Abstract
The occurrence of lymph node metastasis (LNM) and depth of tumour infiltration are significant prognostic factors in oesophageal adenocarcinoma (OAC), however no reliable prognostic biomarkers have been established so far. Aim of this study was to characterize microRNAs (miRs) of OAC patients, who primarily underwent oesophagectomy, in order to identify specific alterations during tumour progression and LNM. MicroRNA array-based quantification analysis of 754 miRs, including tumour specimens of 12 patients with pT2 OAC from three different centres (detection group), was performed. We identified miR-17, miR-19a/b, miR-20a, and miR-106a, showing the best predictive power for LNM. These miRs were validated by quantitative real time-PCR (qRT-PCR) in 43 patients with different tumour stages (pT1: n = 21; pT2: n = 12 and pT3: n = 10) (training group) (p < 0.05), demonstrating that increasing levels of identified miRs were associated with advanced depth of tumour infiltration. These findings were verified in another independent group of 46 pT2 OAC patients (validation group). Quantitative RT-PCR analysis of the miR-panel confirmed these results except for miR-19a (p < 0.05 each). Logistic regression analysis identified miR-17 and miR-20a (p = 0.025 and p = 0.022, respectively) to be independent variables for prediction of LNM. The mathematical prediction model was used in the validation group, and the estimated prognosis was compared to the actual postsurgical follow-up. This comprehensive data demonstrated the importance of miR-17-92 cluster and miR-106a for progression as well as LNM in OAC indicating that those might be feasible prognostic biomarkers.
Similar content being viewed by others
Introduction
Oesophageal adenocarcinoma (OAC) is a severe neoplasia with an overall 5-year survival rate ranging from 15% to 25%1,2. Within western countries, its incidence is still rising3,4,5,6. The occurrence of lymph node metastasis (LNM) is one of the major risk factors resulting in a poor prognosis. Pre-interventional diagnostic methods such as computer tomography (CT) or endosonographic ultrasound (EUS) are used to examine the absence of LNM. Despite the high accuracy in tumour invasion depth, the prediction of LNM is extremely poor since nodal micro-metastasis or extra-nodal lymph node metastasis are phenomena that cannot be detected by these procedures7. Since recent analysis demonstrated that LNM occurs even in early stages of OAC limited to the mucosa (0–8.1% risk) (pT1a) or submucosa (20–30% risk) (pT1b), pretherapeutic prediction of lymph node involvement is mandatory8,9,10,11,12. Nevertheless, there is a lack of pretherapeutic biomarker-based evaluation indicating an aggressive tumour biology to better identify patients at risk before loco-regional or disseminated metastasis becomes clinically evident.
MicroRNAs (miRs) are non-coding, 21–25 nucleotide small RNAs that regulate gene expression by inhibiting translation13. After transcription and splicing within the nucleus and their transfer into cytoplasm, pre-miRs are processed into the mature miRs. Depending on the level of sequence complementarity, the miRs act on specifically targeted messenger RNAs (mRNAs) either by translational repression or mRNA cleavage13,14. More than 1500 miRs have been described as being involved in the tumorigenesis of several malignancies by upregulated oncogenetic or decreased oncosuppressive effects on proliferation, tumour cell survival, migration, metastasis, or angiogenesis14,15,16,17,18. In addition, there is evidence that miRs are feasible predictors for the prognosis of oesophageal cancer19,20,21.
MiR-17, miR-19a/b, and miR-20a are members of the miR-17-92 cluster, which consists of six mature miRs (miR-17, miR-18a, miR-19a, miR-20a, miR-19b-1, and miR-92-1), also known as oncomiR-1. This cluster shows two paralogs within the human genome: miR-106a and miR-106b22. Both are overexpressed in several malignant cancers (including oesophageal cancer) and are suspected to be associated with oesophageal cancer radioresistance23,24.
Since miRs are promising biomarker candidates for prediction and therapeutic response in different neoplasms, this study aimed to characterize miR profiles of OAC patients, not treated with neoadjuvant therapy, to identify putative alterations associated with the progression or occurrence of LNM. The identified miRs were validated regarding to their predictive impact on OAC.
Results
Patient characteristics
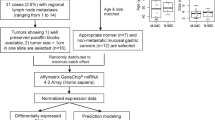
A total of 89 patients who underwent primary oesophagectomy due to OAC were included in this study and divided into three groups: detection, training and validation group. There were 12 pT2 patients with either pN0 (n = 6) or pN+ (n = 6) within the detection cohort while the training cohort included 43 patients with pT1-3 and both, pN0 and pN+ (including those patients of the detection cohort). In the independent validation cohort, only 46 pT2 patients with or without LNM were considered. Our data was comparable to those of all patients who underwent an oesophagectomy at the other three centres and histopathological baseline characteristics of all patients included within the current analysis are summarized within Table 1.
Altered miR profiles depending on the lymphatic invasion of OAC
To identify LNM-indicating candidate miRs, we profiled miR-expression of the 12 pT2 pN0/+ OAC patients (detection cohort) by using PCR-based miR analysis consisting of 754 different miRs. If miRs were detected in at least two independent tumour samples, they were considered for further analysis. 19 miRs showed the most differences in expression between patients with and without LNM (see heatmap in Fig. 1).
MicroRNA Array-based miR profiling within the detection cohort (n = 12). MiRs from 12 formalin-fixed paraffin-embedded (FFPE) samples derived from surgical pT2 oesophageal adenocarcinoma (OAC) specimens underwent miR-profiling of 754 miRs via MicroRNA Array. Intratumoural miR expression of the 19 most differently expressed miRs of OAC patients with and without lymph node metastasis (LNM) including five members of the miR-17-92 cluster (miR-17, miR-19a/b, miR-20a, and miR-106a) was visualized as a heat map.
