Abstract
Keratoconus (KC) is a multi-factorial corneal ectasia with unknown etiology affecting approximately 1:2000 people worldwide. Dysregulated gene expression, using RNA-Seq technology, have been reported in KC corneal tissue. However, the differential expression of genes, in KC corneal stromal cells have been widely ignored. We utilized mRNA-Seq to analyze gene expression in primary human corneal stromal cells derived from five non-Keratoconus healthy (HCF) and four Keratoconus (HKC) donors. Selected genes were further validated using real time PCR (RT-PCR). We have identified 423 differentially expressed genes with 187 down- and 236 up-regulated in KC-affected corneal stromal cells. Gene ontology analysis using WebGestalt indicates the enrichment of genes involved in cell migration, extracellular matrix, adherens junction, and MAPK signaling. Our protein-protein interaction network analysis identified several network seeds, such as EGFR, NEDD4, SNTA1, LGALS3BP, HSPB1, SDC2, MME, and HIF1A. Our work provides an otherwise unknown information on the transcriptional changes in HKCs, and reveals critical mechanisms of the cellular compartment. It also highlights the importance of human-based in vitro studies on a disease that currently lacks strong biomarkers and animal models.
Similar content being viewed by others
Introduction
Keratoconus (KC) is a multi-factorial, complex, idiopathic disease of the cornea that leads to corneal thinning, corneal scarring, and the hallmark “cone-like” shape of the cornea1. Corneal epithelium iron deposition and Descemet membrane rupture are also seen in KC2. The disease usually affects both eyes and can deteriorate very fast leading to potentially blinding conditions to young people, if left untreated. KC affects approximately 1:400 people worldwide and typically begins during puberty, until its arrest in the third or fourth decade3. The reason for this arrest remain a mystery.
KC has a substantial financial burden associated with it, post-diagnosis. Though the monetary costs associated with KC are apparent, effects on more subjective areas, such as quality of life and job outlook, are harder to compute. Since KC is a progressive visual disorder, a diagnosis of KC is considered disqualifying for securing various jobs as well as joining the military4, regardless of best-corrected or even uncorrected visual acuity. Various studies have shown that individuals with KC have lower quality of life that stems from the lack of quality corrected vision5,6,7. The National Eye Institute’s Visual Function Questionnaire data from over 1,100 KC patients over a 7-year period identified a significant decline in quality of life in KC patients8. Therefore, efforts to improve methods of treatment are critical.
KC has been associated with various genetic and other exogenous degenerative factors9,10. However, despite significant advancements, the pathophysiology of KC remains poorly understood. A lack of an animal model has severely hampered our progress in unravelling the KC roots. Except for one case report of KC in a rhesus monkey11, KC has not been reported in animals. In addition, we have yet to see the development of a transgenic animal with the KC phenotype. These limitations are the main reason that key molecular mechanisms responsible for the disease onset and progression have not been identified. Thus, it is urgent to develop a global understanding of how changes occur, in KC, at the cellular level.
Gene expression profile using disease-affected tissues have been applied to identify disease-related genes and pathways using different high throughput technologies, including microarray and next generation sequencing (NGS). Using PCR-based subtractive hybridization, Wentz-Hunter et al. (2001) identified elevated expression of HSP90, decorin, fibronectin, ferritin heavy chain, and keratocan in KC-affected corneal stromal layer when compared to the normal/healthy stromal layer12. More gene expression studies using whole cornea, epithelium, or tears from individuals with or without KC have identified a number of differentially expressed genes and KC-related pathways13,14,15,16,17,18,19,20,21,22,23,24. Recently, three RNA-Seq-based expression profiling studies using whole cornea or corneal epithelium have identified many differentially expressed genes involved in TGFβ, WNT, and PI3K/AKT signaling in KC pathogenesis13,21,25.
We have previously shown that human corneal stromal cells isolated from KC patients, termed HKCs, maintain characteristics of the disease phenotype in vitro and secrete and assemble a thinner extracellular matrix (ECM) high in the pro-fibrotic marker, collagen type III, compared to their healthy counterparts26,27. Furthermore, consistent with tear analysis of KC patients in vivo28,29,30, we have found that HKCs maintain altered cellular metabolism31 correlating to increased oxidative stress32. In this study, for the first time, we have investigated the transcriptome profiles of primary human corneal stromal cells from Healthy and KC donors using RNA-Seq to identify global changes in transcript levels that may contribute to the disease phenotype. Our bioinformatics analysis and RT-PCR validation implicated the potential involvement of genes in several KC-related pathways. If in vitro studies are to lead the way towards future KC therapeutic targets, it is critical to examine the primary corneal cells used. Our study provides, otherwise unknown, information to our understanding of the disease mechanisms.
