Abstract
Streptococcus thermophilus is a lactic acid bacterium widely used by the dairy industry for the manufacture of yogurt and specialty cheeses. It is also a Gram-positive bacterial model to study phage-host interactions. CRISPR-Cas systems are one of the most prevalent phage resistance mechanisms in S. thermophilus. Little information is available about other host factors involved in phage replication in this food-grade streptococcal species. We used the model strain S. thermophilus SMQ-301 and its virulent phage DT1, harboring the anti-CRISPR protein AcrIIA6, to show that a host gene coding for a methionine aminopeptidase (metAP) is necessary for phage DT1 to complete its lytic cycle. A single mutation in metAP provides S. thermophilus SMQ-301 with strong resistance against phage DT1. The mutation impedes a late step of the lytic cycle since phage adsorption, DNA replication, and protein expression were not affected. When the mutated strain was complemented with the wild-type version of the gene, the phage sensitivity phenotype was restored. When this mutation was introduced into other S. thermophilus strains it provided resistance against cos-type (Sfi21dt1virus genus) phages but replication of pac-type (Sfi11virus genus) phages was not affected. The mutation in the gene coding for the MetAP induces amino acid change in a catalytic domain conserved across many bacterial species. Introducing the same mutation in Streptococcus mutans also provided a phage resistance phenotype, suggesting the wide-ranging importance of the host methionine aminopeptidase in phage replication.
Similar content being viewed by others
Introduction
Bacteriophages are one of the most important drivers of bacterial evolution as they exert a constant selective pressure1. Bacteria inevitably evolve to acquire phage resistance, which is often associated with a fitness cost2,3,4,5. This evolutionary arms race underscores the complex network of phage-host interactions6,7,8. Comprehensive knowledge of how phages interact with their host components is available for only some Escherichia coli phage-host systems as the literature is very sparse for other bacterial species.
To protect themselves against these viral invaders, bacterial hosts have acquired and evolved numerous phage resistance systems6,8. These anti-phage systems are generally classified based on the step of the infectious cycle they interfere with, for example, inhibition of phage adsorption, preventing DNA entry, DNA degradation, and abortive infection, among others6. However, our knowledge of the bacterial arsenal against phages is partial as new defense systems are still being discovered9,10,11,12.
Streptococcus thermophilus is a lactic acid bacterium extensively used in milk fermentation, second only to Lactococcus lactis in its widespread usage. Phages infecting S. thermophilus have been historically divided into two large groups, the so-called cos- and pac-type phages, also named the Sfi21dt1virus and Sfi11virus genera, respectively. This grouping is based on comparative genome analyses, DNA packaging strategy and the number of major structural proteins13. In 2011, S. thermophilus phage 5093, which did not share the common features of either group, was characterized, prompting the creation of the eponymous taxon14. Five years later, four members of a new S. thermophilus phage taxon (987) were reported15. Still the cos- and pac-type phages are by far the most common S. thermophilus phages worldwide and are responsible for most milk fermentation failures16,17,18,19.
Only a few phage resistance mechanisms have been identified in S. thermophilus. Among the most common phage defense systems encoded by S. thermophilus genomes there are restriction-modification systems, as reflected by the number of different entries in REBASE (http://rebase.neb.com). Two superinfection exclusion (Sie) mechanisms encoded by prophages were also identified in S. thermophilus20,21. Sie systems interfere with the phage genome ejection into the host cytoplasm. However, it is the CRISPR-Cas systems that seem to be the most dominant defense mechanism in this bacterial species9,22. CRISPR-Cas systems are bacterial adaptive immune systems that protect the cell from invading nucleic acids via targeted cleavage23. The bacterial cell accumulates short sequences, named spacers, from the phage genomes into CRISPR arrays that are composed of these spacers interspersed by short repeats9,24. These spacers act as the cell’s memory of previous encounters and serve as guides for specific cleavage of invading DNA. The adaptive nature and efficiency of this system may explain why only a few other phage resistance mechanisms have been identified in S. thermophilus. However, the recent discovery of anti-CRISPR proteins (AcrIIA5 and AcrIIA6) in S. thermophilus virulent phages illustrates the ongoing armsrace between phages and their hosts, and suggests that additional mechanisms are at play25,26.
Recently, phage-immunity systems in S. thermophilus that are linked to cryptic, non-inducible prophages were discovered11. Another study demonstrated that transient inactivation of the CRISPR-Cas system allows isolation of non-CRISPR-mediated bacteriophage insensitive mutants (BIMs)27. Characterization of these non-CRISPR BIMs offers the possibility of identifying new bacterial factors that can be mutated to reduce phage infection such as the newly described mutations in cell wall glycans28.
Here, we used the cos-type virulent phage DT129 and its host strain SMQ-301 to identify an additional host factor involved in phage infection. This phage encodes an anti-CRISPR protein which is naturally disabling most of the CRISPR1-Cas system of S. thermophilus26, one of the two type II-A systems active in S. thermophilus SMQ-301. Using this phage-host system, spontaneous BIMs selected after phage challenge either arise from acquisition of new spacer within the CRISPR3 array24,30 or from unknown mechanism(s). Sequencing the genome of S. thermophilus SMQ-301 BIMs that had not acquired new spacers led to the identification of the methionine aminopeptidase (MetAP) as a key host factor for the replication of S. thermophilus cos-type phages.
