat for this ciliate. Such aposymbiotic growth forms were not found on sulfide-emitting wood (MB pers. obs.). Whether host plasticity still retains the option to live aposymbiotically in nature or only in the lab by reverting to the growth form that they share with other species of clade 2 from oxic environments remains to be further investigated. The preferential recruitment behavior of the swarmer to abundant sulfide-emitting habitats and biological interactions such as competition and predation pressure may exclude the prevalence of aposymbiotic phenotypes when sulfide is produced during the warm seasons. They might be crucial, however, when temperature drops in late fall and ensure survival during the cold season.
The different growth of aposymbiotic morphs grown under oxic versus oxic, sulphide supplemented conditions suggests that sulphide stresses the aposymbiotic host and impacts growth. The mitochondria of invertebrates and protists from sulphidic environments detoxify sulphide to thiosulphate in the presence of oxygen78,79. Our experiments point to a trade-off between growth and longevity. Growth was fast and led to larger sizes but shorter life spans under oxic conditions. Under oxic, sulphide supplemented conditions the energy-consuming processes of sulphide detoxification may slow growth and extend the life span.
The aposymbiotic and symbiotic morphs of Zoothamnium niveum grown in experimental chambers differ remarkably in morphological traits used in the classification of species in this genus. Without experimental proof these two highly different morphs would be classified as different species. The 18S rRNA gene sequences from one colony collected from the field and resembling the aposymbotically grown morph in the lab as well as the 18S rRNA gene sequence from the symbiotic colony from the field were identical and allowed us to identify them as Z. niveum. Both colonies from the Adriatic Sea have 99.5% similarity in their 18S rRNA gene sequence with the sequence from USA49. In contrast, Z. alternans also from geographically distant locations and considered morphologically identical53 exhibit 96.7% sequence similarity45. This rather points to different species51 when applying a threshold between 97 to 99%51 or 99% sequence similarity80. Even more so, Z. plumula gene sequences from two relatively nearby locations in China46,47 fall in different clades46. Overall, the presence of cryptic species in the genus Zoothamnium and the discovered polyphenism in Z. niveum warrants to call for the addition of possible symbiont descriptions on morphological and molecular level as well as host gene sequences applied when new species are described and when population from previously unknown locations are studied.
The closely related species49 of Zoothamnium niveum in the clade 2 grow predominantly in oxic environments51,53,54,55,57,59. They show a wide range of either symbiotic or aposymbiotic life styles but share the diagnostic characteristic of a growth form with alternating branches49. They resemble, at first glance, the aposymbiotic morph with relatively long branches51,53,54,55,57,59 (Fig. 5). We could confirm the similarity of GFIs of aposymbiotic Z. niveum (0.47) and symbiotic Z. ignavum (0.49), but for other species data are not yet available. Visually Z. alternans53 highly resembles both aposymbiotic Z. niveum and Z. ignavum. Also relatively long branches are found in symbiotic Z. pelagicum that is in fact a “pseudo-colony“, composed of several ‘short and wide’ colonies growing on each other54,55,56 and in aposymbiotic Z. plumula with additional secondary branches57. Future research will help to determine whether specific growth forms can be related to specific environmental conditions.
Conclusion
Our study revealed a mechanism of mutualism breakdown. Host development with and without the symbiont led to a polyphenism i.e. to discrete alternative colony growth forms triggered by the chemical conditions in the environment that the swarmer encounters. Prolonged oxic conditions lead to symbiont loss inducing the development of an aposymbiotic morph with reduced fitness, whereas symbionts were maintained during sulphidic or brief oxic conditions leading to the development of the symbiotic morph with high fitness. Whether aposymbiotic host populations play indeed a role in nature and if so, how the aposymbiotic host regains its symbiont to guarantee connectivity from aposymbiotic to symbiotic host populations remains to be studied.
Methods
Swarmers during dispersal
Colonies were collected from submerged wood at Sv. Jernej, Adriatic Sea, 2013 (replicate 1) and 2014 (replicate 2) (Supplementary Table S1). Colonies were cut from the substrate and transferred to embryo dishes filled with oxic seawater (24.5 °C ± 0.7 °C, salinity 33.3 ± 0.9, hereafter abiotic factors expressed as mean ± standard deviation). Release of swarmers was monitored after 4 h, 24 h, and 48 h (n = 2). Released swarmers were fixed for scanning electron microscopy (SEM) (Supplementary Table S1).
Swarmer recruitment in preference chambers
Colonies were sampled from concrete blocks surrounded by seagrass debris in 2003 (Supplementary Table S2). For each experiment (n = 6), 30 to 50 released swarmers from about 50 colonies kept in oxic seawater (24 °C ± 1 °C, salinity 38 ± 1) were transferred into the central plastic cube (8 × 8 × 7 cm) of the preference chamber filled with oxic seawater. On each side of the cube, one vial was mounted and sealed with a dialysis polyethylene membrane (2.6 cm in diameter) permeable for small molecules to allow diffusion into the central chamber81 (Supplementary Fig. S1). The vials were filled with 30 mL seawater containing 1) ~1.5 mmol L−1 sulphide, 2) ~2.5 mmol L−1 sulphide, 3) 10 mmol L−1 thiosulphate, and 4) continuously N2-bubbled, anoxic seawater. After 24 h, the settlement of swarmers on the four membranes and on the inner cube’s surface was counted using a dissection microscope. Medians and quartiles were calculated and the nonparametric Wilcoxon-Mann-Whitney test was performed to assess differences between swarmer recruitment to high and low sulphide (R82 version 3.5.1, package Coin, v1.2-2).
