Abstract
The fungal pathogen Leptosphaeria maculans causes blackleg disease on canola and rapeseed (Brassica napus) in many parts of the world. A B. napus cultivar, ‘Quinta’, has been widely used for the classification of L. maculans into pathogenicity groups. In this study, we confirmed the presence of Rlm1 in a DH line (DH24288) derived from B. napus cultivar ‘Quinta’. Rlm1 was located on chromosome A07, between 13.07 to 22.11 Mb, using a BC1 population made from crosses of F1 plants of DH16516 (a susceptible line) x DH24288 with bulked segregant RNA Sequencing (BSR-Seq). Rlm1 was further fine mapped in a 100 kb region from 19.92 to 20.03 Mb in the BC1 population consisting of 1247 plants and a F2 population consisting of 3000 plants using SNP markers identified from BSR-Seq through Kompetitive Allele-Specific PCR (KASP). A potential resistance gene, BnA07G27460D, was identified in this Rlm1 region. BnA07G27460D encodes a serine/threonine dual specificity protein kinase, catalytic domain and is homologous to STN7 in predicted genes of B. rapa and B. oleracea, and A. thaliana. Robust SNP markers associated with Rlm1 were developed, which can assist in introgression of Rlm1 and confirm the presence of Rlm1 gene in canola breeding programs.
Similar content being viewed by others
Introduction
Blackleg (stem canker), caused by the ascomycete fungus Leptosphaeria maculans (Desmaz.) Ces. et De Not. [anamorph: Phoma lingam (Tode ex Fr.) Desmaz.], is one of the most important diseases of canola and rapeseed in many parts of the world1,2. It has been suggested that L. maculans can cause severe infection in a field and reduce the seed yield by more than 50%3. Improving resistance to L. maculans is one of the major objectives in canola breeding programs worldwide, especially due to the fact that the disease is difficult to control with fungicides4.
Efforts have been made to identify and map blackleg R genes in B. napus. The R genes LEM15, LmFr16, LmR1/CLmR17,8, Rlm1, Rlm3, Rlm4, Rlm7 and Rlm96,7,8,9 have been mapped to the B. napus linkage group A07, and Rlm2 to A109. Four additional R genes, LepR1, LepR2, LepR3 and LepR4, originating from an accession of wild B. rapa subsp. sylvestris, have been identified in B. napus or progenies from B. napus x B. rapa subsp. sylvestris. LepR1 and LepR2 were mapped to B. napus linkage groups A02 and A10, respectively10. LepR3, derived from B. rapa subsp. sylvestris and present in the B. napus cultivar Surpass 400, was mapped to the B. napus linkage group A1011. LepR3 and Rlm2 have been cloned with a map‐based cloning strategy, and encode a receptor-like protein (RLP)12,13. Blackleg R genes previously mapped in B. napus are generally the dominant type, except LepR4 (Brassica A-genome), which was mapped to the B. napus linkage group A0614. The R genes BLMR1 and BLMR2 were mapped to the B. napus linkage group A10 in the B. napus cultivar Surpass 40015. All these blackleg R genes were identified through genetic mapping with a marker system, such as RFLPs, Simple Sequence Repeats (SSRs), Amplified Fragment Length Polymorphisms (AFLPs), Cleaved Amplified Polymorphic Sequences (CAPSs), or Sequence Related Amplified Polymorphisms (SRAPs). Developing and genotyping these types of molecular marker can be laborious and time consuming.
Bulk segregant analysis (BSA), which consists of genotyping two bulks of individual plants with extreme phenotypes, was developed to rapidly identify genetic markers linked to genomic regions associated with selected phenotypes16. Presently, the pipelines of BSA and high throughput sequencing have been developed to locate and identify candidate genes associated with selected phenotypes, using BSA RNA sequencing (BSR-Seq)17,18,19.
Canola is the most important oilseed crop in Canada. ‘Quinta’, a winter oilseed rape originating from Europe, was found to be highly resistant to lots of Canadian isolates and was widely used as a differential cultivar for the classification of pathogenicity groups20,21,22,23,24. The first genetic evidence for a gene-for-gene interaction between Brassica and L. maculans was observed between the resistance (R) gene Rlm1 in ‘Quinta’ and the corresponding avirulence gene AvrLm1.25. Delourme et al. (2004) roughly mapped Rlm1 in ‘Quinta’ to linkage group A07. The R gene was further mapped using SSR and CAPS markers in a large genetic interval26.
In western Canada, AvrLm1 was detected in 46% of the L. maculans isolates collected from infested canola stubble between 1997 and 200527. Although recent data showed lower presence of AvrLm1 in L. maculans populations28,29 due to use of Rlm1 in canola cultivars extensively30, Rlm1 could still be valuable for effective control of blackleg disease in Canada as a strategy through rotation of R genes that has been recently implemented.
The objectives of the current study were to: (i) identify genome wide DNA variants to map Rlm1 through BSR-Seq; (ii) determine the precise location of Rlm1; (iii) identify the most probable candidates for Rlm1; and (iv) develop SNP markers tightly linked to Rlm1 in use of marker assisted selection. In this study, Rlm1 was located on chromosome A07 using a modified popoolation2 pipeline withBSR-Seq. To the best of our knowledge, this is the first report on mapping of a blackleg resistance gene using BSR-Seq in Brassica spp..
