Abstract
Biological and regulatory mechanisms underlying many multi-gene expression-based disease biomarkers are often not readily evident. We describe an innovative framework, NeTFactor, that combines network analyses with gene expression data to identify transcription factors (TFs) that significantly and maximally regulate such a biomarker. NeTFactor uses a computationally-inferred context-specific gene regulatory network and applies topological, statistical, and optimization methods to identify regulator TFs. Application of NeTFactor to a multi-gene expression-based asthma biomarker identified ETS translocation variant 4 (ETV4) and peroxisome proliferator-activated receptor gamma (PPARG) as the biomarker’s most significant TF regulators. siRNA-based knock down of these TFs in an airway epithelial cell line model demonstrated significant reduction of cytokine expression relevant to asthma, validating NeTFactor’s top-scoring findings. While PPARG has been associated with airway inflammation, ETV4 has not yet been implicated in asthma, thus indicating the possibility of novel, disease-relevant discovery by NeTFactor. We also show that NeTFactor’s results are robust when the gene regulatory network and biomarker are derived from independent data. Additionally, our application of NeTFactor to a different disease biomarker identified TF regulators of interest. These results illustrate that the application of NeTFactor to multi-gene expression-based biomarkers could yield valuable insights into regulatory mechanisms and biological processes underlying disease.
Similar content being viewed by others
Introduction
Biological and regulatory mechanisms underlying most multi-gene expression-based disease biomarkers are often not readily evident. Using RNA sequencing (RNAseq)1 and machine learning2 in a well-characterized cohort of asthmatics and controls, we recently identified a nasal brush-based biomarker of asthma3. This biomarker consists of 90 genes, whose expression is interpreted through a logistic regression function3. Although our nasal biomarker of asthma produced accurate (AUC 0.994) and specific classification of asthma3, the biological and regulatory mechanisms underlying its performance were not readily evident. For instance, although the genes in the biomarker had a higher tendency to be differentially expressed (Kolmogorov-Smirnov statistic = 0.289, FDR = 2.73 × 10−37), only non-specific pathways such as defense response (fold change = 2.86, FDR = 0.006) were enriched in these genes, and only a minority have been previously studied in the context of asthma3. A gene expression-based biomarker like ours is expected to include genes known to associate with the target disease. However, it is also possible and even likely, given our incomplete understanding of complex diseases such as asthma, that genes not previously associated with the disease can provide information that is useful to the classification, and perhaps to the disease process itself. Indeed, such an approach has led to important results in other disease areas such as cancer4,5,6,7, illustrating the idea that RNA traits can serve as sensitive sensors of one state (e.g. disease) relative to another (e.g. healthy) beyond known associations with established disease-related pathways. It is intriguing to consider what further dissection of our asthma and other biomarkers could yield as insights into biologic mechanisms relevant to asthma and other diseases.
Here we describe a novel framework that combines network analyses with RNA sequence (RNAseq) data to identify transcription factors (TFs) significantly regulating a disease biomarker. This framework, named NeTFactor (Network-identified Transcription Factor), uses a computationally inferred context-specific gene regulatory network (GRN)8 to guide the analysis. Such a GRN consists of directed edges denoting interactions between regulators (e.g. TFs) and their target(s) (e.g. gene(s) they regulate). NeTFactor utilizes the structure and constituents of such a GRN to identify the regulators, specifically TFs, that most significantly regulate the genes underlying the biomarker. To illustrate the utility of our framework, we applied NeTFactor to identify the most significant TF regulators of our nasal gene expression-based asthma biomarker3 and then experimentally validated the identified regulators using silencing RNA (siRNA)9 in airway epithelial cell line models. Further, we show that that NeTFactor’s results are robust when the gene regulatory network and biomarker are derived from independent data and additionally demonstrate application of NeTFactor to a different disease biomarker.
Biomolecular networks, including GRNs, have been widely used to glean useful insights into biological processes and how the dysregulation of the constituent interactions may lead to disease8,10,11,12. In particular, network analyses have been used to identify disease-related genes and regulators, often connected through interactions in the network, representing a subnetwork or module13,14,15. Master Regulator Analysis (MRA)16 and its variants17 represent such an approach where a GRN is used to directly identify TF regulators that are expected to be associated with the target disease or phenotype. In parallel, similar to our asthma biomarker, multi-gene expression-based biomarkers have been developed in other disease areas, e.g., breast cancer prognosis4,18. The goal of this study was to analyze a GRN to identify the most significant set of key TF regulators of the set of genes constituting a separately identified biomarker, namely our asthma biomarker. This is complementary to investigating the constituent genes of the biomarker individually, as well as only identifying TF regulators associated with the target disease or phenotype using methods like MRA. In other words, we used computational and systems biology principles19,20,21 to develop a novel framework that integrates machine learning- and network-based analyses of complex biomolecular data.
Results
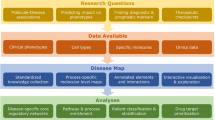
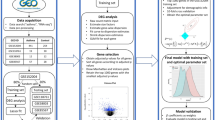
Our study comprised multiple steps (Fig. 1), including the application of NeTFactor to construct a context-specific gene regulatory network (Box 1) and identify TF regulators of the biomarker (Box 2), followed by experimental validation of the inferred TF regulators (Box 3).
Study flow for the identification and validation of transcription factor (TF) regulators of a gene expression-based biomarker of asthma3 using the proposed NeTFactor framework. Box 1 denotes the first step of NeTFactor, namely the inference of gene regulatory networks (GRNs) from the datasets that yielded the original biomarker. Box 2 represents steps 2–4 of NeTFactor which identify the most significant set of likely TF regulators, which are themselves active in the disease and regulate a significant fraction of genes constituting the biomarker. Box 3 depicts siRNA-mediated knock-down experiments in an airway epithelial cell line model employed to experimentally validate the identified regulators.