MiR-17-92 cluster and miR-106a and their correlation with lymphatic invasion
Among the eight most significantly different markers, five miRs were members of the miR-17-92 cluster or its paralogue, miR-106a. Therefore, we focused on miR-17, miR-19a/b, miR-20a, and miR-106a for further analysis.
Single quantitative real time-PCR (qRT-PCR) examinations were performed to specifically validate those five miRs within the training cohort. Both, pT1 and pT2 OAC patients, showed a significant increase of miR levels in occurrence of LNM (p < 0.05 each, see Fig. 2a,b). However, there was only one patient with pN0 among ten pT3 patients so that reliable correlation between miR expression and status of nodal metastasis was not possible considering this tumour stage (data not shown).
Intratumoural miR expression of miR-17, miR-19a/b, miR-20a and miR-106a in correlation to lymph node status within the training cohort (n = 43). Further examination of miR-17, miR-19a/b, miR-20a, and miR-106a was performed within an enlarged training cohort of 43 pT1-3 patients (pN0 versus pN+) who underwent primary oesophagectomy. Quantitative real time-PCR (qRT-PCR) analysis of the intratumoural miR expression (a) for patients with pT1 OAC and (b) for those with pT2 OAC tumours were compared to the corresponding pN status. MiR-17, miR-19a/b and miR-20a were significantly upregulated in pT1pN+ tumours (each p < 0.05) while all five miRs showed significant increase in pT2pN+ patients (each p < 0.05).
For 43 patients with pT1-pT3 OAC of the training cohort, logistic regression analysis was performed including the five selected miRs to optimize the prediction of LNM. This statistical calculation model with backward elimination of non-significant markers showed a highly significant overall model fit (p < 0.0001) with logit (p) = −0.367 + 4.148 * miR17a −0.397 * miR20a. Independent variables for prediction of LNM were miR-17 and miR-20a (p = 0.025, respectively p = 0.022). The “area under the curve” (AUC) within the receiver operating characteristic (ROC) analysis was AUC = 0.908 (95% confidence interval: 0.782–0.974) demonstrating a very close correlation between these two miRs and nodal metastasis within the cohort (Fig. 3). The sensitivity of this model was 88.89% and the specificity 92.31% for a cut-off value of 0.632.
Design of a prediction model for lymph node metastasis (LNM) in the training cohort based on miR-17 and miR-20a. A prediction model was designed from logistic regression analysis with elimination of non-significant parameters in the training cohort (n = 43) utilizing miR-17 and miR-20a as independent predictors for pN+ (p = 0.025 and p = 0.022). ROC analysis showed AUC = 0.908 (95% confidence interval: 0.782–0.974). Sensitivity was 88.89% and specificity 92.31% for a cut-off value of 0.632. Limits of the confidence intervals were illustrated as dotted lines within the graphic illustration.
MiR-17-92 cluster and miR-106a and their correlation with depth of tumour infiltration
Analysing the results of the qRT-PCR of the training cohort revealed, that the panel of miR-17, miR19a/b, miR-20a, and miR-106a was significantly lower within early OAC limited to the mucosa or submucosa (pT1) compared to locally advanced stages such as pT2 or pT3 (see Fig. 4) (p < 0.05). Therefore, an upregulation of those miRs was associated with proceeding depth of tumour infiltration.
Intratumoural miR expression of miR-17, miR-19a/b, miR-20a and miR-106a in correlation to pT status within the training cohort (n = 43). Relative miR expression of miR-17, miR-19a/b, miR-20a and miR-106a was analysed in correlation to the pT-status within the training cohort (including pT1-3 tumours) using qRT-PCR and normalized to snoU6. MiR-17, miR-19a/b, miR-20a, and miR-106a were significantly decreased in pT1 OAC compared to locally advanced tumours (pT2-3) (each p < 0.05).
Validation of the results in an independent group of patients with pT2 OAC
An independent cohort of 46 consecutive patients with pT2 OAC (validation cohort) was used for further validation of the miR-17-92 cluster. Quantitative RT-PCR analysis of these specific miRs confirmed the results for all miRs of this panel except miR-19a (p < 0.05 each) demonstrating elevated intratumoural miR expression in patients with LNM (see Fig. 5a).
Intratumoural miR expression of miR-17, miR-19a/b, miR-20a and miR-106a in correlation to pN status within the validation cohort (n = 43). (a) Correlation between relative miR expression of miR-17, miR-19a/b, miR-20a and miR-106a and pN status of the independent 46 pT2 patients of the validation cohort confirmed upregulation of all miRs except for miR-19a in pN+ tumours using qRT-PCR (each p < 0.05). (b) Using the established prediction model for LNM on the validation cohort (n = 46) resulted in an AUC = 0.781 (95% confidence interval: 0.634–0.889) within the ROC analysis (p = 0.001). Sensitivity was 61.3% and specificity 100.0% with an optimal cut-off value of 0.725. Using the cut-off value of the training group (0.632), sensitivity was 67.7% and specificity 83.4%. Limits of the confidence intervals were illustrated as dotted lines within the graphic illustration.
In the second step, the above-mentioned prediction model of the training cohort for pN+ was applied to the validation cohort by calculating the predictive value for pN+ using the same equation as in the training group. Here, the AUC was 0.781 (95% confidence interval: 0.634–0.889) within the ROC analysis, p = 0.001 (see Fig. 5b). The sensitivity of this model was 61.3% and the specificity 100.0% with an optimal cut-off value of 0.725. The sensitivity of the model was 67.7% and the specificity 83.4% for the cut-off value of the training group of 0.632.
