Abstract
Recently, the search for novel therapeutic agents against Acanthamoeba species has been focused on the evaluation of natural resources. Among them, marine microorganisms have risen as a source of bioactive compounds with the advantage of the ability to obtain unlimited and constant amounts of the compounds in contrast to other natural sources such as plants. Furthermore, marine actinomycetes have recently been reported as highly rich in bioactive agents including salinosporamides, xiamycines, indolocarbazoles, naphtyridines, phenols, dilactones such as antimycines and macrolides among others. In this study, staurosporine (STS) was isolated from a strain of Streptomyces sanyensis and tested against Acanthamoeba to characterize the therapeutic potential of STS against this protozoan parasite. We have established that STS is active against both stages of the Acanthamoeba life cycle, by the activation of Programmed Cell Death via the mitochondrial pathway of the trophozoite. We have also established that STS has relatively low toxicity towards a macrophage cell line. However, previous studies have highlighted higher toxicity levels induced on other vertebrate cell lines and future research to lower these toxicity issues should be developed.
Similar content being viewed by others
Introduction
Acanthamoeba genus belongs to the Free-Living Amoebae (FLA) group and includes some strains which are able to cause opportunistic infections in humans and other animals such as encephalitis and keratitis1. Current therapeutic approaches against Acanthamoeba infections involve the use of a combination of drugs which is highly toxic for the patient and not fully effective. In the case of Acanthamoeba Keratitis (AK), most of the commonly used topical agents such as biguanides are used to treat this pathogen1,2 but these compounds are unfortunately toxic to human corneal cells. Other compounds used against AK have shown activity but are not available worldwide, hence the problem of treatment continues.
In addition to the therapeutic issues mentioned above, Acanthamoeba is able to form a highly resistant cyst stage which complicates the development of fully effective therapeutic agents against this pathogenic protozoa3,4,5.
The search for novel compounds against Acanthamoeba species has been lately focused in the evaluation of natural sources presenting anti-Acanthamoeba activity2,6,7. Among these sources, recent reports have focused on bioactive compounds from marine microorganisms. Mangrove ecosystems in particular, have attracted recently the attention of the scientific community due to the discovery of new drugs with pharmaceutical potential. Mangroves are singular ecosystems, characterized for their high biodiversity and the feasibility to collect samples. More than 200 endophytic fungi and over 2000 actinomycetes have been collected from mangroves, from which a wide array of identified secondary metabolites have displayed a variety of pharmacological properties including antimicrobial, anticancer and antiviral ones among others8.
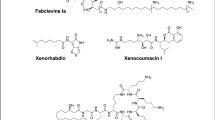
Many secondary metabolites have been isolated from marine actynomycetes such as salinosporamides, xiamycines, indolocarbazoles, naphtyridines, phenols, dilactones such as antimycines, macrolides, etc9,10. Among actinomycetes, the genus Streptomyces has gained particular attention in this research field for its richness in bioactive molecules. In particular, the species Streptomyces sanyensis, isolated from a mangrove sediment11, was first identified in 2011. Cloning and chemical studies confirmed S. sanyensis as a prolific producer of indolocarbazole (ICZ) alcaloids12,13. The large family of ICZs is characterized by their strong inhibitory effect against many cancer cell lines, platelet aggregation inhibitor, antibacterial, antiviral and neuroprotective properties14,15. Staurosporine (STS, Fig. 1), isolated from Streptomyces staurosporeus14 in 1977, was the first reported ICZ. In addition, STS is a potent affinity inhibitor of protein kinases which blocks the ATP-binding site of the enzimes16 and it has been demonstrated that STS induces apoptosis by activation of caspase-317 in higher eukaryotes.
In the present work, we have identified and developed cultures of the strain Streptomyces sanyensis PBLC04, a high-yield producer of STS, and tested the activity of this alkaloid against Acanthamoeba genus with the aim to characterize its therapeutic potential.
Results
Culture and molecular identification of Streptomyces sp. PBL04
The strain Streptomyces sp. PBLC04 showed a high growth rate in 75% seawater-based media, with concave and dry colonies presenting grey aerial mycelia with smooth borders when cultured on plate (Fig. 2). Microscopic examination revealed pseudomycelia and gram-positive rod-shaped bacteria and supported the identification at genus level. Metabolic production of the strain was detected after 7 days of cultivation at 30 °C, 120 rpm, in a flask containing 75% seawater-based media. DNA sequencing of the obtained PCR fragments of Streptomyces sp. PBLC04, yielded two sequences which showed 100% similarity, for this reason only one was included in the phylogenetic analysis (Genbank Accession Number MK639686). Phylogenetical analyses demonstrated that Streptomyces sp. PBLC04 was included in one clade with bootstrap values 100 NJ/87 ML containing the type sequence 219820 of Streptomyces sanyensis. The pairwise distance for both sequences corresponds to 0.37%. The sequences from Strain PBLC04 compared with the other sequences into this clade corresponded to ≤0.22% (1360 bp base pair compared). The alignments used can be obtained from TreeBASE (http://www.treebase.org/) under accession number S24696.