Development of NeTFactor and its application to nasal RNAseq data and the asthma biomarker
Generation of a context-specific gene regulatory network (GRN)
The first step of NeTFactor is the derivation of a base GRN that reflects the biological context, such as the same tissue of origin, of the target biomarker. For this, in our study, the application of the ARACNE algorithm22,23,24 to nasal RNAseq data from a case-control asthma cohort (n = 150) (Supplementary Table 1) yielded a base GRN consisting of 56976 interactions between 132 TFs and 11049 genes. Since this network was inferred from nasal gene expression data, it is expected to be directly relevant to our nasal brush-based asthma biomarker as well as to asthma overall, given shared biology between the nasal and bronchial airways3,25,26. Applying ARACNE with 1000 bootstraps instead of the default value of 100 generated a much larger but fully encompassing GRN (Fig. 2A), indicating that the core network was preserved between these variations of the algorithm. Although there were no set criteria for selecting the size of the final GRN, we observed that the base network was the closest in size to the total number (66883) of curated TF → target gene interactions in MSigDB27,28 version 5.1, which was also the source of TFs used to derive the ARACNE networks. To capture the extent of our current knowledge of GRNs, we used the 100 bootstrap base GRN for further analyses. However, due to the general lack of knowledge about human TFs and their putative target genes, this network only included 78 of the 90 (87%) genes in the asthma biomarker, placing an upper limit on how many of these genes could be regulated by the TFs in the GRN.
Derivation of context-specific gene regulatory networks (GRNs) from and application of the VIPER algorithm30 to a nasal RNAseq data set. (A) Venn diagram showing the overlap between the TF→target gene interactions constituting GRNs generated by applying the ARACNE algorithm23 to a nasal RNAseq dataset using 100 (N100) and 1000 (N1000) bootstraps shows that the former is completely contained in the latter. (B) Venn diagram showing the statistically significant overlap of the TF→target gene interactions constituting the N100 network and the GRN of the same size inferred using the GENIE3 algorithm29 from the same dataset (Fisher’s exact test p < 2.2e-16). (C) Transcriptional activities of the 12 transcription factors most significantly differentially active in asthma. The first column indicates VIPER FDR value for differential activity (FDR ≤ 0.05) of TFs active in asthma. All genes in the nasal RNAseq data are graphically summarized in the second column, where each vertical line represents a gene, and the genes are rank-sorted left to right from most down-regulated to most up-regulated in asthma vs normal subjects. Blue and red bars indicate negative and positive regulation, respectively, of each gene by the TF shown in the third column. The fourth column shows VIPER-inferred differential activity of TFs, with red and blue entries indicating more and less activity of the TF in asthma, respectively. (D) Heatmap of VIPER-inferred activity scores of TFs constituting the N100 and N1000 networks. The scores are very highly correlated (Pearson’s correlation coefficient = 0.987, p < 2.2e-16). (E) Heatmap of VIPER-inferred activities of TFs constituting the N100 network and the GRN of the same size inferred using the GENIE3 algorithm. The scores are very highly correlated (Pearson’s correlation coefficient = 0.969, p < 2.2e-16).
We also generated a GRN containing the same number of edges as the base ARACNE network using the GENIE3 algorithm29, and found that the two networks overlapped significantly (Fig. 2B; Fisher’s exact test p < 2.2e-16). This supported the robustness of using ARACNE within NeTFactor and its resulting GRN.
VIPER identifies 12 asthma-associated TFs in the GRN
NeTFactor next examines if a TF is differentially active in the disease under consideration, such as asthma in our application study, for the TF to be considered a regulator of the target biomarker under consideration. To identify such TFs, we applied the VIPER algorithm30 to the base GRN and the RNAseq data to identify TFs that are differentially active between asthmatic and control subjects. The output of VIPER consisted of a normalized enrichment score (NES), which was positive for TFs that were more active in asthma and negative for TFs more active in control subjects, as well as the associated false discovery rate (FDRVIPER) value for each NES. Figure 2C shows the 12 TFs found to be differentially active in asthma (FDRVIPER ≤ 0.05). These TFs included HSF1, which has been reported to affect airway hyperresponsiveness and airway inflammation in mice with asthma31.
We also tested the sensitivity of the VIPER results to the choice of the input GRN and the algorithm used to infer it. For this, we compared the TF activity scores (NESs) inferred from the base GRN with those inferred from the ARACNE 1000 bootstrap and GENIE3 networks. As shown in Fig. 2D (Pearson’s correlation coefficient = 0.99, p < 2.2e-16) and 2E (Pearson’s correlation coefficient = 0.97, p < 2.2e-16), the scores from these alternative GRNs were highly correlated with those from the base GRN. These results strongly support that the TF activity scores and downstream analyses utilizing them would be robust to the choice of the network inference algorithm and the resulting GRN.
Context-specific regulators of the asthma biomarker
The next step of NeTFactor assesses if a TF significantly regulates the genes constituting the target biomarker. For this, we calculated the likelihood that a particular TF regulated our asthma biomarker by conducting a Fisher’s exact test32 for the statistical significance of the overlap between the set of genes regulated by each TF in the base nasal GRN (i.e. its regulon) and the member genes of the asthma biomarker. This was followed by correction for multiple hypothesis testing using the Benjamini-Hochberg procedure33, which yielded a regulation likelihood for each TF (FDRBIOMARKER) that was used in subsequent analyses (Supplementary Table 2). The most significant of these regulators (FDRBIOMARKER ≤ 0.05) included XBP1, which modulates endoplasmic reticulum stress in type 2 airway inflammation34 and mucin production35 that may relate to how the well-replicated asthma locus ORMDL affects asthma36.
Convex optimization identifies the most significant set of asthma-active TFs that most significantly and non-redundantly regulate the asthma biomarker
The previous two steps of NeTFactor identified TFs that were disease-active and likely to regulate the target biomarker. The final step of the framework aims to identify the most significant set of TFs that scores highly on both these aspects, but has as little redundancy as possible among the sets of biomarker genes they regulate, thus maximizing the coverage of the biomarker. We considered several approaches for determining this most significant set. The greedy approach incrementally selects TFs ranked by the number of biomarker genes they target in the GRN, not taking the redundancy among target biomarker gene sets into account. In contrast, the LASSO-based convex optimization approach calculates a global weight for the TFs that incorporates the non-redundancy of their target biomarker gene sets, in addition to the FDRVIPER and FDRBIOMARKER likelihoods calculated above. The LASSO approach covers a higher total number of biomarker genes with fewer TFs than the greedy approach due to LASSO’s better control of redundancy (Supplementary Fig. 1). For instance, with 40 selected TFs, the LASSO approach covered approximately 15 more biomarker genes than the greedy approach.