Materials and Methods
Bacterial growth and phage propagation
Bacterial strains, phages and plasmids used in this study are listed in Table 1 and Table 2. S. thermophilus was grown at 37 °C for the pre-cultures or 42 °C for the assays in M17 medium (Oxoid) supplemented with 0.5% lactose (LM17). When needed, chloramphenicol (Sigma) was added to a final concentration of 5 µg/mL for growth and selection of S. thermophilus strains containing pNZ12331 and derivatives. Agar was added to a final concentration of 1% for solid media. Escherichia coli was grown with agitation at 37 °C in Luria Broth (LB) supplemented with 20 µg/mL chloramphenicol for selection. Streptococcus mutans HER 1503 was grown at 37 °C and 5% CO2 in Brain Heart Infusion (BHI, Difco), supplemented with 10 µg/mL of chloramphenicol when selecting for cells carrying pNZ123. Phages were propagated as previously described32. Phage adsorption assays were performed as described elsewhere33 and phage DNA replication assays were performed as reported34.
Bacteriophage-insensitive mutant isolation
BIMs were isolated using the soft agar overlay assay as detailed elsewhere32. Briefly, the wild-type host SMQ-301 was challenged with the lytic phage DT1. The bacterial host was grown in LM17 at 42 °C until the optical density at 600 nm (OD600nm) reached 0.6. Then, 300 µL of the culture was added to 3 mL of molten LM17 soft agar supplemented with calcium chloride (10 mM) and phage DT1 was added to achieve a multiplicity of infection (MOI) of 0.1. Plates were incubated at 42 °C for 16 h. The resulting colonies were streaked on LM17 + agar and screened for spacer acquisition in CR1 and CR3 loci using primers CR1-fwd and CR1-revLong for the CR1 locus, and primers CR3-fwd and CR3-rev for the CR3 locus32 (Table S1). When spacer acquisition was not detected in CRISPR arrays, BIMs were conserved for genome sequencing.
DNA isolation, sequencing and bioinformatics analysis
The genomic DNA of the selected BIMs was extracted as previously described35. The genomes were sequenced using the Illumina MiSeq platform. The libraries were prepared with the Nextera XT DNA sample preparation kit according to the manufacturer’s instructions and sequenced using MiSeq reagents (2 × 250 nt paired-end). The average coverage ranged from 6.8- to 80.1-fold. The DNA reads obtained for the genome of nine BIMs were aligned on the S. thermophilus SMQ-301 wild-type genome36 using Novoalign (http://www.novocraft.com) with the default settings. The mutations were extracted from the alignment file using SAMTools37. In-house Python scripts were used to map the mutations in the bacterial genome and to determine their impact in coding sequences. The mutations with the highest score as provided by SAMTools were considered first.
Complementation assays
The BIM S. thermophilus SMQ-301:metAPH206Q was complemented with the wild-type metAP. First, metAP was cloned into pNZ123 using Gibson assembly38. The vector pNZ123 was linearized with XbaI (Roche) according to the manufacturer’s instructions. The insert was amplified with the primers SJL154 and SJL155 (Table S1) with Q5 high-fidelity DNA polymerase (NEB). Both primers had 30-nt extensions complementing the 3′- and 5′-ends, respectively, of the linearized vector. An insert/vector ratio of 3:1 was used for assembly. The master mixture for Gibson assembly was prepared as described previously38. The resulting construction was transformed into E. coli NEB5α according to the manufacturer’s instructions and the clones were selected on LB medium supplemented with agar and 20 µg/mL chloramphenicol. One clone was confirmed by Sanger sequencing (ABI 3730xl) at the Plateforme de séquençage et de génotypage des génomes at the Université Laval. Plasmid DNA was extracted using a QIAprep Spin Miniprep kit (Qiagen) and transformed into S. thermophilus32. The clones were selected by spreading the transformation mixture on LM17 supplemented with agar and 5 µg/mL of chloramphenicol.
Proteomic analysis of the phage-infected S. thermophilus DGCC7796 cells
Phage infection and protein analyses were performed as previously described39. Briefly, S. thermophilus DGCC7796 and S. thermophilus DGCC7796:metAPH206Q were each grown to an OD600nm of 0.5 at 42 °C. The cultures were infected with cos-type phage D4090 or pac-type phage M5876 at a MOI of 5. The infections were arrested after 20 min by harvesting the infected cells by centrifugation and the cell pellet was flash-frozen at −80 °C. The cell pellet was quickly thawed and resuspended in 200 µL of lysis buffer (0.5% sodium deoxycholate, 50 mM ammonium bicarbonate, 5 mM DL-Dithiothreitol (DTT), Complete Protease Inhibitor Cocktail (1 µL for 12.5 mL of buffer, Roche)). An equal volume of acid-washed glass beads was added to the cell suspension. The mixture was vortexed in a Mini-Beadbeater-8 cell (BioSpec Products) five times for 1 min with 1 min intervals on ice. The final lysate was centrifuged and the soluble fraction was kept at −80 °C until analysis. The LC-MS/MS analysis was done using a QTRAP 6500 at the Centre de Protéomique de l′Est du Québec. The spectra were analyzed using the complete peptide database, a peptide database that contained only the N-terminal peptides, and a database that contained only the N-terminal peptides without the N-terminal methionine. The latter two databases allowed the distinction of post-translational processed proteins from native proteins that still had the N-terminal methionine. The final analysis was done with Scaffold v4.