Materials and Methods
Plant materials
Two doubled haploid lines, DH24288 (resistant, R) from the B. napus cultivar ‘Quinta’ and DH16516 (susceptible, S) from B. napus cultivar ‘Topas’, were kindly provided by Dr. G Séguin-Swartz at Saskatoon Research and Development Centre, Agriculture and Agri-Food Canada. A DH24288 plant (male) was crossed with a DH16516 plant (female) to produce a F1 population. A F1 plant was backcrossed with a DH16516 plant to obtain a BC1 population. The F1 plant was self-pollinated to produce a F2 population.
Plant growth conditions, preparation of L. maculans isolates and plant inoculations
L. maculans inoculum and plant inoculation followed the protocols described previously10. The parents, F1, F2, and BC1 were inoculated with isolate L. maculans SC006 (carrying AvrLm1). The reaction to inoculation on cotyledon was rated 10 to 14 days post-inoculation (dpi) using the 0 to 9 scale described by Williams31. Disease ratings of 0 to 5 and 6 to 9 were considered as resistant and susceptible interactions, respectively. Segregation for resistance and susceptibility in the BC1 population was analyzed using the Chi-square (χ2) test for goodness-of-fit32. B. napus susceptible cultivar ‘Westar’ was used as a susceptible check in this study.
RNA isolation and RNA-seq
The BC1 population was used for RNA-Seq. At 10 dpi, leaf samples from 30 R plants and 30 S plants were combined to form R and S bulks, respectively; each bulk was treated as one biological replicate. Three replicates, consisting of a total of 90 R or 90 S plants, were used. RNA from each sample replicate was isolated using an RNeasy Plant Mini Kit (Qiagen; Toronto, ON) with on-column deoxyribonuclease (DNase) digestion using a Qiagen RNase-Free DNase kit, following manufacturer’s instructions. The RNA concentration and quality were checked using a NanoDrop 2000c Spectrophotometer (Thermo Scientific; Waltham, MA) and an Agilent Bioanalyzer 2100 (Agilent Technologies; Mississauga, ON), to ensure that the RNA integrity number (RIN) was >8 for each sample. The preparation of cDNA library and RNA-seq were performed at Databio2 LLC (Ames, IA).
Sequence alignment, SNP calling and filtration
Raw RNA-Seq reads were trimmed to remove low-quality nucleotides with FASTX-Toolkit (Version 0.0.13)33. GSNAP (Genomic Short-read Nucleotide Alignment Program, version 2016-11-07)34 was performed to map the trimmed reads to the B. napus reference genome35, allowing gap alignment including intron-spanning alignment. SNP calling and filtration for BSR-Seq was performed following the same parameters as previously described by Liu et al.17. The SNP discovery pipeline (123SNP) was downloaded from http://schnablelab.plantgenomics.iastate.edu/software/123SNP/. Reads of the R and S bulks were more than 90% of the total reads aligned to each SNP site. Each SNP site had ≥3 reads with a quality score of SNP base ≥15, and the reads accounted for 20% of the total reads at that SNP site. Moreover, each SNP site for the BSR-Seq analysis was required to have at least five sequencing reads in both the R and S bulks. In addition, two methods for SNP calling were used as previously described18; the single sample alignment (SSA), in which short reads from three biological replicates of R and S plants were aligned to a reference genome. The other was called the pooled sample alignment (PSA), in which short reads from a pool of three R bulks and a pool of three S bulks were aligned to the reference genome as described by Yu et al.18.
Identification of SNP markers tightly linked to Rlm1
In order to verify SNP markers and the target region, an improved PoPoolation236 was employed to analyze the linked SNP markers. (Fig. 1A). Firstly, the same files of R and S bulks were obtained as previously described using GSNAP34. Then, Picard tools (Version 1.96, http://broadinstitute.github.io/picard/) was performed to remove duplicate and ambiguously mapped reads. Finally, the allele frequencies of R and S bulks were compared using PoPoolation236. All analyses used the SSA and PSA methods described previously18 with reference B. napus Darmor-bzh genome (v4.1,http://www.genoscope.cns.fr/brassicanapus/data/)35.
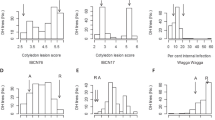
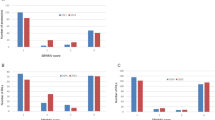
Schematic flowchart and disease ratings. (A) Schematic flowchart of the experimental procedure. (B) Average disease ratings of B. napus cultivars ‘Quinta’ and its DH line DH24288. Westar is the susceptible control. The L. maculans isolate SC006 carries AvrLm1. The disease reaction on cotyledons was rated at 10 dpi using the 0 to 9 scale31. Mini bars indicate the standard deviation from three biological replicates of three independent repetitions (n = 9). Letters indicate significant difference based on the Least Significant Difference (LSD) (P < 0.05).
Identification of Rlm1 candidate genes
In order to identify the candidates for Rlm1 in the target region, all genes in this region were analyzed using the following steps:
-
1.
We annotated all proteins to the target region using InterProScan37.
-
2.
The variation of R genes in the target region was identified using samtools38 and SNPeff 4.139. The variant frequency of Rlm1 was approximately 0.5 in the R bulks and 1 in S bulks, since the resistance genotype would be Rr and susceptible genotype be rr in the BC1 population.
-
3.
Resistance gene prediction
The plant resistance genes were identified using RGAugury pipeline40 with reference B. napus Darmor-bzh genome (v4.1,http://www.genoscope.cns.fr/brassicanapus/data/)35,41.