Isolation and characterization of STS from Streptomyces sanyensis PBLC04
Culture of Streptomyces sanyensis PBLC04 produced STS in great amounts (134 mg), which represents 1.06% of the total content of the biomass extract. The isolation of STS was carried out through three chromatographic steps which involved: gel-filtration on Sephadex LH-20, flash reversed phase C18 with 5 mM ammonium acetate (NH4OAc) buffer at pH 6.0 and Si-gel chromatography. Pure STS was separated and crystallized (28.4 mg) as yellow needles from dichloromethane (DCM).
(+)-Staurosporine possesses the molecular formula C28H24N4O3 as determined from the HRESIMS peak found at m/z 489.1908 (calc. for 489.1897 [M + Na]+), and [α]D20 + 43 (c 0.3, CHCl3). Its UV spectrum (CHCl3) λmax (log ε) 297 (4.48) nm, showed characteristic peaks of the ICZ chromophore18. The IR absorption bands at 2925, 2360, 2341, 1681, 1455, 1353 and 1318 cm−1 agreed the presence of an aromatic nucleus, a carbonyl group belonging to the indolocarbazole and a lactam ring18. Comparison of 1H and 13C NMR spectral data (see supplementary material) with those previously reported confirmed the structure of STS19,20.
STS eliminates Acanthamoeba trophozoites and cysts at low concentrations and presents low cytotoxicity levels
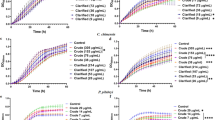
Staurosporine was able to kill Acanthamoeba trophozoites when incubated in vitro in a dose dependent way. The obtained results using the mentioned colorimetric assay in the material and methods section allowed us to establish IC50 and IC90 values of 0.265 ± 0.057 and 1.27 ± 0,007 µg/mL (0.568 ± 0.122 µM and 2.70 ± 0.015 µM) respectively (Fig. 3). Interestingly, the effects of this compound against the trophozoite stage were seen even at 15 min post incubation with the drug (Fig. 4).
Acanthamoeba castellanii Neff trophozoite incubated with various concentrations of STS: 2.5 µg/ml (5.3 µM) (A 20X, C 40X) and 1.25 µg/ml (2.6 µM) (B 20X). Images are representative of the cell population observed in the performed experiments. Negative control (Amoebae alone) (D 20X) Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA. Acanthamoeba castellanii Neff cysts incubated with 25 µg/ml (53.5 µM) of STS (A), 3.125 µg/ml (6.7 µM) (B) and 0.781 µg/ml (1.6 µM) (C) compared to the negative control (D). Images (20X) are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA.
In the case of cysticidal activity, STS was also active at low concentrations and dose-dependent, being the calculated IC50 of 0.771 ± 0.008 µg/mL (1.653 ± 0.017 µM) (Fig. 5).
Acanthamoeba castellanii Neff cysts incubated with 25 µg/ml (53.5 µM) of STS (A), 3.125 µg/ml (6.7 µM) (B) and 0.781 µg/ml (1.6 µM) (C) compared to the negative control (D). Images (20X) are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA.
Moreover, amoebic forms were observed to present dramatic morphological changes such enlargement of the cytoplasm and loss of cellular structure (lack of amoeboid form), again after 15 min of incubation with the compound (Fig. 6).
Observed morphological changes in Acanthamoeba castellanii Neff trophozoites incubated with 20 µg/ml (42.8 µM) of STS at 15 min (A) and 1 h (B) compared to the negative control (Amoebae alone) (C). Images are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA (40X). Acanthamoeba castellanii Neff trophozoites incubated with IC90 of STS and the evolution of chromatin condensation observed for 30 min, 24 h, 48 h, 72 h and 96 h. Hoechst stain is different in control cells, where uniformly faint-blue nuclei are observed, and in treated cells, where the nuclei are bright blue. Red fluorescence corresponds to the propidium iodide stain. Images (10X) are showing chromatin condensation (blue) in Acanthamoeba treated cells. Images (10X) are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA.
Furthermore, toxicity assays against the murine macrophage J774.A1 cell line yielded an CC50 of 4.076 ± 0.335 (8.737 ± 0.718 µM). Therefore, the toxicity was considered low when compared to the obtained Acanthamoeba inhibitory concentrations of STS.