Based on these observations, we examined the results of the LASSO approach in greater detail, which revealed that there was substantial variation in the weights of the TFs, especially due to differences in FDRVIPER and FDRBIOMARKER. Details of the top seven weighted TFs and all 132 TFs in the base nasal GRN are provided in Table 1 and Supplementary Table 2 respectively. In particular, ETV4 and PPARG were the only TFs that met three criteria: (1) highly weighted, (2) significantly differentially active in asthma (FDRVIPER ≤ 0.05), and (3) significantly regulating the asthma biomarker (FDRBIOMARKER ≤ 0.05) (Table 1). Just these two TFs regulated 24 (27%) of the asthma biomarker genes, including SERPINE237 and CDHR338, asthma-associated genes co-regulated by ETV4 and PPARG. Gene Ontology enrichment analysis, conducted using MSigDB)27,28, of the GRN subnetwork regulated by ETV4 and PPARG (Fig. 3) included terms highly relevant to disease processes in asthma, including response to corticosteroid (FDR = 2.94 × 10−5), regulation of immune system process (FDR = 8.92 × 10−4), and innate immune response (FDR = 5.89 × 10−3). Given these results, we focused on ETV4 and PPARG in our experimental validation efforts.
Subnetwork of the base nasal GRN consisting of the seven regulator TFs listed in Table 1, and their target asthma biomarker genes, denoted by squares and small filled circles respectively. The most significant regulators of the biomarker, ETV4 and PPARG, and the genes regulated exclusively by each of them are shown in green and pink respectively, and those regulated by both TFs are shown in yellow. Other TFs and target genes are shown in grey, with the exception of ESR1, which is itself regulated by PPARG. Figure made using CytoScape71.
Experimental validation of NeTFactor findings
To test the results from NeTFactor, we chose to investigate how knockdown of the regulatory TFs prioritized by NeTFactor (ETV4 and PPARG) in nasal epithelial cell line models would affect the production of inflammatory cytokines involved in asthma39,40,41,42. We chose to employ a nasal epithelial cell line to optimize context-specific validation, given the asthma biomarker is based on nasal gene expression, and we also used nasal RNAseq data as input for NeTFactor. Recognizing the limitations of cell lines as a model for in vivo inflammation, we first conducted a series of pilot experiments to assess the baseline responsiveness of a commercially available nasal epithelial cell line (HNEpC) to inflammatory stimulation. In response to the immunostimulant polyinosinic:polycytidylic acid (poly(I:C)), this cell line produced detectable amounts of IL6, IL8, TGFβ, CCL2, TSLP, and CCL17, with the highest levels noted for IL6 and IL8 (Supplementary Table 3). There was no detectable production of IL25 or IL33, and the cell line was less responsive to stimulation with lipopolysaccharide (LPS) or cytosine–phosphate–guanosine (CpG). With these pilot results as background, we designed our experimental validation of NeTFactor findings to include the measurement of IL8 and IL6 following stimulation with poly(I:C) in the nasal epithelial cell line model with and without siRNA knockdown of ETV and PPARG.
Figure 4 shows that both at baseline and in response to inflammatory stimulation with poly(I:C), the nasal epithelial cell line with ETV4 knocked down by siRNA (siETV4+) produced significantly smaller quantities of IL8 (Fig. 4A) and IL6 (Fig. 4B) compared to the negative siRNA control with intact ETV4. Similarly, the nasal epithelial cell line with PPARG knocked down by siRNA (siPPARG) yielded significantly lower IL8 (Fig. 4A) and IL6 (Fig. 4B) compared to negative siRNA control. These findings supported our expectation that levels of IL8 and IL6 would be repressed both at baseline and more significantly with knock down of ETV4 and PPARG based on NeTFactor’s results and our understanding of these cytokines in inflammation.
Experimental Validation of NeTFactor Predictions. Nasal epithelial cell lines with siRNA-mediated knockdowns of PPARG (siPPARG) and ETV4 mRNA (siETV), as well as negative siRNA control (Negative siRNA), were stimulated with Poly (I:C) and also left unstimulated. The concentrations of IL8 (Panel A) and IL6 (Panel B) at 24 hours are shown.
NeTFactor’s performance when the biomarker and GRN are derived from different gene expression datasets
Up to this point, we assessed NeTFactor’s performance when the GRN in its first step was derived from the same gene expression dataset from which the target biomarker was also identified. To assess NeTFactor’s performance when the GRN and biomarker are derived from different datasets, we derived a GRN from an independent nasal gene expression dataset from a cohort of asthmatic children and controls43 different from the primary dataset from which the 90-gene biomarker was identified. Application of NeTFactor’s steps to the GRN derived from the independent cohort yielded a ranked list of TF regulators that largely overlapped with the ranked list obtained from when the GRN and biomarker were derived from the same dataset (Supplementary Table 4). Specifically, the top 10 ranked TFs from the two applications of NeTFactor significantly overlapped (Fisher’s exact test p = 0.0019), indicating consistency between the top-ranked regulators identified by the framework when distinct GRNs are used. The top two ranked regulators from the primary application of NeTFactor, ETV4 and PPARG, were the second and third ranked regulators when NeTFactor was applied to the independently-derived GRN. Due to clinical differences between the primary3 and additional independent cohort43 used to derive the two GRNs, most prominently differences in age given the original cohort consisted only of adult subjects while the independent cohort included only children, we expectedly did not find the exact same ranking of regulators, although as a group, the top ranked regulators were significantly consistent.
Application of NeTFactor to a different disease biomarker
To assess NeTFactor’s ability to generalize to biomarkers of other diseases and/or phenotypes, we applied NeTFactor to identify TF regulators of a biomarker of peanut allergic reactions44. In this study, peanut allergic children underwent double-blind, placebo-controlled oral challenges to peanut where peripheral blood samples for whole blood transcriptome profiling were obtained during each challenge (i.e. peanut challenge and placebo challenge)44. All of the children reacted to peanut and none reacted to placebo44. In this scenario, peanut allergic reaction was the target phenotype, and the two classes were reaction (i.e. which occurred when peanut was given during peanut challenge), and no reaction (i.e. no reaction when placebo was given)44. In the primary study of this cohort44, a series of analyses on the subjects’ whole blood transcriptome profiles were performed that identified 26 key driver genes of peanut allergic reaction (Supplementary Table 4 of ref.44). These 26 key driver genes were considered the target biomarker for NeTFactor.