Gene expression analysis using RNA-Seq
All trimmed RNA-seq reads were aligned18 with reference B. napus Darmor-bzh genome (v4.1,http://www.genoscope.cns.fr/brassicanapus/data/)35 using Tophat 242,43. Tophat2 alignment parameters were set to allow a maximum of two mismatches and exclude reads mapping to more than one position on the reference. Transcriptome assembly and differential expression analysis for RNA-Seq were conducted using cufflink, with FDR < 0.05 and minimum fold change ≥2.
Genotyping SNP and InDel markers
Selected SNPs identified in the target region were confirmed using the Kompetitive Allele Specific PCR (KASP) method (http://www.lgcgroup.com), following manufacturer’s instructions. The primer sequences of KASP assay were presented in Table S1. The PCR reaction was performed using StepOne Plus Real Time PCR system (Applied Biosystem, Mississauga, ON).
Phylogenetic tree construction and sequence alignment
The protein sequences of STN7 were searched using Blastp with default parameters in the nr database of NCBI. Alignment was performed using MAFFAT44 with distance correction off, gaps excluded. Then the alignment sequence computed the residue-wise confidence scores and extracted well-aligned residues using GUIDANCE245. The neighbor-joining (NJ) clustering method46 was performed to build a phylogeny tree with bootstrap 1000. Visualization was performed using Interactive tree of life (iTOL) v4 (https://itol.embl.de/)47.
Results
Identification of Rlm1 in the B. napus DH line DH24288
DH24288 was derived from a single microspore of the cultivar ‘Quinta’ (B. napus). Its resistance to the L. maculans isolate SC006 (AvrLm1) was compared with its donor (Quinta) and the susceptible cultivar ‘Westar’ (Fig. 1B) based on cotyledon inoculation. As expected, ‘Westar’ was highly susceptible whileDH24288 and ‘Quinta’ showed a high level of resistance, implying that DH24288 carries Rlm1.
RNA-Seq and sequence alignment
In order to identify the resistance gene in the DH line, we tested the BC1 segregating population and their parental lines with the isolate SC006. DH16516 was highly susceptible (Fig. 2). The BC1 population consisted of 571 R and 596 S plants, which would fit a 1:1 ratio (χ2 = 0.54, P = 0.46). Furthermore, the F2 population showed 2,216 R and 696 S plants, which would fit a 3:1 ratio (X2 = 1.88, P = 2.37), confirming that Rlm1 is a single dominant gene in DH24288. A total of 180 plants with a clear R (DR ≤ 2) and S (DR ≥ 8) designation, consisting of 90 R and 90 S (Fig. 2) were selected from the population for mapping of Rlm1 using BSR-Seq.
Frequency distributions of resistance phenotypes in the BC1 population derived from DH16516 x DH24288/’Quinta’. Parents and BC1 plants were inoculated with L. maculans isolate SC006. The symptoms on cotyledons was rated at 10–14 dpi using the 0 to 9 scale31. Disease ratings of 0 to 4 and 5 to 9 were considered as resistant and susceptible interactions, respectively. The lines in the red boxes were selected as R and S bulks for RNA-seq analysis.
More than 90 million 101 paired-end reads were obtained, including 47 million from R bulks and 45 million from S bulks. After quality control and clipping of adapters, more than 99% of the reads were retained for both bulks, with the average length reduced to about 98 bp (Table S2).
The average short read counts and accumulated short read lengths were 30.6 million (M) and 3003 Mb for R bulks, and 29.6 M and 2897 Mb for S bulks, using the SSA method (Table S3). This provided an average depth of coverage for the transcripts of reference genome at 30 folds in R bulks and 29 folds in S bulks. More short reads were aligned to the longer chromosomes A09, C03, C04, and C09, while fewer short reads were assembled in the shorter chromosomes A08 and A10 (Fig. 3). Short reads from three R bulks and three S bulks were further aligned using PSA. As observed previously with the SSA method, more sequences were aligned to A09, C03, C04, and C09, and fewer to A08 and A10 (Fig. 3). A total of 91.9 M short reads were aligned to the B. napus reference genome, resulting in an alignment of 9008 Mb in length and a coverage of 90 folds of transcripts of the reference genome from the pool of three R bulks, and 88.6 M short reads were aligned, resulting in an alignment 8691 Mb in length and an 87-fold coverage from the pool of three S bulks (Table S3).
Counts of short reads aligned to chromosomes of the B. napus reference B. napus Damor-bzh genome35 in single sample alignment (SSA) and pooled sample alignment (PSA), respectively.
Identification of variants in R and S bulks
About 106K SNPs and 9K InDels were identified in R bulks and 116K SNPs and 9K InDels in S bulks were identified using the SSA method (Table S4). The counts of SNPs and InDels identified using PSA were higher than those using the SSA method, with 288K SNPs and 20K InDels in R bulks and 281K SNPs and 19K InDels in the S bulks (Table S4). There was a strong positive correlation between the numbers of SNPs and InDels (r = 0.97) within R and S bulks using either SSA or PSA. The number of SNPs and InDels varied among chromosomes (Fig. 4), with chromosomes A03 and C03 carrying more variants than others. However, SNPs and InDels seemed to show a similar distribution pattern on different chromosomes among the R and S bulks based on both SSA (Fig. S1) and PSA (Fig. S2).