Prognostic relevance of the miR-17-92 cluster
LNM was significantly associated with poor prognosis in the validation cohort resulting in a 5-year survival rate of 35% in pN+ OAC compared to 65% in pN0 patients within this group (p = 0.024) (see Fig. 6a).
Prognostic impact of the miR-17 and miR-20a for lymph node metastasis (LNM) within the validation cohort (n = 46). (a) Kaplan-Meier analysis demonstrated that LNM was significantly associated with poor postsurgical outcome in those pT2 patients of the validation cohort (p = 0.024). (b) Applying the prediction model for LNM based on miR-17 and miR-20a led to a pN+ status in 24 patients versus 22 patients with pN0 within the validation cohort. Resulting Kaplan-Meier analysis of predicted postsurgical prognosis showed worse, but nonsignificant changes in survival rates for patients with estimated LNM (p = 0.068).
Applying our mathematical prediction model to the validation cohort for calculating the predictive value of pN+ subdivided the group into 22 patients without suspected LNM and another 24 patients with suspected LNM. The corresponding 5-year survival rates were estimated to be 35% for pN+ and 60% for pN0 patients. Although the survival curves using this mathematical prediction were not significantly different (p = 0.068), the results were comparable to those derived from the histopathologically verified LNM of the patients’ actual postsurgical follow-up (see Fig. 6b).
Discussion
The current study assesses the putative miR expression patterns in patients who underwent radical primary oesophagectomy due to OAC. We were able to demonstrate that the differently regulated expression of members of the miR-17-92 cluster (miR-17, miR-19a/b and miR-20a) as well as its paralogue, miR-106a, were associated with either depth of tumour infiltration or presence of LNM. Particularly, miR-17 and miR-20a were independent predictors of LNM in a mixed (pN0 and pN+) cohort of patients with pT1-pT3 OAC. Additionally, these results were confirmed in a further cohort of patients with pT2 OAC. Therefore, our findings support the role of these particular miRs as possible biomarkers for OAC. The influence of the miR-17-92 cluster has been reported in several solid and hematogenous malignancies interfering in apoptosis, angiogenesis, tumour metabolism or proliferation22,25,26,27,28,29. However, only a small amount of data exists regarding the effects of the miR-17-92 cluster in oesophageal cancer and this data mostly focuses on Asian cohorts and oesophageal squamous cell carcinoma (OSCC).
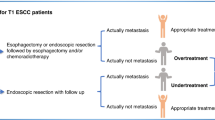
For therapeutic decision, knowledge about lymphatic infiltration is of high interest as the therapy is directly affected by a precise staging. In early local invasive cancers, an endoscopic resection can be applied in assumption of a very low risk of LNM (including early tumour stages up to pT1 sm1). However, even technically feasible, more invasive tumours (>pT1, sm2) are not considered for endoscopic resection, not because of technical aspects, but due to an uncalculated risk of LNM. A similar situation is given in advanced tumour stages (pT3 or higher) whenever a neoadjuvant treatment is recommended. With a more precise preoperative staging, in some of those patients an aggressive, potential toxic multimodal therapy could be prevented in case of exact knowledge of the lymph node status before treatment. Nowadays, imaging methods such as endoscopic ultrasound (EUS), computer tomography (CT) with/without combined positron-emission tomography (PET) or magnetic resonance imaging (MRI) are utilized for pretherapeutic evaluation. Nevertheless, all these procedures show a lack of accuracy in detecting local nodal metastases which consecutively leads to the dilemma how to clearly identify patients at risk. A recent study analysing 112 patients who either underwent primary surgery (n = 41) or neoadjuvant therapy (n = 71) due to OAC revealed a significant underestimation of the risk of LNM for EUS (p < 0.001), CT (p < 0.001) and PET/CT (p < 0.001). Accuracy, sensitivity and specificity of lymphatic metastases were 55.4%, 42.6% and 75% for EUS, 54.5%, 39.7% and 77.3% for CT and 57.1% 35.3%, and 90.9% for combined PET/CT30. Utilizing MRI technique for lymph node detection showed comparable accuracy for lymphatic involvement (66%)31. In contrast to this, Zhang et al. recently described high efficacy of EUS for detection of suspicious lymph nodes before surgical resection within their retrospective study on 112 OSCC patients32. The authors postulated an AUC value of 0.801 for EUS compared to the histopathological findings of the surgical specimens. However, EUS detection referred to morphological nodal aspects such as longitudinal diameter or echo pattern32 and therefore did not consider histological phenomena like extra-nodal metastases or nodal micro-metastases7. But not only evaluation of LNM is inaccurate by current methods, also detection of infiltration depth remains a problem especially among those patients under surveillance due to premalignant lesions such as Barrett oesophagus. A meta-analysis published by Qumseya et al. in 2018 including a total of 895 Barrett oesophagus patients from 11 studies demonstrated that the pooled false positive rate for estimating an advanced disease (pT1a/b) via EUS was 9.1% ([6.5–12.5%], p < 0.001) while the pooled false negative rate was 9.2% ([95%CI: 4.7–17.3%], p < 0.01)33. That means that almost twenty percent of Barrett oesophagus patients were wrongly evaluated and therefore received the non-optimal treatment significantly reducing the pooled accuracy of EUS to 74.6% ([58.7–85.8%], p = 0.004)33. Other studies supported these findings34,35,36. Additional (bio)markers are urgently needed. Therefore, we used miR-17 and miR-20a as independent predictors of LNM in this cohort of patients with OAC. The survival of the group with estimated LNM within the validation cohort was comparable to the actual follow-up of the group with histological proven LNM. The calculated parameters of our prediction model were as follows: AUC was 0.781 (95% confidence interval: 0.634–0.889) with p = 0.001. Specificity was 100% and sensitivity was 61.3%. Thus, our miR-based prediction model demonstrated comparable results to those clinical procedures during everyday routine.