Table 2 shows the top-ranked TF regulators of the peanut allergy biomarker identified by NeTFactor (full ranking of TFs in Supplementary Table 5). Specifically, these TFs were identified using the LASSO procedure in NeTFactor’s last step and also satisfied the conditions of having both FDRVIPER and FDRBIOMARKER ≤ 0.05, supporting both their significant association with disease and biomarker relevance. These top-ranked regulators included STAT6, a TF known to have a central role in allergy through its modulation of Th2 cell differentiation, cell surface marker expression, and class-switching of immunoglobulins45, as well as, NFIL3, which is induced by STAT6 and regulates IgE production, an immunoglobulin central to allergy46. STAT3 plays a pivotal role in immune responses, regulating B cells and CD4+ and CD8+ T cells47, and whose dysregulation has been linked to aberrent IgE production and allergy48. SPI1 regulates follicular B cell development and germinal center responses49, and VDR is the receptor for vitamin D3, where studies have suggested that vitamin D is associated with allergy outcomes50,51.
Discussion
With rapid advances in genomic technology, several multi-gene expression-based biomarkers have been identified for diseases like asthma3, breast cancer52, stroke53 and Alzheimer’s disease54. Although some of these biomarkers are already used in clinical practice, such as MammaPrint and Oncotype DX for breast cancer prognosis4,18, their biological interpretation beyond examination of their individual constituent genes or enriched Gene Ontology terms or pathway is not commonly undertaken. In this paper, we have proposed the NeTFactor framework, which is designed to identify the most significant set of transcription factors (TFs) most likely to regulate such a biomarker. This is complementary to investigating the constituent genes individually or only identifying regulators associated with the target disease or phenotype using methods like MRA16. Based on NeTFactor’s findings when applied to our nasal brush-based biomarker of asthma3, we knocked down the identified regulator TFs in a nasal epithelial cell line model, finding that cytokine output was appropriately repressed as expected given our understanding of the role of these cytokines in inflammation and asthma. We also demonstrated that NeTFactor can be used to identify TF regulators of a peanut allergy biomarker44. Our findings demonstrate that NeTFactor can be successfully applied to identify TF regulators of multi-gene expression-based biomarkers, yielding valuable insights into disease-relevant biological processes and allowing us to gain more from biomarkers beyond their main role as classifiers or predictors.
NeTFactor requires as input a GRN, preferably a reliable context-specific one, the inference of which generally requires a sizeable gene expression data set. Our results show that the GRN may be derived from the same or different disease-relevant data set from which the biomarker is derived. However, in applications where such data may not be available, one can still use the NeTFactor algorithm by providing a generic network, such as the set of TF → target gene interactions in MSigDB27,28, as input. However, this may result in reduced sensitivity of the results, as NeTFactor will not have access to the biological context expected from the GRN. Another requirement of NeTFactor is a set of TFs that are needed for inferring the GRN, which we obtained from MSigDB in this study. If this set is not reliable or comprehensive, the resulting GRN and downstream analyses, especially the assessment of the constituent TFs’ disease-relevant activity (FDRVIPER) and enrichment of the biomarker genes among their targets (FDRBIOMARKER), may be adversely affected.
The final step of NeTFactor adopts a novel LASSO-based convex optimization approach to determine the most significant set of regulator TFs that maximizes the coverage of the biomarker genes. This approach has the benefit of collectively optimizing the relevant factors, namely FDRVIPER, FDRBIOMARKER, and non-redundancy among the target biomarker genes, for determining the most significant TF regulators of the biomarker. However, in certain cases, the solution of such an optimization problem may result in local optima, and consequently potentially false discoveries. More generally, due to the computational nature of NeTFactor, the possibility of false positive and negative results from the framework cannot be ruled out. The reliability of the results can be partially addressed by experimentally validating the results, as was done in this study, but that necessitates resources for this validation. These potential issues also indicate the possibility of developing alternative regulator prioritization methods that may improve the results of the LASSO approach used in this work.
Application of NeTFactor to a cohort of asthmatics and controls indicated that PPARG and ETV4 were the most likely regulators of an asthma biomarker. PPARG has been reported in some contexts to exert anti-inflammatory effects through the regulation of signaling in immune cells, including monocytes/macrophages, platelets, lymphocytes, and dendritic cells (DCs), as well as in epithelial, endothelial, and smooth muscle cells55,56,57. However, its role in asthma is more controversial with heterogeneous findings. While activation of PPARG has been associated with anti-inflammatory effects on airway58,59,60,61,62,63, PPARG signaling has more recently been shown to be critical for IL33–driven Th2 effector function in type-2 allergic airway responses, suggesting a contrasting pro-inflammatory role63. Furthermore, upregulation of PPARG in lung-resident CD11b+ DCs enhances migration to draining lymph nodes and Th2 priming capacity63.
Distinct from PPARG, far less is known about ETV4 (ETS variant 4) in the context of asthma. ETV4 belongs to the PEA3 (polyomavirus enhancer activator 3) subfamily of a larger E26 transformation-specific gene family of transcription factors63,64,65,66. ETV4 promotes morphogenesis of epithelial organs including lung during embryogenesis and plays a role in cell proliferation, growth, migration and apoptosis64. Our findings indicate a novel role of this transcription factor in asthma.
Our application of NeTFactor to a biomarker of peanut allergy44 revealed top-ranking TF regulators with established roles in immune regulation and allergy45,46,47,48,49,50,51. NeTFactor’s results suggest that their specific roles in peanut allergy in particular may be worth further study.
To conclude, we propose NeTFactor, an innovative framework that combines gene expression data and network analyses to identify the most significant set of transcription factor regulators of a multi-gene expression-based biomarker. Such regulators can yield valuable insights into regulatory mechanisms and disease-relevant biological processes related to their biomarkers, extending their utility beyond being mainly used as classifiers or predictors.