Comparison and annotation of variants in R and S bulks
Common or monomorphic (mono) variants which were present in both R and S bulks, and unique or polymorphic (poly) variants observed only in R or S bulks were identified. The percentage of mono and poly variants identified using PSA was more than those of SSA on each chromosome, although the ratio of mono and poly variants identified with the two methods were similar (Fig. 5). The mono variants comprised of 51.7% (48.1–54.9%) of the variants identified across the B. napus genome using PSA, while poly variants comprised of 48.3% (45.1–51.9%) (Table S5). There were no differences among chromosomes in terms of the number of mono and poly variants found in R and S bulks except on chromosome A07, where more poly variants were found in R bulks than in S bulks (Fig. 5B)
In order to investigate the effects caused by the variants, all low-quality sequence reads were filtered out using the B. napus reference genome35. Only the variants identified with PSA were annotated using SNPeff because they included all variants identified through SSA. In total, 18,857 InDels and 272,837 SNPs were annotated in R bulks and 18,816 InDels and 266,772 SNPs in S bulks. The percentage of variant effect by gene regions was almost the same between R and S bulks (Fig. 6); more than 60% of variants were identified in regions down- and up-stream of the gene Rlm1, with about 10% of InDels and 20% of SNPs in the gene exon region. Additionally, around 10% of the variants were located in the intergenic region and several variants were in the gene intron region.
Refining the interval of Rlm1 using SNP markers identified through BSR-Seq
Rlm1 was located on chromosome A07 within a region of 13.07 Mb to 22.11 Mb based on significant SNPs (P value < 10−8; Fst value > 0.2) using modified popoolation2 (Figs 7A,B, Fig. S5). A total of 18,035 variants were identified within this region using PSA, including 16,967 SNPs, 563 insertions and 599 deletions.
Rlm1 was located on chromosome A07 using Popoolation2 with window slide (window size = 1000 bp, slide size 100 bp). (A) Significant variants by substantial Fisher’s exact values in whole genome (-log (P value) > 8)). (B) Significant variants by substantial Fisher’s exact values in chromosome A07 (-log (P value) > 8). Green dots indicate the significant variants in the Rlm1 interval. (C) Fine mapping of Rlm1 between 19.93 Mb and 20.03 Mb on chromosome A07 using KASP assays with the BC1 population (a total of 1,022 plants) and F2 population (a total of 2,912 plants). (D) BnaA07g27460D gene structure and variants.
In order to narrow down the interval of Rlm1, a total of 1,022 BC1 plants and 2,912 F2 plants, including the 90 R and 90 S plants used for RNA-Seq, were inoculated with the isolate SC006 and genotyped with two flanking SNP markers designed based on SNP information in the Rlm1 region (Fig. 7C). A total of 13 recombinant lines were identified, and these recombinants were further confirmed to be within the genetic interval with an additional 13 SNP markers (Fig. 7C and Table S1). As a result, Rlm1 was narrowed down to a 100 Kb region (19.93–20.03 Mb) flanked by markers SNP_A07_12_19.93 and SNP_A07_11_20.03 on chromosome A07 (Fig. 7C). A total of 152 variants (Table S7) could be annotated to 19 B. napus genes (Table S6) in the Rlm1 region, including 62 variants identified in both R and S bulks. The variants in the Rlm1 region were examined using the Integrative Genomics Viewer (IGV)48. One gene (BnaA07g27460D) showed similar frequencies of DNA variants at about 50% in R bulks and 100% in S bulks (Fig. S5 and Table S7). A total of 11 polymorphic SNPs in BnaA07g27460D were identified, including 1 downstream, 9 synonymous, and two missense SNPs. BnaA07g27460D (19,986,679–19,989,481 bp) encodes Serine/threonine dual specificity protein kinase with a catalytic domain. The full-length protein sequence was used to search for homologous proteins in the NCBI database, which was aligned using MAFFT and used to construct a neighbor-joining tree with UPGMA clustering (Fig. S6). The results indicated that BnaA07g27460D was homologous to STN7 in predicted genes of B. rapa and B. oleracea, and A. thaliana (Fig. S6).
Analysis of differential expression genes (DEGs) in R and S bulks
The expression of genes in the Rlm1 region may provide additional molecular evidence for the confirmation of the candidate R gene. A total of 153 DEGs were identified in comparing R bulks with S bulks using cufflink with FDR < 0.05 (Table S8), and18 of them were located within the Rlm1 region. However, the gene expression level for BnaA07g27460D was not significantly different between R and S bulks.
Discussion
In this study, a BSR-Seq strategy was used to map Rlm1 to B. napus chromosome A07 in a 9-Mb region. More than 18,000 variants were identified in this region. Selected SNP loci in the interval were analyzed using KASP assay. A total of 1,022 plants in BC1 and and 2,912 plants in F2) were analyzed with 13 KASP markers. The Rlm1 interval was eventually narrowed within an interval of 100 kb through linkage analysis. Previously, Rlm1 was predicted to be in an interval of approximately 920 kb between a SSR marker (sN9539) and a CAPS marker (Ind07-02) though Blast search26. The results indicate that BSR-Seq in combination with KASP assay is much more powerful technology than using marker systems for fine mapping of blackleg resistance genes49,50,51,52,53,54,55, which marker systems, such as RFLPs56, AFLPs57, SSRs58, CAPSs59, and SRAPs60, are laborious, time consuming, and low efficiency for identification of QTLs or for marker-assisted selection (MAS). BSR-seq has been used for fine mapping of several resistance genes to clubroot in Brassica species18,61,62. However, the application and efficacy of the modified popoolation2 comparing with other mapping by sequencing methods still needs to be determined.