Methods
Ethics and inclusion criteria
Primary human corneal stromal cells were isolated from five healthy/normal and four KC individuals. The Institutional Review Board at the University of Oklahoma Health Science Center – Dean McGee Eye Institute approved the protocol and studies, prior to initiation. This study adhered to the tenets of the Declaration of Helsinki. Prior to collection of corneas, written permission was obtained from all subjects following corneal transplantation. None of the KC donors was from a vulnerable population and all donors or next of kin provided written informed consent that was freely available. Patients who had any other ocular or systemic disease, or previously underwent collagen crosslinking, were excluded from this study. All donors were examined comprehensively using Pentacam HR, refraction, and slit lamp to confirm KC diagnosis. Stabilization of corneal thinning or KC progression was not reported by the clinician prior to tissue isolation and remains unknown.
Healthy corneal tissues with no history of ocular or systemic diseases was provided by, the National Development and Research Institutes (NDRI).
Cell isolation and expansion
Primary corneal stromal fibroblasts were isolated as previously described33,34. KC corneal tissue was provided from the Dean McGee Eye Institute in Oklahoma City, OK. Corneal stromal cells were isolated, by removing both the epithelial and endothelial layers with a sterile surgical scalpel. Tissues were then cut into small pieces (~2 × 2 × 2 mm), and incubated in sterile flasks to promote adhesion. Explants were supplemented with EMEM containing 10% fetal bovine serum (FBS, Atlanta biologicals, Flowery Branch, GA) and antibiotic/antimycotic (anti/anti, Life Technologies, Grand Island, NY), and incubated at 37 °C/5% CO2 for 2–3 weeks until cells migrated from the explant (~70–80% confluence).
Both Healthy Corneal Fibroblasts (HCFs) and Human Keratoconus cells (HKCs) were cultured on T75 tissue culture flasks. Cells were seeded at 5,000 cells/cm2 and maintained at 37 °C with 5% CO2 in a humidified incubator. Cells used in all experiments were between 2nd and 4th passages.
RNA extraction
Total RNA was extracted using the Ambion RNA mini extraction kit (Ambion TRIzol® Plus RNA Purification Kit: Life technologies, Carlsbad, CA). Briefly, TriReagent® (Life Technologies Corporation, Carlsbad, CA), was added to the cell layer after aspiration of medium from the culture and brief washing with Phosphate Buffered Saline (PBS). The cellular layer was mechanically disrupted from tissue culture plate using gentle pipetting. Phase separation was conducted with chloroform and the total RNA contained in the aqueous phase was purified using RNeasy® mini kit column (QIAGEN, Hilden, Germany), according to the manufacturer’s protocol. Three extractions were carried out for each sample and pooled at the end of the RNeasy protocol. The purity and quantity of total RNA were evaluated using an ultraviolet spectrometer (CLARIOstar, BMG LABTECH, Cary, NC).
RNA-Seq and bioinformatics analysis
RNA sequencing library was prepared using TruSeq RNA Library Prep kit, from Illumina with sample-specific indexes at the Oklahoma Medical Research Foundation (OMRF) Genomics facility. Briefly, 1 µg of total RNA was used to purify and fragment mRNA, followed by first and second strand cDNA synthesis. After purification and ligation of Illumina adapters and sample-specific indexes, sequencing libraries were validated using Agilent DNA 100 kit with Agilent Bio analyzer 2100. The libraries were normalized, and pooled for sequencing, with Illumina NextSeq 550 system using the paired end 75 bp with the high-output kit.
The data analysis for RNA-Seq was done using our established pipeline as described previously13. Briefly, after quality check and quality control with all the sequencing reads, demultiplexed reads were aligned by TopHat in paired-end reading with the approximation of the median library size13. Counts of sequencing reads were normalized using Cufflinks in fragments per kilo bases and millions reads (FPKM)13. Normalized sequencing read counts were analyzed by Cuffdiff13, with a transcript file from Ensembl database for the annotations at the gene as well as the isoform levels in a group comparison manner such as 4 cases vs 5 controls in this experimental design. Missing expression data in specific samples was replaced with a value of 0.001 to enable the differential analysis between cases and controls. Without such replacement of missing data, the related mRNAs would have been excluded from the differential expression analysis. In the final gene list with the positions at the corresponding chromosomes, it showed comparative fold changes and adjusted P values (false discovery rate, FDR) through the resulting analysis of four cases vs. five controls. Differentially expressed genes were defined to have an FDR value ≤ 0.05 and a |fold change| ≥ 2.
The differentially expressed genes were loaded into WEB-based Gene Set Analysis Toolkit (WebGestalt)35,36, for bioinformatics analysis to identify enriched gene ontologies, pathways, network modules, and associations with phenotypes/disease/drug. Potential targeted miRNAs were identified for the differentially expressed genes using WebGestalt.