In 2017, Liu et al. published a meta-analysis including 26 studies, which detected miR alterations in tissue, blood samples or both in a great variety of different human cancer types. They described that a high expression of the miR-17-92 cluster predicted poor overall survival in Asian patients with OSCC. Within the six Caucasian cohorts no correlation was found37. A possible explanation for those results might be the focus on different subtypes of oesophageal cancer (OSCC versus OAC) and the relatively small percentage of Caucasian patients within the meta-analysis of Liu et al. In contrast to this, we demonstrated that upregulation of members of the miR-17-92 cluster correlated with LNM in our OAC Caucasian study group with patients with pT2 OAC. The occurrence of LNM was associated with worse prognosis.
In 2014, Xu et al. found miR-17, miR-18a and miR-19a to be relevant predictive biomarkers in OSCC after analysing 105 surgical specimens and corresponding normal tissue utilizing qRT-PCR38. Concordant to our results, miR-17 overexpression correlated with the occurrence of LNM (p = 0.035) and clinical stage (p = 0.022), while miR-19a upregulation was positively associated with tumour size (p = 0.005), clinical stage (p = 0.011), and LNM (p = 0.040)38.
Summarizing those studies, there is still a lack of information considering the miR-17-92 cluster and its influence in OAC. In 2013, Wu et al. conducted a quantitative profiling of 754 human miRs in 35 normal epithelium, 34 Barrett’s oesophagus, and 36 OAC tissues, using the miR array approach39. They identified that an increase of miR-106b-3p, miR-18, miR-18-3p, miR-20b and miR-92a-1-3p were only found in OAC and was not detected in Barrett’s oesophagus tissues39. This leads to the conclusion, that the miR-17-92 cluster might drive the OAC progression, supporting our results and further demonstrating upregulation within advanced tumour stages. However, there was no comparison to normal oesophageal mucosa within this current study.
To the best of our knowledge, this is the first study illustrating miR-106a to be associated with LNM and depth of tumour infiltration within OAC, although its role in tumorigenesis has been described within other malignancies before29. Interestingly, miR-106a has been shown to be upregulated in both gastric cancer FFPE tissue samples and gastric cancer cell lines40. In vivo miR-106a expression was positively associated with LNM (p = 0.002), vascular invasion (p = 0.017), and depth of infiltration (p = 0.009). Moreover, miR-106a promoted human gastric cancer cell migration and invasion in vitro. Furthermore, the tissue inhibitor of metalloproteinase-2 (TIMP-2) was identified to be the direct downstream target of miR-106a40. A knockdown of TIMP-2 resulted in increased cell proliferation, migration, and invasion simulating the inhibitory effects of miR-106a on the expression of TIMP-2.
It is still uncertain which cellular pathways are affected by the dysregulation of the miR-17-92 cluster within OAC. Some hints revealed that Proto-oncogene serine/threonine-protein kinase Pim-1 (PIM-1) is supposed to be involved in the transcriptional regulation of the miR-17-92 cluster since silencing of this proto-oncogene caused decreased miR17-92 expression in HeLa cells41. Additionally, overexpressing miR-17-92 members resulted in the suppression of downstream targets such as tumour necrosis factor-α (TNF-α)42. Both miR-17 and miR-20a have been shown to suppress cell migration and invasion of OSCC cells by modulating the TGF-β/ITGB6 pathway43.
A few studies demonstrated the predictive value of circulating miR-18a, miR-19a and miR-20a within serum samples from OSCC patients44,45,46. Bai and co-workers analysed blood samples of healthy subjects in comparison to those patients with OSCC by qRT-PCR. Additionally, the authors compared the pre- versus postsurgical miR-19a levels showing high levels of miR-19a in cancer patients compared to normal probands and decreasing postsurgical miR-19a serum levels44. The findings of Hirajima et al. and He et al. confirmed the impact of the miR-17-92 cluster in OSCC as non-invasive biomarkers45,46. Hirajima et al. initially examined miR-18a expression in human OSCC primary tissue, OSCC cell lines, and fibroblast cell lines by performing qRT-PCR assays. They described an increased miR-18a expression in both in vivo (p = 0.0020) and in vitro (p = 0.0121) OSCC samples. In the next step, serum samples of 106 OSCC patients were compared to 54 healthy volunteers, verifying these results. When performing an ROC curve analysis, considering miR-18a for detecting OSCC patients, the AUC was 0.944946. For miR-20a, He and co-workers showed that plasma levels of miR-20a were significantly higher in OSCC patients (n = 70) than in healthy controls (n = 40), with an AUC of 0.76745. Contrary to the results of the current study, there was no significant correlation between miR-20a and the pathologic stage45. On the other hand as mentioned before, we did not include normal oesophageal tissue within our analysis and therefore cannot draw any conclusions considering the direct comparison between healthy mucosa and OAC. However, it should not be ignored that there is a limited comparability to the current study since both entities of oesophageal cancer (OAC and OSCC) seem to be biologically different.