STS treated amoebae stained positive in the double stain assay
After performance of the double staining protocol, it was observed that STS at both concentrations of IC50 and IC90 could induce chromatin condensation. Treated amoebae showed bright-blue stained nuclei even at 24 h after incubation with the compound as shown in Fig. 7.
Acanthamoeba castellanii Neff trophozoites incubated with IC50 (B) and IC90 (A) of STS and the evolution of chromatin condensation observed for 30 min, 24 h, 48 h, 72 h and 96 h. Hoechst stain is different in control cells (C), where uniformly faint-blue nuclei are observed, and in treated cells, where the nuclei are bright blue. Red fluorescence corresponds to the propidium iodide stain. Images (40X) are showing chromatin condensation (blue) in Acanthamoeba treated cells. Images (40X) are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA.
STS caused plasma membrane permeability in treated cells
Amoebae treated with the IC90 of STS induced plasmatic membrane damage after 24 h of incubation as illustrated in Fig. 8. Moreover, it is also important to mention that even though the membrane was damaged, cell integrity was maintained.
Permeation of the Acanthamoeba Neff plasma membrane to the vital dye SYTOX green caused by addition of STS IC90 after 24 h (A). Negative Control (B). Images (40X) are representative of the cell population observed in the performed experiments. Images were obtained using an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA.
STS induced amoebic mitochondrial malfunction
Figures 9 and 10, show STS induced changes on the mitochondrial potential since the JC-1 dye remained in the cytoplasm in its monomeric form, shown as green fluorescence. Furthermore, the mitochondrial damage was also checked by measuring the ATP level generated in 24 h. The observed results showed that STS IC90 treated cells presented a 20% decrease of ATP levels when compared to untreated cells.
The effect of IC50 (B) and IC90 (A) on the mitochondrial potential, JC-1 dye accumulates in the mitochondria of healthy cells as aggregates (C) (red fluorescence) in cells treated with the IC90 of STS for 24 h, due to collapse of mitochondrial potential, the JC-1 dye remained in the cytoplasm in its monomeric form, green fluorescence. (Images are representative of the population of treated amoeba 20X).
STS increases Reactive Oxygen Species (ROS) levels in Acanthamoeba
STS treated amoebae caused increased levels of ROS when treated with IC90 after 24 h of incubation (Fig. 11).
Increased levels of ROS in Acanthamoeba Neff caused by addition of STS IC90 after 24 h (A). Negative Control (B) cells exposed to CellROX Deep Red in absence of STS. Images (63X) are representative of the cell population observed in the performed experiments. Cells were observed in a Leica TSC SPE- confocal microscope.
Discussion
Marine microorganisms, in particular actinobacteria, are responsible for the production of approximately half of the known bioactive secondary metabolites: mainly antibiotics, antitumor agents, and immunosuppressants21. The biotechnological potential of marine microorganisms provides the additional advantage in providing a continuous supply of the compound of interest, and to avoid drawbacks such as seasonal occurrence of metabolites, usually linked to other natural sources such as seaweeds or marine invertebrates. In the present study, the strain Streptomyces sanyensis PBLC04, isolated from a mangroove ecosystem in Ecuador, has allowed us the isolation of STS as the major compound when cultured under laboratory conditions. Although STS was first identified and isolated from a strain of S. staurosporeus in 197714, to the best of our knowledge, this is the first time that this molecule is assayed against Acanthamoeba genus.
Regarding the obtained activities against both the trophozoite and cyst stage of Acanthamoeba, it is important to highlight that STS was active at low concentrations when compared to other commonly used drugs against Acanthamoeba such as chlorhexidine (IC50 = 1.10 ± 0.01 µM) amphotericin B (IC50 = 65.98 ± 5.91 µM) or PHMB (0.02%)22,23. Moreover, as it was mentioned in the results section, toxicity values when tested against a macrophage cell line were low in comparison to the obtained IC50 and IC90 with values 4-fold higher than the STS IC50 against cysts. However, previous research has shown higher toxicity values after 12 h of exposure to STS24,25,26.
Programmed Cell Death (PCD) has been previously reported in our laboratory in Acanthamoeba genus after incubation with compounds such as statins and voriconazole among other molecules2,27,28. The process of PCD involves multiple changes such as chromatin condensation, cell shrinkage and loss of mitochondrial potential among other phenomena29.
In the present study, STS was shown to induce plasma membrane damage, chromatin condensation, collapse of ATP level and mitochondrial membrane potential as well as increased levels of ROS. Since these effects were observed even after 15 min of incubation with the compound, it is probable that STS induces damages at the cellular membrane level, entering the cytoplasm without necrotic effects. We argue STS induces apoptosis in Acanthamoeba through the intrinsic pathway since the mitochondrial potential was collapsed even at 24 h (as well as ATP levels) (Fig. 12). PCD is also indicated by the generation of ROS species and condensation of DNA.