Validation using Real-time PCR
To validate the differential expression of selected genes, we used RT-PCR with all the donors/samples, as previously described32,37,38. The cDNA synthesis was followed using a SuperScript™ III First-Strand Synthesis SuperMix (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. The TaqMan gene expression assays (Applied Biosystems, Foster City) for GAPDH (Hs99999905_m1) and 18S (Hs99999901_s1) were used as the reference assays to normalize target gene expression. We selected the following genes for validation: KRT7 (Hs00559840_m1), MME (Hs00153510_m1), ANKRD1 (Hs00173317_m1), ERG1 (Hs00152928_m1), IL1B (Hs01555410_m1) CXCL1 (Hs00236937_m1), and GDF15 (Hs00171132_m1). Furthermore, 10 ng of cDNA was used for initiating the PCR reaction for a 20-µl reaction mixture containing our desired probes and the TaqMan Fast Advanced MasterMix (Applied Biosystems, Life technologies, Foster City, CA). Amplification of samples was performed using the StepOnePlusTM real-time PCR system (Life Technologies) in accordance with the manufacture’s protocol. Graph Pad Prism 7 and MS-Excel were used for data analysis.
Statistical analysis
GraphPad Prism 7.02 was used to determine statistical significance using ANOVA or t-test, where appropriate. A p ≤ 0.05 was considered statistically significant.
Results
Clinical phenotypes
Our study included HCF cells from 5 postmortem donors (3 males/2 females) with postmortem delay less than 24 hours. All four HKC cells were derived from surgically removed keratoconic corneas immediately after surgery. It included 1 male and 3 females. The average age was 47.2 years for controls and 60.8 years for cases.
HCFs and HKCs – transcriptional profiles
We performed RNA-Seq with primary human corneal stromal fibroblast cells derived from five unaffected controls (HCFN21, HCFN20, HCFN23, HCFN22, and HCFN4) and four KC patients (HKCWV1, HKCWV2, HKCDM1, and HKCDM2). Individual information for all the samples is provided in Table 1.
All nine RNA samples were sequenced with 33.7–47.9 million paired-end reads of 75 nucleotides. All the samples had at least 30 million paired sequence reads aligned. Sequence reads with multiple alignments were removed during quality control process. Out of the total 15,159 genes, 11,540, 11,637, 11,568, 11,577, and 11,740 genes were expressed in the individual HCF cells with FPKM ≥ 1.0 and 9,870 genes were expressed in all five primary HCF cells with FPKM ≥ 1.0. 11,791, 11,768, 11,751, 12,122, and 11,544 genes were expressed in the individual HKC cells with FPKM ≥ 1.0 and 9,923 genes were expressed in all five primary HKC cells with FPKM ≥ 1.0. With all corneal stromal fibroblast cells from 10 donors, 9,302 genes were expressed with FPKM ≥ 1.0.
Our differential analysis using Cuffdiff identified 423 differentially expressed genes in HKC cells with at least 2 fold change and false discovery rate q value ≤ 0.05 (supplemental Table 1). There were 187 down-regulated and 236 up-regulated genes in KC-affected corneal stromal cells. After applying a minimum expression of 10 normalized reads in either cases or controls, a total of 208 genes (121 down and 87 up-regulated) were differentially expressed in HKC fibroblast cells (supplemental Table 2). The top 20 up- and down-regulated genes with largest fold changes are listed in Table 2.
Diverse and dominant pathways in HKCs
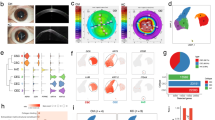
Gene ontology analysis using all 423 genes indicated the significant enrichment of genes coding for proteins involved in or related with cell migration, collagen-containing ECM, adherens junction, intrinsic to plasma membrane, cytokine and growth factor activity, growth factor binding, and kinase activity. KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analysis identified the enriched pathways including MAPK signaling and Rap1 signaling. Reactome analysis indicated the involvement of PTK6 promoting HIF1A stabilization, non-integrin membrane-ECM interactions, ECM organization, and syndecan interactions. Wikipathway analysis suggested the enrichment of genes in MAPK signaling, differentiation, and focal adhesion-PI3K-Akt-mTOR-signaling pathway. Seven genes (DAPK1, ELK4, HIF1A, MEF2C, RPS6KA3, and SGK1) are targets of MAPK7 kinase activity. Many of the differentially expressed genes are enriched targets of miR-26a/b, miR-203, miR-380-3p, and miR-96 with 28, 23, 12, and 22 target genes respectively. Our miRNA profiling in seven normal human corneal tissues39 indicated the expression of these miRNAs, suggesting their potential role in corneal function. Protein network analysis identified a subnetwork with several connection hubs or seeds, including EGFR, NEDD4, SNTA1, LGALS3BP, HSPB1, SDC2, MME, HIF1A, CBL, and ERRFI1 (Fig. 1).