Moreover, the miR-17-92 cluster does not only seem to be feasible for evaluation of tumour progression and occurrence of LNM, but also could influence therapeutic decisions. Within an in vitro OAC/OSCC model of acquired chemotherapy resistance, miR profiles in cisplatin or 5-fluorouracil (5-FU) resistant variants versus chemotherapy sensitive control cells were compared using microarray and qRT-PCR47. The authors demonstrated that 5-FU resistant OAC as well as OSCC cells showed downregulated levels of miR-18a-3p (among other dysregulated miRs). Additionally, miR-18a-3p was decreased in 5-FU resistant OAC cells47. Interestingly, Hummel et al. reported that these findings were consistent with the expression of the putative target mRNA: While miR-18a-3p was decreased, its targeted mRNA KRAS was increased within 5-FU resistant OSCC cells47. Recently, Lynam-Lennon et al. analysed the resistance to neoadjuvant chemoradiation therapy in OAC by utilizing a radioresistant OAC cell line (subline of OE33) and endoscopic biopsies of patients prior to neoadjuvant treatment23. Their data suggested that miR-17-5p was decreased in radioresistant cells and that upregulation of miR-17-5p sensitized those radioresistant OAC cells to radiation. In vivo miR-17-5p was significantly decreased in pretherapeutic biopsies of those patients who showed poor response to neoadjuvant chemoradiation23.
Our analysis surely has its limitations being performed retrospectively and considering only a relatively small number of patients included within the examinations. Additionally, the study patients were recruited from three centres of upper gastrointestinal surgery resulting in a certain degree of heterogeneity. Another possible limitation lies in the lack of blood samples for evaluation of plasma levels of the identified miRs, especially according the chronological sequence including pre- and postsurgical samples. Nevertheless, all histopathological results were validated by an experienced gastrointestinal pathologist to guarantee a consistent assessment. Further, follow-up was worked out comprehensively over a relatively long period of time. In the future, we recommend further validation of our results by establishing a large prospective observational trial, recruiting a larger number of patients, and implementing both liquid biopsies, as well as FFPE tumour specimens. Besides patients who undergo primary surgery, patients who recieve different neoadjuvant therapies should also be considered. Additional in vitro experiments silencing those miRs or putative downstream targets might complement ex vivo examinations.
In conclusion, upregulation of miR-17, miR-19a/b and miR-20a as members of the miR-17-92 cluster, as well as increased expression of their paralogue, miR-106a, were identified by comprehensive expression analysis to be associated with tumour progression and occurrence of LNM in human OAC, and therefore, might be feasible prognostic biomarkers for the future.
Methods
Patients
Surgical specimens analysed within the current study were derived from patients who underwent oesophagectomy between 2000 and 2010 in three centres for surgery of the upper gastrointestinal tract: (1) Department of General, Visceral and Cancer Surgery of the University Hospital of Cologne, (2) Department of General, Visceral and Transplant Surgery at the University Medical Centre of the Johannes Gutenberg-University Mainz, and (3) Department of Surgery, Horst Schmitt Kliniken Wiesbaden. The selection criteria for patients included were histologic proven OAC and primary surgery without neoadjuvant treatment. Patients with loco-regional LNM (pN+) and without (pN0) were included. The study was approved by the Institutional Ethics Committee of the University Hospital of Cologne and written informed consent was obtained from all patients. All experiments were performed in accordance with relevant guidelines and regulations.
Detection, training and validation cohort
Detection group
The initial detection cohort was comprised of 12 patients with pT2 OAC who underwent primary oesophagectomy and from whom formalin-fixed paraffin-embedded (FFPE) tumour specimen samples were collected. This group of patients (pN0: n = 6 vs pN+: n = 6) was recruited for the miR array-based approach.
Training group
The miR data of the detection cohort was tested using qRT-PCR data of 43 patients forming the training cohort (including the 12 pT2 patients of the initial detection group). Additionally, this cohort was used for calculation of a prediction model.
Validation group
The miRs detected were used for validation using qRT-PCR in an independent group of 46 consecutive OAC patients with stage pT2 who underwent oesophagectomy at the Department of General, Visceral and Cancer Surgery of the University Hospital of Cologne. The prediction model established within the training cohort was also verified on the validation group.
Staging and surgery
Routine staging diagnostics included esophagogastroduodenoscopy, endoscopic ultrasound, as well as a spiral contrast computer tomography of the thorax and abdomen.
The standard surgical procedure was laparotomic or minimally invasive gastrolysis and right transthoracic en bloc Ivor-Lewis oesophagectomy, including two-field lymphadenectomy of mediastinal and abdominal lymph nodes with gastric pull-up and intrathoracic anastomosis. Lymph nodes were removed from the resected oesophageal specimens for further histopathological examination according to a standardized protocol. The surgical procedure and lymph node preparation has been described previously9.
The patients’ outcome was followed up by regular visits within the local outpatient clinics of the participating centres.
Histopathological procedure
Oesophageal adenocarcinoma specimens and resected lymph nodes were fixed within 5% formaldehyde and embedded in paraffin afterwards. Five µm thick slides were used for haematoxylin and eosin (HE) staining. If necessary, further staining with periodic acid/Schiff of the oesophagus specimens was performed to better evaluate the depth of tumour infiltration. All specimens were histopathologically analysed and classified by an experienced gastrointestinal pathologist according to the seventh edition of the Union for International Cancer Control/TNM-classification of malignant tumours including tumour localization, depth of tumour infiltration, grading, residual tumour as well as total number of resected and infiltrated lymph nodes48.
Isolation of RNA from paraffin-embedded tissue
HE stained FFPE specimens were examined by an experienced pathologist (U.D. and A. Q.) for the presence of malignant tumour. Macrodissection was performed and a total of 60–80 µm of paraffin-embedded tissue per specimen was used for total RNA extraction. Paraffin extraction was performed by a 3-minute incubation in 320 µl of deparaffinization solution at 56 °C, followed by the addition of 240 µl of PKD buffer (Qiagen, Hilden, Germany) and 10 µl of proteinase K (Qiagen, Hilden, Germany). MiRs were isolated using miRNeasy FFPE Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. RNA quantity was estimated by A260-measurement using the ND-1000 NanoDrop spectrophotometer (NanoDrop, Wilmington, USA). 12 µl of total RNA (including miR) of 200–800 ng was isolated.