Materials and Methods
Primary study population and RNAseq data
In the primary study, we applied NeTFactor to the same development set of RNAseq-derived nasal gene expression data from 150 subjects with and without asthma that was used to identify the asthma biomarker in our earlier study3. The baseline characteristics of this study population are shown in Supplementary Table 1. This is one of the largest publicly available datasets of nasal gene expression from a well-characterized asthma cohort. The use of this rich data set for complementary analyses underlying our biomarker and NeTFactor is expected to improve our ability to identify regulatory network-derived explanations for the strong performance of our biomarker.
NeTFactor
Below, we describe the four main steps of NeTFactor, which finds the most significant set of TF regulators for a given biomarker, explained in terms of its main application to our asthma biomarker and the associated nasal RNAseq dataset mentioned above:
-
(1)
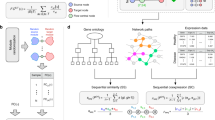
Reverse engineering a context-specific Gene Regulatory Network (GRN): We applied the well-established and publicly available ARACNE algorithm22,23,24 to the RNAseq dataset to infer a GRN specific to the nasal tissue. Briefly, ARACNE22,24 uses mutual information67, a symmetric information theoretic similarity measure that accounts for possible non-linear relationship(s) between two entities (here, genes). Given a \(sample\times gene\) expression matrix, ARACNE calculates mutual information between all pairs of genes (columns of the matrix), representing an initial fully connected network. Next, augmented with a list of putative TFs provided, ARACNE uses the data processing inequality (DPI)67 to remove interactions whose constituent information is sufficiently captured by the rest of the interactions, which generates the resulting GRN consisting of TF → target gene interactions24. Specifically, for all the cliques consisting of three gene \(\leftrightarrow \,\)gene edges, where some of the genes represent the putative TFs, ARACNE removes the edge with the lowest value of mutual information in accordance with DPI. The only exceptions to this process are the cases where this removal will retain a non-TF ↔ non-TF edge, which is not considered a transcriptional interaction, and is removed instead. Repeating this process throughout the original mutual information matrix generates the resulting GRN consisting of TF → target gene interactions.
The Adaptive Partitioning (AP) version of ARACNE23 improves its computational efficiency and robustness using multiple bootstraps of the gene expression data and averaging the resulting network to create the final GRN. We applied this version with the default number of bootstraps (100) in the publicly available implementation23 to create the nasal GRN for further analysis, but also tested other numbers of bootstraps (e.g., 1000) to test the GRN’s dependence on this parameter’s value. 221 putative TFs for running ARACNE were obtained from the Molecular Signature Database (MSigDB)27,28 version 5.1 (accessed June 3rd, 2016).
-
(2)
Inferring disease-related activity of TFs in the GRN: To enhance the functional relevance of candidate regulator TFs, NeTFactor requires such TFs to be differentially active under disease conditions. For this, we applied the VIPER algorithm30 to the nasal GRN, along with the asthma/no-asthma status of the expression profiles used to derive the GRN. Specifically, VIPER was run with the sample-based permutation option to build the null hypothesis. The output of VIPER consists of a normalized enrichment score, which is positive and negative for TFs that are more active in asthma and no-asthma respectively. VIPER also produces the associated False Discovery Rate values (FDRVIPER), which were used in NeTFactor as a measure of the disease-related activity of the TFs constituting the GRN. This is also the goal of methods like MRA16 and its variants17.
-
(3)
Calculating the likelihood of a TF regulating the biomarker gene set: Even if a TF is determined to be active in a disease in the above step, it may not necessarily regulate the target biomarker, as it may be regulating other genes. Therefore, to determine the likelihood of a TF regulating the biomarker gene set, we used the Fisher’s exact test32 to assess the statistical significance of the number of genes in the overlapping the set of genes regulated by the TF in the GRN (its regulon) and the biomarker. We then used the Benjamini-Hochberg procedure33 to correct for multiple hypothesis testing and used the resulting FDRBIOMARKER as the final likelihood of the TF regulating the biomarker gene set for further analysis.
-
(4)
Using convex optimization to find the most significant set of regulators: Since a gene in the biomarker may be regulated by multiple TFs, there may be redundancy (overlap) among the targets of the regulator TFs identified. To enhance NeTFactor’s ability to identify a set of regulators that reveal complementary aspects of disease biology, this step of the framework also takes into account the sets of biomarker genes each regulator targets. A possible greedy approach to this task is to incrementally select TFs by the number of biomarker genes they target in the corresponding GRN, until a certain fraction, say 80%, of all the biomarker genes has been covered. However, this approach does not tackle the redundancy issue, as several of the selected regulators may still regulate highly overlapping sets of biomarker genes. Therefore, NeTFactor adopts a convex optimization method for prioritizing candidate regulators based on the GRN inferred in Step 1 and the likelihoods calculated in Steps 2 and 3. Mathematically, this optimization problem can be formulated as
where \({\rm{A}}\) is the (#TFs in GRN)X(#biomarker genes) matrix, with the A(i, j) entry representing the likelihood that TF i is active and regulating gene j in the biomarker set, which is defined as (1 − FDRVIPER) × (1 − FDRBIOMARKER) if j is a target of i in the GRN, and 0 otherwise.
The purpose of solving the above optimization problem stated is to find the value of the vector x, which contains one entry for each TF, such that there are as few non-zero entries in x as possible (purpose of the L0 norm) and each gene in the biomarker would be targeted/covered by at least one TF (denoted by the \({\rm{A}}x\,\ge 1\) condition) in the GRN. However, since this problem is known to be computationally intractable68, we relaxed it by using the L1 norm, thus changing the optimization problem to
This problem is a derivative of the well-known LASSO algorithm, and thus can be solved using standard methods69 and packages like CVXR70. The decreasing magnitudes of the values constituting \(x\) (LASSO weights) that are obtained by solving the above problem were used to rank the TFs in terms of their likelihood of non-redundantly regulating the biomarker.