Directed and random mutagenesis
The metAP gene with the desired mutation was amplified by PCR using the primers SJL128 and SJL130 (Table S1). At least 10 PCR reactions of 50 µL were done for each assay to obtain enough DNA for natural transformation. All reactions were pooled and precipitated by adding 1.5 mL of 95% ethanol and 75 µL of sodium acetate 3 M (pH 5.2). The tubes were centrifuged at 25,000 × g for 20 min at 4 °C. The DNA pellet was washed twice with 1 mL of 70% ethanol and let dry for 10 min at room temperature to remove traces of ethanol. Finally, the pellets were dissolved in 150 µL of water. The linear DNA fragments were introduced into S. thermophilus by natural transformation40,41. Briefly, S. thermophilus strains SMQ301, DGCC7710, DGCC7796 and DGCC782 were grown overnight at 37 °C in 1 mL of LM17. The cultures were centrifuged at 17,000 × g for 1 min, and the bacterial cells were washed twice with chemically defined medium (CDM)40,41 and recovered in 1 mL of CDM. The final cultures were diluted 30-fold to obtain an OD600nm of 0.05 and incubated 75 min at 37 °C before storing at −20 °C. For the transformation, 1 µM of the ComS peptide and 1 µg of linear DNA were added to 300 µL of naturally competent cells. The mixture was incubated for 3 h at 37 °C. The cells were serially diluted and spread on non-selective LM17 agar to obtain isolated colonies. The colonies were screened using a PCR approach specifically designed to detect clones with the desired mutation. One of the primers used in this protocol included the desired mutation at its 3′-end. Thus, the PCR amplification was only positive if the mutation was present in the clone screened. Primers SJL150 and SJL151 (Table S1) were used for screening the transformants. A similar approach was used for random mutagenesis, but we amplified the wild-type metAP gene using an error-prone PCR amplification. The proof-reading activity of the DNA polymerase was reduced by adding 10 µM of MnSO4.. Phage DT1 was used for the selection of resistant clones.
Directed metAP mutagenesis in Streptococcus mutans
The metAP gene was amplified from S. mutans HER 1503 with two sets of primers containing the desired mutation (MetAPH206Q). The two sets of primers (SJL160 and SJL161 as well as SJL162 and SJL163; Table S1) generated two overlapping amplicons, with the mutation in the overlapping region. The two fragments were purified using the QIAquick PCR purification kit (Qiagen) and fused by Gibson Assembly38. A PCR was then performed on the Gibson Assembly sample using primers SJL160 and SJL163 (Table S1) to amplify the fused fragments. After purification using the QIAquick PCR purification kit, the mutated metAP gene was transformed into S. mutans HER 1503 by natural transformation. The Competence Stimulating Peptide (CSP) ordered from Biomatik was added to 500 μl of an exponentially growing culture of S. mutans (OD600nm of 0.1) at a concentration of 1 μM along with 1 μg of purified mutated metAP amplicon42. The culture was incubated overnight at 37 °C with 5% CO2, then mixed with the virulent phage M102AD43 in BHI supplemented with 0.7% agar and plated onto BHI supplemented with agar. The metAP gene in the surviving BIMs was amplified by PCR using primers SJL160 and SJL163 (Table S1) and sequenced to confirm the mutation.
For the complementation of the S. mutans BIM HER 1503:metAPH206Q, the wild-type S. mutans metAP was cloned into pNZ123. pNZ123 was linearized with XhoI and EcoRI to remove the 24 base pairs between the two sites and the resulting plasmid was purified using the QIAquick Gel Extraction kit (Qiagen). The metAP gene was amplified from S. mutans HER 1503 using primers CM145 and CM146 (Table S1) to generate the gene, flanked by XhoI and EcoRI restriction sites. The PCR sample was purified using the QIAgen PCR purification kit, cut by XhoI and EcoRI and ligated into the linearized pNZ123 using T4 DNA ligase. The molar insert/vector ratio used during the ligation was 3:1. The ligated plasmid was transformed into E. coli NEB5α and plated onto solid LB medium supplemented with 20 µg/mL chloramphenicol. Amplification of the metAP gene by primers CM145 and CM146 and sequencing confirmed the sequence cloned in the complementation plasmid. Plasmid DNA was extracted using a QIAprep Spin Miniprep kit and transformed into the S. mutans HER 1503:metAPH206Q. The strain was grown in BHI until an OD600nm of 0.1 was reached, an aliquot of 500 µL was exposed to 1 µM CSP and transformed with 1 µg of the complementation plasmid. After a 2.5-hour incubation period at 37 °C with 5% CO2, the culture was plated onto BHI + agar supplemented with 10 µg/mL of chloramphenicol.
Growth curves of S. thermophilus strains
Bacterial strains were grown overnight at 37 °C. M17 medium (Nutri-Bact) supplemented with 0.5% lactose (LM17) were inoculated at 1% with the pre-cultures and incubated at 42 °C. The OD600nm was taken every 30 min for 8 hours. Generation time was calculated based on three biological triplicates and from the linear section of the semi-logarithm graph of the OD600nm as a function of time.
Mutation stability test
To test the stability of the mutation H206Q, we inoculated (1%) the strains S. thermophilus SMQ-301 and DGCC7796 and their respective mutants in 10 mL of M17 medium (Nutri-Bact) and in reconstituted milk (10% nonfat dry milk). The strains were incubated at 42 °C during the day. Then, an inoculum of 1% was transferred to fresh media or milk and incubated at 37 °C for the night. After 9 transfers (5 days, 4 nights) corresponding to approximately 60 generations, we randomly selected 10 colonies from each of the four strains and tested them for sensitivity to phage DT1 (cos) for S. thermophilus SMQ-301 and S. thermophilus SMQ-301:metAPH206Q, and phages D4090 (cos) and D4274 (pac) for S. thermophilus DGCC7796 and S. thermophilus DGCC7796:metAPH206Q. Sequencing of the metAP gene and CR1 was done for SMQ-301 and SMQ-301:metAPH206Q colonies with primers SJL128/SJL130 and CR1-fwd/CR1-revLong, respectively, to confirm both the presence of the mutation after the transfers and the identity of the strains.