With the relatively highly expressed 208 differential genes, gene ontology analysis indicated the enrichment of genes involved with cell migration/motility, focal adhesion, endoplasmic reticulum lumen, and ECM, similar to the bioinformatics findings with all 423 genes.
RT-PCR validation
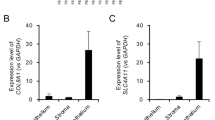
We further validated the differential gene expression by selecting seven genes from the top 20 genes identified by transcriptomics. We focused on novel pathways associated with proliferation, wound healing, and pro-inflammatory pathways. We compared our RNA-Seq results with RNA expression using RT-PCR. Of the genes tested, ANKRD1 and KRT7 were significantly down-regulated in HKCs when compared to controls (Fig. 2), consistent with our RNA-Seq findings. In contrast, EGR1, IL1B, GDF15, MME, and CXCL1 RNA expression was not significantly different between HKCs and Controls, where RNA-Seq analysis showed significant up-regulation in HKCs (Fig. 2).
Real-time PCR validation of selected genes for their differential expression from RNA-Seq. The expression of (a) ANKRD1, and (b) Keratin7 was successfully validated using RT-PCR with significant differential expression with p values < 0.0001, and 0.0365 respectively. The expression of (c) ERG1, (d) GDF15, (e) MME1, (f) IL1B, and (g) CXCL1 was not validated for their differential expression from RNA-Seq (p value > 0.05).
Discussion
Previous reports applying transcript and gene analysis approaches in KC have identified altered expression ranging from a diverse group of pathways, including expression of keratocan, ECM proteins, Wnt signaling, lysyl oxidase, and inflammatory genes12,13,14,15,16,17,18,19,20,21,22,23,24. These studies have highlighted the heterogeneity of the KC condition and the need to identify common biomarkers of KC. In this study, we investigated whether gene expression in corneal stromal cells is altered by KC. For the first time, RNA-Seq was used to compare the global gene expression of the corneal stroma derived cells from KC and healthy controls. Bioinformatics were used to delineate critical pathways and highly modulated genes were validated by RT-PCR. Our analysis indicated the potential involvement of several pathways in KC pathogenesis via stromal cells, including cell migration, cellular adherence, and ECM.
The transcriptional factor, ANKRD1, is known to function in wound healing responses in cardiac tissue40 via regulation of the ERK and TGF-β pathways, among others41,42, though its expression and role in the cornea has yet to be reported. Furthermore, knockout of ANKRD1 in the mouse has shown delayed wound healing in skin tissue, due to defects in interactions between dermal fibroblasts and the surrounding ECM43. The interplay of ANKRD1 expression and the TGF-β pathway has also been reported44,45, further establishing a functional role in wound healing46. Interestingly, in our study, we have identified a downregulation of ANKRD1 in HKCs under homeostatic conditions validated in both the RNA-Seq and RT-PCR-data, which supports the hypothesis of an abnormal wound healing phenotype in KC47,48. TGF-β is known to also regulate Keratin749. Keratin 7 is a protein coding gene, which plays a role in DNA synthesis. Krenzer and Freddo reported Keratin 7 expression in the conjunctiva50. Elder et al. described Keratin 7 in the basal and suprabasal epithelial cells of the central cornea51. However, it was not found in the central corneal epithelium52.
While Keratin 7 is not normally reported in the corneal stromal layer or in KC studies, our data shows downregulation of Keratin 7 in HKCs. This finding could indicate a role in the context of wound healing and loss of corneal transparency. Keratin 7 isoform polymerizes to form the intermediate cytoskeletal filaments that provide structure and stability to corneal epithelial cells. Thus, it is associated with cytoskeletal signaling and remodeling pathways49. Through many factors and specifically TGF-β regulation, Keratin 7 has also been implicated in cellular stress responses to wound healing and tissue repair53,54. Acting through the recruitment and activation of SMADs to regulate gene expression. We, and others, have reported the role of SMADs in KC55,56,57,58,59. Given that KC has previously been associated with an altered response to TGF-β1 and TGF-β3 ligands57,60, the downregulation of both ANKRD1 and Keratin 7 in HKCs may contribute to this complex interplay favoring a pro-fibrotic phenotype in HKCs compared to healthy controls. The functional relevance of our findings to the KC condition may be related to these downstream pathways regulated by TGF-β signaling, a key pathway important in wound healing and ECM deposition in corneal biology61. Further studies to determine the functional effects of decreased gene expression in the context of KC development or progression are warranted. Moreover, validation of protein level changes are needed to determine that altered gene expression identified in our study contributes to protein level changes.