Finally, to facilitate deeper examination of the top-ranking regulators, such as experimental validation in this study, we were most interested in determining the most significant subset of these regulator TFs that maximizes coverage of the biomarker. For this, we only selected TFs that were both significantly relevant to the disease (FDRVIPER \(\le \,\) 0.05) and the biomarker (FDRBIOMARKER ≤ 0.05) for further study.
Testing the robustness of NeTFactor’s results to the choice of the GRN inference algorithm
To test this aspect, we applied NeTFactor to the nasal RNAseq data by replacing the constituent ARACNE algorithm with GENIE329, another well-established algorithm to infer GRNs. We used GENIE3’s publicly available R implementation (https://bioconductor.org/packages/release/bioc/html/GENIE3.html), setting the number of potential regulators of each gene, a required input parameter, to the default value of the square root of the total number of genes in the nasal RNAseq dataset. To compare the results obtained with the ARACNE-based ones, we selected the same number of top-weighted edges in the inferred GENIE3 network as the original nasal ARACNE GRN.
Experimental validation
We used a human nasal epithelial primary cell line (PromoCell (Heidelberg, Germany)) at second passage for our experimental work, based on the rationale that a nasal epithelial cell line is from tissue closest in nature to the nasal brush samples that yielded the nasal RNAseq data used to develop the asthma biomarker. The cell line was grown according to manufacturer instructions in the presence of Primocin (Millipore Sigma, St Louis, MO, USA) at 1:500 dilution in Opti-MEM™ Reduced Serum Medium (Thermo Fisher Scientific, Waltham, MA, USA). Cells were cultured to a confluence of 70–90% in a 24-well plate. For pilot studies to determine optimal stimulant concentration, stimulations were done with lipopolysaccharide (LPS) (Millipore Sigma, St Louis, MO, USA) at 1 mg/mL, polyinosinic:polycytidylic acid (Poly (I:C)) (InvivoGen, San Diego, CA, USA) at 20 and 50 mg/mL, and immunostimulatory cytosine–phosphate–guanosine (CpG) (InvivoGen, San Diego, CA, USA) at 2.5 mM. Supernatants were collected at 6, 24, 48 and 72 hours and stored frozen at −80 °C for further experiments.
siRNA-mediated knockdown of PPARG and ETV4 mRNA was performed according to the protocol and supplies from ThermoFisher Scientific (RNAi Handbook, ThermoFisher Scientific, Waltham, MA, USA; thermofisher.com/RNAi). Cells were seeded at the sixth passage on a 24-well plate at ~2 × 104 cell per well. At 48 hours, cell media were refreshed. siRNA for PPARG and ETV4, as well as the corresponding negative siRNA controls were each mixed with Lipofectamine RNAiMAX reagent and added to the well at 5 pmol/0.5 mL medium/well. At 72 hours following addition of the transfection mix, cells were stimulated with Poly(I:C) or media control for 24 hours. Cytokine and chemokine levels in supernatants were then measured using X Multiplex Human Cytokine/chemokine assay kits (Millipore Sigma, St. Louis, MO, USA).
Testing the generalizability of NeTFactor to scenarios when the GRN and biomarker are derived from different data sets
To assess the generalizability of NetTFactor’s performance to instances when the GRN and biomarker are derived from different data sets, we also applied NeTFactor to a GRN derived from an independent dataset from a distinct cohort of children with asthma and controls43. This independent nasal transcriptome dataset was generated from 225 asthmatics and controls recruited separately from the original dataset. All subjects were recruited as part of an IRB-approved study at the Mount Sinai Health System, New York, NY and provided written informed consent.
Specifically, we constructed a new GRN using nasal transcriptome data from the independent cohort by applying ARACNE with 100 bootstraps and the seed set of 132 MSigDB TFs–the same settings used to derive the original asthma GRN. We then applied the same four steps of NetFactor as had been applied to the original GRN to this independently-derived GRN to identify potential regulators of the 90-gene asthma biomarker.
Application of NeTFactor to a different disease biomarker
To assess NeTFactor’s ability to generalize to biomarkers of other diseases and/or phenotypes, we applied NeTFactor to identify TF regulators of a biomarker of peanut allergic reactions44. The data for GRN construction included whole blood transcriptome profiles from 40 peanut allergic children undergoing double-blind, placebo-controlled oral challenges to peanut. Longitudinal peripheral blood samples for whole blood transcriptome profiling were obtained during each challenge44. Here, peanut allergic reaction was the target phenotype, and the two classes were reaction (i.e. captured by samples obtained during peanut challenge), and no reaction (i.e. captured by samples obtained during placebo challenge)44. In the primary study of this cohort44, 26 key driver genes of peanut allergic reaction were identified (Supplementary Table 4 of this ref.44) and were considered the biomarker for NeTFactor.
We applied the NeTFactor framework to the gene expression data from the peanut allergy cohort and the biomarker of peanut allergic reactions44. For GRN inference using ARACNE, we used 100 bootstraps and the seed set of 221 MSigDB TFs, the same settings used to derive the original asthma GRN. The other steps of NeTFactor were applied exactly as in the original asthma case study, with the exception that VIPER was run with the gene-based permutation option to build the null hypothesis due to the fact that each subject had whole blood transcriptome profiles from longitudinal samples obtained during both peanut and placebo challenges, and samples were thus not independent within or between the classes.
Ethics approval and consent to participate
The institutional review boards of Brigham & Women’s Hospital and the Icahn School of Medicine at Mount Sinai approved the study protocols. Written informed consent was obtained from all subjects and all research was performed in accordance with relevant guidelines and regulations.
Software Availability
The code for NeTFactor implementation is available at https://github.com/GauravPandeyLab/NeTFactor.
Data Availability
RNAseq data from the asthma cohort that yielded the primary nasal gene expression data set used in this study are available at https://www.synapse.org/#!Synapse:syn9878922/files/ (https://doi.org/10.7303/syn9878922). The RNAseq data for the peanut allergy cohort can be found at https://www.synapse.org/#!Synapse:syn10212437/files/ (https://doi.org/10.7303/syn10212437).
References
Ozsolak, F. & Milos, P. M. RNA sequencing: advances, challenges and opportunities. Nat Rev Genet 12, 87–98 (2011).
Libbrecht, M. W. & Noble, W. S. Machine learning applications in genetics and genomics. Nat Rev Genet 16, 321–332 (2015).
