Abstract
Complexes of closely related species provide key insights into the rapid and independent evolution of adaptive traits. Here, we described and studied Anopheles fontenillei sp.n., a new species in the Anopheles gambiae complex that we recently discovered in the forested areas of Gabon, Central Africa. Our analysis placed the new taxon in the phylogenetic tree of the An. gambiae complex, revealing important introgression events with other members of the complex. Particularly, we detected recent introgression, with Anopheles gambiae and Anopheles coluzzii, of genes directly involved in vectorial capacity. Moreover, genome analysis of the new species allowed us to clarify the evolutionary history of the 3La inversion. Overall, An. fontenillei sp.n. analysis improved our understanding of the relationship between species within the An. gambiae complex, and provided insight into the evolution of vectorial capacity traits that are relevant for the successful control of malaria in Africa.
Similar content being viewed by others
Introduction
Species at earlier speciation stages provide unique insights into the evolutionary forces involved in the origin of new species before signal blurring by demographic and selective processes. However, when closer to the first signals of divergence, it is harder to define the species concept, and to predict whether this process will lead to speciation1. Complexes of species and closely related taxa where the species boundaries are uncertain offer a precious opportunity to study the “speciation continuum”1,2. Unfortunately, the incomplete reproductive isolation cannot fully prevent introgression between taxa, thus hindering the true phylogenetic relationships3. On the other hand, genetic exchanges in backcrossed hybrids can favour adaptation4. Indeed, advantageous alleles can be selected in one species and introgressed in another, thus favouring, for instance, range expansion5, altitudinal adaptation6, and insecticide resistance7.
Most of the major malaria vectors worldwide belong to species complexes that include also other non-vector species8, providing a compelling opportunity to understand the rapid and independent evolution of their vectorial capacity9,10. Indeed, malaria mosquitoes exhibit wide ecological plasticity, preference for feeding on humans, and large population size. Conversely, non-vector species display narrower geographical range, zoophilic host preference, and strong seasonal dependence or reduced population size11. In Africa, three of the six major malaria vectors belong to the Anopheles gambiae complex: Anopheles gambiae, Anopheles coluzzii, and Anopheles arabiensis12. This complex includes eight cryptic species13,14,15 that differ in many ecological aspects, particularly in host feeding preference, breeding sites, feeding behaviour, and role in malaria transmission13,16. Most of the An. gambiae species live in natural habitats, and do not have any or only a secondary role in malaria transmission. Indeed, adaptation to anthropogenic habitats, and therefore a role in human malaria transmission, is an exception rather than the rule within this complex16. The An. gambiae complex is an example of speciation with gene flow where species exhibit extensive genomic introgression (an indication of permeable gene flow barriers among them3,10) that is sustained by heterogenic patterns of reproductive isolation17. Consequently, pervasive introgression has hindered the elucidation of the correct phylogenetic relationships18. Moreover, gene exchanges between species in the complex have modulated their local adaptation capacity. For instance, An. arabiensis ability to live in desiccating environments has been conferred by introgression of the 2La inversion from An. gambiae/An. coluzzii10,19. An. coluzzii has developed resistance to insecticide treatments due introgression of the kdr mutation from An. gambiae7. Introgression of genes linked to insecticide resistance has repeatedly occurred in the last decades20,21. Thus, introgression has accelerated local adaptation and range expansion within the complex.
Complexes of closely related species also offer a compelling opportunity to study locally adapted alleles. Comparative genomic analysis of recent species radiations allowed unravelling the genetic basis of the traits involved in their ecological, behavioural and genetic divergence22. In Anopheles, these comparative studies have contributed to elucidate some traits involved in vectorial capacity that in turn could be used to improve vector control strategies9. For instance, comparison of the antennal transcriptome profiles of An. gambiae and An. quadriannulatus has provided genomic insights into host preference evolution towards humans23. Moreover, comparison of genome-wide data of one fresh water (An. gambiae) and one salt water (Anopheles melas) species allowed identifying genomic regions involved in salinity tolerance24. These data can be used to develop alternative malaria control strategies by targeting genes involved in reducing vectorial capacity traits, such as human bite rates or local adaptation10,23,24. Therefore, understanding the origin and the mechanisms underlying vectorial capacity within the An. gambiae complex is crucial for the successful control of malaria in Africa16,25.
In the present study, we carried out an exploratory survey at La Lopé National Park (Gabon) in 2014. In this survey, we discovered mosquitoes that we initially morphologically identified as An. gambiae. However, additional bio-ecological, behavioural, taxonomic, cytogenetic, and preliminary molecular analyses suggested the existence of a new taxon in the An. gambiae complex. Then, genome-wide phylogenetic analysis placed this new taxon in the An. gambiae complex as a sister species of Anopheles bwambae, and in the same clade as An. quadriannulatus, An. arabiensis, and An. melas. Comparative genomic analysis indicated the existence of recent introgression events between the potential new species and An. gambiae/An. coluzzii. These events concerned genes involved in detoxification, desiccation, and olfactory perception functions that are directly linked to local adaptation and host preference. These analyses also elucidated the evolutionary history of the 3La inversion within the complex. Overall, the discovery of this new taxon demonstrates the importance of new species for understanding the evolutionary relationships among the An. gambiae complex species, with potential implications for elucidating vectorial capacity traits and consequently malaria control.
Results
In this study, all specimens that were initially morphologically identified as An. gambiae in the 2014 exploratory survey belonged to a unknown taxon hereafter called Anopheles fontenillei n.sp. This species is dedicated to our colleague Didier Fontenille, medical entomologist who greatly contributed to the study of mosquitoes and the development of medical entomology in Africa.
Bio-ecology of Anopheles fontenillei sp.n
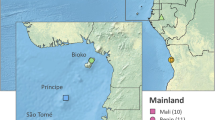
We prospected 22 sites in the National Park of La Lopé in Gabon: 17 sites in the park and 5 sites in the village of La Lopé, 10–15 km away from the park sites. In total, we collected more than 1,500 mosquitoes, belonging to 13 different species. Among these samples, 45 adults and 2 larvae that we morphologically identified as An. gambiae presented an unexpected DNA band in the PCR assay used for species identification26. In Gabon, only three species of the An. gambiae complex have been recorded, and all of them can be identified based on specific PCR bands27,28. The individuals of the unknown species came from six natural sites across the park, away from any human activity or presence (Fig. 1, Table S1). Specifically, these sites were situated at the edge of forest patches and close to natural marshes frequented by wild animals (e.g., African forest buffalos and other ungulates). In the same breeding sites, we collected also An. maculipalpis that breeds in sympatry with An. gambiae29 in sun-exposed, low oxygen, and generally stagnant water. This larval habitat typology is very similar to that of An. gambiae, An. coluzzii, and An. arabiensis16, but different from that of the other complex members, such as An. merus and An. melas (mangrove swamps) or An. bwambae (hot thermal springs). This places the new taxon in the fresh-water group of species within the An. gambiae complex16. Although we did not capture any blood-fed mosquito, we assumed a preference for feeding on animals (zoophily) due to the lack of human hosts in these sylvatic sites. However, as mosquitoes were sampled using BG® traps baited with BG-lure (a source of CO2)30 and Human Landing Catches –HLC– (Fig. 1B), the new species may also feed on humans. Our collections in La Lopé village revealed the presence of two other members of the complex (An. gambiae and An. coluzzii), while. We did not find any An. fontenillei specimen in the village (HLC and larva prospections).
Overview of An. fontenillei bionomic characteristics. (A) Red points indicate the mosquito collection sites in the National Park of La Lopé where An. fontenillei specimens were captured. The bottom right corner shows a photograph of the breeding site where one larva of the new species was found. (B) Mean number (black dots) of An. fontenillei collected using human landing catch (human, red) vs. BG traps (trap, green) in the six sylvatic sites (A) Table S1). (C) Morphological features of An. fontenillei: dorsal view of the wing, maxillary palpus and hindleg with femur, tibia and tarsomeres. (D) Images of polytene chromosomes from ovarian nurse cells of An. fontenillei obtained with a contrast-phase microscope (specimen n. 23). Chromosomal arm karyotypes are indicated following the classical nomenclature34. The paracentric inversions are designed by lines (red and blue) above the 3 R(b) and 3 L(a) arms, respectively.