Our study also has a few limitations. Firstly, the sample size is relatively small. Large sample size of both cases and controls with similar age range will definitely increase the statistical power to detect additional signaling pathways and networks. Secondly, we have used primary corneal stromal fibroblast cells instead of the isolated stromal layer tissue. The culture process may influence the expression profile. However, studies from our lab27,57 and others62,63 have consistently supported maintenance of an altered disease phenotype by KC-derived stromal cells cultured in vitro with characteristics parallel to the in vivo condition, including altered ECM expression and regulation9. Thirdly, the identified KC-associated signaling pathways and networks could either be the disease outcome or may actively contribute to KC pathogenesis, which is difficult to discern in a snapshot of transcript levels. A systematic approach to determine the role of specific pathways in KC development from onset to further progression to the late-stage disease is required to validate disease causation. However, this is difficult with the lack of tissue/cell isolation from early-stage KC patients. Lastly, the age differences between control and KC patients was ~13.6 years with an average age of 47.2 ± 18.3 years for controls and 60.8 ± 11.6 years for KC. The effects of age on gene expression has previously been reported with downregulation associated with collagen64 and elastin expression by fibroblasts of the skin65. Likewise, aging influences corneal structure with increased keratocyte senescence and altered ECM rigidity66. Thus, the age gap in our study between groups may have contributed to differential gene expression independent of the KC disease.
In summary, using RNA-Seq technology, we have successfully profiled the differential gene expression in primary human corneal stromal cells derived from patients with KC. This data will advance our molecular understanding of the pathogenesis of KC.
Data Availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Rabinowitz, Y. S. Keratoconus. Surv. Ophthalmol. 42, 297–319 (1998).
Khaled, M. L. et al. Molecular and histopathological changes associated with keratoconus. Biomed. Res. Int. 2017, 7803029, https://doi.org/10.1155/2017/7803029 (2017).
Godefrooij, D. A., de Wit, G. A., Uiterwaal, C. S., Imhof, S. M. & Wisse, R. P. L. Age-specific incidence and prevalence of keratoconus: a nationwide registration study. Am. J. Ophthalmol. 175, 169–172, https://doi.org/10.1016/j.ajo.2016.12.015 (2017).
U.S. Department of Defense. DoD Instruction 6130.03 Medical standards for appointment, enlistment, or induction into the military services. Available at https://www.esd.whs.mil/DD/ (2018).
Kymes, S. M., Walline, J. J., Zadnik, K. & Gordon, M. O. Quality of life in keratoconus. Am. J. Ophthalmol. 138, 527–535, https://doi.org/10.1016/j.ajo.2004.04.031 (2004).
Aydin Kurna, S., Altun, A., Gencaga, T., Akkaya, S. & Sengor, T. Vision related quality of life in patients with keratoconus. J. Ophthalmol. 2014, 694542, https://doi.org/10.1155/2014/694542 (2014).
Sahebjada, S. et al. Impact of keratoconus in the better eye and the worse eye on vision-related quality of life. Invest. Ophthalmol. Vis. Sci. 55, 412–416, https://doi.org/10.1167/iovs.13-12929 (2014).
Kymes, S. M., Walline, J. J., Zadnik, K., Sterling, J. & Gordon, M. O. Changes in the quality-of-life of people with keratoconus. Am. J. Ophthalmol. 145, 611–617, https://doi.org/10.1016/j.ajo.2007.11.017 (2008).
Bykhovskaya, Y., Margines, B. & Rabinowitz, Y. S. Genetics in Keratoconus: where are we. Eye Vis. (Lond.). 3, 16 (2016).
Sharif, R., Bak-Nielsen, S., Hjortdal, J. & Karamichos, D. Pathogenesis of keratoconus: the intriguing therapeutic potential of prolactin-inducible protein. Prog. Retin. Eye Res. 67, 150–167, https://doi.org/10.1016/j.preteyeres.2018.05.002 (2018).
Peiffer, R. L. Jr., Werblin, T. P. & Patel, A. S. Keratoconus in a rhesus monkey. J. Med. Primatol. 16, 403–406 (1987).
Wentz-Hunter, K., Cheng, E. L., Ueda, J., Sugar, J. & Yue, B. Y. Keratocan expression is increased in the stroma of keratoconus corneas. Mol. Med. 7, 470–477 (2001).
Khaled, M. L. et al. Differential Expression of Coding and Long Noncoding RNAs in keratoconus-affected corneas. Invest. Ophthalmol. Vis. Sci. 59, 2717–2728, https://doi.org/10.1167/iovs.18-24267 (2018).