There were more DNA variants identified in chromosomes A03 and C03 than the other chromosomes. The number of variants is usually associated with chromosome length18. C03 is the longest chromosome with 60.6 Mb in length in the sequencedreference B. napus Darmor-bzh genome35. In addition, these two chromosomes carry higher numbers of genes although A03 is only 29.8 Mb in length (Table S2).Therefore, it is not surprising that more DNA variants were identified based on BSR-Seq.
In this study, Rlm1 was located within a range of 100 Kb on chromosome A07. A total of eight blackleg R genes (LEM15, LmFr16, LmR1/CLmR17,8, Rlm1, Rlm3, Rlm4, Rlm7 and Rlm96,7,8,9) have been reported previously on chromosome A07 (linkage group N7),. LmR1/CLmR1 was found to co-segregate with three molecular markers, which were located around the 15.8 Mb of chromosome A07, by blasting the marker sequences against the B. napus genome database8. The homologous region of LEM1 and LmFr1 cannot be identified based on the genomic information available. The genetic position of Rlm3, Rlm4, Rlm7, and Rlm9 were clearly different from that of Rlm19. In our study, Rlm1 was fine mapped to an interval between 19.93 Mb and 20.03 Mb with flanking markers SNP9_A07_12_19.93 and SNP8_A07_11_20.03 on chromosome A07, indicating that Rlm1 is unlikely allelic to LmR1/CLmR1.
In the mapping interval, no RLP genes described as blackleg resistance genes35 were identified. DNA variants were only identified in gene BnaA07g27460D (Fig. S5 and Table S7). Hence, it is possible that BnaA07g27460D was a candidate for Rlm1. BnaA07g27460D is homologous to STN7 in predicted genes of B. rapa and B. oleracea, and A. thaliana. Earlier studies on Arabidopsis clearly demonstrated that thylakoid phosphorylation is predominantly mediated by the protein kinases STN763,64 and STN865. However, several recent studies have revealed that light intensity or quality induces the reduction/oxidation (redox) state of the photosynthetic electron chain66. STN7 has been verified to operate in retrograde signaling through controlling redox balance in the electron transfer chain67. These confirmations indicate that STN7 is involved in plant defense reactions by regulating the excitation of energy distribution between PSII and PSI via the phosphorylation of thylakoid membrane to control cell signaling and systemic response via ROS-induced, hormone-mediated signaling networks67,68. In this study, although BnaA07G27460D was considered as the candidate gene for Rlm1, other candidates may exist. Firstly, the coverage of the sequence reads for several genes in the Rlm1 region was insufficient for definitive inference of Rlm1 based only on BSR-Seq. Secondly, the genomic sequence of Darmor-bzh was not complete in the Rlm1 region. Four gaps were found between 19.93 Mb and 20.03 Mb, form 19,922,209 to 19923154 bp, from 19,949,953 to19,960,559 bp, from 19,968,763 to 19,974,252 bp, and from 20,017,902 to 20,020,342. Thirdly, the sequence of Darmor-bzh does not carry Rlm1 gene69. Hence, gene cloning and transformation are necessary to confirm the candidate gene of Rlm1, but that is out of the scope of the current study. More biochemical and molecular confirmations can be used to confirm the involvement of STN7 in blackleg resistance.
A total of 153 DEGs were identified in comparing R bulks with S bulks, and18 of them were located within the Rlm1 region. However, the gene expression level for BnaA07g27460D was not significantly different between R and S bulks although DNA variants were identified in the gene. The reason for this is to be determined. No significant difference in R gene expression was previously observed during fungal infection between R and S materials18.
Breeding for blackleg resistance in canola is one of the most important objectives in many parts of the world. Use of highly specific markers in MAS could be useful to accurate assess the reaction of lines under controlled conditions. KASP offers cost-effective and scalable flexibility in applications. A large number of SNP sites associated with Rlm1 were identified based on BSR-Seq. In total, 13 robust SNP markers were confirmed to be associated with Rlm1 using KASP assay. The closely linked markers shown in Table S7 could be used in MAS for Rlm1.
References
Fitt, B. D. L., Brun, H., Barbetti, M. J. & Rimmer, S. R. World-Wide Importance of Phoma Stem Canker (Leptosphaeria maculans and L. biglobosa) on Oilseed Rape (Brassica napus). European Journal of Plant Pathology 114, 3–15, https://doi.org/10.1007/s10658-005-2233-5 (2006).
Rimmer, S. R. Resistance genes to Leptosphaeria maculans in Brassica napus. Canadian. Journal of Plant Pathology 28, S288–S297, https://doi.org/10.1080/07060660609507386 (2006).
Gugel, R. K. & Petrie, G. A. History, occurrence, impact, and control of blackleg of rapeseed. Canadian Journal of Plant Pathology 14, 36–45, https://doi.org/10.1080/07060669209500904 (1992).
Kutcher, H. R., Yu, F. & Brun, H. Improving blackleg disease management of Brassica napus from knowledge of genetic interactions with Leptosphaeria maculans. Canadian. Journal of Plant Pathology 32, 29–34, https://doi.org/10.1080/07060661003620961 (2010).