Anopheles (Cellia) fontenillei sp.n
We preserved five An. fontenillei specimens for taxonomic purposes (Table S1). The holotype female number LOP3 was collected at La Lopé National Park, site LOP 43. The four females paratypes LOP473, LOP781, LOP1 and LOP2, were also collected at La Lopé National Park, sites LOP 40, LOP 40, LOP 40 and LOP 16, respectively. All the specimens are deposited in the IRD museum collections in Montpellier, France. Overall, An. fontenillei presented the classical morphotype of species within the An. gambiae complex15,31,32: three white-scaled bands on the maxillary palpus, irregularly shaped speckling on femora and tibiae, and a pale interruption in the third main dark area of wing vein R1 (Fig. 1C) (for further taxonomic details see Text S1). However, small differences can be observed among species of the complex. Indeed, we found that the maxillary palpus exhibited a large white-scaled band covering completely palpomere 5 and part of palpomere 4 (Fig. 1C). This morphological trait has been also described in An. bwambae33, although, it is not a discriminant trait with regard to the other members of the complex.
Cytogenetic analysis
Fixed chromosomal inversions have been used to differentiate species within the An. gambiae complex since the 1970’s34. To confirm the species status and its phylogenetic relationships within the An. gambiae complex, we then collected 270 sylvatic Anopheles specimens for cytogenetic analyses at the LOP 39 site (Fig. 1A). Among the 40 mosquitoes that belonged to the An. gambiae complex (morphological analysis), only four specimens displayed the new PCR band associated with the new species (see below). Three of these specimens survived to attain the correct stage (half-gravid) to observe polytene chromosomes. According to the classical nomenclature for chromosomal rearrangements in the An. gambiae complex34, all three specimens exhibited the X chromosome and the 2 L arm standard arrangements, and the inversions 3Rb and 3La were fixed. Conversely, the 2Rl inversion was polymorphic: present in one and absent in the other two specimens (Figs 1D, S1). We then used the available molecular karyotyping test35 to confirm the presence of the 2La inversion in five additional specimens35. All specimens revealed a PCR-band consistent with the 2La standard arrangement, confirming the cytogenetic karyotype. Our data indicate that An. fontenillei karyotype is similar to that of An. bwambae33. However, the 3Rb inversion might be fixed in the new taxon, while it is polymorphic in An. bwambae. Further cytogenetic studies in a larger number of individuals will be necessary to confirm the inversion polymorphisms of this species.
Phylogenetic analysis of the Anopheles gambiae complex including An. fontenillei sp.n
We first carried out preliminary phylogenetic analysis using 16 specimens to obtain sequences for the nuclear ITS2 and IGS non-coding spacers and the mitochondrial ND5 and COI regions, which are routinely used for Anopheles phylogenetic studies36. We could amplify and sequence the ITS2 and ND5 regions in nine and five specimens, respectively (Table S1). All regions exhibited low diversity with a unique haplotype, except for the COI gene that displayed five haplotypes. The phylogenetic trees showed that An. fontenillei sequences always clustered with An. bwambae within a monophyletic clade (Fig. S2), corroborating the cytogenetic results, but in contrast with the ecological observations. Among the four sequenced regions, ITS2 and ND5 revealed differences between An. fontenillei and An. bwambae (Fig. S2). These results are in agreement with previous studies showing that most of the classical molecular markers do not discriminate among species in the complex due to their extensive introgression10.
Overall, the new taxon revealed important similarities with An. bwambae, a thermal spring breeding species from a forested area of Uganda (Semliki valley), at the taxonomical (large band in the palpomeres 4 and 5), cytogenetic (chromosomal inversions), and molecular (sequence divergence) levels. On the other hand, ecological (fresh-water marshes vs thermal springs) and geographical (allopatric distribution: Gabon vs Uganda) results clearly discriminated between An. fontenillei and An. bwambae. Thus, additional studies are needed to determine the true phylogenetic place of An. fontenillei within the An. gambiae complex.
We then performed a genome-wide analysis to accurately locate the new species in the phylogenetic tree of the An. gambiae complex. According to previous studies10, we considered that the true species tree is mainly observed in the X chromosome. Hence, we initially focused on the X chromosome, and made a genome assembly of one An. fontenillei individual sequenced at high coverage (~112X) (Table S2). This assembly was nearly complete (96% of BUSCO complete genes), but made of highly fragmented contigs (N50 = 21 kb) (see Methods, Table S3B and Table S3C). We then added our An. fontenillei assembly and the highest coverage publicly available An. bwambae individual (see Methods) to the available multiple alignment file (MAF) based on six described An. gambiae complex species10. We built maximum likelihood (ML) phylogenetic trees for each non-overlapping 50 kb windows (see Methods, Table S4). Following this approach, the relationship among species observed in the X chromosome (Fig. 2) showed that similarly to previous studies, the relative position of the An. merus and An. coluzzii-An. gambiae node was not clearly determined due to incomplete linage sorting10. We found that An. fontenillei appeared as the sister species of An. bwambae in 83% of the trees (264 out of 319). However, there was some ambiguity in determining the closest taxon of the clade. Indeed, the clade branched with An. quadriannulatus in 78 of 319 windows (Fig. 2), and with An. arabiensis in 59 of 319 windows (Fig. S3). If the X chromosome shows the true species tree, we could hypothesize that An. quadriannulatus or An. arabiensis shares a common ancestor with An. fontenillei and An. bwambae.
Most common phylogenetic tree topology for the analysed X chromosome sequences of different An. gambiae complex species. 78 windows in the X chromosome show this tree topology with a weak disagreement in the most basal branch. Black numbers represent bootstrapping values, and red numbers the divergence (Ma, million years ago) estimated based on the pairwise distances of the ML phylogeny and assuming a substitution rate of 11 × 10−9 per site, per generation, and 10 generation per year38.
To determine whether we could find a stable distinction between An. fontenillei and An. bwambae, we repeated the analysis creating a new MAF including the sequences of three more An. fontenillei and two more An. bwambae specimens (see Methods). Among the 343 analysed windows, 278 (81%) showed trees where An. fontenillei and An. bwambae clustered together but were always separated, indicating that they are different populations and/or species (Fig. S4).
We also estimated the pairwise genetic distance between An. fontenillei and An. bwambae and compared it with that between An. coluzzii and An. gambiae, the most recently diverged species within the complex (Fig. S5A)10,37. The pairwise genetic distance was significantly larger in the An. fontenillei - An. bwambae clade than in the An. gambiae – An. coluzzii clade (bootstrapping analysis, median 0.0117 and 0.0067 respectively, Fig. S5B, Table S5). Assuming a substitution rate of 1.1 × 10−9 per site, per generation, and 10 generations per year38, we calculated that the An. fontenillei - An. bwambae clade split ~ 0.53 million years ago (Ma), and the An. gambiae – An. coluzzii clade diverged ~ 0.31 Ma. This result and the clear ecological distinction between An. fontenillei and An. bwambae, suggested that An. fontenillei is a new species in the An. gambiae complex rather than an An. bwambae sub-population.
We then extended our analysis to the whole genome, and estimated the closest species to An. fontenillei, before and after speciation with An. bwambae (Fig. 3). In 84% of the analysed genome, An. bwambae was the closest species to An. fontenillei, forming the An. fontenillei - An. bwambae (FB) clade, (Fig. 3, after speciation –A–, Fig. S6, Table S5). This proportion was similar in each chromosome arm (from 78.4% in the 3 R to 86.6% in the 3 L chromosome arm), and was similar to the 82.8% observed in the X chromosome, suggesting that An. fontenillei has not extensively introgressed with other members of the complex in a recent period.