Bykhovskaya, Y., Gromova, A., Makarenkova, H. P. & Rabinowitz, Y. S. Abnormal regulation of extracellular matrix and adhesion molecules in corneas of patients with keratoconus. Int. J. Keratoconus Ectatic Corneal Dis. 5, 63–70, https://doi.org/10.5005/jp-journals-10025-1123 (2016).
You, J., Wen, L., Roufas, A., Madigan, M. C. & Sutton, G. Expression of SFRP family proteins in human keratoconus corneas. PloS one 8, e66770, https://doi.org/10.1371/journal.pone.0066770 (2013).
Joseph, R., Srivastava, O. P. & Pfister, R. R. Downregulation of beta-actin gene and human antigen R in human keratoconus. Invest. Ophthalmol. Vis. Sci. 53, 4032–4041, https://doi.org/10.1167/iovs.11-9062 (2012).
Joseph, R., Srivastava, O. P. & Pfister, R. R. Differential epithelial and stromal protein profiles in keratoconus and normal human corneas. Exp. Eye Res. 92, 282–298, https://doi.org/10.1016/j.exer.2011.01.008 (2011).
Hao, X. D. et al. Decreased integrity, content, and increased transcript level of mitochondrial DNA are associated with keratoconus. PloS one 11, e0165580, https://doi.org/10.1371/journal.pone.0165580 (2016).
Shetty, R. et al. Attenuation of lysyl oxidase and collagen gene expression in keratoconus patient corneal epithelium corresponds to disease severity. Mol. Vis. 21, 12–25 (2015).
Shetty, R. et al. Elevated expression of matrix metalloproteinase-9 and inflammatory cytokines in keratoconus patients is inhibited by cyclosporine A. Invest. Ophthalmol. Vis. Sci. 56, 738–750, https://doi.org/10.1167/iovs.14-14831 (2015).
Kabza, M. et al. Collagen synthesis disruption and downregulation of core elements of TGF-beta, Hippo, and Wnt pathways in keratoconus corneas. Eur. J. Hum. Genet. 25, 582–590, https://doi.org/10.1038/ejhg.2017.4 (2017).
Mootha, V. V., Kanoff, J. M., Shankardas, J. & Dimitrijevich, S. Marked reduction of alcohol dehydrogenase in keratoconus corneal fibroblasts. Mol. Vis. 15, 706–712 (2009).
Rabinowitz, Y. S., Dong, L. & Wistow, G. Gene expression profile studies of human keratoconus cornea for NEIBank: a novel cornea-expressed gene and the absence of transcripts for aquaporin 5. Invest. Ophthalmol. Vis. Sci. 46, 1239–1246, https://doi.org/10.1167/iovs.04-1148 (2005).
Pahuja, N. et al. Differential molecular expression of extracellular matrix and inflammatory genes at the corneal cone apex drives focal weakening in keratoconus. Invest. Ophthalmol. Vis. Sci. 57, 5372–5382, https://doi.org/10.1167/iovs.16-19677 (2016).
You, J., Corley, S. M., Wen, L. & Hodge, C. RNA-Seq analysis and comparison of corneal epithelium in keratoconus and myopia patients. Sci. Rep. 8, 389, https://doi.org/10.1038/s41598-017-18480-x (2018).
McKay, T. B. et al. Quercetin attenuates lactate production and extracellular matrix secretion in keratoconus. Sci. Rep. 5, 9003, https://doi.org/10.1038/srep09003 (2015).
Karamichos, D. et al. Novel in vitro model for keratoconus disease. J. Funct. Biomater. 3, 760–775, https://doi.org/10.3390/jfb3040760 (2012).
Goncu, T. et al. Tear film and serum prolidase activity and oxidative stress in patients with keratoconus. Cornea 34, 1019–1023, https://doi.org/10.1097/ico.0000000000000510 (2015).
Karamichos, D. et al. Tear metabolite changes in keratoconus. Exp. Eye Res. 132, 1–8, https://doi.org/10.1016/j.exer.2015.01.007 (2015).
Saijyothi, A. V. et al. Tear fluid small molecular antioxidants profiling shows lowered glutathione in keratoconus. Exp. Eye Res. 103, 41–46, https://doi.org/10.1016/j.exer.2012.07.010 (2012).
McKay, T. B., Sarker-Nag, A., Lyon, D., Asara, J. M. & Karamichos, D. Quercetin modulates keratoconus metabolism in vitro. Cell Biochem. Funct. 33, 341–350, https://doi.org/10.1002/cbf.3122 (2015).
Karamichos, D. et al. In vitro model suggests oxidative stress involved in keratoconus disease. Sci. Rep. 4, 4608, https://doi.org/10.1038/srep04608 (2014).
Karamichos, D. et al. TGF-beta3 stimulates stromal matrix assembly by human corneal keratocyte-like cells. Invest. Ophthalmol. Vis. Sci. 54, 6612–6619, https://doi.org/10.1167/iovs.13-12861 (2013).