Because abiotic parameters were not measured during this experiment in 2003, we repeated the experiment without swarmers to measure temperature, pH, salinity, oxygen, and sulphide in the chamber and oxygen and sulphide at all four membranes in October 2018 using a Fibox 4 (PreSenS, Germany) for oxygen and temperature, Multi 340i (WTW, Germany) for salinity and pH, and the Cline assay for ΣH2S83 (Supplementary Table S2).
Swarmer recruitment in flow-through chambers
Colonies were collected from diverse submerged wood in the Adriatic Sea in 2013 and 2014 (Supplementary Table S3). Flow-through chambers68 (n = 17; Supplementary Fig. S1) were filled with 17 mL of oxic seawater containing sulphide (147 ± 35 µmol L−1). Between 90 and 227 pale, aposymbiotic and white, symbiotic swarmers were added to each chamber. The time for settlement was kept between 2 and 23 h during which the pumps were switched off to avoid swarmers being flushed out of the chamber. Afterwards, settled pale and white swarmers were counted, sulphide was measured, and flow through the chambers was established to follow colony growth (Supplementary Table S3). The covariance test (R package Energy v1.7-5) was used to show independence of time of sulphide exposure and number of settled swarmers.
Effects of sulphide and food supply on aposymbiotic host traits
Colonies were collected from wood in Sv. Jernej in 2014 (Supplementary Table S4). For continuous flow through the chambers, we used an 8-channel peristaltic pump (Minipuls 3, Gilson International, Austria) set at 80 mL h−1 flow for oxic seawater and a syringe pump (KD Scientific, Inc., USA) set between 1.0 and 1.5 mL h−1 for sulphide supplementation in 50 mL syringes (between 1.5 and 1.8 mmol L−1 sulphide in Milli-Q water) (Supplementary Fig. S1). Three chambers were kept at oxic conditions, three others supplemented with sulphide. Two of these chambers (oxic and sulphidic) were fed with 32 µm filtered seawater to exclude eukaryote predation but to simulate the in situ microbial abundances (MA) commonly found in the northern Adriatic Sea during July84. To reduce the MA, we filtered the 32 µm pre-filtered seawater using the filter cartridge systems Polygard and Express SHC (Millipore, USA), with a final filtration of 0.2 µm pore size. To enhance the prokaryote density, we quadruple-concentrated 32 µm pre-filtered seawater using a Vivaflow 200 tangential flow module (Vivascience, Germany). Water and syringes were changed daily.
Abiotic parameters (flow, temperature, salinity, pH, oxygen, sulphide) and MA in the outflow water were monitored daily. Outflow water was filtered through polycarbonate filter membranes (Millipore GTBP02500 Isopore). Filters were fixed in 2% formalin for 24 h, air dried, and stored at −20 °C until staining with DAPI (4′,6-diamidino-2-phenylindole). MA was estimated by counting on an Axio Imager A1 epifluorescence microscope (Zeiss, Germany).
We followed survival as well as colony growth by counting the number of colonies and their respective branches under a dissection microscope daily. After 7 d the chambers were opened and colonies were fixed for SEM and fluorescence in situ hybridisation (FISH) (Supplementary Table S4). Shapiro-Wilk tests indicated deviations from normality for several of the experiments. Wilcoxon-Mann-Whitney tests were performed to assess differences of colony size between oxic and sulphidic conditions at low MA, in situ MA, and high MA. Linear regressions to quadratic polynomial parabolas (least squares fit) with Pearson R-square coefficients, and p-values were applied using R, because parabolas best reflected the growth and degenerative phase described for this ciliate68. Confidence intervals for parabolas were approximated by 10,000 random bootstrap re-samplings using the program ‘MUBOQB’.
Growth of symbiotic and aposymbiotic Z. niveum colonies
Using colonies collected at Sv. Jernej in 2014, both white symbiotic and pale aposymbiotic settled swarmers (Supplementary Table S3, experiment 68) grew together into white symbiotic and pale aposymbiotic colonies, respectively, in one flow-through chamber under in situ MA and sulphidic conditions for seven days (Supplementary Table S4, experiment 68). Differences in colony size of aposymbiotic and symbiotic morphs after seven days were assessed using the Wilcoxon-Mann-Whitney test.