Results and Discussion
A mutation in the gene coding for the methionine aminopeptidase provides phage resistance
We isolated BIMs of S. thermophilus SMQ-301 that were resistant to phage DT1. Most of the BIMs had acquired new spacers targeting the phage DT1 genome in their CRISPR3 locus (data not shown). Since we were interested in the discovery of other host factors involved in phage infection, we selected nine SMQ-301 BIMs that had not acquired any new spacer, conjecturing they had a mutation elsewhere in their chromosome to provide phage resistance. We sequenced the genomes of these nine non-CRISPR BIMs. By comparing these sequences with that of the wild-type strain SMQ-301 (accession CP011217.1)36 we found several mutations (data not shown). Although many of these mutations may potentially be involved in phage infection, the same transversion at position 1,426,980 occurred in the genomes of three of the nine BIMs (designated BIM #2, #3, and #5). This mutation resides within the gene coding for methionine aminopeptidase (MetAP; locus_tag SMQ301_1544, T618G) and induces an amino acid substitution from a histidine to a glutamine at position 206 (H206Q) in the protein. BLAST searches indicated that this histidine is conserved in all MetAP sequences analyzed, including those deduced from all S. thermophilus sequences currently available in GenBank (29 complete and 27 draft genomes; data not shown). While the MetAP of S. thermophilus has not been previously studied, it shares 89% identity and 96% similarity with the MetAP of Streptococcus pneumoniae TIGR4 for which the structure has been determined (PDB 4KM3)44. Although the histidine at position 206 was not annotated as part of the active site (Fig. 1), this residue was shown to be a substrate-binding site in the MetAP of E. coli45. While this site is not essential for the catalytic activity of E. coli MetAP, replacing the corresponding histidine at position 178 with an alanine led to an impairment of the catalytic activity by 70–190 fold46.
Protein alignment of S. thermophilus SMQ-301 MetAP with 4 other MetAP for which the structure is available. The WebLogo was only kept when the residue was conserved in all sequences. The secondary structures are represented over each sequence and diamonds indicate active sites and substrate binding sites. Red boxes highlight the mutations that provide phage resistance listed in Table 4. When linked, they occurred in the same MetAP mutant of S. thermophilus SMQ-301. Uniprot accession number of the protein sequences: E. coli K12 (P0AE18), Rickettsia prowazekii Madrid E (Q9ZCD3), S. pneumonia TIGR4 (B2IQ22) and M. tuberculosis H37Rv (P9WK19).
A mutation in the metAP gene confers phage resistance in S. thermophilus
To confirm that the phage resistance phenotype was due to the MetAPH206Q mutation – and not to other mutations in the genome – the mutated gene metAPH206Q was amplified and transformed into wild-type strain SMQ-301 using natural competence. Selecting with phage DT1 is very efficient, but it also increases the odds of selecting CRISPR BIMs as well as other mutations that confer phage resistance. Instead, we designed a PCR strategy to detect the transformants that integrated the PCR product with the mutation into their genome, in the absence of phage selective pressure. We used the primers SJL150 in combination with SJL151 with the desired mutation at its 3′-end (Table S1) to specifically detect the mutation in the bacterial genome. To avoid false positives due to the potential presence of residual linear DNA from the transformation inside the bacterial cytoplasm, we designed the primer SJL150 that matched the flanking genomic region of the transformed DNA fragment. A total of 282 clones were screened for the desired mutation and four positive clones were obtained (MetAPH206Q). We randomly selected one of these clones (designated SMQ-301:metAPH206Q) and tested its phage resistance phenotype. No plaque was visible on the bacterial lawn of SMQ-301:metAPH206Q using various titers of the phage DT1, indicating a very strong phage resistance phenotype (Table 3). It was possible to observe a lysis zone at high phage titers, likely due to lysis from without or to the endolysin activity in the phage lysate. We also tried to isolate DT1 phage mutants that would overcome the effect of the MetAPH206Q mutation using various experimental conditions, but to no avail. We examined incubation temperature, anaerobic conditions, added different concentrations of glycine to the medium to weaken the bacterial cell wall, replaced agar with different concentrations of agarose and even using liquid medium in case the lysis plaque were not visible enough. Overall, we were unable to obtain phage DT1-derivatives able to propagate on SMQ-301:metAPH206Q.
Complementation with the wild-type allele restores phage sensitivity
We cloned the wild-type metAP gene from strain SMQ-301 into the expression vector pNZ123. The resulting construct was transformed into S. thermophilus SMQ-301:metAPH206Q to complement the mutation in trans. Sensitivity to phage DT1 was completely restored for the two of the three spontaneous BIMs complemented with plasmid pNZ123:metAP (Table 3), confirming that the metAPH206Q mutation was solely responsible for the phage resistance phenotype. Of note, the sensitivity to phage DT1 was not completely restored (EOP = 0.02) for the spontaneous BIM #3, suggesting that other mutations are at play (Table 3). The mutated SMQ-301:metAPH206Q metAP gene was also cloned into pNZ123 and transformed into the wild-type strain SMQ-301. There was no difference in EOP between the strain with pNZ123 when compared to the strain with pNZ123:metAPH206Q, suggesting that the mutation does not produce a dominant phenotype.