Karamichos, D., Guo, X. Q., Hutcheon, A. E. & Zieske, J. D. Human corneal fibrosis: an in vitro model. Invest. Ophthalmol. Vis. Sci. 51, 1382–1388, https://doi.org/10.1167/iovs.09-3860 (2010).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 41, W77–83, https://doi.org/10.1093/nar/gkt439 (2013).
Zhang, B., Kirov, S. & Snoddy, J. WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res. 33, W741–748, https://doi.org/10.1093/nar/gki475 (2005).
Karamichos, D., Hutcheon, A. E. K. & Zieske, J. D. Transforming growth factor-β3 regulates assembly of a non-fibrotic matrix in a 3D corneal model. J. Tissue Eng. Regen. Med. 5, e228–e238, https://doi.org/10.1002/term.429 (2011).
Priyadarsini, S. et al. Gross cystic disease fluid protein-15/prolactin-inducible protein as a biomarker for keratoconus disease. PloS one 9, e113310, https://doi.org/10.1371/journal.pone.0113310 (2014).
Drewry, M., Helwa, I., Allingham, R. R., Hauser, M. A. & Liu, Y. miRNA Profile in Three different normal human ocular tissues by miRNA-Seq. Invest. Ophthalmol. Vis. Sci. 57, 3731–3739, https://doi.org/10.1167/iovs.16-19155 (2016).
Samaras, S. E. et al. 26S proteasome regulation of Ankrd1/CARP in adult rat ventricular myocytes and human microvascular endothelial cells. Biochem. Biophys. Res. Commun. 425, 830–835, https://doi.org/10.1016/j.bbrc.2012.07.162 (2012).
Ling, S. S. M., Chen, Y. T., Wang, J., Richards, A. M. & Liew, O. W. Ankyrin Repeat Domain 1 Protein: A Functionally Pleiotropic Protein with Cardiac Biomarker Potential. Int. J. Mol. Sci. 18, https://doi.org/10.3390/ijms18071362 (2017).
Moulik, M. et al. ANKRD1, the gene encoding cardiac ankyrin repeat protein, is a novel dilated cardiomyopathy gene. J. Am. Coll. Cardiol. 54, 325–333, https://doi.org/10.1016/j.jacc.2009.02.076 (2009).
Samaras, S. E., Almodovar-Garcia, K., Wu, N., Yu, F. & Davidson, J. M. Global deletion of Ankrd1 results in a wound-healing phenotype associated with dermal fibroblast dysfunction. Am. J. Pathol. 185, 96–109, https://doi.org/10.1016/j.ajpath.2014.09.018 (2015).
Labbe, E. et al. Transcriptional cooperation between the transforming growth factor-beta and Wnt pathways in mammary and intestinal tumorigenesis. Cancer Res. 67, 75–84, https://doi.org/10.1158/0008-5472.can-06-2559 (2007).
Belgrano, A. et al. Multi-tasking role of the mechanosensing protein Ankrd2 in the signaling network of striated muscle. PloS one 6, e25519, https://doi.org/10.1371/journal.pone.0025519 (2011).
Shi, Y. et al. CARP, a cardiac ankyrin repeat protein, is up-regulated during wound healing and induces angiogenesis in experimental granulation tissue. Am. J. Pathol. 166, 303–312, https://doi.org/10.1016/s0002-9440(10)62254-7 (2005).
Cheung, I. M., McGhee, C. N. & Sherwin, T. A new perspective on the pathobiology of keratoconus: interplay of stromal wound healing and reactive species-associated processes. Clin. Exp. Optom. 96, 188–196, https://doi.org/10.1111/cxo.12025 (2013).
Cheung, I. M., McGhee, C. & Sherwin, T. Deficient repair regulatory response to injury in keratoconic stromal cells. Clin. Exp. Optom. 97, 234–239, https://doi.org/10.1111/cxo.12118 (2014).
Ranganathan, P. et al. Expression profiling of genes regulated by TGF-beta: differential regulation in normal and tumour cells. BMC Genomics. 8, 98–98, https://doi.org/10.1186/1471-2164-8-98 (2007).
Krenzer, K. L. & Freddo, T. F. Cytokeratin expression in normal human bulbar conjunctiva obtained by impression cytology. Invest. Ophthalmol. Vis. Sci. 38, 142–152 (1997).
Elder, M. J., Hiscott, P. & Dart, J. K. Intermediate filament expression by normal and diseased human corneal epithelium. Hum. Pathol. 28, 1348–1354 (1997).
Moroi, S. E. et al. Clinicopathologic correlation and genetic analysis in a case of posterior polymorphous corneal dystrophy. Am. J. Ophthalmol. 135, 461–470 (2003).