Ferreira, M., Rimmer, S., Williams, P. & Osborn, T. Mapping loci controlling Brassica napus resistance to Leptosphaeria maculans under different screening conditions. Phytopathology 85, 213–217 (1995).
Dion, Y., Gugel, R. K., Rakow, G. F., Seguin-Swartz, G. & Landry, B. S. RFLP mapping of resistance to the blackleg disease [causal agent, Leptosphaeria maculans (Desm.) Ces. et de Not.] in canola (Brassica napus L.). Theor Appl Genet 91, 1190–1194, https://doi.org/10.1007/BF00220928 (1995).
Mayerhofer, R., Good, A. G., Bansal, V. K., Thiagarajah, M. R. & Stringam, G. R. Molecular mapping of resistance to Leptosphaeria maculans in Australian cultivars of Brassica napus. Genome 40, 294–301 (1997).
Mayerhofer, R. et al. Complexities of chromosome landing in a highly duplicated genome: toward map-based cloning of a gene controlling blackleg resistance in Brassica napus. Genetics 171, 1977–1988, https://doi.org/10.1534/genetics.105.049098 (2005).
Delourme, R. et al. A Cluster of Major Specific Resistance Genes to Leptosphaeria maculans in Brassica napus. Phytopathology 94, 578–583, https://doi.org/10.1094/PHYTO.2004.94.6.578 (2004).
Yu, F., Lydiate, D. J. & Rimmer, S. R. Identification of two novel genes for blackleg resistance in Brassica napus. Theor Appl Genet 110, 969–979, https://doi.org/10.1007/s00122-004-1919-y (2005).
Yu, F., Lydiate, D. J. & Rimmer, S. R. Identification and mapping of a third blackleg resistance locus in Brassica napus derived from B. rapa subsp. sylvestris. Genome 51, 64–72, https://doi.org/10.1139/g07-103 (2008).
Larkan, N. J. et al. The Brassica napus blackleg resistance gene LepR3 encodes a receptor-like protein triggered by the Leptosphaeria maculans effector AVRLM1. New Phytol 197, 595–605, https://doi.org/10.1111/nph.12043 (2013).
Larkan, N. J., Ma, L. & Borhan, M. H. The Brassica napus receptor-like protein RLM2 is encoded by a second allele of the LepR3/Rlm2 blackleg resistance locus. Plant Biotechnol J 13, 983–992, https://doi.org/10.1111/pbi.12341 (2015).
Yu, F., Gugel, R. K., Kutcher, H. R., Peng, G. & Rimmer, S. R. Identification and mapping of a novel blackleg resistance locus LepR4 in the progenies from Brassica napus x B. rapa subsp. sylvestris. Theor Appl Genet 126, 307–315, https://doi.org/10.1007/s00122-012-1919-2 (2013).
Long, Y. et al. Identification of two blackleg resistance genes and fine mapping of one of these two genes in a Brassica napus canola cultivar ‘Surpass 400’. Theor Appl Genet 122, 1223–1231, https://doi.org/10.1007/s00122-010-1526-z (2011).
Michelmore, R. W., Paran, I. & Kesseli, R. V. Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88, 9828–9832 (1991).
Liu, S., Yeh, C. T., Tang, H. M., Nettleton, D. & Schnable, P. S. Gene mapping via bulked segregant RNA-Seq (BSR-Seq). PloS one 7, e36406, https://doi.org/10.1371/journal.pone.0036406 (2012).
Yu, F. et al. Identification of Genome-Wide Variants and Discovery of Variants Associated with Brassica rapa Clubroot Resistance Gene Rcr1 through Bulked Segregant RNA Sequencing. PloS one 11, e0153218, https://doi.org/10.1371/journal.pone.0153218 (2016).
Huang, Z. et al. Fine Mapping of a Clubroot Resistance Gene in Chinese Cabbage Using SNP Markers Identified from Bulked Segregant RNA Sequencing. Frontiers in Plant Science 8, https://doi.org/10.3389/fpls.2017.01448 (2017).
Koch, E., Song, K., Osborn, T. & Williams, P. Relationship between pathogenicity and phylogeny based on restriction fragment length polymorphism in Leptosphaeria maculans. Molecular Plant-Microbe Interactions 4, 341–349 (1991).
Kutcher, H. R., Keri, M., McLaren, D. L. & Rimmer, S. R. Pathogenic variability of Leptosphaeria maculans in western Canada. Canadian. Journal of Plant Pathology 29, 388–393, https://doi.org/10.1080/07060660709507484 (2007).
Mengistu, A., Rimmer, S., Koch, E. & Williams, P. H. Pathogenicity grouping of isolates of Leptosphaeria maculans on Brassica napus cultivars and their disease reaction profiles on rapid-cycling Brassicas. Plant Dis 75, 1279–1282 (1991).
M, K. Genetic studies of host – pathogen interaction between Brassica napus and Leptosphaeria maculans. Ph. D. Thesis, University of Manitoba (1999).
Mahuku, G. S., Goodwin, P. H., Hall, R. & Hsiang, T. Variability in the highly virulent type of Leptosphaeria maculans within and between oilseed rape fields. Canadian Journal of Botany 75, 1485–1492, https://doi.org/10.1139/b97-863 (1997).