The MetAP mutation has a broad range of action against cos-type phages
To determine if the resistance provided by the MetAPH206Q mutation is phage-dependent, we tested the cos-type phage MD247, which is also capable of infecting strain SMQ-301. Phage MD2 was also severely inhibited by the mutated MetAP (EOP <1 × 10−6). To show that this mutation is not bacterial strain-dependent, we introduced the MetAPH206Q mutation into several S. thermophilus strains using directed mutagenesis. We introduced the mutation into strain DGCC782, which is sensitive to cos-type phages N1032, N1117, N1119, N1169, N1358 and N3782 (Table 2). The mutation was also introduced into the strain S. thermophilus DGCC7710, which is sensitive to cos- (D5691, D5913, D6037 and D6215) and pac-type phages (858, 2972, D3288, 4259, D4752, D4754 and D939) (Table 2). Finally, the mutation was introduced into S. thermophilus DGCC7796, which is also sensitive to cos- (D4090, D5821 and D4807) and pac-phages (D2765, D4274, D5787 and M5876) (Table 2). Although the mutation was less efficient against all four cos-type phages infecting S. thermophilus DGCC7710 (EOP 10−4), the mutation provided resistance against all cos-type phages tested. The introduction of MetAPH206Q into S. thermophilus DGCC7796 and DGCC782 led to a reduction in the EOP of at least 10−6 for all cos-type phages (Table 2), which corresponded to the limit of detection. No phage mutant could be recovered from these assays. However, none of the S. thermophilus pac-type phages were affected by the mutation, suggesting that MetAP is not key for the replication of these phages.
To determine if the MetAPH206Q mutation would provide resistance to another streptococcal species, we introduced it into the S. mutans strain HER 1503, which is sensitive to the cos-type virulent phage M102AD43. The methionine aminopeptidase gene of S. mutans HER 1503 has the same length (861 bp) as that of S. thermophilus. The two genes share 74% identity while the two MetAP share 85% identity. The histidine residue found at position 206 is present in S. mutans. The MetAPH206Q mutation inserted into S. mutans provided complete resistance to phage M102AD (EOP <1 × 10−7). Complementation of the mutated strain with the wild-type metAP restored phage sensitivity. Our data suggest a relatively widespread role of MetAP in phage replication.
Other mutations in MetAP affect replication of phage DT1
To determine if other mutations in the metAP gene provide phage resistance, we used an error-prone PCR amplification approach. The resulting PCR products were transformed into S. thermophilus SMQ-301 using natural competence. Since we were not introducing an antibiotic resistance marker, we used phage DT1 to select the clones. The transformation assay with the positive control (MetAPH206Q mutation) resulted in many more BIMs than with the negative control (no DNA) (Fig. S1). Fewer BIMs were obtained with the PCR product containing random mutations than with the positive control (Fig. S1). The metAP gene was sequenced in 10 randomly selected clones and all of them had various mutations. Seven mutants had a single mutation in the metAP gene resulting in an amino acid substitution. We tested phage sensitivity of these mutants and they were all resistant to phage DT1. We complemented the 10 resistant strains with pNZ123:metAP and as previously observed, the wild-type MetAP restored phage sensitivity, confirming that other mutations in the MetAP can confer phage resistance (Table 4; Fig. 1).
Phage adsorption and phage DNA replication are not affected
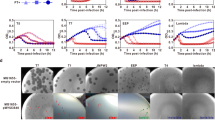
To understand the molecular mechanism underlying the phage resistance provided by this mutation, we dissected the lytic cycle of a phage infecting a mutated strain. First, we tested if phage DT1 could still adsorb to S. thermophilus SMQ-301:metAPH206Q (Table 3). The viral particles adsorbed at a slightly higher level on the wild-type phage-sensitive host (93%) than on the spontaneous BIMs #2, #3, and #5 (80% to 86%). The adsorption of DT1 was more reduced on S. thermophilus SMQ-301:metAPH206Q (68%). However, this reduced adsorption cannot explain the strong phage resistance phenotype (EOP of 10−8), suggesting that another step of the lytic cycle is impeded by the MetAPH206Q mutation. Since phage DT1 can adsorb to the BIM surface, we tested whether phage DNA was replicated inside the bacterial cell. Both S. thermophilus SMQ-301 and SMQ-301:metAPH206Q were infected with phage DT1 using a MOI of 5 and total DNA was extracted from the infected cells at different time points, digested with a restriction enzyme and migrated on an agarose gel (Fig. 2). The DNA fragments corresponding to DT1 digested genome were visible after 20 min of infection in both the MetAPH206Q mutant and the wild-type cells, indicating that the phage DNA entered the bacterial cell and was replicated (Fig. 2). Lysis of the culture was only observed with S. thermophilus SMQ-301.
The MetAPH206Q mutation affects N-terminal methionine processing
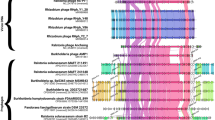
Since the MetAPH206Q mutation does not affect phage DNA replication, we investigated whether protein expression was affected by the mutation. To compare the activity of MetAP, we used S. thermophilus DGCC7796, a strain sensitive to cos- and pac-type phages. Using mass spectrometry (LC-MS/MS), we analyzed the complete proteome of the wild-type and mutant DGCC7796 after 20 min of infection with phages D4090 (cos-type) or M5876 (pac-type) using a MOI of 5. During the replication of phage D4090, although we observed a slight difference in protein expression, there was no significant trend in protein expression between mutant and wild-type strains (Fig. 3, Supp. Table 1). Peptides of 10 and 12 phage proteins were detected in the wild-type and mutant strains, respectively, and their relative abundance was similar in both strains. The difference in the number of proteins detected can be explained by the very low level of detection of the corresponding proteins in the wild-type strain.