Growth form index (GFI)
To estimate the relationship between cell population size (number of zooids) and colony size (number of branches), we selected micrographs of 96 Z. niveum (51 symbiotic and 45 aposymbiotic) from the above-described experiments and collections of white colonies at Sv. Jernej in 2014. For comparison we took photographs of 30 Z. ignavum colonies sampled at Sv. Jernej in 2014. A colony growth form index (GFI, exponent of the power law relating zooids and branches) was obtained from aposymbiotic and symbiotic Z. niveum and Z. ignavum as the slope of the log-log linear regression of the number of branches and zooids. This approach is similar to a study in which the topology of plant roots was characterized by comparing the ‘altitude’ (longest path length of root system, equivalent to number of branches) and the ‘magnitude’ (the number of root tips, equivalent to the number of zooids)85. The GFI describes the (log-transformed) number of branches that will develop from the addition of one (log-transformed) new zooid. The significance of the difference between the GFIs was assessed with a covariance analysis86 using R and by calculating of confidence intervals approximated by 10,000 random bootstrap re-samplings per analysis using ‘MUBOQB’.
Scanning electron microscopy (SEM)
Colonies sampled at Sv. Jernej in 2014 (Supplementary Table S4) and swarmers collected in 2013 and 2014 (Supplementary Table S1) were fixed in Trump’s fixative (2.5% glutaraldehyde, 2% paraformaldehyde in 0.1 mol L−1 sodium cacodylate buffer 1100 mOsm L−1, pH 7.2) for up to 12 h, rinsed in 0.1 mol L−1 sodium cacodylate buffer, dehydrated in an ethanol series, transferred to 100% acetone, chemically dried with hexamethyldisilazane (EMS), coated with gold using a Sputter Coater 108 (Agar, United Kingdom), and observed on a IT 300 scanning electron microscope (JEOL, Tokyo, Japan). We distinguished symbiotic hosts with full coverage and partial coverage, and aposymbiotic hosts with less than 10 symbionts in 1,000 µm2.
Fluorescence in situ hybridisation (FISH)
Colonies sampled at Sv. Jernej in 2014 (Supplementary Table S4) were fixed in 100% ethanol, embedded in the medium grade LR White resin, polymerized under nitrogen atmosphere at 40 °C for three days, and several semi-thin (0.5 µm) sections were placed in four drops of MilliQ water on gelatin-coated slides and air dried. Hybridisation was carried out according to Volland et al.66 using the symbiont-specific oligonucleotide probe ZNS196_mod labelled with Cy3 and the EUBmix probes EUB338-I, EUB338-II, EUB338-III87 together with the Arch915 probe88 labelled with Cy5 to target most bacteria and archaea on sections placed into two drops. The sections of the other two drops of each slide were used for the nonsense probes labelled in Cy3 and Cy5 to control for false positives and no signals were observed.
Host 18S rRNA and symbiont 16S rRNA genes sequencing and phylogenetic analysis
Colonies resembling the pale, aposymbiotic growth form of Z. niveum (#4697/2-7) were collected from minimally degraded, most likely oxic parts of wood in July 2018. For comparison, also white, symbiotic Z. niveum (#4577) was collected nearby but from highly degraded, most likely sulphide emitting parts of wood. Samples were fixed in 100% ethanol, and DNA was extracted with DNeasy blood tissue kit (Quiagen, Germany). DNA content was measured with a Nanodrop 2000 spectrophotometer. The primers 82F89 and MedlinB90 were used for PCR amplification of the 18S rRNA gene. Gel electrophoresis was performed on a 1% Agarose gel in TBE buffer for 50 minutes at 90 Volt. For the 16S rRNA gene the primers 27F91 and 1492R92 were used. Bidirectional Sanger sequencing was performed (Microsynth AG, Switzerland). Sequences were analyzed with Geneious v. 11.1. (Biomatters, New Zealand).
To test the close affiliation between Z. niveum from Fort Pierce, USA and Z. niveum from Piran, Slovenia the nucleotide sequences of the small subunit rRNA genes of all members of clade 246,51 were obtained from NCBI, aligned with MAFFT93 v7.407 and trimmed to the shortest sequence length obtained from the collected pale phenotype (1,466 bp) using SeaView94 version 4 and ape95 v5.2 package of R: Z. niveum DQ868350, Fort Pierce, FL, USA49; Z. alternans DQ868352, Fort Pierce, FL USA49; Z. alternans DQ662855, Shazikou, China47; Z. alternans DQ662850, Haibohe, China47; Z. pelagicum DQ868351 Topsail Sound, NC, USA49; Z. plumula DQ662854, Quindao, China (published under the name Z. pluma)47; Z. ignavum. KX669262, Piran, Slovenia51. Z. plumula KY675162, Yantai, China46 was used as outgroup. The 18S rRNA eukaryote gene sequences of both collected colonies were deposited in the GenBank database under accession number MN535886 (Aposymbiotic_Zoothamnium_niveum_str._Piran_4697/5) and MN535887 (Symbiotic_Zoothamnium_niveum_str._Piran_4577).