Acanthamoeba castellaniii Neff trophozoite (63X) stained with Hoechst and CellROX Deep Red after 24 h incubation of amoebae with IC90 of staurosporine (A). A bright condensed nucleus is observed as well as bright blue staining. Moreover, red dots mark regions of the cytoplasm where ROS species were generating. Blue channel for Hoechst (B). Red channel for CellROX Deep Red (C). Cells were observed in a Leica TSC SPE- confocal microscope.
The family of proteins kinases are considered to be the major target of STS. Furthermore, this molecule has demonstrated antiproliferative activity in several human cancer cell lines, inducing apoptosis by activation of caspase-330. These findings in higher eukaryotic cells support the conclusions of our study. The genome of this amoeba31 is known to be rich in kinase genes and this may be the reason why it is so sensitive.
Conclusion
In conclusion, Streptomyces sanyensis strain PBLC04 produces STS as the major metabolite. Evaluation of the activity of this compound against Acanthamoeba trophozoites and cysts, have shown a high activity of this compound. Moreover, STS-treated amoebae started apoptosis via the mitochondrial pathway. Therefore, STS is presented as a novel, highly active, PCD inducer and low toxic anti-Acanthamoeba compound at least in vitro against the used macrophage cell line which should be exploited for the development of novel therapeutic agents. However, due to the observed toxicity levels in previous studies using other cell lines such as human endothelial cell lines and even the eye, future research should focus on the development of improved delivery of STS in the host tissue which allow to lower toxicity.
Methods
General experimental procedures
Optical rotations were measured in CH2Cl2 on a PerkinElmer 241 polarimeter by using a Na lamp. NMR spectra were recorded on a Bruker AVANCE 600 MHz instrument equipped with a 5 mm TCI inverse detection cryoprobe. NMR spectra were obtained dissolving samples in CD2Cl2 (99.9%) and chemical shifts are reported relative to solvent (δH 5.32 and δC 54.0 ppm). Standard Bruker NMR pulse sequences were utilized. HR-ESI-MS data were obtained on an LCT Premier XE Micromass spectrometer. IR spectra were recorded on a Perkin-Elmer Spectrum BX spectrometer. UV spectra was recorded in a Jasco V-560 UV/Vis spectrophotometer. EnSpire Multimode Reader (Perkin Elmer) using absorbance values of Alamar Blue reagent. TLC (Thin layer chromatography) (was visualized by UV light (254 and 365 nm).
Materials
Ammonium acetate (NH4OAc, LiChropur, Merck, Germany), Methylene chloride-d2 (Euriso-top, 99.90%D, UK), Methanol-d4 (water < 0.03%, euriso-top), RP18-prepacked cartridge (25–40 µm 70 g, Götec-Labortechnik GmbH), Sephadex LH-20 (Sigma-Aldrich), Starch, yeast extract and Proteose peptone from DIFCO, Calcium carbonate (Merck), Potassium bromide (Merck), Ferric sulfate (Sigma-Aldrich), TLC plates Silica gel G60 F254-Merck.
Collection and characterization of Streptomyces sanyensis PBLC04 strain
Streptomyces sanyensis PBLC04 strain was collected in Jambelí mangrove (3°15′792″S, 80°00′739″W - 03°17′711″S, 80°01′924″W), Ecuador. The strain is part of the microbial collection of Universidad Técnica Particular de Loja (UTPL, Loja-Ecuador) and the methodology for extraction, culturing and isolation of pure strains from mangrove sediments was performed as described by Cartuche L. et al.32.
DNA isolation, PCR and phylogenetic analysis
The DNA was extracted with PureLink Genomic DNA Mini Kit (Invitrogen) for gram-positive bacteria, according to manufacturing specifications. Partial 16S rDNA region was amplified by using the universal primers for bacteria: 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492R (I) (5′-GGTTACCTTGTTACGACTT-3′)33. The PCR protocol used was: initial denaturation at 98 °C for 30 s, followed by 30 cycles of 98 °C denaturation for 10 s, annealing at 60 °C for 20 s, extension at 72 °C for 30 s and a final extension at 72 °C for 10 min. The volume of reaction was 20 μL, 1 ρmol each primer and 1U of Finnzymes’ Phusion High-Fidelity DNA Polymerase of final concentration. PCR products were visualized on 1% (w/v) agarose gels stained with 5% (v/v) GelRed (Biotium, Hayward, USA). The purified products were sequenced at Macrogen (Seoul – Republic of Korea).