Relationships of An. fontenillei with other species in the An. gambiae complex according to the phylogenetic trees in 50 kb non-overlapping windows along each chromosome arm. For each chromosomal arm, (A) refers to after speciation and (B) refers to before speciation of An. fontenillei with An. bwambae.
However, the relationship of the FB clade with its closest species and with other clades showed a very different pattern between the X chromosome and the autosomes (Fig. 3, after speciation –A– with An. bwambae). In the X chromosome, most windows showed the species tree, as previously described10. Accordingly, the FB clade was closely related to An. quadriannulatus (27.5%) and An. arabiensis (24.5%). In autosomes, most windows showed the recent introgression between An. arabiensis and the An. gambiae – An. coluzzii clade (A(GC) clade)10. Moreover, the FB clade branched with the A(GC) clade in 27.6% of windows (Figs 3, S6, Table S6). The next more frequent topology (16.4%) showed the FB clade with An. quadriannulatus as the closest species. However, when we did not take into account the 2La inversion (see below), this proportion decreased to 9.6%. On the basis of the X chromosome analysis, we could not determine whether An. quadriannulatus or An. arabiensis is the closest species to the FB clade, due to the similar number of windows showing one or the other topology (Figs 2, S3). Conversely, in autosomes, the FB clade branched more frequently with the A(GC) clade.
Most of the windows that did not show the FB clade were close to the centromere ends. (~10–11 Mb, Fig. 3, after speciation –A–). In these regions, the proportion of windows showing the FB clades was smaller than in the rest of the chromosome. Moreover, the proportion of trees showing An. fontenillei close to the GC clade or An. gambiae or An. coluzzii was substantially higher close to centromeres than in the rest of the chromosome. Specifically, in regions close to centromeres, the FB clade proportion was ~40% for 2 R and 3 R, and, 51% for the 2 L chromosome arm, while in the rest of the three chromosome arms, the FB clade proportion was >80%. The proportion of trees showing An. fontenillei close to the GC clade or An. gambiae or An. coluzzi in regions close to the centromere was ~20% for the 2R and 3R, and 7% for the 2 L chromosome arm, while on the rest of the three chromosome arms this proportion was <1%. Although the aligned regions were shorter (Fig. S7, Methods), the number of informative positions (16,482 on average) and the alignment quality (proportion of gaps and alignment fragmentation) were higher than in other regions.
While we cannot rule out the hypothesis that these regions are a consequence of incomplete lineage sorting, it is difficult to explain why the FB clade appeared repeatedly close to the GC clade. When we removed the FB clade from the analysis, we did not observe any difference in these regions compared with the rest of the genome. Therefore, these windows might indicate a very recent introgression between An. fontenillei and An. gambiae or An. coluzzii, or both.
Recent introgressed genes are mainly involved in metabolic detoxification, desiccation, and olfactory perception
We then analysed the gene content of windows where An. fontenillei instead of branching with An. bwambae (its closer species), clustered with the major malaria vectors An. gambiae and An. coluzzii, or the GC clade. As these species occur in sympatry in the studied area, they could share DNA through secondary contacts. We analysed the three ML tree topologies related with this possible recent introgression separately because the presence of the 2 La polymorphic inversion may affect the results. Indeed, the inversion breaks the more frequently observed GC clade, because the An. coluzzii individuals used for this study carried predominantly the inversion, while the An. gambiae individuals harboured mainly the standard arrangement.
We found that 64 windows harboured 198 genes where An. fontenillei branched with An. gambiae. Functional enrichment analysis of these genes with DAVID using An. gambiae genome as background39,40 showed four significant clusters (Table 1). The first three clusters were related to cuticle proteins, membrane transporter activity, peptidases and proteases. All these protein families had been linked to metabolic detoxification of insecticides in high-throughput genome-wide studies in several mosquito species (reviewed in41). Moreover, cuticle proteins are critical for desiccation tolerance in embryos42. Interestingly, the GO terms of the peptidases and proteases cluster have been previously related to high evolutionary rates9. The last cluster, was related to heat shock protein 70, a conserved protein related to heat stress, oxidative stress, and detoxification43,44. The InterPro domain in this cluster also indicates a rapid evolving gene family (Table 1)9.
There were 25 windows that contained 62 genes where An. fontenillei clustered with An. coluzzii. However, none of these genes was significantly enriched for a particular functional term. Finally, 35 autosomal windows contained 89 genes in which An. fontenillei branched with the GC clade. If these windows were regions of recent introgression, this would mean that these genes were introgressed between An. fontenillei and the common ancestors of An. gambiae and An. coluzzii. We found two enriched clusters (Table 1): the most significant cluster was enriched in flavin monooxygenase. This protein shows function similarity with the cytochrome P450-monooxygenases45 that belong to one of the main protein families related to metabolic detoxification of insecticides in mosquito species (reviewed in41). The other significant cluster was related to olfaction. Three of the four GO terms in the cluster (GO:0050911, GO:0004984 and GO:0005549) and the InterPro domain (Table 1) suggest high evolutionary rates9.
Following a reverse complementary approach, we also checked whether known mutations that confer resistance to insecticides or to some infections (two traits relevant for malaria transmission) were present in An. fontenillei. Specifically, we checked 42 mutations in 14 genes related to insecticide resistance20 and five mutations in one gene related to immunity and infection resistance46. By mapping the four An. fontenillei individuals to the reference genome (AgamP3) with bwa-mem (Text S2.4), we found only two mutations in two glutathione S-transferase genes related to insecticide resistance (Table S7): GSTE6 (E89D mutation) and GSTE3 (N73I mutation). Finally, we also checked whether these mutations were present in the other members of the complex. All the available genome references (An. gambiae Pimperena, An. coluzzii, An. quadriannulatus, and An. arabiensis) showed the wild type alleles (thus, susceptible to insecticides). However, in the MAF made with wild specimens, all the species that could be mapped in those regions (An. gambiae, An. coluzzii and the three An. bwambae individuals) showed the mutant alleles, as did the four An. fontenillei individuals. This result suggests the presence of ancestral polymorphisms within the complex.
Chromosome inversions reveal putative introgression events in the Anopheles gambiae complex
There are two main inversions in the An. gambiae complex that shaped its chromosomal evolution and that emerged in our cytogenetic and phylogenetic analysis: the 2La inversion and the 3La inversion10,47. The 2La inversion is present in An. arabiensis, An. gambiae, and An. coluzzii34, and absent in An. bwambae and An. fontenillei., with regard to the 9 specimens cytogenetically and molecularly karyotyped. Consequently, in this region of the 2L chromosome arm, the FB clade was closer to An. quadriannulatus, defining a well-determined different block that could be easily distinguished in Fig. 3 (line A). According to the four specimens karyotyped, the 3La inversion is fixed in An. fontenillei as well as in An. bwambae and An. melas, as shown by the cytogenetic results. In the 3L chromosome arm, the inverted region could be easily identified because in these windows, the FB clade was closely related to An. melas (Fig. 3, line A). The inferred breakpoints based on the ML tree topology of the 2La and 3La inversions were inside the known cytological breakpoint ranges, except for the 2L telomeric breakpoint, which was 400 kb shorter (Table S834, VectorBase.org).