Maximum Likelihood and Maximum Parsimony phylogenies with bootstrap support were calculated with ape and phangorn96 v2.4.0 packages in R, under a GTR + I nucleotide substitution model (with best corrected AIC value). Bayesesian inference phylogeny with posterior probability support was generated with MrBayes97 v3.2.7a with 1,750,000 generations and a burn-in of 25% of the length. The obtained tree was combined with colony drawings of a symbiotic colony43 and a aposymbiotically grown colony of Z. niveum (this publication), Z. alternans53, Z. pelagicum54, Z. plumula57, and Z. ignavum51 and data on life style43,51,53,54,55,57,59. Reproduction of Z. plumula57 drawing was granted by Magnolia Press, of Z. alternans53 by the Instytut Biologii Doświadczalnej im. M. Nenckiego, of Z. niveum43 by Elsevier, and Z. pelagicum54 by CNRS Éditions.
Data availability
The datasets generated during and/or analysed during the current study are included in this published article and its Supplementary Information Files and are available in the FigShare repository, https://figshare.com/s/98e63972a493c272930e.
References
Gilbert, S. F. & Epel, D. Ecological developmental biology: integrating epigenetics, medicine, and evolution. (Sinauer Associates Inc., 2009).
Bronstein, J. L. Mutualism. (Oxford University Press, 2015).
Douglas, A. E. The symbiotic habit. (Princeton University Press, 2010).
West-Eberhard, M. J. Developmental plasticity and evolution. (Oxford University Press, 2003).
Fraune, S. & Bosch, T. C. G. Why bacteria matter in animal development and evolution. BioEssays 32, 571–580, https://doi.org/10.1002/bies.200900192 (2010).
Gilbert, S. F. Ecological developmental biology: environmental signals for normal animal development. Evolution & Development 14, 20–28, https://doi.org/10.1111/j.1525-142X.2011.00519.x (2012).
Gilbert, S. F., Sapp, J. & Tauber, A. I. A symbiotic view of life: we have never been individuals. The Quarterly Review of Biology 87, 325–341, https://doi.org/10.1086/668166 (2012).
McFall-Ngai, M. Unseen forces: the influence of bacteria on animal development. Developmental Biology 242, 1–14, https://doi.org/10.1006/dbio.2001.0522 (2002).
McFall-Ngai, M. et al. Animals in a bacterial world, a new imperative for the life sciences. Proceedings of the National Academy of Sciences 110, 3229–3236, https://doi.org/10.1073/pnas.1218525110 (2013).
Bennett, G. M. & Moran, N. A. Heritable symbiosis: The advantages and perils of an evolutionary rabbit hole. Proceedings of the National Academy of Sciences 112, 10169–10176, https://doi.org/10.1073/pnas.1421388112 (2015).
Gilbert, S. F. et al. Symbiosis as a source of selectable epigenetic variation: taking the heat for the big guy. Philosophical Transactions of the Royal Society B 365, 671–678 (2010).
Gilbert, S. F., Bosch, T. C. G. & Ledón-Rettig, C. Eco-Evo-Devo: developmental symbiosis and developmental plasticity as evolutionary agents. Nature Reviews Genetics 16, 611, https://doi.org/10.1038/nrg3982 (2015).
Pradeu, T. A mixed self: the role of symbiosis in development. Biological Theory 6, 80–88, https://doi.org/10.1007/s13752-011-0011-5 (2011).
Szathmáry, E. & Smith, J. M. The major evolutionary transitions. Nature 374, 227–232, https://doi.org/10.1038/374227a0 (1995).
Kiers, E. T. & West, S. A. Evolving new organisms via symbiosis. Science 348, 392–394, https://doi.org/10.1126/science.aaa9605 (2015).
West, S. A., Fisher, R. M., Gardner, A. & Kiers, E. T. Major evolutionary transitions in individuality. Proceedings of the National Academy of Sciences 112, 10112–10119, https://doi.org/10.1073/pnas.1421402112 (2015).
Bright, M. & Lallier, F. H. The biology of vestimentiferan tubeworms. Oceanography and Marine Biology: An Annual Review 48, 213–266, https://doi.org/10.1201/EBK1439821169-c4 (2010).
Raikov, I. Bactéries épizoiques et mode de nutrition du cilié psammophile Kentrophoros fistulosum Fauré-Fremiet (étude au microscope électronique). Protistologica 7, 365–378 (1971).
Seah, B. K. B. et al. Specificity in diversity: single origin of a widespread ciliate-bacteria symbiosis. Proceedings of the Royal Society B: Biological Sciences 284, 764, https://doi.org/10.1098/rspb.2017.0764 (2017).
Heckmann, K., Hagen, R. T. & Görtz, H.-D. Freshwater Euplotes species with a 9 type 1 cirrus pattern depend upon endosymbionts. The Journal of Protozoology 30, 284–289, https://doi.org/10.1111/j.1550-7408.1983.tb02917.x (1983).
Heckmann, K. & Schmidt, H. J. Polynucleobacter necessarius gen. nov., sp. nov., an obligately endosymbiotic bacterium living in the cytoplasm of Euplotes aediculatus. International Journal of Systematic and Evolutionary Microbiology 37, 456–457, https://doi.org/10.1099/00207713-37-4-456 (1987).