The sequence chromatograms were verified using the software Codon Code Aligner 5.1.4 (CodonCode Corporation, Centerville, MA, USA). The obtained sequences were aligned with sequences available at the International Nucleotide Sequence Database Collaboration (INSDC) (http://www.insdc.org/). All 16S rDNA sequences were aligned using the G-INS-i strategy as implemented in MAFFT V734. Three phylogenetic trees were built using a Neighbour-Joining (NJ) analysis with BIONJ modification of the NJ algorithm35, Maximum Likelihood (ML) based on 1000 bootstrap replications36 and GTRMIX as DNA substitution model37 implemented in MEGA v.7 software38.
Fermentation conditions and culture extraction
Streptomyces sanyensis PBLC04 strain was cultured in 30 L of a modified seawater-based medium (A1) consisting of 75% seawater containing 10 g starch, 4 g yeast extract, 2 g proteose peptone, 1 g calcium carbonate, supplemented with 5 mL/L of a solution of potassium bromide (67 mM) and ferric sulfate (20 mM)39. Fernbach flasks with 1 L each were kept at 30 °C and mixed in an orbital shaker at 200 rpm for 7 days.
Cultures of S. sanyensis PBLC04 were centrifuged at 5000 rpm for 10 min to separate supernatant from cell biomass. Amberlite XAD7-HP resin (20 g/L) were added to the supernatant and stirred for 3 h to adsorb the excreted metabolites. After filtration, both the resin and the biomass pellet were separately macerated with a mixture of MeOH:AcOEt:Acetone (2:7:3) for 12 h at 120 rpm. Then, the two resulting extracts were filtered and dried at reduced pressure at 30 °C in a rotary evaporator.
Isolation of STS from biomass extract
The salt-free biomass extract (12.6 g) was fractionated by size-exclusion chromatography on Sephadex-LH-20 with methanol as eluent to yield 10 fractions that were finally gathered in four final fractions according to TLC and 1H-NMR analysis. Fractions 3 and 4 exhibited characteristic signals for ICZs in the 1H-NMR spectrum (MeOH-d4)20. Fraction 4 (716.0 mg) was separated by flash chromatography on a RP18 prepacked cartridge (25–40 µm, 70 g, Götec-Labortechnik GmbH) with a step gradient elution system of H2O:MeOH, 5 mM NH4OAc, (20% to 100% MeOH) at 4 mL/min flow and UV detection at 254 nm, to afford seven fractions according to TLC and 1H-NMR analysis. The active fraction (124.0 mg) was further purified by elution on Si-60 open column (230–400 mesh, 60 Å) using CHCl3:MeOH (9:1) as elution system to obtain 106.0 mg. Pure STS was separated and crystallized (28.4 mg) as yellow needles using DCM.
In vitro drug sensitivity assay
Strain used
The anti-Acanthamoeba activity of STS was evaluated against the Acanthamoeba castellanii Neff (ATCC 30010) type strain from the American Type Culture6,7. This strain was grown axenically in PYG medium (0.75% (w/v) proteose peptone, 0.75% (w/v) yeast extract and 1.5% (w/v) glucose) containing 40 μg gentamicin ml−1 (Biochrom AG, Cultek, Granollers, Barcelona, Spain).
In vitro effect against the trophozoite stage of acanthamoeba
The anti-Acanthamoeba activities of the tested compound was determined by the Alamar Blue assay as previously described2,27. Briefly, Acanthamoeba strains were seeded in duplicate on a 96-well microtiter plate with 50 μL from a stock solution of 104 cells/mL. Amoebae were allowed to adhere for 15 min and 50 μL of serial dilution series of the tested compound was added. Finally, the Alamar Blue Assay Reagent (Bioresource, Europe, Nivelles, Belgium) was added into each well at an amount equal to 10% of the medium volume. The plates were then incubated for 96 h at 28 °C with a slight agitation and the emitted fluorescence was examined with an Enspire microplate reader (PerkinElmer, Massachusetts, USA) at 570/585 nm.
In vitro effect against the cyst stage of acanthamoeba
The cysticidal activity of STS was determined by the Alamar Blue assay and confirmed visually by inverted microscopy. Cysts of A. castellaniii Neff were prepared as previously described40. Briefly, trophozoites were transferred from PYG medium based cultures (trophozoite medium) to Neff´s encystment medium (NEM; 0.1 M KCl, 8 mM MgSO4·7H2O, 0.4 mM CaCl2·2H2O, 1 mM NaHCO3, 20 mM ammediol [2-amino-2-methyl-1,3-propanediol; Sigma Aldrich Chemistry Ltd., Madrid, Spain], pH 8.8, at 25 °C) and were cultured in this medium with gently shaking for a week in order to obtain mature cysts. After that, mature cysts were harvested and washed twice using PYG medium.