Pandey, G. et al. A Nasal Brush-based Classifier of Asthma Identified by Machine Learning Analysis of Nasal RNA Sequence Data. Scientific Reports 8, 8826 (2018).
van ’t Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
van de Vijver, M. J. et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347, 1999–2009 (2002).
Badal, B. et al. Transcriptional dissection of melanoma identifies a high-risk subtype underlying TP53 family genes and epigenome deregulation. JCI Insight 2 (2017).
Rykunov, D. et al. A new molecular signature method for prediction of driver cancer pathways from transcriptional data. Nucleic Acids Res 44, e110 (2016).
Schlitt, T. & Brazma, A. Current approaches to gene regulatory network modelling. BMC Bioinformatics 8, S9 (2007).
Dana, H. et al. Molecular Mechanisms and Biological Functions of siRNA. Int J Biomed Sci 13, 48–57 (2017).
de la Fuente, A. From ‘differential expression’ to ‘differential networking’ - identification of dysfunctional regulatory networks in diseases. Trends Genet 26, 326–333 (2010).
Hasty, J., McMillen, D., Isaacs, F. & Collins, J. J. Computational studies of gene regulatory networks: in numero molecular biology. Nat Rev Genet 2, 268–279 (2001).
Vidal, M., Cusick, M. E. & Barabasi, A. L. Interactome networks and human disease. Cell 144, 986–998 (2011).
Huang, J. K. et al. Systematic Evaluation of Molecular Networks for Discovery of Disease Genes. Cell Systems 6, 484–495.e485 (2018).
Wang, X., Gulbahce, N. & Yu, H. Network-based methods for human disease gene prediction. Briefings in Functional Genomics 10, 280–293 (2011).
Barabási, A.-L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nature Reviews Genetics 12, 56 (2010).
Lim, W. K., Lyashenko, E. & Califano, A. Master regulators used as breast cancer metastasis classifier. Pac Symp Biocomput, 504–515 (2009).
Califano, A. & Alvarez, M. J. The recurrent architecture of tumour initiation, progression and drug sensitivity. Nat Rev Cancer 17, 116–130 (2017).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351, 2817–2826 (2004).
Bunyavanich, S. & Schadt, E. E. Systems biology of asthma and allergic diseases: A multiscale approach. The Journal of allergy and clinical immunology (2014).
Kitano, H. Computational systems biology. Nature 420, 206–210 (2002).
Kitano, H. Systems biology: a brief overview. Science 295, 1662–1664 (2002).
Margolin, A. A. et al. ARACNE: An Algorithm for the Reconstruction of Gene Regulatory Networks in a Mammalian Cellular Context. BMC Bioinformatics 7, S7 (2006).
Lachmann, A., Giorgi, F. M., Lopez, G. & Califano, A. ARACNe-AP: gene network reverse engineering through adaptive partitioning inference of mutual information. Bioinformatics 32, 2233–2235 (2016).
Margolin, A. A. et al. Reverse engineering cellular networks. Nature Protocols 1, 662 (2006).
Yii, A. C. A. et al. Precision medicine in united airways disease: A “treatable traits” approach. Allergy (2018).
Fazlollahi, M. et al. The nasal microbiome in asthma. The Journal of allergy and clinical immunology (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Huynh-Thu, V. A., Irrthum, A., Wehenkel, L. & Geurts, P. Inferring regulatory networks from expression data using tree-based methods. PLoS One 5 (2010).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat Genet 48, 838–847 (2016).
Wang, J., Li-Hong, X., Cheng, W., Wang, Z. & Zhang, W. Effect of heat shock factor 1 on airway hyperresponsiveness and airway inflammation in mice with allergic asthma. Chinese Journal of Contemporary Pediatrics 19, 222–228 (2017).
Sprent, P. Fisher Exact Test. In: International Encyclopedia of Statistical Science (ed^(eds) (2011).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B (Methodological) 57, 289–300 (1995).
Bhakta, N. R. et al. IFN-stimulated Gene Expression, Type 2 Inflammation, and Endoplasmic Reticulum Stress in Asthma. Am J Respir Crit Care Med 197, 313–324 (2018).
Martino, M. B. et al. The ER stress transducer IRE1beta is required for airway epithelial mucin production. Mucosal Immunol 6, 639–654 (2013).
Liu, Y. P. et al. Association of ORMDL3 with rhinovirus-induced endoplasmic reticulum stress and type I Interferon responses in human leucocytes. Clin Exp Allergy 47, 371–382 (2017).
Himes, B. E. et al. Association of SERPINE2 with asthma. Chest 140, 667–674 (2011).
Bochkov, Y. A. et al. Cadherin-related family member 3, a childhood asthma susceptibility gene product, mediates rhinovirus C binding and replication. Proc Natl Acad Sci USA 112, 5485–5490 (2015).
Gandhi, V. D. & Vliagoftis, H. Airway epithelium interactions with aeroallergens: role of secreted cytokines and chemokines in innate immunity. Front Immunol 6, 147 (2015).
Divekar, R. & Kita, H. Recent advances in epithelium-derived cytokines (IL-33, IL-25, and thymic stromal lymphopoietin) and allergic inflammation. Curr Opin Allergy Clin Immunol 15, 98–103 (2015).
Kumar, R. K., Herbert, C. & Foster, P. S. Expression of growth factors by airway epithelial cells in a model of chronic asthma: regulation and relationship to subepithelial fibrosis. Clin Exp Allergy 34, 567–575 (2004).
Wang, Y., Bai, C., Li, K., Adler, K. B. & Wang, X. Role of airway epithelial cells in development of asthma and allergic rhinitis. Respir Med 102, 949–955 (2008).
Do, A. N. et al. Network analysis reveals causal key driver genes of severe asthma in children. Journal of Allergy and Clinical Immunology 143, AB186 (2019).
Watson, C. T. et al. Integrative transcriptomic analysis reveals key drivers of acute peanut allergic reactions. Nat Commun 8, 1943 (2017).
Junttila, I. S. Tuning the Cytokine Responses: An Update on Interleukin (IL)-4 and IL-13 Receptor Complexes. Front Immunol 9, 888 (2018).