Ansan-Melayah, D. et al. Genes for race-specific resistance against blackleg disease in Brassica napus L. Plant Breeding 117, 373–378, https://doi.org/10.1111/j.1439-0523.1998.tb01956.x (1998).
Larkan, N. J. et al. Gene Introgression Lines for Accurate Dissection of the Brassica - Leptosphaeria Pathosystem. Front Plant Sci 7, 1771, https://doi.org/10.3389/fpls.2016.01771 (2016).
Kutcher, H. R. et al. Frequency of avirulence genes in Leptosphaeria maculans in western Canada. Canadian. Journal of Plant Pathology 32, 77–85, https://doi.org/10.1080/07060661003594109 (2010).
Zhang, X. et al. Breakdown of Rlm3 resistance in the Brassica napus–Leptosphaeria maculans pathosystem in western Canada. European Journal of Plant Pathology 145, 659–674, https://doi.org/10.1007/s10658-015-0819-0 (2016).
Liban, S. H., Cross, D. J., Kutcher, H. R., Peng, G. & Fernando, W. G. D. Race structure and frequency of avirulence genes in the western Canadian Leptosphaeria maculans pathogen population, the causal agent of blackleg in brassica species. Plant Pathology 65, 1161–1169, https://doi.org/10.1111/ppa.12489 (2016).
Hubbard, M. & Peng, G. Quantitative resistance against an isolate of Leptosphaeria maculans (blackleg) in selected Canadian canola cultivars remains effective under increased temperatures. Plant Pathology 67, 1329–1338, https://doi.org/10.1111/ppa.12832 (2018).
Williams, P. H. The Crucifer Genetics Cooperative. Plant Molecular Biology Reporter 3, 129–144, https://doi.org/10.1007/bf02885592 (1985).
Sokal, R. R. & Rohlf, F. J. The principles and practice of statistics in biological research. (WH Freeman and company San Francisco, 1969).
Gordon, A. & Hannon, G. Fastx-toolkit. FASTQ/A short-reads preprocessing tools (unpublished), http://hannonlab cshl. edu/fastx_toolkit (2010).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881, https://doi.org/10.1093/bioinformatics/btq057 (2010).
Chalhoub, B. et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 950–953, https://doi.org/10.1126/science.1253435 (2014).
Kofler, R., Pandey, R. V. & Schlotterer, C. PoPoolation2: identifying differentiation between populations using sequencing of pooled DNA samples (Pool-Seq). Bioinformatics 27, 3435–3436, https://doi.org/10.1093/bioinformatics/btr589 (2011).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Research 33, W116–W120, https://doi.org/10.1093/nar/gki442 (2005).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993, https://doi.org/10.1093/bioinformatics/btr509 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92, https://doi.org/10.4161/fly.19695 (2012).
Li, P. et al. RGAugury: a pipeline for genome-wide prediction of resistance gene analogs (RGAs) in plants. BMC Genomics 17, 852, https://doi.org/10.1186/s12864-016-3197-x (2016).
Sun, F. et al. The high-quality genome of Brassica napus cultivar ‘ZS11’ reveals the introgression history in semi-winter morphotype. Plant J 92, 452–468, https://doi.org/10.1111/tpj.13669 (2017).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome biology 14, R36, https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111, https://doi.org/10.1093/bioinformatics/btp120 (2009).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform, https://doi.org/10.1093/bib/bbx108 (2017).
Sela, I., Ashkenazy, H., Katoh, K. & Pupko, T. GUIDANCE2: accurate detection of unreliable alignment regions accounting for the uncertainty of multiple parameters. Nucleic Acids Res 43, W7–14, https://doi.org/10.1093/nar/gkv318 (2015).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4, 406–425, https://doi.org/10.1093/oxfordjournals.molbev.a040454 (1987).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Research 44, W242–W245, https://doi.org/10.1093/nar/gkw290 (2016).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14, 178–192, https://doi.org/10.1093/bib/bbs017 (2013).
Fu, F.-Y. et al. Localization of QTLs for seed color using recombinant inbred lines of Brassica napus in different environments. Genome 50, 840–854, https://doi.org/10.1139/G07-068 (2007).
Ferreira, M. E., Williams, P. H. & Osborn, T. C. RFLP mapping of Brassica napus using doubled haploid lines. Theoretical and Applied Genetics 89, 615–621, https://doi.org/10.1007/bf00222456 (1994).
Parkin, I. A. P., Sharpe, A. G., Keith, D. J. & Lydiate, D. J. Identification of the A and C genomes of amphidiploid Brassica napus (oilseed rape). Genome 38, 1122–1131, https://doi.org/10.1139/g95-149 (1995).
Sharpe, A. G., Parkin, I. A. P., Keith, D. J. & Lydiate, D. J. Frequent nonreciprocal translocations in the amphidiploid genome of oilseed rape (Brassica napus). Genome 38, 1112–1121, https://doi.org/10.1139/g95-148 (1995).
Lombard, V. & Delourme, R. A consensus linkage map for rapeseed (Brassica napus L.): construction and integration of three individual maps from DH populations. Theoretical and Applied Genetics 103, 491–507, https://doi.org/10.1007/s001220100560 (2001).
Lowe, A. J., Moule, C., Trick, M. & Edwards, K. J. Efficient large-scale development of microsatellites for marker and mapping applications in Brassica crop species. Theoretical and Applied Genetics 108, 1103–1112, https://doi.org/10.1007/s00122-003-1522-7 (2004).