A serial dilution of the tested molecules was made in PYG. The in vitro susceptibility assay was performed in sterile 96-well microtiter plates (Corning™). To these wells, containing 50 µL of drug dilutions, 5·104 mature cysts of Acanthamoeba/mL were added. After 7 days of incubation with the drugs, the plate was centrifuged at 3000 rpm for 10 min. The supernatant was removed and replaced with 100 µL of fresh medium PYG in each well. Finally, 10 μL of the Alamar Blue Assay Reagent (Biosource, Europe, Nivelles, Belgium) was placed into each well, and the plates were then incubated for 144 h at 28 °C and the emitted fluorescence was periodically examined with an Enspire microplate reader (PerkinElmer, Massachusetts, USA) at 570/585 nm.
Cytotoxicity assays
Cytotoxicity of STS was evaluated after 24 h incubation of murine macrophage J774.A1 cell line (ATCC # TIB-67) with different concentration of the tested compound at 37 °C in a 5% CO2 humidified incubator. The viability of the macrophages was determined with the Alamar Blue assay as previously described2.
Double-stain assay for programmed cell death determination
A double-stain apoptosis detection kit (Hoechst 33342/PI) (GenScript, Piscataway, NJ, USA) and an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA were used. The experiment was carried out by following the manufacturer’s recommendations, and 105 cells/well were incubated in a 24-well plate for 24 h with the previously calculated IC90. The double-staining pattern allows the identification of three groups in a cellular population: live cells will show only a low level of fluorescence, cells undergoing PCD will show a higher level of blue fluorescence (as chromatin condenses), and dead cells will show low-blue and high-red fluorescence (as the Propidium Iodide stain enters the nucleus).
Intracellular ROS production using CellROX deep red staining
The generation of intracellular ROS was detected using the CellROX Deep Red fluorescent probe (Invitrogen). The cells were treated with IC90 of STS for 24 h and exposed to CellROX Deep Red (5 μM, 30 min) at 26 °C in the dark. Cells were observed in a Leica TSC SPE- confocal microscope equipped with inverted optics at λexc = 633 and λem = 519 nm.
Analysis of mitochondrial membrane potential
The collapse of an electrochemical gradient across the mitochondrial membrane during apoptosis was detected with the JC-1 mitochondrial membrane potential detection kit (Cell Technology. After being treated with IC90 of the test solution for 24 h, the cells were centrifuged (1000 r.p.m. ×10 min) and resuspended in JC-1 buffer. Images were taken on an EVOS FL Cell Imaging System AMF4300, Life Technologies, USA from Life Technologies (Madrid, Spain). The staining pattern allows the identification of two groups in a cellular population: live cells will show only red fluorescence; cells with low mitochondrial potential, (undergoing PCD) will show a higher level of green and red fluorescence.
Measurement of ATP levels
ATP level was measured using a CellTiter-Glo Luminescent Cell Viability Assay. The effect of the drug on the ATP production was evaluated by incubating (105) of cells/ml with the previously calculated IC50 and IC90 of STS.
Plasma membrane permeability
The SYTOX Green assay was performed to detect alterations of the membrane permeability in treated cells. Briefly, 105 trophozoites were washed and incubated in saline solution with the SYTOX Green at a final concentration of 1 μM (Molecular Probes) for 15 min in the dark. Subsequently STS solution was added (IC90). After 24 h of treatment, cells were observed in a Leica TSC SPE- confocal microscope equipped with inverted optics at λexc = 482 nm and λem = 519 nm9.
References
Lorenzo-Morales, J., Naveed, A. K. & Walochnik, J. An Update on Acanthamoeba Keratitis: Diagnosis, Pathogenesis and Treatment. Parasite. 22, 10, https://doi.org/10.1051/parasite/2015010 (2015).
Sifaoui, I. et al. Programmed cell death in Acanthamoeba castellanii Neff induced by several molecules present in olive leaf extracts. PloS one. 12, e0183795, https://doi.org/10.1371/journal.pone.0183795 (2017).
Khan, N. A. Pathogenesis of Acanthamoeba infections. Microbial pathogenesis 34, 277–285, https://doi.org/10.1016/S0882-4010(03)00061-5 (2003).
Khan, N. A. Acanthamoeba invasion of the central nervous system. International Journal for Parasitology 37, 131–138, https://doi.org/10.1016/j.ijpara.2006.11.010 (2007).
Lorenzo-Morales, J. et al. Acanthamoeba keratitis: an emerging disease gathering importance worldwide? Trends in parasitology 29, 181–187, https://doi.org/10.1016/j.pt.2013.01.006 (2013).
Chiboub, O. et al. In vitro amoebicidal and antioxidant activities of some Tunisian seaweeds. Experimental parasitology 183, 76–80, https://doi.org/10.1016/j.exppara.2017.10.012 (2017).