Genetic alignment of phages D4090 and DT1. Each protein-coding gene is represented by an arrow. When two deduced proteins share 70% identity or more, they are represented with the same color, otherwise they are shown in white. The boxes between the two genomes represent the expression modules. Phage D4090 expression modules were extrapolated from DT1 experimental data. The graph above the alignment represents the relative abundance of the proteins when phage D4090 infects the wild-type strain DGCC7796 and mutant DGCC7796:metAPH206Q.
We then used the proteomic datasets to verify the activity of the mutant and wild-type MetAP. We generated two distinct databases of peptides to search against. The first database was composed of only N-terminal peptides after tryptic digestion. The second database was essentially the same as the first one but we removed the N-terminal methionine of each peptide. This allowed us to determine if the protein underwent N-terminal post-translational processing. The complete database included the 2,051 proteins encoded by the host genome and the 41 proteins encoded by the phage genome. Following in silico tryptic digestion, the N-terminal peptide database included 579 peptides of 8–30 amino acids that have a molecular mass that could be detected with the setting used to conduct the mass spectrometry analysis. Using LC-MS/MS, we detected a total of 134 from 579 N-terminal peptides across all samples analyzed (Table S2). We did not observe any difference in post-translational processing of the N-terminal methionine between the uninfected and infected samples nor between the cos- and pac-type phage-infected wild-type cells, suggesting that phage infection does not affect MetAP activity. Then, we compared the data with the MetAP mutated cells and classified the results into four groups: A) 28 proteins were unprocessed by MetAP in both wild-type and mutant strains, B) 34 proteins were processed by MetAP in both the wild-type and mutant strains, C) 22 proteins were processed by MetAP in the wild-type strain only, and D) only one protein was processed by MetAP in the mutated strain (Fig. 4).
Proteomic analysis of the N-terminal peptides from the proteome of the wild-type strain DGCC7796 and mutant DGCC7796:metAPH206Q. Post-translational processing of the N-terminal methionine was detected in both proteomes but it was less abundant in the mutant strain. N represents unprocessed peptides while Y stands for processed peptides.
E. coli methionine aminopeptidase is more specific for protein with a penultimate amino acid (a.a.) with small side chain (e.g. Ala, Gly, Pro, Ser, Thr, or Val)49,50. Thus, it is not surprising that most proteins (Thr is the only exception out of 28) found in group A (non-processed) have penultimate a.a. with long side chain (Ile, Asn, Leu, Glu, Tyr, Gln and Asp). Most of the penultimate a.a. of proteins found in groups B and C have short side chain (Ala, Gly, Pro, Ser, Thr, or Val) with only two exceptions found in group C (Cys and Lys). Since there are no significant difference in the penultimate a.a. of groups B and C, the findings that some proteins were not processed by the mutated MetAP (group C) suggest that its activity is impaired to some degree. Analyzing functions of the above proteins and metabolic pathways in which they are involved did not reveal why phage infection was blocked in presence of the mutated MetAP (Supp. Table 2).
The MetAPH206Q mutation affects growth of the bacterial strains
Growth of the mutant strains was also evaluated (Fig. 5). Surprisingly, the MetAPH206Q mutation had different impact depending of the S. thermophilus strains. The growth rate difference between the wild-type strains and their mutants was not significant, except for S. thermophilus DGCC7796 and its mutant for which the growth rates were respectively 36.2 ± 3.6 min and 53.5 ± 3.3 min (Table 5). We also observed a latent period longer with S. thermophilus DGCC7796:metAPH206Q than with the other strains tested (Fig. 5).
The MetAPH206Q mutation is stable
One of the important features for industrial fermentation is the stability of the phage resistance phenotype. Thus, we verified the stability of the mutation MetAPH206Q over 60 generations of S. thermophilus SMQ-301 and DGCC7796 as well as their mutants in milk and LM17. While all selected colonies of the wild -type strains remained phage sensitive, all 10 colonies from each mutant remained phage resistant, indicating that the MetAPH206Q mutation is stable.
Conclusion
While cell wall proteins and polysaccharides were identified as host factors needed for phage infection, the literature is sparse about cytoplasmic proteins. MetAP are cytoplasmic enzymes found in all living organisms51. They are a unique class of proteases that remove the N-terminal residue from nascent proteins and play a central role in the synthesis and maturation of proteins in prokaryotes and eukaryotes52. The essentiality of this post-translational processing is underscored by the lethality of metAP gene inactivation in E. coli53, Salmonella typhimurium54, and Saccharomyces cerevisiae55. This dependence now extends to some of their parasites.
In this study, we showed that phage resistance can be acquired through mutations in the metAP gene of S. thermophilus and S. mutans. We were unable to isolate phage mutants that overcame MetAPH206Q mutation, confirming that such mutation can provide a robust phage resistance. Early steps (adsorption, DNA replication, protein expression) of the phage lytic cycle occurred in phage-infected S. thermophilus SMQ-301:metAPH206Q, suggesting that this mutation acts lately during the lytic cycle, perhaps at the virion assembly step. Since the catalytic activity of MetAP is at the core of living cells, we expected that the MetAPH206Q mutation provided a broad phage resistance. Although the replication of cos-type phages was affected by MetAP mutation, the pac-type phages were not. As the enzymatic activity of MetAP:H206Q was less effective than the wild-type MetAP, it appears that cos-type phages require efficient cleavage of the N-terminal methionine of some of their proteins and/or of their host to complete their lytic cycle.