MiR profiling of the detection cohort using MicroRNA Arrays
Comparative RT-PCR-based TaqMan low-density miR arrays A and B were applied for profiling of 3 μl samples containing 1000 ng of RNA. Reverse transcription reaction of the extracted RNA was performed using Megaplex RT Primer A and B and TaqMan™ microRNA RT Kit (Thermo Fisher Scientific, Darmstadt, Germany) as previously described49. 1 µg of cDNA was loaded on TaqMan™ Human MicroRNA Arrays Cards V2.0 (Set A and B) for miR profiling of 754 miRs with snoU6 as endogenous control (Thermo Fisher Scientific, Darmstadt, Germany). Quantification was performed by TaqMan™ ABI PRISM 7900HT Sequence Detection System (Thermo Fisher Scientific, Darmstadt, Germany). A total of 100 μl master mix containing 100 ng cDNA was loaded into each of the eight ports. The distribution into 48 reaction cavities per port was carried out by two short centrifugation steps (1 min 1200 rpm in a swinging bucket rotor, Heraeus-Multifuge-3S, Langensebold, Germany). Cross contamination was avoided by individual sealing of the 384 reaction cavities. Cycling conditions were as followed: Activation step at 92 °C for 10 min, followed by 40 PCR cycles at 95 °C for 1 sec and 60 °C for 20 sec.
MiR quantification of the validation cohort by single real-time PCR
Candidate miRs identified by microRNA profiling were validated using qRT-PCR. 10 ng of whole RNA have been reverse-transcribed using cDNA by the TaqMan™ microRNA RT kit (Thermo Fisher Scientific, Darmstadt, Germany) in separate reactions made up of 3 µl of the specific primer miR-17 (ID002308), miR-19a (ID000395), miR-19b (ID000396), miR-20a (ID000580), miR-26a (ID000405), miR-106a (ID002169) and snoU6 snRNA (ID001973) as calibrator (all by Thermo Fisher Scientific, Darmstadt, Germany), 7 µl of the master mix (including 10X RT buffer, dNTP mix w/dTTp (100 M total), RNase inhibitor (20U/µL), MultiScribe™ RT enzyme (50U/µL) and nuclease-free water to scale the reaction up to a final volume of 15 µl. Parallel to this, 1 µg universal RNA from normal human tissues (# R4234565-1) (BioChain, Newark, USA) has been reverse-transcribed the same way. Reverse transcription was performed according to the instructions given by the manufacturer. Reaction conditions were 16 °C for 30 min, 42 °C for 30 min, 85 °C for 5 min and 4 °C until the end of the reaction.
Expression of miR-17, miR-19a/b, miR-20a and miR-106a was quantified using the QuantStudio 7 Flex Real-Time PCR System (Applied Biosystems, Darmstadt, Germany). The following reagents were used in single assays: miR-17 (ID002308), miR-19a (ID000395), miR-19b (ID000396), miR-20a (ID000580), miR-26a (ID000405), miR-106a (ID002169), the snoU6 snRNA (ID001973) in TaqMan Universal PCR Master Mix without AmpErase® UNG (Thermo Fisher Scientific, Darmstadt, Germany) and nuclease-free water for scaling up the volume to 20 µl. MiR levels were measured by their threshold cycle (Ct) and normalized by snoU6 RNA as a calibrator. Therefore, cycle threshold (Ct) values were calculated into relative expression by standard curves for each specific primer in 1:2 serial dilutions of the reverse-transcribed universal RNA (# R4234565-1) (BioChain, Newark, USA). 1 µl containing 1 ng of sample cDNA was used in each 20 µl qRT-PCR assay. Real-time PCR quantification of miR-17, miR-19a/b, miR-20a, miR-106a and miR-26a was carried out in triplicates according to the manufacturer’s recommendations. Thermal cycling conditions were as followed: 50 °C for 2 min, 95 °C for 10 min, followed by 40 PCR cycles at 95 °C for 15 sec and 60 °C for 1 min.
Statistical analysis
MicroRNA array data was quantified and normalized via comparative Ct-method using snoU6 as an endogenous control. Relative miR expression was calculated as ΔCt (ΔCt = Cttarget miR − CtsnoU6) for each group (pN0 versus pN+) and fold changes were computed using the 2−ΔΔCt function. QIucore Omics Explorer software V 3.2 (QIucore, Lund, Sweden) was utilized to find variables that were best in separating pN+ versus pN0, presented as heatmap. Two-group comparison (two-sided T-test) was used, identifying data with a fold change of at least 1.5 and a p-value of <0.05.
Quantitative RT-PCR of the training and validation cohort was performed using single assays in triplicates, analysed via standard curve method. Relative quantification of each target miR (R) was calculated as the ratio of the target miR related to snoU6 according to each cDNA sample.
Results of the miR expression profiles comparing pN+ vs. pN0 or pT-categories were graphically presented with Box-plots with a line representing the mean value. Correlation between nominal or ordinal parameters was tested using Chi-square test or Fisher’s Exact test. T-test or Mann–Whitney–test was utilized for comparison of two independent samples. Kaplan-Meier curves, with the log-rank test, were used for the analysis of survival prognosis. The level of statistical significance in all experiments was set to a p-value < 0.05.
Prediction of lymph node metastasis within the validation cohort was calculated by utilizing logistic regression analysis with backwards elimination of non-significant predictors for pN+ in those 43 patients with pT1, pT2 or pT3 OAC, including 18 cases with pN+ and 25 cases with pN0. Variables were either included if p < 0.05 or removed if p > 0.1. The statistical analysis was performed using the SPSS statistic program version 23 (SPSS, Chicago, USA). The graphical presentation of the designed predictive model and follow-up data was done with MedCalc Version 18.5 (MedCalc Software, Ostend, Belgium).