García-Davis, S. et al. Anti-Acanthamoeba Activity of Brominated Sesquiterpenes from Laurencia johnstonii. Mar. Drugs 16(11), 443, https://doi.org/10.3390/md16110443 (2018).
Gomes, N., Cleary, D., Calado, R. & Costa, R. Mangrove bacterial richness. Commun. Integr. Biol. 4, 419–423, https://doi.org/10.4161/cib.4.4.15253 (2011).
Xu, D. B., Ye, W. W., Han, Y., Deng, Z. X. & Hong, K. Natural Products from Mangrove Actinomycetes. Mar. Drugs. 12, 2590–2613, https://doi.org/10.3390/md12052590 (2014).
Ancheeva, E., Daletos, G. & Proksch, P. Lead Compounds from Mangrove-Associated Microorganisms. Mar Drugs. 16, 319, https://doi.org/10.3390/md16090319 (2018).
Sui, J. L. et al. Streptomyces sanyensis sp. nov., isolated from mangrove sediment. Int J Syst Evol Microbiol. 61, 1632–1637, https://doi.org/10.1099/ijs.0.023515-0 (2011).
Li, T. et al. Cloning, Characterization and Heterologous Expression of the Indolocarbazole Biosynthetic Gene Cluster from Marine-Derived Streptomyces sanyensis FMA. Marine Drugs. 11, 466–488, https://doi.org/10.3390/md11020466 (2013).
Fu, P. et al. Streptocarbazoles A and B, two novel indolocarbazoles from the marine-derived actinomycete strain Streptomyces sp. FMA. Org. Letters 14, 2422–2425, https://doi.org/10.1021/ol3008638 (2012).
Omura, S. et al. A new alkaloid AM-2282 OF Streptomyces origin. Taxonomy, fermentation, isolation and preliminary characterization. J. Antibiot. 30, 275–282, https://doi.org/10.7164/antibiotics.30.275 (1977).
Syuichi, O., Mikiji, K., Hiroyuki, T., Noboru, T. & Hideo, S. Staurosporine, a Potent Platelet Aggregation Inhibitor from a Streptomyces Species. Agric Biol Chem 50, 2723–2727, https://doi.org/10.1080/00021369.1986.10867821 (1986).
Karaman, M. W. et al. A quantitative analysis of kinase inhibitor selectivity. Nat. Biotechnol. 26, 127–132, https://doi.org/10.1038/nbt1358 (2008).
Chae, H. J. et al. Molecular mechanism of staurosporine-induced apoptosis in osteoblasts. Pharmacological Research 42, 373–381, https://doi.org/10.1006/phrs.2000.0700 (2000).
Meksuriyen, D. & Cordell, G. A. Biosynthesis of Staurosporine, 2. Incorporation of Tryptophan. J. Nat. Prod. 51(5), 893–899, https://doi.org/10.1021/np50059a013 (1988).
Yang, S.-W. & Cordell, G. A. Biosynthesis of Staurosporine: Incorporation of Glucose. J. Nat. Prod. 59(9), 828–833, https://doi.org/10.1021/np960109d (1996).
Meksuriyen, D. & Cordell, G. A. Biosynthesis of Staurosporine, 1. 1H- and 13C-NMR Assignments. J. Nat. Prod. 51(5), 884–892, https://doi.org/10.1021/np50059a012 (1988).
Dewapriya, P. & Kim, S. K. Marine microorganisms: An emerging avenue in modern nutraceuticals and functional foods. Food Research International 56, 115–125 (2014).
Taravaud, A., Loiseau, P. M. & Pomel, S. In vitro evaluation of antimicrobial agents on Acanthamoeba sp. and evidence of a natural resilience to amphotericin B. Int J Parasitol Drugs Drug Resist 7, 328–336, https://doi.org/10.1016/j.ijpddr.2017.09.002 (2017).
Lim, N. et al. Comparison of polyhexamethylene biguanide and chlorhexidine as monotherapy agents in the treatment of Acanthamoeba keratitis. Am J Ophthalmol 145, 130–135, https://doi.org/10.1016/j.ajo.2007.08.040 (2008).
Rock, N. & Chintala, Y. N. Mechanisms regulating plasminogen activators in transformed retinal ganglion cells. Exp Eye Res. 86, 492–499 (2008).
Andersson, M. et al. Caspase and proteasome activity during staurosporin-induced apoptosis in lens epithelial cells. Invest Ophthalmol Vis Sci. 41, 2623–2632 (2000).
Thuret, G. et al. Mechanisms of staurosporine induced apoptosis in a human corneal endothelial cell line. Br J Ophthalmol. 87, 346–352 (2003).
Martín-Navarro, C. M. et al. Statins and voriconazole induce programmed cell death in Acanthamoeba castellanii. Antimicrob Agents Chemother 59, 2817–2824, https://doi.org/10.1128/AAC.00066-15 (2015).