Vannini, C., Petroni, G., Verni, F. & Rosati, G. Polynucleobacter bacteria in the brackish-water species Euplotes harpa (Ciliata Hypotrichia). Journal of Eukaryotic Microbiology 52, 116–122, https://doi.org/10.1111/j.1550-7408.2005.04-3319.x (2005).
Vannini, C. et al. Endosymbiosis in statu nascendi: close phylogenetic relationship between obligately endosymbiotic and obligately free-living Polynucleobacter strains (Betaproteobacteria). Environmental Microbiology 9, 347–359, https://doi.org/10.1111/j.1462-2920.2006.01144.x (2007).
Vannini, C., Ferrantini, F., Ristori, A., Verni, F. & Petroni, G. Betaproteobacterial symbionts of the ciliate Euplotes: origin and tangled evolutionary path of an obligate microbial association. Environmental Microbiology 14, 2553–2563, https://doi.org/10.1111/j.1462-2920.2012.02760.x (2012).
Hahn, M. W., Schmidt, J., Pitt, A., Taipale, S. J. & Lang, E. Reclassification of four Polynucleobacter necessarius strains as representatives of Polynucleobacter asymbioticus comb. nov., Polynucleobacter duraquae sp. nov., Polynucleobacter yangtzensis sp. nov. and Polynucleobacter sinensis sp. nov., and emended description of Polynucleobacter necessarius. International Journal of Systematic and Evolutionary Microbiology 66, 2883–2892, https://doi.org/10.1099/ijsem.0.001073 (2016).
Boscaro, V. et al. Parallel genome reduction in symbionts descended from closely related free-living bacteria. Nature Ecology &. Evolution 1, 1160–1167, https://doi.org/10.1038/s41559-017-0237-0 (2017).
Mayr, E. Animal species and evolution. (Harvard University Press, 1963).
Pfennig, D. W. et al. Phenotypic plasticity’s impacts on diversification and speciation. Trends in Ecology & Evolution 25, 459–467, https://doi.org/10.1016/j.tree.2010.05.006 (2010).
Whitman, D. W. & Agrawal, A. A. In Phenotypic plasticity of insects (eds D. W. Whitman & T. N. Ananthakrishnan) Ch. 1, 1–63 (Science Publishers: Enfield, NH, 2009).
Nijhout, H. F. Development and evolution of adaptive polyphenisms. Evolution & Development 5, 9–18, https://doi.org/10.1046/j.1525-142X.2003.03003.x (2003).
Sachs, J. L., Gano, K. A., Hollowell, A. C. & Regus, J. U. The legume-rhizobium symbiosis, http://www.oxfordbibliographies.com/view/document/obo-9780199830060/obo-9780199830060-0095.xml (2013).
Weis, V. M. & Levine, R. P. Differential protein profiles reflect the different lifestyles of symbiotic and aposymbiotic Anthopleura elegantissima, a sea anemone from temperate waters. The Journal of Experimental Biology 199, 883 (1996).
Schrallhammer, M., Castelli, M. & Petroni, G. Phylogenetic relationships among endosymbiotic R-body producer: Bacteria providing their host the killer trait. Systematic and Applied Microbiology 41, 213–220, https://doi.org/10.1016/j.syapm.2018.01.005 (2018).
Grosser, K. et al. More than the “killer trait”: infection with the bacterial endosymbiont Caedibacter taeniospiralis causes transcriptomic modulation in Paramecium host. Genome Biology and Evolution 10, 646–656, https://doi.org/10.1093/gbe/evy024 (2018).
Decelle, J., Colin, S. & Foster, R. A. In Marine Protists: Diversity and Dynamics (eds Susumu Ohtsuka et al.) Ch. 19, 465–500 (Springer Japan, 2015).
Tonooka, Y. & Watanabe, T. A natural strain of Paramecium bursaria lacking symbiotic algae. European Journal of Protistology 38, 55–58, https://doi.org/10.1078/0932-4739-00846 (2002).
McFall-Ngai, M. J. The importance of microbes in animal development: lessons from the squid-Vibrio symbiosis. Annual Review of Microbiology 68, 177–194, https://doi.org/10.1146/annurev-micro-091313-103654 (2014).
Fenchel, T. O. M. & Finlay, B. J. Endosymbiotic methanogenic bacteria in anaerobic ciliates: significance for the growth efficiency of the host. The Journal of Protozoology 38, 18–22, https://doi.org/10.1111/j.1550-7408.1991.tb04788.x (1991).
Fisher, R. M., Henry, L. M., Cornwallis, C. K., Kiers, E. T. & West, S. A. The evolution of host-symbiont dependence. Nature Communications 8, 15973, https://doi.org/10.1038/ncomms15973 (2017).
Keeling, P. J. & McCutcheon, J. P. Endosymbiosis: The feeling is not mutual. Journal of Theoretical Biology 434, 75–79, https://doi.org/10.1016/j.jtbi.2017.06.008 (2017).
Lowe, C. D., Minter, E. J., Cameron, D. D. & Brockhurst, M. A. Shining a light on exploitative host control in a photosynthetic endosymbiosis. Current Biology 26, 207–211, https://doi.org/10.1016/j.cub.2015.11.052 (2016).