In 45% of windows in the 3L chromosome arm, the three known An. gambiae complex species with the 3La inversion (An. fontenillei, An. bwambae, and An. melas) were together and separated from the species without the inversion (An. arabiensis, An. quadriannulatus, An. merus, An. gambiae, and An. coluzzii) (Figs 4, S6). This topology suggested two introgression events: i) An. arabiensis with the common ancestor of An. gambiae and An. coluzzii, and ii) An. merus with An. quadriannulatus (Fig. 4). To date the 3La inversion, we estimated the pairwise distances between An. fontenillei and An. quadriannulatus in the 3L chromosome arm outside and inside the inversion (3L: 14.5–35.9 Mb ± the 500 Kb flanking region). Outside the inversion, An. fontenillei and An. quadriannulatus diverged 1.4 Ma (±0.91), similar to the estimate based on the more common phylogenetic tree of the X chromosome (1.25 Ma ± 0.54) (Fig. S8, Table S9). The divergence between An. bwambae and An. quadriannulatus outside the inversion was also similar to the one estimated for the more common phylogenetic tree of the X chromosome: 1.24 Ma (±0.6). However, the divergence dates estimated inside the inversion between An. fontenillei and An. quadriannulatus, and An. bwambae and An. quadriannulatus were 2.53 Ma (±0.97) and 2.23 Ma (±0.76), respectively. These estimates were in the range of the estimated date of An. gambiae complex origin (around 2 ± 0.64 Ma ago). We obtained similar results by repeating the analysis using An. arabiensis instead of An. quadriannulatus (Table S9). We could not accurately date the 3La inversion with this method due to the high uncertainty, but we could show that the inversion was at least older than the An. melas, An. arabiensis, An. quadriannulatus, An. bwambae, and An. fontenillei group. However, An. arabiensis and An. quadriannulatus showed the standard karyotype of the 3La inversion. Therefore, on the basis of the phylogenetic trees, we hypothesized that the ancestral karyotype of the group includes the 3La inversion and that An. quadriannulatus lost the inversion in the introgression from An. merus, as already suggested by Fontaine et al.10, and An. arabiensis during the introgression with An. gambiae/An. coluzzii.
Species topology estimated from the X chromosome sequences compared with the topology of the 3La inversion. An. christyi was used as outgroup species. Green colour: possible introgression between An. arabiensis and the An. gambiae – An. coluzzii clade common ancestor. Purple colour: species that share the 3La inversion. Yellow colour: possible introgression event between An. quadriannulatus and An. merus.
We found some interesting windows in the 3La inversion were An. fontenillei was more closely related to An. melas than to An. bwambae (Fig. 3, before speciation). Functional enrichment analysis of these windows showed that there was only one enriched cluster containing genes related to the stage-specific breakdown of the larval midgut during metamorphosis (Table 1).
Finally, the 3Rb inversion and the 2Rl polymorphic inversions revealed by the karyotyping of An. fontenillei individuals did not leave a clear pattern in our genomic analysis (see Fig. 3). Both inversions were only shared by An. bwambae, the closest species to An. fontenillei, and therefore, big differences are not expected in these regions.
Discussion
In 1975, the English entomologist G. B. White wrote: “As time passes, it becomes increasingly less likely that other sibling species of this complex (An. gambiae) will be found”48. Indeed, during the last 40 years, only one new species, An. quadriannulatus B (recently named An. amharicus), was discovered15,49, and An. coluzzii was separated from An. gambiae, its sister species15. Our field work of 2014 led to the discovery of a new species of the An. gambiae species complex that we named An. fontenillei sp.n. Although, we found An. fontenillei at La Lopé park, a mosaic savanna-forest area of Gabon, this species could also inhabit other mosaic-savanna parts of Gabon and/or Central Africa. Moreover, we cannot exclude its presence in deeper forested areas. This study should warn scientists about the possible presence of this mosquito species when sampling in this part of Africa. On the basis of our collection sites, we hypothesize that this new mosquito species breeds in rain-dependent, sunlit, and open pools, showing similar larval ecology as other fresh-water species within the complex16. Although there are not hot spring water or salt-dependent breeding sites at La Lopé, our hypothesis about fresh-water species is limited by the low number of larvae collected (only two). A more extensive larval collection should be carried out to better characterize other potential breeding sites. According to its geographical distribution, we hypothesized a zoophilic host preference (Fig. 1B). This behaviour has already been found in other members of the complex, such as An. quadriannulatus50, and it seems an ancestral character. Although, An. fontenillei might feed on humans, we did not capture any blood-fed mosquito; therefore, we cannot make strong inferences about its trophic preferences. Nevertheless, the fact that it could be captured with different types of traps suggests a generalist feeding habit with potential consequences on parasite transfer between humans and animals51. Indeed, the ancient and recent history of La Lopé provides multiple opportunities for An. fontenillei to adapt to humans52. In the Neolithic age, La Lopé was commonly colonized by nomad tribes for hunting, and in the last century there was a forestry industry in the park. However, more investigations are needed to determine whether this trait is ancestral, or recently acquired (i.e., by introgression, see below) (Table 1).
To determine its phylogenetic position within the complex, we sequenced and de novo assembled An. fontenillei genome. Its analysis allowed us to determine that An. fontenillei and An. bwambae are sister species. Pairwise comparisons revealed a higher divergence time between An. fontenillei and An. bwambae than between An. gambiae and An. coluzzii (Fig. 2)37, corroborating the geographical and ecological assumption of two different species (Fig. 1). The An. fontenillei - An. bwambae clade was placed together with An. quadriannulatus, An. arabiensis and An. melas, and An. quadriannulatus or An. arabiensis were the closest species to the clade (Fig. 2). This is, to date, the most exhaustive phylogenetic tree of the An. gambiae complex and includes eight of the nine described species (no genome sequence is available for An. amharicus).
Consistent with Fontaine et al.10, we found pervasive evidence of introgression in An. fontenillei, confirming the permeable species boundaries in the An. gambiae complex37,53. Introgression events within species complexes are common in nature, challenging the possibility to trace the evolutionary history of species3. We observed patterns of recent introgression between An. fontenillei and the An. gambiae-An. coluzzi clade, particularly in centromeric regions (20% of the phylogenetic trees). These last two species were found in La Lopé village that is close to the sylvatic sites where An. fontenillei was sampled (Fig. 1A), indicating a potential contact among them. The introgressed genomic windows were mostly enriched in genes associated with detoxification, desiccation tolerance, and olfactory perception (Table 1), traits that have been linked to enhanced vectorial capacity9. Indeed, they allow species to live/breed in a broader range of habitats, and blood-feed on different hosts. The evidence of recent gene exchanges between An. gambiae-An. coluzzii and other species of the complex may influence the evolution of these two major malaria vectors, with potential consequences for malaria transmission (i.e., adaptation to sylvatic habitats and/or preference for feeding on animals). However, this patterns of recent introgression events in centromeric regions could be affected by the low recombination rate in these areas that could help to protect introgressed haplotypes for longer time compared with other genomic regions54.
Finally, we analyzed the evolution of the 3La inversion in the An. gambiae complex. While, this inversion was thought to be present in the ancestor of An. melas and An. bwambae, we estimated that its origin predated the An. gambiae complex radiation33. Moreover, we hypothesized that the inversion was independently lost by An. arabiensis and An. quadriannulatus (Figs 3 and 4). Although the 3La inversion has not been associated with any trait yet, we observed functional enrichment in larval midgut histolysis genes in recently introgressed regions between An. melas and An. fontenillei (Table 1). Again, these two species are present in Gabon, and potential gene exchanges could have occurred between them. Chromosomal rearrangements have modulated the evolution of multiple species by affecting local adaptation or speciation5,55,56,57,58,59,60. In our genomic analysis (Fig. 3), we also observed the genomic signature of the 2La inversion that affects the phylogenetic relationship between An. fontenillei, An. arabiensis, and An. quadriannulatus, highlighting the impact of fixed inversions in chromosome evolution within the complex.
Despite the titanic collection efforts led in Africa during the last century, the rainforest of Central Africa has carefully hidden a new piece of the jigsaw of the An. gambiae complex. The discovery of this new species has provided new insights into genome evolution (i.e., the 3La inversion) and local adaptation (i.e., salinity tolerance) in this group of closely related species. Moreover, the new species has been an active actor in An. gambiae-An. coluzzii evolution, through the exchange of genes involved in vectorial capacity. These introgression events bring new questions about how local populations of An. gambiae and An. coluzzii, the major malaria vectors, have been affected. Indeed, adaptation to rainforest habitats, host preference or resting behaviour could have been modified at La Lopé. New studies may provide important information on how vectorial traits have evolved from wild to domestic populations within the complex, with a direct impact for future malaria control strategies.