Rothman, P. B. The transcriptional regulator NFIL3 controls IgE production. Trans Am Clin Climatol Assoc 121, 156–171; discussion 171 (2010).
Deenick, E. K., Pelham, S. J., Kane, A. & Ma, C. S. Signal Transducer and Activator of Transcription 3 Control of Human T and B Cell Responses. Front Immunol 9, 168 (2018).
Lyons, J. J. et al. ERBIN deficiency links STAT3 and TGF-beta pathway defects with atopy in humans. J Exp Med 214, 669–680 (2017).
Wang, H. et al. Transcription factors IRF8 and PU.1 are required for follicular B cell development and BCL6-driven germinal center responses. Proc Natl Acad Sci USA 116, 9511–9520 (2019).
Yepes-Nunez, J. J. et al. Vitamin D supplementation in primary allergy prevention: Systematic review of randomized and non-randomized studies. Allergy 73, 37–49 (2018).
Bunyavanich, S. et al. Prenatal, perinatal, and childhood vitamin D exposure and their association with childhood allergic rhinitis and allergic sensitization. The Journal of allergy and clinical immunology 137, 1063–1070 e1062 (2016).
Carlson, J. J. & Roth, J. A. The impact of the Oncotype Dx breast cancer assay in clinical practice: a systematic review and meta-analysis. Breast Cancer Res Treat 141, 13–22 (2013).
Adamski, M. G. et al. Expression profile based gene clusters for ischemic stroke detection. Genomics 104, 163–169 (2014).
Arisi, I. et al. Gene expression biomarkers in the brain of a mouse model for Alzheimer’s disease: mining of microarray data by logic classification and feature selection. J Alzheimers Dis 24, 721–738 (2011).
Kim, J.-H., Song, J. & Park, K. W. The multifaceted factor peroxisome proliferator-activated receptor γ (PPARγ) in metabolism, immunity, and cancer. Arch Pharm Res 38, 302–312 (2015).
Croasdell, A. et al. PPARγ and the Innate Immune System Mediate the Resolution of Inflammation. PPAR Res 2015, 549691 (2015).
da Rocha Junior, L. F. et al. PPARγ Agonists in Adaptive Immunity: What Do Immune Disorders and Their Models Have to Tell Us? PPAR Res 2013, 519724 (2013).
Wang, A. C., Dai, X., Luu, B. & Conrad, D. J. Peroxisome proliferator-activated receptor-gamma regulates airway epithelial cell activation. Am J Respir Cell Mol Biol 24, 688–693 (2001).
Trifilieff, A. et al. PPAR-alpha and -gamma but not -delta agonists inhibit airway inflammation in a murine model of asthma: in vitro evidence for an NF-kappaB-independent effect. Br J Pharmacol 139, 163–171 (2003).
Woerly, G. et al. Peroxisome proliferator-activated receptors alpha and gamma down-regulate allergic inflammation and eosinophil activation. J Exp Med 198, 411–421 (2003).
Zhao, Y. et al. Rosiglitazone, a peroxisome proliferator-activated receptor-γ agonist, attenuates airway inflammation by inhibiting the proliferation of effector T cells in a murine model of neutrophilic asthma. Immunol Lett 157, 9–15 (2014).
Lee, H. Y. et al. Effect of intranasal rosiglitazone on airway inflammation and remodeling in a murine model of chronic asthma. Korean J Intern Med 31, 89–97 (2016).
Nobs, S. P. et al. PPARγ in dendritic cells and T cells drives pathogenic type-2 effector responses in lung inflammation. J Exp Med 214, 3015–3035 (2017).
Oh, S., Shin, S. & Janknecht, R. ETV1, 4 and 5: an oncogenic subfamily of ETS transcription factors. Biochim Biophys Acta 1826, 1–12 (2012).
Tyagi, N. et al. ETV4 Facilitates Cell-Cycle Progression in Pancreatic Cells through Transcriptional Regulation of Cyclin D1. Mol Cancer Res 16, 187–196 (2018).
Zeng, S. et al. ETV4 collaborates with Wnt/β-catenin signaling to alter cell cycle activity and promote tumor aggressiveness in gastrointestinal stromal tumor. Oncotarget 8, 114195–114209 (2017).
Cover, T. M. & Thomas, J. A. Elements of information theory. John Wiley & Sons (2012).
Jin, J., Gu, Y.-T. & Mei, S.-L. An Introduction to Compressive Sampling and Its Applications. Journal of electronics information & technology 32, 470–475 (2010).
Tibshirani, R. Regression shrinkage and selection via the lasso: a retrospective. J R Stat Soc Series B Stat Methodol 73, 273–282 (2011).
Fu, A., Narasimhan, B. & Boyd, S. CVXR: An R Package for Disciplined Convex Optimization. arXiv preprint arXiv:171107582 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504 (2003).
Acknowledgements
This study was enabled in part by computational resources provided by Scientific Computing at the Icahn School of Medicine at Mount Sinai. We thank Dr. Madhan Masilamani for sharing his perspectives on the design of cell-line experiments, and Dr. Avner Schlessinger for his advice on the preparation of this manuscript. This work was supported by NIH grants R01AI118833 and K08AI093538 to SB, and R01GM114434 to GP, a pilot project grant from the Mount Sinai Mindich Child Health and Development Institute to SB and GP, and an IBM Faculty Award to GP. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
G.P., S.B., G.S. and M.E.A. conceived the study. G.P. and S.B. supervised the work. M.E.A. and Y.C. executed the computational components of the study, while A.G. and G.G. executed the experimental aims. All the authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahsen, M.E., Chun, Y., Grishin, A. et al. NeTFactor, a framework for identifying transcriptional regulators of gene expression-based biomarkers. Sci Rep 9, 12970 (2019). https://doi.org/10.1038/s41598-019-49498-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49498-y
This article is cited by
-
Pancancer network analysis reveals key master regulators for cancer invasiveness
Journal of Translational Medicine (2023)
-
The regulatory role of AP-2β in monoaminergic neurotransmitter systems: insights on its signalling pathway, linked disorders and theragnostic potential
Cell & Bioscience (2022)
-
Expression quantitative trait loci for ETV4 and MEOX1 are associated with adult asthma in Japanese populations
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.