Dusi, E. et al. Vertically transmitted symbiont reduces host fitness along temperature gradient. Journal of Evolutionary Biology 27, 796–800, https://doi.org/10.1111/jeb.12336 (2014).
Bauer-Nebelsick, M., Bardele, C. F. & Ott, J. A. Redescription of Zoothamnium niveum (Hemprich & Ehrenberg, 1831) Ehrenberg, 1838 (Oligohymenophora, Peritrichida), a ciliate with ectosymbiotic, chemoautotrophic bacteria. European Journal of Protistology 32, 18–30 (1996).
Bauer-Nebelsick, M., Bardele, C. F. & Ott, J. A. Electron microscopic studies on Zoothamnium niveum (Hemprich & Ehrenberg, 1831) Ehrenberg 1838 (Oligohymenophora, Peritrichida), a ciliate with ectosymbiotic, chemoautotrophic bacteria. European Journal of Protistology 32, 202–215 (1996).
Li, L., Ma, H. & Al-Rasheid, K. A. S. Monophyly or polyphyly? Possible conflict between morphological and molecular interpretations of the well-known genus Zoothamnium (Ciliophora, Peritrichia). Chinese Journal of Oceanology and Limnology 33, 490–499, https://doi.org/10.1007/s00343-015-4083-0 (2015).
Zhuang, Y., Clamp, J. C., Yi, Z. & Ji, D. Phylogeny of the families Zoothamniidae and Epistylididae (Protozoa: Ciliophora: Peritrichia) based on analyses of three rRNA-coding regions. Molecular Phylogenetics and Evolution 118, 99–107, https://doi.org/10.1016/j.ympev.2017.09.023 (2018).
Li, L. et al. Reconsideration of the phylogenetic positions of five peritrich genera, Vorticella, Pseudovorticella, Zoothamnopsis, Zoothamnium, and Epicarchesium (Ciliophora, Peritrichia, Sessilida), based on small subunit rRNA gene sequences. Journal of Eukaryotic Microbiology 55, 448–456 (2008).
Sun, P., Clamp, J., Xu, D., Huang, B. & Shin, M. K. An integrative approach to phylogeny reveals patterns of environmental distribution and novel evolutionary relationships in a major group of ciliates. Scientific Reports 6, 21695, https://doi.org/10.1038/srep21695 (2016).
Clamp, J. C. & Williams, D. A molecular phylogenetic investigation of Zoothamnium (Ciliophora, Peritrichia, Sessilida). Journal of Eukaryotic Microbiology 53, 494–498, https://doi.org/10.1111/j.1550-7408.2006.00132.x (2006).
Ji, D. et al. Two new species of Zoothamnium (Ciliophora, Peritrichia) from Korea, with new observations of Z. parahentscheli Sun et al. 2009. Journal of Eukaryotic Microbiology 62, 505–518, https://doi.org/10.1111/jeu.12205 (2015).
Schuster, L. & Bright, M. A novel colonial ciliate Zoothamnium ignavum sp. nov. (Ciliophora, Oligohymenophorea) and its ectosymbiont Candidatus Navis piranensis gen. nov., sp. nov. from shallow-water wood falls. PLoS One 11, e0162834, https://doi.org/10.1371/journal.pone.0162834 (2016).
Fauré-Fremiet, E. Images électroniques d’une mirobiocénose marine. Cahiers de Biologie Marine 4, 61–64 (1963).
Ji, D., Song, W. & Warren, A. Redescriptions of three marine peritrichous ciliates, Zoothamnium alternans Claparède et Lachmann, 1859, Z. sinense Song, 1991 and Z. commune Kahl, 1933 (Ciliophora, Peritrichia), from North China. Acta Protozoologica 45, 27–39 (2006).
Laval, M. Zoothamnium pelagicum du Plessis. Cilié péritriche planctonique: morphologie, croissance et comportament. Protistologica 4, 333–363 (1968).
Gómez, F. Motile behaviour of the free-living planktonic ciliate Zoothamnium pelagicum (Ciliophora, Peritrichia). European Journal of Protistology 59, 65–74, https://doi.org/10.1016/j.ejop.2017.03.004 (2017).
Dragesco, J. Sur la biologie du Zoothamnium pelagicum (du Plessis). Bulletin de la Société Zoologique de France 73, 130–134 (1948).
Ji, D. et al. Redescriptions of five species of marine peritrichs, Zoothamnium plumula, Zoothamnium nii, Zoothamnium wang, Pseudovorticella bidulphiae, and Pseudovorticella marina (Protista, Ciliophora). Zootaxa 2930, 47–59, https://doi.org/10.5281/zenodo.278023 (2011).
Song, W.-B., Al-Rasheid, K. A. S. & Hu, X.-Z. Notes on the poorly-known marine peritrichous ciliate, Zoothamnium plumula Kahl, 1933 (Protozoa: Ciliophora), an ectocommensal organism from cultured scallops in Qingdao, China. Acta Protozoologica 41, 163–168 (2002).