Methods
Research and ethics statements
This study and the methods employed for mosquito sampling were approved by CENAREST (national research authorization AR0013/16/MESRS/CENAREST/CG/CST/CSAR). Specimen collection in the National Parks was approved by a national park entry authorization AE16008/PR/ANPN/SE/CS/AEPN. Finally, the National Research Ethics Committee of Gabon (0031/2014/SG/CNE) authorized the use of the human-landing catch (HLC) method. All volunteers signed the appropriate informed consent documents.
Mosquito sampling and species identification
Mosquitoes were sampled in the National Park of La Lopé in Gabon, Central Africa, in an exploratory survey in November 2014. Since then, several collections have been carried out in June 2015, February 2016, and November 2016. (Fig. 1, Table S1). Adults were collected using BG traps with BG-lure and a source of CO2, and HLC, while larvae were sampled with the dipping method61. Collected Anopheles mosquitoes were taxonomically identified according to standard morphological features31,32. Then, they were individually stored in 1.5 mL tubes at −20 °C and sent to CIRMF for molecular analysis. Total genomic DNA from specimens that had been morphologically identified as belonging to the An. gambiae complex was extracted using the DNeasy Blood and Tissue Kit (Qiagen), according to the manufacturer’s instructions. Genomic DNA was eluted in 100 μL of TE buffer. A first molecular analysis (PCR-based) performed to identify species within the complex26 highlighted the presence of an unspecific fragment of 700 bp. This band did not correspond to any of the species identified using the PCR-RFLP diagnostic test26.
Mosquito karyotyping
Half-gravid females were sampled in November 2016 (Table S1) in forest sites where we previously found specimens belonging to the unspecified taxon. Females were collected by HLC and fed to complete their blood-meal. Mosquitoes were allowed to develop follicles for 25 h at field temperature. Then, ovaries were dissected and stored in Carnoy’s fixative solution (100% ethanol: glacial acetic acid, 3:1 by volume). At the CIRMF, ovaries were squashed in a drop of 50% propionic acid to obtain the polytene chromosomes62. The banding patterns of polytene chromosomes were examined using a Leica DM2000 microscope equipped with a Leica DFC 450 camera system (Leica Microsystems GmbH, Wetzlar, Germany). Chromosomal arms and inversions were recorded and scored according to the An. gambiae chromosome map63.
Preliminary sequencing analysis
To obtain further information about the new 700 bp PCR band, three genomic regions previously employed for phylogenetic studies in the complex were sequenced following the authors’ instructions: internal transcribed spacer subunit 2 (ITS2 ~490 bp64), NADH dehydrogenase subunit 5 (ND5 ~300 bp65,66), and cytochrome c oxidase subunit I (COI ~495 bp67). Moreover, a new set of primers was designed to amplify a fragment of the intergenic spacer gene (IGS ~267 bp): IGSKPF 5′-CTCTTGTGAGAGCAAGAGTGT-3′ and IGSKPR 5′-ATCAAGACAATCAAGTCGAGA-3′. These primers were used also for species identification in the complex. For the IGS gene, PCR reactions were carried out in 25 µl reaction volume than included 1X Qiagen PCR buffer (Qiagen, France), 1.5 mM MgCl2, 200 µM each dNTP (Eurogentec, Belgium), 10 pmol of each primer, 2.5 U Taq DNA polymerase (Qiagen, France) and 1–20 ng of template DNA. Amplifications were performed using a Mastercycler Gradient thermocycler (Eppendorf) with the following conditions: an initial step at 94 °C for 5 minutes followed by 35 cycles of 30 seconds at 94 °C, 30 seconds at 54 °C, 1 minute at 72 °C, and a final elongation step of 10 minutes at 72 °C. Five microliters of the PCR product were visualized by electrophoresis on 1.5% agarose gels containing 0.5 µl/ml ethidium bromide and photographed under UV light.
The sequences obtained for the four regions were analysed using Geneious R1068. The consensus sequences of each gene were aligned with randomly chosen sequences of each species within the complex. Unique haplotypes were selected to be included in the phylogenetic analysis. The best substitution model for each gene was identified using SMS69. Phylogenetic trees were then built using the maximum likelihood (ML) method and PhyML70, with nearest neighbour interchange (NNI) for tree searching and approximate likelihood-ratio test (aLRT SH-like,71) for branch support. Trees were visualized with iTOL v.3.4.372.
Genome sequencing and assembly
The genome of four individuals of the unknown species was sequenced using the Illumina platform at the CNAG (Barcelona). To make a de novo genome assembly of this species, the genome of one specimen was deeply sequenced to ~112X. The other three specimens were sequenced with an average coverage of ~29X. All reads were paired-end 126 bp long (Table S2).
The genome of the more deeply sequenced An. fontenillei specimen was assembled at the Bioinformatics Unit, CRG (Barcelona) (Tables S2, S3A). Reads were trimmed and filtered using Skewer, version 0.2.273, to remove the adapter sequence and the low quality part. A FastQC analysis was performed to check the quality of the trimmed reads. Analysis of contaminants using a Kraken database that includes complete bacterial, archaeal, and viral genomes in RefSeq74 highlighted the presence of the enterobacteria phage phiX as the only contaminant (Table S3B). Then, the trimmed reads were assembled with the Platanus software version 1.2.475 to produce contigs and scaffolds using the paired-end information. To join the contigs within the same scaffolds, stretches of N needed to be added. These gaps were filled with the Platanus gap_close function using the original reads (Table S3A). Then, the scaffolding of the assembled genome was improved using the proteins described for the AgamP4 reference in VectorBase (www.vectorbase.org). Blat76 was used to map the proteins to the assembled scaffolds and to reorder and join scaffolds accordingly with PEP_scaffolder77. For the second round of gap filling, and due to format incompatibilities, the GapCloser tool from the SOAPdenovo package78 was used (Table S3A). To evaluate the assembly quality, the presence of conserved genes among the Diptera order was assessed using the BUSCO software79. The 2,799 gene models conserved among Diptera species were classified as: i) completely found in a single sequence, ii) fragmented in different sequences, or iii) completely missing. Most of the BUSCO genes (96%) were completely found in a single sequence (Table S3C). Finally, a polishing step was performed by removing the scaffolds that mapped to previously found contaminants (Table S3A).
Phylogenetic analysis
The available multiple alignment file (MAF) for six species of the An. gambiae complex, including two outgroup species (An. christyi and An. epiroticus10) was used to make the genome-wide phylogenetic tree by window analysis. This MAF represented the alignment formed by whole genome sequences from population samples of multiple individuals of An. gambiae, An. coluzzii, An. merus, An. melas, An. quadriannulatus, and An. arabiensis. The An. gambiae PEST v3 (AgamP3) reference genome obtained from VectorBase (www.vectorbase.org) was also included. Fontaine et al.10 made a whole multiple genome alignment using ROAST80 that represents approximately 40% of the euchromatic genome. The resulting MAF based on field-collected samples was downloaded from http://datadryad.org/resource/doi:10.5061/dryad.f411410. Then, this MAF was added to our An. fontenillei assembly, and the highest coverage An. bwambae genome sequences available (see below).
Anopheles fontenillei sp.n
First, a database was generated with the scaffolds of the An. fontenillei assembly. Then, blastn was run for each region in the MAF using AgamP3 as query sequence against the An. fontenillei scaffold database. This blastn analysis was repeated using other species of the MAF regions as query: An. arabiensis, An. quadriannulatus, An. melas, and An. merus (Text S2.1.1–Text S2.1.4), but not An. coluzzii and An. gambiae, due to their similarity to the reference genome AgamP3, or the two outgroup species because they are too divergent. Then, the MAF regions that gave a single hit in any of the species were selected, which represented 63.2% of all the MAF regions for the eight species (Table S10). For the additional MAF region that gave more than one hit, the multiple hits with e –value >10−4 or with ≤40% of the query covered for each region in each species were excluded (Text S2.1.5), and the sequences that became a unique hit after this filtering were recovered (Table S10). In total, An. fontenillei could be included in 75.2% of the previous MAF regions, which represents ~30% of the euchromatic genome. For each of these MAF regions, the scaffolds were cut according to the blast result information (Text S2.1.6). Then, these sequences were added to the corresponding MAF region using MAFFT as aligner (v7.221,81). The function–add was used to modify as less as possible the initial MAF82. Finally, each region of the MAF was joined to generate the new MAF that included the An. fontenillei genome (Text S2.2).