Sifaoui, I. et al. Toxic effects of selected proprietary dry eye drops on Acanthamoeba. Sci. Rep 8, 8520, https://doi.org/10.1038/s41598-018-26914-3 (2018).
Debrabant, A., Lee, N., Bertholet, S., Duncan, R. & Nakhasi, H. L. Programmed cell death in trypanosomatids and other unicellular organisms. Int J Parasitol 33, 257–267, https://doi.org/10.1016/S0020-7519(03)00008-0 (2003).
Antonsson, A & Persson, J. L. Induction of Apoptosis by Staurosporine Involves the Inhibition of Expression of the Major Cell Cycle Proteins at the G2/M Checkpoint Accompanied by Alterations in Erk and Akt Kinase Activities. Anticancer Res, 29, 2893–2898, PMID: 19661292 (2009).
Clarke, M. et al. Genome of Acanthamoeba castellanii highlights extensive lateral gene transfer and early evolution of tyrosine kinase signaling. Genome Biol. 14, R11, https://doi.org/10.1186/gb-2013-14-2-r11. (2013).
Cartuche, L., Cruz, D., Ramírez, M. I., Bailón, N. & Malagón, O. Antibacterial and cytotoxic activity from the extract and fractions of a marine derived bacterium from the Streptomyces genus. Pharm Biol 53, 1826–1830, https://doi.org/10.3109/13880209.2015.1010739 (2015).
Turner, S., Pryer, K. M., Miao, V. P. W. & Palmer, J. D. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J. Eukaryot Microbiol. 46, 327–338, https://doi.org/10.1111/j.1550-7408.1999.tb04612.x (1999).
Katoh, K., Misawa, K., Kuma, K., & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30, 3059–3066 PMID: 12136088 (2002).
Gascuel, O. BIONJ: An improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 14, 685–695, https://doi.org/10.1093/oxfordjournals.molbev.a025808 (1997).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evol 39, 783–791, https://doi.org/10.1111/j.1558-5646.1985.tb00420.x (1985).
Stamatakis, A. RAxML-VI-HPC: Maximum Likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690, https://doi.org/10.1093/bioinformatics/btl446 (2006).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol 33, 1870–1874, https://doi.org/10.1093/molbev/msw054 (2016).
Hu, Y. et al. Chromomycin SA analogs from a marine-derived Streptomyces sp. Bioorg Med Chem 19, 5183–5189, https://doi.org/10.1016/j.bmc.2011.07.013 (2011).
Lorenzo-Morales, J. et al. Glycogen phosphorylase in Acanthamoeba spp.: determining the role of the enzyme during the encystment process using RNA interference. Eukaryotic Cell 7, 509–517, https://doi.org/10.1128/EC.00316-07 (2008).
Acknowledgements
This research was funded by PI18/01380 from Instituto de Salud Carlos III, Spain and CTQ2014-55888-C03-01/R (MINECO). MRB: RICET [RD16/0027/0001 project, from Programa Redes Temáticas de Investigación Cooperativa, FIS (Ministerio Español de Salud, Madrid, Spain) and by Universidad Técnica Particular de Loja, Ecuador through doctoral general budgets of teachers appliants. LC thanks to UTPL by the schorlaship given for his doctoral research. ALA, IS, ARDM thank to Programa Agustín de Bethancourt, Cabildo de Tenerife, Spain.
Author information
Authors and Affiliations
Contributions
L.C., A.R.D.M. and J.J.F. conducted the isolation, culturing, phytochemical and spectral analysis of the chemical compounds. D.C. contributed to the molecular characterization and phylogenetic analysis. J.L.M., A.L.A., M.R.B., J.E.P. and I.S. contributed with the amoebicidal activity and their interpretation assays, Programmed Cell Death analysis and biological data compilation. All authors contributed equally to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
All authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cartuche, L., Sifaoui, I., Cruz, D. et al. Staurosporine from Streptomyces sanyensis activates Programmed Cell Death in Acanthamoeba via the mitochondrial pathway and presents low in vitro cytotoxicity levels in a macrophage cell line. Sci Rep 9, 11651 (2019). https://doi.org/10.1038/s41598-019-48261-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-48261-7
This article is cited by
-
Energetic metabolic reprogramming in Jurkat DFF40-deficient cancer cells
Molecular and Cellular Biochemistry (2022)
-
Indispensable role of microbes in anticancer drugs and discovery trends
Applied Microbiology and Biotechnology (2022)
-
High oxygen concentrations inhibit Acanthamoeba spp.
Parasitology Research (2021)
-
PI3K/Akt/mTOR Signaling as Targets for Developing Anticancer Agents from Marine Organisms
Journal of Ocean University of China (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.