Piquemal, J. et al. Construction of an oilseed rape (Brassica napus L.) genetic map with SSR markers. Theoretical and Applied Genetics 111, 1514–1523, https://doi.org/10.1007/s00122-005-0080-6 (2005).
Tanksley, S. D., Young, N. D., Paterson, A. H. & Bonierbale, M. W. RFLP Mapping in Plant Breeding: New Tools for an Old Science. Bio/Technology 7, 257, https://doi.org/10.1038/nbt0389-257 (1989).
Vos, P. et al. AFLP: a new technique for DNA fingerprinting. Nucleic Acids Research 23, 4407–4414, https://doi.org/10.1093/nar/23.21.4407 (1995).
Powell, W., Machray, G. C. & Provan, J. Polymorphism revealed by simple sequence repeats. Trends in Plant Science 1, 215–222, https://doi.org/10.1016/1360-1385(96)86898-1 (1996).
Konieczny, A. & Ausubel, F. M. A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. The Plant Journal 4, 403–410, https://doi.org/10.1046/j.1365-313X.1993.04020403.x (1993).
Li, G. & Quiros, C. F. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theoretical and Applied Genetics 103, 455–461, https://doi.org/10.1007/s001220100570 (2001).
Dakouri, A. et al. Analysis of genome-wide variants through bulked segregant RNA sequencing reveals a major gene for resistance to Plasmodiophora brassicae in Brassica oleracea. Sci Rep 8, 17657, https://doi.org/10.1038/s41598-018-36187-5 (2018).
Huang, Z. et al. Fine Mapping of a Clubroot Resistance Gene in Chinese Cabbage Using SNP Markers Identified from Bulked Segregant RNA Sequencing. Front Plant Sci 8, 1448, https://doi.org/10.3389/fpls.2017.01448 (2017).
Bellafiore, S., Barneche, F., Peltier, G. & Rochaix, J. D. State transitions and light adaptation require chloroplast thylakoid protein kinase STN7. Nature 433, 892–895, https://doi.org/10.1038/nature03286 (2005).
Pesaresi, P. et al. Arabidopsis STN7 kinase provides a link between short- and long-term photosynthetic acclimation. Plant Cell 21, 2402–2423, https://doi.org/10.1105/tpc.108.064964 (2009).
Schonberg, A. et al. Identification of STN7/STN8 kinase targets reveals connections between electron transport, metabolism and gene expression. Plant J 90, 1176–1186, https://doi.org/10.1111/tpj.13536 (2017).
Dietzel, L. et al. Identification of Early Nuclear Target Genes of Plastidial Redox Signals that Trigger the Long-Term Response of Arabidopsis to Light Quality Shifts. Mol Plant 8, 1237–1252, https://doi.org/10.1016/j.molp.2015.03.004 (2015).
Tikkanen, M., Gollan, P. J., Suorsa, M., Kangasjarvi, S. & Aro, E. M. STN7 Operates in Retrograde Signaling through Controlling Redox Balance in the Electron Transfer Chain. Front Plant Sci 3, 277, https://doi.org/10.3389/fpls.2012.00277 (2012).
Goral, T. K. et al. Visualizing the mobility and distribution of chlorophyll proteins in higher plant thylakoid membranes: effects of photoinhibition and protein phosphorylation. Plant J 62, 948–959, https://doi.org/10.1111/j.0960-7412.2010.04207.x (2010).
Balesdent, M. H. A. A., D. Ansan-Melayah, R. Delourme, M. Renard, and T. Rouxel. Genetic Control and Host Range of Avirulence Toward Brassica napus Cultivars Quinta and Jet Neuf in Leptosphaeria maculans. Phytopathology 91, 70–76, https://doi.org/10.1094/phyto.2001.91.1.70 (2001).
Acknowledgements
Authors gratefully acknowledge the funding for this research provided by the SaskCanola. We also acknowledge NSERC Discovery grant funding awarded to Dilantha Fernando.
Author information
Authors and Affiliations
Contributions
F.Y., D.F. and G.P. conceived the study. F.Y.F. performed data mining, bioinformatics analysis, fine mapping and drafted the manuscript. X.L. and C.Z. isolated RNA and performed RNA-Seq. R.W. performed greenhouse experiments. All authors wrote, reviewed the manuscript and approved the final draft.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Authors contributions:
F.Y., D.F. and G.P. conceived the study. F.Y.F. performed data mining, bioinformatics analysis, fine mapping and drafted the manuscript. X.L. and C.Z. isolated RNA and performed RNA-Seq. R.W. performed greenhouse experiments. All authors wrote, reviewed the manuscript and approved the final draft.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fu, F., Liu, X., Wang, R. et al. Fine mapping of Brassica napus blackleg resistance gene Rlm1 through bulked segregant RNA sequencing. Sci Rep 9, 14600 (2019). https://doi.org/10.1038/s41598-019-51191-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-51191-z
This article is cited by
-
Identification and fine mapping of a major QTL (qRtsc8-1) conferring resistance to maize tar spot complex and validation of production markers in breeding lines
Theoretical and Applied Genetics (2022)
-
Genome-wide transcriptome reveals mechanisms underlying Rlm1-mediated blackleg resistance on canola
Scientific Reports (2021)
-
Identification of genomic regions associated with resistance to blackleg (Leptosphaeria maculans) in canola using genome wide association study
European Journal of Plant Pathology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.