Anopheles bwambae
The three individual sequences of An. bwambae available at NCBI were downloaded with fastq-dump: i) An. bwambae 1, SRR1255391, SRR1255392, and SRR1255303, ii) An. bwambae 3, SRR1255390, and iii) An. bwambae 4, SRR1255325. Then, the SRR files for An. bwambae 1 were joined (Text S2.3). For each individual, the read quality was assessed with fastQC and reads were trimmed using cutadapt (v. 1.8.383)(Text S2.4.1–Text S2.4.3). After trimming, the quality per base was always higher than 24. Then, the trimmed reads were mapped to the AgamP3 reference genome using bwa-mem84. Several post-mapping steps were performed, including marking duplicates and realigning around indels using Picard (v.1.109; http://picard.sourceforge.net), samtools (v. 1.385), and GATK (v3.4–4686)(Text S2.4.4–Text S2.4.8). Among the three available An. bwambae individuals, the one with the highest coverage (An. bwambae 1 with 33.2X) was added to the MAF (coverage for the other two: An. bwambae 3, 11.7X and An. bwambae 4, 11.2X). A consensus sequence of the An. bwambae 1 reads mapping to the AgamP3 sequence of every MAF region with the nine species (An. gambiae, An. coluzzii, An. merus, An. melas, An. quadriannulatus, An. arabiensis, An. fontenillei, An. christyi, and An. epiroticus) was created using SAMtools mpileup (Text S2.5). If An. bwambae reads did not exist for a MAF region, gaps were added to keep the same number of MAF regions as before. This had a marginal effect because it only concerned 0.06% of all MAF regions. Finally, MAFFT aligner, with the function –add, was used to add each An. bwambae 1 consensus sequence to the MAF regions, and then all these regions were joined in a new MAF (Text S2.2).
MAF with four An. fontenillei sp.n. individuals and three An. bwambae individuals
To check the phylogenetic relationship between An. fontenillei and An. bwambae, an additional MAF was created that included the eight previously available species, four An. fontenillei individuals, and three An. bwambae individuals. Each of the seven individual An. fontenillei and An. bwambae genomes was mapped to the AgamP3 reference genome, as described previously for An. bwambae (Text S2.4). Then, a consensus sequence was generated for each of the new individuals for each of the MAF regions using SAMtools mpileup (Text S2.5). Then, each new sequence was sequentially added to the available multiple alignment regions using the MAFFT –add function as aligner (v7.22181,82, Text S2.2). Finally, all these sequences were joined in a new MAF file.
Window-based phylogenies
From the MAF, 50 kb genome-wide non-overlapping windows were generated (Text S2.6.1). For each window, a ML phylogenetic tree was created using RAxML (v8.2.4,87) with the GTRGAMMA model and bootstrapping for 1,000 replicates (Text S2.6.2)10. The closer related species, An. christyi, was used as outgroup because Fontaine et al.10 already showed that the outgroup choice does not substantially influence the results. Windows with less than 10% of informative base pairs (e.g., <5,000 bp) were excluded (following10). The different topologies obtained were sorted, counted, and analysed using ad-hoc perl scripts (Text S2.6.3).
Pairwise distance and bootstrapping
The R package ‘APE’ (v4.188) was used to estimate pairwise genetic distances based in the ML phylogenetic trees. Then, the bootstrap analysis was performed using the ‘boot’ package in R89 (Text S2.7).
Centromeric region alignment quality
For each chromosomal arm, 30 windows were randomly chosen from centromeric regions, and 30 from other genomic regions. Centromeric regions were defined based on the observed ancestry pattern in Fig. 3: 2L: 0 to 10 Mb, 2R: 50 to 61.3 Mb, 3L: 0 to 10 Mb, 3R: 40 to 53.1 Mb, and X: 15 to 20.2 Mb. For these 60 windows, the alignment length, the alignment length without completely undetermined characters and gaps, the proportion of gaps, and the alignment patterns were extracted from the RAxML information file.
Data analysis
R v3.2.5 (R Development Core Team, http://cran.r-project.org/) was used to perform all the statistical analyses. Figures were prepared with the Inkscape software (https://inkscape.org).
Data Availability
DNA sequences have been deposited to GenBank under accessions numbers MN172532 - MN172595. The four An. fontenillei sp. n. genomes and the genome assembly have been deposited under the bioproject number PRJNA508319.
References
Coyne, J. & Orr, H. Speciation. (Sinauer Associates, 2004).
Feder, J. L., Egan, S. P. & Nosil, P. The genomics of speciation-with-gene-flow. Trends in Genetics 28, 342–350, https://doi.org/10.1016/j.tig.2012.03.009 (2012).
Mallet, J., Besansky, N. & Hahn, M. W. How reticulated are species? BioEssays 38, 140–149, https://doi.org/10.1002/bies.201500149 (2016).
Hedrick, P. W. Adaptive introgression in animals: examples and comparison to new mutation and standing variation as sources of adaptive variation. Molecular Ecology 22, 4606–4618, https://doi.org/10.1111/mec.12415 (2013).
Kirkpatrick, M. & Barrett, B. Chromosome inversions, adaptive cassettes and the evolution of species’ ranges. Molecular Ecology 24, 2046–2055, https://doi.org/10.1111/mec.13074 (2015).
Huerta-Snchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197, https://doi.org/10.1038/nature13408 (2014).
Weill, M. et al. The kdr mutation occurs in the Mopti form of Anopheles gambiae s.s. through introgression. Insect molecular biology 9, 451–455 (2000).
Sinka, M. E. et al. The dominant Anopheles vectors of human malaria in Africa, Europe and the Middle East: occurrence data, distribution maps and bionomic précis. Parasites & Vectors 3, 117, https://doi.org/10.1186/1756-3305-3-117 (2010).
Neafsey, D. E. et al. Highly evolvable malaria vectors: The genomes of 16 Anopheles mosquitoes. Science 347, https://doi.org/10.1126/science.1258522 (2015).
Fontaine, M. C. et al. Extensive introgression in a malaria vector species complex revealed by phylogenomics. Science (New York, N.Y.) 347, 1258524, https://doi.org/10.1126/science.1258524 (2015).
Cohuet, A., Harris, C., Robert, V. & Fontenille, D. Evolutionary forces on Anopheles: what makes a malaria vector? Trends in Parasitology 26, 130–136 (2010).
Sinka, M. E. et al. A global map of dominant malaria vectors. Parasites & Vectors 5, 69, https://doi.org/10.1186/1756-3305-5-69 (2012).
Davidson, G. Anopheles gambiae complex. Nature 196, 907 (1962).
Davidson, G. & Hunt, R. H. The crossing and chromosome characteristics of a new 6th species in the Anopheles gambiae complex. Parassitologia 15, 121–128 (1973).
Coetzee, M., Hunt, R. H. & Wilkerson, Ra Anopheles coluzzii and Anopheles amharicus, new members of the Anopheles gambiae complex. Zootaxa 3619, 246–274 (2013).
White, B. J., Collins, F. H. & Besansky, N. J. Evolution of Anopheles gambiae in Relation to Humans and Malaria. Annual Review of Ecology, Evolution, and Systematics 42, 111–132, https://doi.org/10.1146/annurev-ecolsys-102710-145028 (2011).
Pombi, M. et al. Dissecting functional components of reproductive isolation among closely related sympatric species of the Anopheles gambiae complex. Evolutionary Applications 10, 1102–1120, https://doi.org/10.1111/eva.12517 (2017).
Fontaine, M. C. et al. Data from: Extensive introgression in a malaria vector species complex revealed by phylogenomics. Science, https://doi.org/10.5061/dryad.f4114 (2014).