Change history
11 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Cobián Güemes, A. G. et al. Viruses as winners in the game of life. Annu Rev Virol 3, 197–214 (2016).
Lennon, J. T., Khatana, S. A., Marston, M. F. & Martiny, J. B. Is there a cost of virus resistance in marine cyanobacteria. ISME J 1, 300–312 (2007).
Koskella, B. & Brockhurst, M. A. Bacteria-phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol Rev 38, 916–931 (2014).
Avrani, S. & Lindell, D. Convergent evolution toward an improved growth rate and a reduced resistance range in Prochlorococcus strains resistant to phage. Proc Natl Acad Sci USA 112, E2191–200 (2015).
Vale, P. F. et al. Costs of CRISPR-Cas-mediated resistance in Streptococcus thermophilus. Proc Biol Sci 282, 20151270 (2015).
Labrie, S. J., Samson, J. E. & Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 8, 317–327 (2010).
Buckling, A. & Brockhurst, M. Bacteria-virus coevolution. Adv Exp Med Biol 751, 347–370 (2012).
Samson, J. E., Magadán, A. H., Sabri, M. & Moineau, S. Revenge of the phages: defeating bacterial defences. Nat Rev Microbiol 11, 675–687 (2013).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Kronheim, S. et al. A chemical defence against phage infection. Nature 564, 283–286 (2018).
da Silva Duarte, V. et al. A cryptic non-inducible prophage confers phage-immunity on the Streptococcus thermophilus M17PTZA496. Viruses 11, 1 (2018).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, 6379 (2018).
Le Marrec, C. et al. Two groups of bacteriophages infecting Streptococcus thermophilus can be distinguished on the basis of mode of packaging and genetic determinants for major structural proteins. Appl Environ Microbiol 63, 3246–3253 (1997).
Mills, S. et al. A new phage on the ‘Mozzarella’block: bacteriophage 5093 shares a low level of homology with other Streptococcus thermophilus phages. International dairy journal 21, 963–969 (2011).
McDonnell, B. et al. Identification and analysis of a novel group of bacteriophages infecting the lactic acid bacterium Streptococcus thermophilus. Appl Environ Microbiol 82, 5153–5165 (2016).
McDonnell, B. et al. Global survey and genome exploration of bacteriophages infecting the lactic acid bacterium Streptococcus thermophilus. Front Microbiol 8, 1754 (2017).
Achigar, R., Magadán, A. H., Tremblay, D. M., Julia Pianzzola, M. & Moineau, S. Phage-host interactions in Streptococcus thermophilus: Genome analysis of phages isolated in Uruguay and ectopic spacer acquisition in CRISPR array. Sci Rep 7, 43438 (2017).
Lavelle, K. et al. A decade of Streptococcus thermophilus phage evolution in an Irish dairy plant. Appl Environ Microbiol 84, 10 (2018).
Lavelle, K. et al. Biodiversity of Streptococcus thermophilus phages in global dairy fermentations. Viruses 10, 577 (2018).
Ali, Y. et al. Temperate Streptococcus thermophilus phages expressing superinfection exclusion proteins of the Ltp type. Front Microbiol 5, 98 (2014).
Sun, X., Göhler, A., Heller, K. J. & Neve, H. The ltp gene of temperate Streptococcus thermophilus phage TP-J34 confers superinfection exclusion to Streptococcus thermophilus and Lactococcus lactis. Virology 350, 146–157 (2006).
Horvath, P. et al. Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol 190, 1401–1412 (2008).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J Bacteriol 190, 1390–1400 (2008).
Hynes, A. P. et al. An anti-CRISPR from a virulent streptococcal phage inhibits Streptococcus pyogenes Cas9. Nat Microbiol 2, 1374–1380 (2017).
Hynes, A. P. et al. Widespread anti-CRISPR proteins in virulent bacteriophages inhibit a range of Cas9 proteins. Nat Commun 9, 2919 (2018).
McDonnell, B., Mahony, J., Hanemaaijer, L., Kouwen, T. R. H. M. & van Sinderen, D. Generation of bacteriophage-insensitive mutants of Streptococcus thermophilus via an antisense RNA CRISPR-Cas silencing approach. Appl Environ Microbiol 84, 4 (2018).
Szymczak, P. et al. Cell wall glycans mediate recognition of the dairy bacterium Streptococcus thermophilus by bacteriophages. Appl Environ Microbiol 84, 23 (2018).
Tremblay, D. M. & Moineau, S. Complete genomic sequence of the lytic bacteriophage DT1 of Streptococcus thermophilus. Virology 255, 63–76 (1999).
Magadán, A. H., Dupuis, M. È., Villion, M. & Moineau, S. Cleavage of phage DNA by the Streptococcus thermophilus CRISPR3-Cas system. PLoS One 7,, e40913 (2012).
De Vos, W. M. Gene cloning and expression in lactic streptococci. FEMS Microbiology Letters 46, 281–295 (1987).
Hynes, A. P. et al. Detecting natural adaptation of the Streptococcus thermophilus CRISPR-Cas systems in research and classroom settings. Nat Protoc 12, 547–565 (2017).
Garvey, P., Hill, C. & Fitzgerald, G. F. The lactococcal plasmid pNP40 encodes a third bacteriophage resistance mechanism, one which affects phage DNA penetration. Appl Environ Microbiol 62, 676–679 (1996).