Ethics approval and consent to participate
The authors state that they have obtained appropriate institutional review board approval for using the surgical specimens and have followed the principles outlined in the Declaration of Helsinki for all human or animal experimental investigations. All subjects provided written informed consent.
Data Availability
The datasets generated and/or analysed during this current study are available from the corresponding author on reasonable request.
References
Thrift, A. P. The epidemic of oesophageal carcinoma: Where are we now? Cancer. Epidemiology 41, 88–95 (2016).
Lagergren, J., Smyth, E., Cunningham, D. & Lagergren, P. Oesophageal cancer. The Lancet 390, 2383–2396 (2017).
Gupta, B. & Kumar, N. Worldwide incidence, mortality and time trends for cancer of the oesophagus. Eur. J. Cancer Prev. 26, 107–118 (2017).
Rubenstein, J. H. & Shaheen, N. J. Epidemiology, diagnosis, and management of esophageal adenocarcinoma. Gastroenterology 149, 302–317.e1 (2015).
Arnold, M., Laversanne, M., Brown, L. M., Devesa, S. S. & Bray, F. Predicting the Future Burden of Esophageal Cancer by Histological Subtype: International Trends in Incidence up to 2030. Am. J. Gastroenterol. 112, 1247–1255 (2017).
Xie, S. H., Mattsson, F. & Lagergren, J. Incidence trends in oesophageal cancer by histological type: An updated analysis in Sweden. Cancer Epidemiol. 47, 114–117 (2017).
Plum, P. S. et al. Molecular markers predicting lymph node metastasis in early esophageal cancer. Histol. Histopathol. 30, 1193–1202 (2015).
Bollschweiler, E. et al. High rate of lymph-node metastasis in submucosal esophageal squamous-cell carcinomas and adenocarcinomas. Endoscopy 38, 149–56 (2006).
Hölscher, A. H. et al. Prognostic Impact of Upper, Middle, and Lower Third Mucosal or Submucosal Infiltration in Early Esophageal Cancer. Ann. Surg. 254, 802–808 (2011).
Gertler, R. et al. Prevalence and topography of lymph node metastases in early esophageal and gastric cancer. Ann. Surg. 259, 96–101 (2014).
Dubecz, A., Kern, M., Solymosi, N., Schweigert, M. & Stein, H. J. Predictors of lymph node metastasis in surgically resected T1 esophageal cancer. Ann. Thorac. Surg. 99, 1879–1886 (2015).
Griffin, S. M., Burt, A. D. & Jennings, N. A. Lymph Node Metastasis in Early Esophageal Adenocarcinoma. Ann. Surg. 254, 731–737 (2011).
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Kim, J., Yao, F., Xiao, Z., Sun, Y. & Ma, L. MicroRNAs and metastasis: small RNAs play big roles. Cancer Metastasis Rev. 37, 5–15 (2018).
Su, Y. et al. Small molecule with big role: MicroRNAs in cancer metastatic microenvironments. Cancer Lett. 344, 147–56 (2014).
Koturbash, I., Zemp, F. J., Pogribny, I. & Kovalchuk, O. Small molecules with big effects: the role of the microRNAome in cancer and carcinogenesis. Mutat. Res. 722, 94–105 (2011).
Hayes, J., Peruzzi, P. P. & Lawler, S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol. Med. 20, 460–9 (2014).
Tie, J. & Fan, D. Big roles of microRNAs in tumorigenesis and tumor development. Histol. Histopathol. 26, 1353–1361 (2011).
Hong, L. et al. Prognosis-related microRNAs in esophageal cancer. Expert Opin. Biol. Ther. 14, 483–9 (2014).
Chiam, K. et al. Circulating Serum Exosomal miRNAs As Potential Biomarkers for Esophageal Adenocarcinoma. J. Gastrointest. Surg. 19, 1208–1215 (2015).
Vrana, D. et al. Potential Predictive Role of MicroRNAs in the Neoadjuvant Treatment of Esophageal Cancer. Anticancer Res. 37, 403–412 (2017).
Concepcion, C. P., Bonetti, C. & Ventura, A. The MicroRNA-17-92 Family of MicroRNA Clusters in Development and Disease. Cancer J. 18, 262–267 (2012).
Lynam-Lennon, N. et al. MicroRNA-17 is downregulated in esophageal adenocarcinoma cancer stem-like cells and promotes a radioresistant phenotype. Oncotarget 8, 11400–11413 (2017).
Hemmatzadeh, M., Mohammadi, H., Karimi, M., Musavishenas, M. H. & Baradaran, B. Differential role of microRNAs in the pathogenesis and treatment of Esophageal cancer. Biomed. Pharmacother. 82, 509–519 (2016).
Olive, V., Jiang, I. & He, L. mir-17-92, a cluster of miRNAs in the midst of the cancer network. Int. J. Biochem. Cell Biol. 42, 1348–54 (2010).
Izreig, S. et al. The miR-17∼92 microRNA Cluster Is a Global Regulator of Tumor Metabolism. Cell Rep. 16, 1915–28 (2016).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–33 (2005).
Fuziwara, C. S. & Kimura, E. T. Insights into Regulation of the miR-17-92 Cluster of miRNAs in Cancer. Front. Med. 2, 64 (2015).
Mogilyansky, E. & Rigoutsos, I. The miR-17/92 cluster: a comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ. 20, 1603–14 (2013).
Foley, K. G., Christian, A., Fielding, P., Lewis, W. G. & Roberts, S. A. Accuracy of contemporary oesophageal cancer lymph node staging with radiological-pathological correlation. Clin. Radiol. 72, 693.e1–693.e7 (2017).