Fouet, C., Gray, E., Besansky, N. J. & Costantini, C. Adaptation to Aridity in the Malaria Mosquito Anopheles gambiae: Chromosomal Inversion Polymorphism and Body Size Influence Resistance to Desiccation. PLoS ONE 7, e34841, https://doi.org/10.1371/journal.pone.0034841 (2012).
Miles, A., Harding, N. J. & Bott Genetic diversity of the African malaria vector Anopheles gambiae. Nature 552, 96, 10.1038/nature24995 (2017).
Norris, L. C. et al. Adaptive introgression in an African malaria mosquito coincident with the increased usage of insecticide-treated bed nets. Proceedings of the National Academy of Sciences 112, 815–820 (2015).
Gagnaire, P. A., Pavey, S. A., Normandeau, E. & Bernatchez, L. The genetic architecture of reproductive isolation during speciation-with-gene-flow in lake whitefish species pairs assessed by rad sequencing. Evolution 67, 2483–2497, https://doi.org/10.1111/evo.12075 (2013).
Rinker, D. C. et al. Blood meal-induced changes to antennal transcriptome profiles reveal shifts in odor sensitivities in Anopheles gambiae. Proceedings of the National Academy of Sciences 110, 8260–8265, https://doi.org/10.1073/pnas.1302562110 (2013).
Smith, H. A. et al. Genome-wide QTL mapping of saltwater tolerance in sibling species of Anopheles (malaria vector) mosquitoes. Heredity 115, 471–479, https://doi.org/10.1038/hdy.2015.39 (2015).
Bhatt, S. et al. The effect of malaria control on Plasmodium falciparum in Africa between 2000 and 2015. Nature 526, 207–211, https://doi.org/10.1038/nature15535 (2015).
Fanello, C., Santolamazza, F. & della Torre, A. Simultaneous identification of species and molecular forms of the Anopheles gambiae complex by PCR-RFLP. Medical and veterinary entomology 16, 461–464 (2002).
Elissa, N. et al. Malaria transmission in a region of savanna-forest mosaic, Haut-Ogooué, Gabon. Journal of the American Mosquito Control Association 15, 15–23 (1999).
Mourou, J.-R. et al. Malaria transmission and insecticide resistance of Anopheles gambiae in Libreville and Port-Gentil, Gabon. Malaria journal 9, 321, https://doi.org/10.1186/1475-2875-9-321 (2010).
Hervy, J. F. a. Les Anophèles de la region afrotropicale. (Paris France, 1998).
Pombi, M. et al. Evaluation of a protocol for remote identification of mosquito vector species reveals BG-Sentinel trap as an efficient tool for Anopheles gambiae outdoor collection in Burkina Faso. Malaria journal 14, 161 (2015).
Gillies, M. T. & de Meillon, B. The anophelinae of Africa, south of the Sahara. The South African Institute for Medical Research 54 (1968).
Gillies, M. T. & Coetzee, M. C. A supplement to the Anophelinae of Africa south of the Sahara (Afrotropical Region). Publications of the South African Institute for Medical Research 55, 143 (1987).
White, G. B. Anopheles bwambae, a malaria vector in the Semliki valley, Uganda, and its relationships wiht other sibling species of the An. gambiae complex (Diptera, Culicidae). Systematic Entomology 10, 501–522 (1985).
Coluzzi, M. & Sabatini, A. & della Torre, A. a. A Polytene Chromosome Analysis of the Anopheles gambiae Species Complex. Science 298, 1415–1418, https://doi.org/10.1126/science.1077769 (2002).
White, B. J. et al. Molecular karyotyping of the 2La inversion in Anopheles gambiae. The American journal of tropical medicine and hygiene 76, 334–339 (2007).
Gentile, G., Slotman, M., Ketmaier, V., Powell, J. R. & Caccone, A. Attempts to molecularly distinguish cryptic taxa in Anopheles gambiae s.s. Insect Molecular Biology 10, 25–32, https://doi.org/10.1046/j.1365-2583.2001.00237.x (2001).
Neafsey, D. E. et al. SNP genotyping defines complex gene-flow boundaries among African malaria vector mosquitoes. Science (New York, N.Y.) 330, 514–517, https://doi.org/10.1126/science.1193036 (2010).
Tamura, K., Subramanian, S. & Kumar, S. Temporal Patterns of Fruit Fly (Drosophila) Evolution Revealed by Mutation Clocks. Molecular Biology and Evolution 21, 36–44, https://doi.org/10.1093/molbev/msg236 (2004).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Research 37, 1–13, https://doi.org/10.1093/nar/gkn923 (2009).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57, https://doi.org/10.1038/nprot.2008.211 (2009).
Liu, N. Insecticide Resistance in Mosquitoes: Impact, Mechanisms, and Research Directions. Annual Review of Entomology 60, 537–559, https://doi.org/10.1146/annurev-ento-010814-020828 (2015).
Goltsev, Y. et al. Developmental and evolutionary basis for drought tolerance of the Anopheles gambiae embryo. Developmental Biology 330, 462–470, https://doi.org/10.1016/j.ydbio.2009.02.038 (2009).
Tavaria, M., Gabriele, T., Kola, I. & Anderson, R. L. A hitchhiker’s guide to the human Hsp70 family. Cell stress & chaperones 1, 23–28 (1996).
Morano, K. A. New tricks for an old dog: The evolving world of Hsp70. Annals of the New York Academy of Sciences 1113, 1–14, https://doi.org/10.1196/annals.1391.018 (2007).
Cashman, J. R. Some distinctions between flavin-containing and cytochrome P450 monooxygenases. Biochemical and Biophysical Research Communications 338, 599–604, https://doi.org/10.1016/j.bbrc.2005.08.009 (2005).
White, B. J. et al. Adaptive divergence between incipient species of Anopheles gambiae increases resistance to Plasmodium. Proceedings of the National Academy of Sciences 108, 244, 10.1073/pnas.1013648108/-/DCSupplemental.www.pnas.org/cgi/doi/10.1073/pnas.1013648108 (2011).
Thawornwattana, Y., Dalquen, D. & Yang, Z. H. Coalescent Analysis of Phylogenomic Data Confidently Resolves the Species Relationships in the Anopheles gambiae Species Complex. Molecular Biology and Evolution 35, 2512–2527, https://doi.org/10.1093/molbev/msy158 (2018).
White, G. B. Notes on a Catalogue of Culicidae of the Ethiopian Region. Mosquito Systematics 7 (1975).
Hunt, R. H., Coetzee, M. & Fettene, M. The Anopheles gambiae complex: a new species from Ethiopia. Transactions of the Royal Society of Tropical Medicine and Hygiene 92, 231–235 (1998).
Dekker, T. & Takken, W. Differential responses of mosquito sibling species Anopheles arabiensis and An. quadriannulatus to carbon dioxide, a man or a calf. Medical and Veterinary Entomology 12, 136–140, https://doi.org/10.1046/j.1365-2915.1998.00073.x (1998).
Makanga, B. et al. Ape malaria transmission and potential for ape-to-human transfers in Africa. Proceedings of the National Academy of Sciences 113, 5329–5334, https://doi.org/10.1073/pnas.1603008113 (2016).
Oslisly, R. et al. Climatic and cultural changes in the west Congo Basin forests over the past 5000 years. Philosophical transactions of the Royal Society of London. Series B, Biological sciences 368, 20120304, https://doi.org/10.1098/rstb.2012.0304 (2013).
Besansky, N. J. et al. Semipermeable species boundaries between Anopheles gambiae and Anopheles arabiensis: evidence from multilocus DNA sequence variation. Proceedings of the National Academy of Sciences of the United States of America 100, 10818–10823, https://doi.org/10.1073/pnas.1434337100 (2003).
Cruickshank, T. E. & Hahn, M. W. Reanalysis suggests that genomic islands of speciation are due to reduced diversity, not reduced gene flow. Molecular Ecology 23, 3133–3157, https://doi.org/10.1111/mec.12796 (2014).
Rieseberg, L. H. Chromosomal rearrangements and speciation. Trends in ecology & evolution 16, 351–358 (2001).