Summers, F. M. Some aspects of normal development in the colonial ciliate Zoöthamnium alternans. The Biological Bulletin 74, 117–129, https://doi.org/10.2307/1537891 (1938).
Bright, M., Espada-Hinojosa, S., Lagkouvardos, I. & Volland, J.-M. The giant ciliate Zoothamnium niveum and its thiotrophic epibiont Candidatus Thiobios zoothamnicoli: a model system to study interspecies cooperation. Frontiers in Microbiology 5, 145, https://doi.org/10.3389/fmicb.2014.00145 (2014).
Millero, F. J. Chemical Oceanography. (CRC Press, 1996).
Rinke, C. et al. High genetic similarity between two geographically distinct strains of the sulfur-oxidizing symbiont ‘Candidatus Thiobios zoothamnicoli’. FEMS Microbiology Ecology 67, 229–241, https://doi.org/10.1111/j.1574-6941.2008.00628.x (2009).
Kloiber, U., Pflugfelder, B., Rinke, C. & Bright, M. Cell proliferation and growth in Zoothamnium niveum (Oligohymenophora, Peritrichida) – thiotrophic bacteria symbiosis. Symbiosis 47, 43–50, https://doi.org/10.1007/BF03179969 (2009).
Ott, J. A., Bright, M. & Schiemer, F. The ecology of a novel symbiosis between a marine peritrich ciliate and chemoautotrophic bacteria. P.S.Z.N.: Marine Ecology 19, 229–243, https://doi.org/10.1111/j.1439-0485.1998.tb00464.x (1998).
Vopel, K., Pöhn, M., Sorgo, A. & Ott, J. Ciliate-generated advective seawater transport supplies chemoautotrophic ectosymbionts. Marine Ecology Progress Series 210, 93–99, https://doi.org/10.3354/meps210093 (2001).
Volland, J.-M. et al. NanoSIMS and tissue autoradiography reveal symbiont carbon fixation and organic carbon transfer to giant ciliate host. The ISME Journal, https://doi.org/10.1038/s41396-018-0069-1 (2018).
Nedwell, D. B. & Floodgate, G. D. The effect of microbial activity upon the sedimentary sulphur cycle. Marine Biology 16, 192–200, https://doi.org/10.1007/bf00346941 (1972).
Rinke, C., Lee, R., Katz, S. & Bright, M. The effects of sulphide on growth and behaviour of the thiotrophic Zoothamnium niveum symbiosis. Proceedings - Royal Society of London. Biological sciences 274, 2259–2269, https://doi.org/10.1098/rspb.2007.0631 (2007).
Ott, J. & Bright, M. Sessile ciliates with bacterial ectosymbionts from Twin Cays, Belize. Atoll Research Bulletin 516, 1–7 (2004).
Genkai-Kato, M. & Yamamura, N. Evolution of mutualistic symbiosis without vertical transmission. Theoretical Population Biology 55, 309–323, https://doi.org/10.1006/tpbi.1998.1407 (1999).
Kawato, M., Uematsu, K., Kaya, T., Pradillon, F. & Fujiwara, Y. First report of the chemosymbiotic ciliate Zoothamnium niveum from a whale fall in Japanese waters. Cahiers de Biologie Marine 51, 413–421 (2010).
Agrawal, A. A. Phenotypic plasticity in the interactions and evolution of species. Science 294, 321–326, https://doi.org/10.1126/science.1060701 (2001).
Görtz, H.-D. Towards an understanding of the distribution, dynamics and ecological significance of bacterial symbioses in protists. Denisia 23, 307–311 (2008).
Dziallas, C., Allgaier, M., Monaghan, M. & Grossart, H.-P. Act together—implications of symbioses in aquatic ciliates. Frontiers in Microbiology 3, https://doi.org/10.3389/fmicb.2012.00288 (2012).
Petroni, G., Spring, S., Schleifer, K.-H., Verni, F. & Rosati, G. Defensive extrusive ectosymbionts of Euplotidium (Ciliophora) that contain microtubule-like structures are bacteria related to Verrucomicrobia. Proceedings of the National Academy of Sciences 97, 1813, https://doi.org/10.1073/pnas.030438197 (2000).
Rosati, G., Petroni, G., Quochi, S., Modeo, L. & Verni, F. Epixenosomes: peculiar epibionts of the hypotrich ciliate Euplotidium itoi defend their host against predators. Journal of Eukaryotic Microbiology 46, 278–282, https://doi.org/10.1111/j.1550-7408.1999.tb05125.x (1999).
Giambelluca, M. A. & Rosati, G. Behavior of epixenosomes and the epixenosomal band during divisional morphogenesis in Euplotidium itoi (Ciliata, Hypotrichida). European Journal of Protistology 32, 77–80, https://doi.org/10.1016/S0932-4739(96)80041-1 (1996).
Powell, M. A. & Somero, G. N. Adaptations to sulfide by hydrothermal vent animals: sites and mechanisms of detoxification and metabolism. The Biological Bulletin 171, 274–290, https://doi.org/10.2307/1541923 (1986).