Pekny, M. & Lane, E. B. Intermediate filaments and stress. Exp. Cell Res. 313, 2244–2254, https://doi.org/10.1016/j.yexcr.2007.04.023 (2007).
Russell, D., Andrews, P. D., James, J. & Lane, E. B. Mechanical stress induces profound remodelling of keratin filaments and cell junctions in epidermolysis bullosa simplex keratinocytes. J. Cell Sci. 117, 5233–5243, https://doi.org/10.1242/jcs.01407 (2004).
Lyon, D., McKay, T. B., Sarkar-Nag, A., Priyadarsini, S. & Karamichos, D. Human keratoconus cell contractility is mediated by transforming growth factor-beta isoforms. J. Funct. Biomater. 6, 422–438, https://doi.org/10.3390/jfb6020422 (2015).
Qi, H. et al. Analysis of sphingolipids in human corneal fibroblasts from normal and keratoconus patients. J. Lipid Res. 58, 636–648, https://doi.org/10.1194/jlr.M067264 (2017).
Priyadarsini, S., McKay, T. B., Sarker-Nag, A. & Karamichos, D. Keratoconus in vitro and the key players of the TGF-β pathway. Mol. Vis. 21, 577–588 (2015).
Sharif, R., Hjortdal, J., Sejersen, H., Frank, G. & Karamichos, D. Human in vitro model reveals the effects of collagen cross-linking on keratoconus pathogenesis. Sci. Rep. 7, 12517, https://doi.org/10.1038/s41598-017-12598-8 (2017).
Engler, C. et al. Transforming growth factor-β signaling pathway activation in Keratoconus. Am. J. Ophthalmol. 151, 752–759.e752, https://doi.org/10.1016/j.ajo.2010.11.008 (2011).
Sarker-Nag, A., Hutcheon, A. E. & Karamichos, D. Mitochondrial Profile and Responses to TGF-beta Ligands in Keratoconus. Curr. Eye Res. 41, 900–907, https://doi.org/10.3109/02713683.2015.1078361 (2016).
Tandon, A., Tovey, J. C., Sharma, A., Gupta, R. & Mohan, R. R. Role of transforming growth factor Beta in corneal function, biology and pathology. Curr. Mol. Med. 10, 565–578 (2010).
Foster, J. et al. Transforming growth factor beta and insulin signal changes in stromal fibroblasts of individual keratoconus patients. PloS one 9, e106556, https://doi.org/10.1371/journal.pone.0106556 (2014).
Foster, J. W. et al. Integrated stress response and decreased ECM in cultured stromal cells from keratoconus corneas. Invest. Ophthalmol. Vis. Sci. 59, 2977–2986, https://doi.org/10.1167/iovs.18-24367 (2018).
Chung, J. H. et al. Modulation of skin collagen metabolism in aged and photoaged human skin in vivo. J. Invest. Dermatol. 117, 1218–1224, https://doi.org/10.1046/j.0022-202x.2001.01544.x (2001).
Sephel, G. C. & Davidson, J. M. Elastin production in human skin fibroblast cultures and its decline with age. J. Invest. Dermatol. 86, 279–285 (1986).
Faragher, R. G. A., Mulholland, B., Tuft, S. J., Sandeman, S. & Khaw, P. T. Aging and the cornea. Br. J. Ophthalmol. 81, 814–817, https://doi.org/10.1136/bjo.81.10.814 (1997).
Acknowledgements
We really appreciate all the donors and their families to make this project possible. We also thank the financial support from the National Eye Institute with grants EY023242 and EY028888.
Author information
Authors and Affiliations
Contributions
D.K. and Y.L. conceptualized the study, drafted and edited the paper, provided materials, and obtained funding. R.S., M.L.K., and T.B.M. drafted and edited the paper and compiled and analyzed data. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sharif, R., Khaled, M.L., McKay, T.B. et al. Transcriptional profiling of corneal stromal cells derived from patients with keratoconus. Sci Rep 9, 12567 (2019). https://doi.org/10.1038/s41598-019-48983-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-48983-8
This article is cited by
-
Identification of hub genes and molecular pathways in keratoconus by integrating bioinformatics and literature mining at the RNA level
International Ophthalmology (2024)
-
Corneal stromal regeneration—keratoconus cell therapy: a review
Graefe's Archive for Clinical and Experimental Ophthalmology (2023)
-
Molecular characteristics and spatial distribution of adult human corneal cell subtypes
Scientific Reports (2021)
-
Multi-level consistent changes of the ECM pathway identified in a typical keratoconus twin’s family by multi-omics analysis
Orphanet Journal of Rare Diseases (2020)
-
Omics Analyses in Keratoconus: from Transcriptomics to Proteomics
Current Ophthalmology Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.