Noor, M. A. F., Grams, K. L., Bertucci, L. A. & Reiland, J. Chromosomal inversions and the reproductive isolation of species. Proceedings of the National Academy of Sciences 98, 12084–12088, https://doi.org/10.1073/pnas.221274498 (2001).
Lowry, D. B. & Willis, J. H. A Widespread Chromosomal Inversion Polymorphism Contributes to a Major Life-History Transition, Local Adaptation, and Reproductive Isolation. PLoS Biology 8, e1000500, https://doi.org/10.1371/journal.pbio.1000500 (2010).
Ayala, D., Guerrero, R. F. & Kirkpatrick, M. Reproductive isolation and local adaptation quantified for a chromosome inversion in a malaria mosquito. Evolution 67, 946–958, https://doi.org/10.1111/j.1558-5646.2012.01836.x (2013).
Ayala, D., Ullastres, A. & Gonzlez, J. Adaptation through chromosomal inversions in Anopheles. Frontiers in Genetics 5, 129, https://doi.org/10.3389/FGENE.2014.00129 (2014).
Ayala, D. et al. Chromosome inversions and ecological plasticity in the main African malaria mosquitoes. Evolution 71, 686–701, https://doi.org/10.1111/evo.13176 (2017).
Service, M. W. Mosquito ecology field sampling methods. 2nd edn, (Elsevier Applied Science, 1993).
della Torre, A. The Molecular Biology of Insect Disease Vectors: a Methods Manual. 329–336 (1997).
Pombi, M., Caputo, B. & Simard, F. A. Chromosomal plasticity and evolutionary potential in the malaria vector Anopheles gambiae sensu stricto: insights from three decades of rare paracentric inversions. BMC Evolutionary Biology 8, 309, https://doi.org/10.1186/1471-2148-8-309 (2008).
Kengne, P. et al. Molecular differentiation of three closely related members of the mosquito species complex, Anopheles moucheti, by mitochondrial and ribosomal DNA polymorphism. Medical and Veterinary Entomology 21, 177–182, https://doi.org/10.1111/j.1365-2915.2007.00681.x (2007).
Besansky, N. J. et al. Patterns of mitochondrial variation within and between African malaria vectors, Anopheles gambiae and An. arabiensis, suggest extensive gene flow. Genetics 147, 1817–1828 (1997).
Thelwell, N. J., Huisman, R. A., Harbach, R. E. & Butlin, R. K. Evidence for mitochondrial introgression between Anopheles bwambae and Anopheles gambiae. Insect molecular biology 9, 203–210 (2000).
Simon, C. et al. Evolution, Weighting, and Phylogenetic Utility of Mitochondrial Gene Sequences and a Compilation of Conserved Polymerase Chain Reaction Primers. Annals of the Entomological Society of America 87, 651–701, https://doi.org/10.1093/aesa/87.6.651 (1994).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649, https://doi.org/10.1093/bioinformatics/bts199 (2012).
Lefort, V., Longueville, J.-E. & Gascuel, O. SMS: Smart Model Selection in PhyML. Molecular Biology and Evolution 34, 2422–2424, https://doi.org/10.1093/molbev/msx149 (2017).
Guindon, S. et al. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Systematic Biology 59, 307–321, https://doi.org/10.1093/sysbio/syq010 (2010).
Anisimova, M. & Gascuel, O. Approximate Likelihood-Ratio Test for Branches: A Fast, Accurate, and Powerful Alternative. Systematic Biology 55, 539–552, https://doi.org/10.1080/10635150600755453 (2006).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128, https://doi.org/10.1093/bioinformatics/btl529 (2007).
Jiang, H., Lei, R., Ding, S.-W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinformatics 15, 182, https://doi.org/10.1186/1471-2105-15-182 (2014).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biology 15, R46, https://doi.org/10.1186/gb-2014-15-3-r46 (2014).
Kajitani, R. et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Research 24, 1384–1395, https://doi.org/10.1101/gr.170720.113 (2014).
Kent, W. J. BLAT–The BLAST-Like Alignment Tool. Genome Research 12, 656–664, https://doi.org/10.1101/gr.229202 (2002).
Zhu, B.-H. et al. PEP_ scaffolder: using (homologous) proteins to scaffold genomes. Bioinformatics 32, 3193–3195, https://doi.org/10.1093/bioinformatics/btw378 (2016).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18, https://doi.org/10.1186/2047-217X-1-18 (2012).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Hou, M. TOAST and ROAST (2008).
Katoh, K., Kuma, K.-i, Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Research 33, 511–518, https://doi.org/10.1093/nar/gki198 (2005).
Katoh, K. & Frith, M. C. Adding unaligned sequences into an existing alignment using MAFFT and LAST. Bioinformatics 28, 3144–3146, https://doi.org/10.1093/bioinformatics/bts578 (2012).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10, https://doi.org/10.14806/ej.17.1.200 (2011).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Research 20, 1297–1303, https://doi.org/10.1101/gr.107524.110 (2010).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313, https://doi.org/10.1093/bioinformatics/btu033 (2014).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics (Oxford, England) 20, 289–290 (2004).
Canty, A. & Ripley, B. boot: Bootstrap R (S-Plus) Functions. R package version 1, 3–20 (2017).
Acknowledgements
We are grateful to P. Nosil and G. Lanzaro for comments on the manuscript. We thank the “Agence Nationale de la Preservation de la Nature” (ANPN), the “Station d’Etudes des Gorilles et Chimpanzes” (SEGC) and the “Centre National de la Recherche Scientifique et Technologique of Gabon” (CENAREST) that authorized this study and facilitated the access to the national parks of La Lopé. We specially thank Vincenzo Petrarca for its help interpreting chromosome polymorphisms. Finally, we greatly thank Maureen Coetzee for revising the taxonomic description. Funding was provided by the “Institut de Recherche pour le Developpément”, the “Agence Universitaire de la Francophonie” (grant OKANDA), the “Agence Nationale de Recherche” (ANR-18-CE35-0002-01 – WILDING), the “Consejo Superior de Investigaciones Cientificas” and the “Centre National de la Recherche Scientifique” (PICS ANCESTRAL), and the “Ministerio de Ciencia, Innovación y Universidades/AEI/FEDER” (BFU2017-82937-P) and the “Secretaria d’Universitats i Recerca. Departament d’Economia i Coneixement. Generalitat de Catalunya” (GRC 2017 SGR 880).
Author information
Authors and Affiliations
Contributions
D.A., C.P., M.G.B. and J.G. designed the research; D.A., N.R., M.F.N. and W.T.B. did the field collections, including larva and adults, and taxonomical identification, in the park and in the village. O.A.E. and P.K. performed the molecular karyotyping, genes sequencing, and species delimitation. D.A. performed the preliminary phylogenetic analysis. M.P. did the cytogenetic analysis assisted by D.A. M.G.B. carried out the computational analysis including read mapping, S.N.P. calling and phylogenetic analysis, supported by J.G. and D.A. C.P., F.S. and C.C. helped to supervise the project. M.G.B., J.G. and D.A. wrote the paper. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barrón, M.G., Paupy, C., Rahola, N. et al. A new species in the major malaria vector complex sheds light on reticulated species evolution. Sci Rep 9, 14753 (2019). https://doi.org/10.1038/s41598-019-49065-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49065-5
This article is cited by
-
Contribution of Anopheles gambiae sensu lato mosquitoes to malaria transmission during the dry season in Djoumouna and Ntoula villages in the Republic of the Congo
Parasites & Vectors (2024)
-
The Anopheles coluzzii range extends into Kenya: detection, insecticide resistance profiles and population genetic structure in relation to conspecific populations in West and Central Africa
Malaria Journal (2024)
-
Deep learning and wing interferential patterns identify Anopheles species and discriminate amongst Gambiae complex species
Scientific Reports (2023)
-
Modelling spatiotemporal trends in the frequency of genetic mutations conferring insecticide target-site resistance in African mosquito malaria vector species
BMC Biology (2022)
-
Recurrent co-domestication of PIF/Harbinger transposable element proteins in insects
Mobile DNA (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.