Boucher, I., Emond, E., Dion, E., Montpetit, D. & Moineau, S. Microbiological and molecular impacts of AbiK on the lytic cycle of Lactococcus lactis phages of the 936 and P335 species. Microbiology 146, 445–453 (2000).
Bissonnette, F., Labrie, S., Deveau, H., Lamoureux, M. & Moineau, S. Characterization of mesophilic mixed starter cultures used for the manufacture of aged cheddar cheese. J Dairy Sci 83, 620–627 (2000).
Labrie, S. J. et al. Complete genome sequence of Streptococcus thermophilus SMQ-301, a model strain for phage-host interactions. Genome Announc 3, 3 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6, 343–345 (2009).
Scaltriti, E. et al. Structure and function of phage p2 ORF34(p2), a new type of single-stranded DNA binding protein. Mol Microbiol 73, 1156-1170 (2009).
Gardan, R., Besset, C., Guillot, A., Gitton, C. & Monnet, V. The oligopeptide transport system is essential for the development of natural competence in Streptococcus thermophilus strain LMD-9. J Bacteriol 191, 4647–4655 (2009).
Fontaine, L. et al. Development of a versatile procedure based on natural transformation for marker-free targeted genetic modification in Streptococcus thermophilus. Appl Environ Microbiol 76, 7870–7877 (2010).
Dufour, D., Cordova, M., Cvitkovitch, D. G. & Lévesque, C. M. Regulation of the competence pathway as a novel role associated with a streptococcal bacteriocin. J Bacteriol 193, 6552–6559 (2011).
Delisle, A. L. et al. Biology and genome sequence of Streptococcus mutans phage M102AD. Appl Environ Microbiol 78, 2264–2271 (2012).
Arya, T., Kishor, C., Saddanapu, V., Reddi, R. & Addlagatta, A. Discovery of a new genetic variant of methionine aminopeptidase from Streptococci with possible post-translational modifications: biochemical and structural characterization. PLoS One 8, e75207 (2013).
Lowther, W. T. et al. Escherichia coli methionine aminopeptidase: implications of crystallographic analyses of the native, mutant, and inhibited enzymes for the mechanism of catalysis. Biochemistry 38, 7678–7688 (1999).
Copik, A. J. et al. Kinetic and spectroscopic characterization of the H178A methionyl aminopeptidase from Escherichia coli. Biochemistry 42, 6283–6292 (2003).
Duplessis, M. & Moineau, S. Identification of a genetic determinant responsible for host specificity in Streptococcus thermophilus bacteriophages. Molecular microbiology 41, 325–336 (2001).
Lévesque, C. et al. Genomic organization and molecular analysis of virulent bacteriophage 2972 infecting an exopolysaccharide-producing Streptococcus thermophilus strain. Appl Environ Microbiol 71, 4057–4068 (2005).
Hirel, P.-H., Schmitter, M. J., Dessen, P., Fayat, G. & Blanquet, S. Extent of N-terminal methionine excision from Escherichia coli proteins is governed by the side-chain length of the penultimate amino acid. Proc Natl Acad Sci 86, 8247–8251 (1989).
Xiao, Q., Zhang, F., Nacev, B. A., Liu, J. O. & Pei, D. Protein N-terminal processing: substrate specificity of Escherichia coli and human methionine aminopeptidases. Biochemistry 49, 5588–5599 (2010).
Giglione, C., Boularot, A. & Meinnel, T. Protein N-terminal methionine excision. Cell Mol Life Sci 61, 1455–1474 (2004).
Giglione, C., Fieulaine, S. & Meinnel, T. N-terminal protein modifications: Bringing back into play the ribosome. Biochimie 114, 134–146 (2015).
Chang, S. Y., McGary, E. C. & Chang, S. Methionine aminopeptidase gene of Escherichia coli is essential for cell growth. J Bacteriol 171, 4071–4072 (1989).
Miller, C. G., Kukral, A. M., Miller, J. L. & Movva, N. R. pepM is an essential gene in Salmonella typhimurium. J Bacteriol 171, 5215–5217 (1989).
Zuo, S., Guo, Q., Ling, C. & Chang, Y. H. Evidence that two zinc fingers in the methionine aminopeptidase from Saccharomyces cerevisiae are important for normal growth. Mol Gen Genet 246, 247–253 (1995).
Acknowledgements
We would like to thank Barbara-Ann Conway (Medical Writer & Editor) for editorial assistance. S.M. acknowledges funding from the Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Program. S.M. holds a Tier 1 Canada Research Chair in Bacteriophages.
Author information
Authors and Affiliations
Contributions
S.M. headed the project. S.J.L. and S.M. wrote the manuscript and designed the project. S.J.L. executed the experiments. M.E.D. isolated the natural non-CRISPR BIM, C.M. conducted the experiments with S. mutans and S.L. executed the adsorption test and growth curves. G.M.R. performed the mutation stability test and the growth curves. D.T. sequenced the genomes. P.D. assisted S.J.L. for the MAP experiments. D.A.R., P.H. and C.F. helped with the design of the project. All the authors commented and gave feedback for the manuscript. S.J.L., D.A.R., P.H., C.F. and S.M. are co-inventors on patent applications.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Labrie, S.J., Mosterd, C., Loignon, S. et al. A mutation in the methionine aminopeptidase gene provides phage resistance in Streptococcus thermophilus. Sci Rep 9, 13816 (2019). https://doi.org/10.1038/s41598-019-49975-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49975-4
This article is cited by
-
The kill-switch for CRISPR that could make gene-editing safer
Nature (2020)
-
Therapeutic and biotechnological applications of substrate specific microbial aminopeptidases
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.