Giganti, F. et al. Prospective comparison of MR with diffusion-weighted imaging, endoscopic ultrasound, MDCT and positron emission tomography-CT in the pre-operative staging of oesophageal cancer: results from a pilot study. Br. J. Radiol. 89, 20160087 (2016).
Zhang, Y. et al. Esophageal cancer N staging study with endoscopic ultrasonography. Oncol. Lett. 17, 863–870 (2019).
Qumseya, B. J. et al. High rate of over-staging of Barrett’s neoplasia with endoscopic ultrasound: Systemic review and meta-analysis. Dig. Liver Dis. 50, 438–445 (2018).
Bartel, M. J. et al. Role of EUS in patients with suspected Barrett’s esophagus with high-grade dysplasia or early esophageal adenocarcinoma: impact on endoscopic therapy. Gastrointest. Endosc. 86, 292–298 (2017).
Shridhar, R., Huston, J. & Meredith, K. L. Accuracy of endoscopic ultrasound staging for T2N0 esophageal cancer: a National Cancer Database analysis. J. Gastrointest. Oncol. 9, 887–893 (2018).
Esophageal Cancer Study Group Participating Centers. Predictors of staging accuracy, pathologic nodal involvement, and overall survival for cT2N0 carcinoma of the esophagus. J. Thorac. Cardiovasc. Surg. 157, 1264–1272.e6 (2018).
Liu, F. et al. Prognostic role of miR-17-92 family in human cancers: evaluation of multiple prognostic outcomes. Oncotarget 8, 69125–69138 (2017).
Xu, X. L. et al. MicroRNA-17, MicroRNA-18a, and MicroRNA-19a are prognostic indicators in esophageal squamous cell carcinoma. Ann. Thorac. Surg. 97, 1037–1045 (2014).
Wu, X. et al. MicroRNA expression signatures during malignant progression from Barrett’s esophagus to esophageal adenocarcinoma. Cancer Prev. Res. (Phila). 6, 196–205 (2013).
Zhu, M. et al. MicroRNA-106a targets TIMP2 to regulate invasion and metastasis of gastric cancer. FEBS Lett. 588, 600–607 (2014).
Thomas, M. et al. Analysis of transcriptional regulation of the human miR-17-92 cluster; evidence for involvement of Pim-1. Int. J. Mol. Sci. 14, 12273–12296 (2013).
Liu, M. et al. TNF-α is a novel target of miR-19a. Int. J. Oncol. 38, 1013–22 (2011).
Jing, C. et al. MicroRNA-17/20a impedes migration and invasion via TGF-β/ITGB6 pathway in esophageal squamous cell carcinoma. Am. J. Cancer Res. 6, 1549–62 (2016).
Bai, Y. et al. Plasma microRNA-19a as a potential biomarker for esophageal squamous cell carcinoma diagnosis and prognosis. Biomark. Med. 11, 431–441 (2017).
He, F. C. et al. Expression of circulating microRNA-20a and let-7a in esophageal squamous cell carcinoma. World J. Gastroenterol. 21, 4660–4665 (2015).
Hirajima, S. et al. Clinical impact of circulating miR-18a in plasma of patients with oesophageal squamous cell carcinoma. Br. J. Cancer 108, 1822–9 (2013).
Hummel, R. et al. MicroRNA signatures in chemotherapy resistant esophageal cancer cell lines. World J. Gastroenterol. 20, 14904–12 (2014).
Sobin, L. H., Gospodarowicz, M. K. & Wittekind, C. TNM Classification of Malignant Tumours, 7th Edition. (Wiley-Blackwell, 2009).
Warnecke-Eberz, U., Chon, S. H., Hölscher, A. H., Drebber, U. & Bollschweiler, E. Exosomal onco-miRs from serum of patients with adenocarcinoma of the esophagus: comparison of miRNA profiles of exosomes and matching tumor. Tumor Biol. 36, 4643–4653 (2015).
Acknowledgements
The authors are thankful for excellent technical assistance of Michaela Heitmann, Susanne Neiß, and Anke Wienand-Dorweiler. Furthermore, the authors thank Dolores Müller for proofreading the manuscript.
Author information
Authors and Affiliations
Contributions
P.S.P. conceived, designed and performed the experiments, analysed and interpreted the data and wrote the manuscript. P.S.P., S.-H.C., H.A., I.G. and D.L. provided clinical samples. U.W.-E., R.M. and E.B. participated in the study design and coordination. E.B. performed the statistical analysis. U.D. and A.Q. performed the histopathological analysis of all surgical specimens. A.H.H. and C.J.B. participated in the interpretation of the clinical data. All authors contributed to scientific discussion and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
P. S. Plum is fellow of the Else Kröner Forschungskolleg Cologne “Clonal Evolution in Cancer” (2016-Kolleg-19). This work was funded through the “Köln Fortune” Program of the Medical Faculty Cologne (No. 219/2015 to P.S. Plum.). U. Warnecke-Eberz, U. Drebber, S.-H. Chon, H. Alakus, A.H. Hölscher, A. Quaas, C.J. Bruns, I. Gockel, D. Lorenz, R. Metzger and E. Bollschweiler declare no potential conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Plum, P.S., Warnecke-Eberz, U., Drebber, U. et al. Upregulation of miR-17-92 cluster is associated with progression and lymph node metastasis in oesophageal adenocarcinoma. Sci Rep 9, 12113 (2019). https://doi.org/10.1038/s41598-019-48624-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-48624-0
This article is cited by
-
Decoding the interaction between miR-19a and CBX7 focusing on the implications for tumor suppression in cancer therapy
Medical Oncology (2023)
-
Designing strategies of small-molecule compounds for modulating non-coding RNAs in cancer therapy
Journal of Hematology & Oncology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.