Grieshaber, M. K. & Völkel, S. Animal adaptations for tolerance and exploitation of poisonous sulfide. Annual Review of Physiology 60, 33–53, https://doi.org/10.1146/annurev.physiol.60.1.33 (1998).
Boscaro, V., Syberg-Olsen, M. J., Irwin, N. A. T., del Campo, J. & Keeling, P. J. What can environmental sequences tell us about the distribution of low-rank taxa? The case of Euplotes (Ciliophora, Spirotrichea), including a description of Euplotes enigma sp. nov. Journal of Eukaryotic Microbiology 66, 281–293, https://doi.org/10.1111/jeu.12669 (2019).
Goodwin, L. R., Francom, D., Urso, A. & Dieken, F. P. Determination of trace sulfides in turbid waters by gas dialysis/ion chromatography. Analytical Chemistry 60, 216–219, https://doi.org/10.1021/ac00154a006 (1988).
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. Journal of Computational and Graphical Statistics 5, 299–314, https://doi.org/10.2307/1390807 (1996).
Cline, J. D. Spectrophotometric determination of hydrogen sulfide in natural waters. Limnology and Oceanography 14, 454–458 (1969).
Vojvoda, J. et al. Seasonal variation in marine-snow-associated and ambient-water prokaryotic communities in the northern Adriatic Sea. Aquatic Microbial Ecology 73, 211–224, https://doi.org/10.3354/ame01718 (2014).
Fitter, A. H. An architectural approach to the comparative ecology of plant root systems. New Phytologist 106(Suppl.), 61–77, https://doi.org/10.1111/j.1469-8137.1987.tb04683.x (1987).
Cochran, W. G. Analysis of Covariance: its nature and uses. Biometrics 13, 261–281, https://doi.org/10.2307/2527916 (1957).
Daims, H., Brühl, A., Amann, R., Schleifer, K.-H. & Wagner, M. The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Systematic and Applied Microbiology 22, 434–444, https://doi.org/10.1016/S0723-2020(99)80053-8 (1999).
Stahl, D. A. & Amann, R. In Nucleic acid techniques in bacterial systematics (eds E. Stackebrandt & M. Goodfellow) 205–248 (Wiley & Sons Ltd., 1991).
Elwood, H. J., Olsen, G. J. & Sogin, M. L. The small-subunit ribosomal RNA gene sequences from the hypotrichous ciliates Oxytricha nova and Stylonychia pustulata. Molecular Biology and Evolution 2, 399–410, https://doi.org/10.1093/oxfordjournals.molbev.a040362 (1985).
Medlin, L., Elwood, H. J., Stickel, S. & Sogin, M. L. The characterization of enzymatically amplified eukaryotic 16S-like rRNA-coding regions. Gene 71, 491–499, https://doi.org/10.1016/0378-1119(88)90066-2 (1988).
Lane, D. J. In Nucleic acid techniques in bacterial systematics (eds E. Stackebrandt & M. Goodfellow) 115–175 (John Wiley & Sons Ltd, 1991).
Loy, A. et al. 16S rRNA gene-based oligonucleotide microarray for environmental monitoring of the betaproteobacterial order “Rhodocyclales”. Applied and Environmental Microbiology 71, 1373–1386, https://doi.org/10.1128/AEM.71.3.1373-1386.2005 (2005).
Katoh, K., Misawa, K., Kuma, K. I. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research 30, 3059–3066, https://doi.org/10.1093/nar/gkf436 (2002).
Gouy, M., Guindon, S. & Gascuel, O. SeaView Version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Molecular Biology and Evolution 27, 221–224, https://doi.org/10.1093/molbev/msp259 (2009).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20, 289–290, https://doi.org/10.1093/bioinformatics/btg412 (2004).
Schliep, K. P. Phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593, https://doi.org/10.1093/bioinformatics/btq706 (2011).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574, https://doi.org/10.1093/bioinformatics/btg180 (2003).
Acknowledgements
We thank the Station de Recherches Sous Marine et Oceanographiques, Corse, and the Marine Biological Station Piran, Slovenia, for their hospitality during our fieldwork, Nikolaus Császár, Niels Heindl, Jasmin Löffler, Margit Schraick for the design and accomplishment of the preference experiment during a student course, Michael Stachowitsch for editorial work, and the Core Facility Cell Imaging and Ultrastructure Research of the University of Vienna for its continuous support. This work was funded by the Austrian Science Fund under grant # P24565-B22 and # P 32197 to M.B.
Author information
Authors and Affiliations
Contributions
M.B. designed the research, A.N. designed and J.O. participated in the preference experiment, J.-M.V. led the growth experiments together with M.B., S.E.H., I.K. and L.S., J.D. and J.K. performed SEM, H.C.Z. performed FISH, D.M. and F.S. performed sequencing, S.E.H. and H.L.N. analysed the data and performed the statistical analyses, M.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bright, M., Espada-Hinojosa, S., Volland, JM. et al. Thiotrophic bacterial symbiont induces polyphenism in giant ciliate host Zoothamnium niveum. Sci Rep 9, 15081 (2019). https://doi.org/10.1038/s41598-019-51511-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-51511-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.