Abstract
In this study, 140 cattle STEC isolates belonging to serogroups O157, O26, O145, O121, O103 and O45 were characterized for 38 virulence-associated genes, antimicrobial resistance profiles and genotyped by PFGE. The majority of isolates carried both stx1 and stx2 concurrently, stx2c, and stx2d; plasmid-encoded genes ehxA, espP, subA and saa but lacked katP and etpD and eaeA. Possession of eaeA was significantly associated with the presence of nle genes, katP, etpD, ureC and terC. However, saa and subA, stx1c and stx1d were only detected in eaeA negative isolates. A complete OI-122 and most non-LEE effector genes were detected in only two eaeA positive serotypes, including STEC O157:H7 and O103:H2. The eaeA gene was detected in STEC serotypes that are commonly implicated in severe humans disease and outbreaks including STEC O157:H7, STEC O145:H28 and O103:H2. PFGE revealed that the isolates were highly diverse with very low rates of antimicrobial resistance. In conclusion, only a small number of cattle STEC serotypes that possessed eaeA, had the highest number of virulence-associated genes, indicative of their high virulence. Further characterization of STEC O157:H7, STEC O145:H28 and O103:H2 using whole genome sequencing will be needed to fully understand their virulence potential for humans.
Similar content being viewed by others
Introduction
Shiga toxin-producing Escherichia coli (STEC) are zoonotic food-borne pathogens characterized by mild to severe diarrhea, hemorrhagic colitis (HC) and the hemolytic uremic syndrome (HUS), a leading cause of acute renal failure in young children and the elderly1. Cattle are the major reservoir of STEC, and humans acquire STEC infections through ingestion of contaminated food of cattle origin2. STEC serogroups O26, O45, O103, O111, O121, O145, and O157 are frequently associated with severe illness and outbreaks in humans3, and colloquially termed the “top or big 7”.
The ability of STEC to cause disease in humans depends on a number of virulence factors. Bacteriophage-encoded Shiga toxins (Stx1 and Stx2) are the main STEC virulence factors4. Furthermore, a number of stx1 and stx2 Shiga toxin subtypes (15) have been described, including four stx1 subtypes, (stx1a, stx1c and stx1d, stx1e) and at least 11 stx2 subtypes have been identified: stx2a, stx2b, stx2c, stx2d, stx2e, stx2f, stx2g, stx2h, stx2i, stx2k, stxl5,6,7. (http://old.iss.it/binary/vtec/cont/STEC_2018_Wrap_up.pdf). The stx1a and stx2a genes represent prototypic Stx1 and Stx2 toxins subtypes repectively5.
Another important virulence factor of STEC is intimin (eaeA)8. Intimin is encoded on the locus of enterocyte effacement (LEE), and is responsible for intimate attachment of eaeA-positive STEC strains to the host intestinal mucosa and formation of typical attaching and effacing (A/E) lesions commonly observed in STEC disease8. Intimin is mainly present in a subset of STEC that are involved in severe disease and have been termed enterohemorrhagic E. coli (EHEC).
STEC possess a number of plasmid-encoded virulence-associated genes, including enterohemolysin (ehxA)9, catalase-peroxidase (katP)10, extracellular serine protease (espP)11 and a type II secretion system (etpD)12. Plasmid-encoded virulence-associated genes enhance pathogenicity and contribute to STEC survival in humans. Enterohemolysin is a heat labile pore-forming toxin, which lyses human erythrocytes with subsequent release of iron from heme. Possession of enterohemolysin (ehxA) by a STEC strain has been associated with HUS13. The type II secretion system facilitates protein transport across the outer membrane14. The extracellullar serine protease autotransporter cleaves coagulation factor V and enhances haemorrhage in HC12. Catalase-peroxidase defends the bacterial cell against oxidative damage by host macrophages10. Other plasmid-encoded virulence-associated genes include a subtilase cytotoxin (subA) and the STEC autoagglutinating adhesin (Saa) (Paton et al., 2001). SubA suppresses the host’s immune system15 and facilitates STEC adherence to enterocytes. Both subA and saa are mainly observed in eaeA negative STEC strains16.
Several STEC O-islands (OIs), including OI-122, OI-57, OI-71 OI-36 and OI-43/48 encode genes, which are absent in nonpathogenic E. coli and are considered STEC virulence-associated genes17. These genes have been used in molecular risk assessment studies to classify STEC serotypes into different seropathotypes based on whether a particular serotype has been implicated in mild, severe illness or no disease at all in humans18,19. OI-122 carries pagC (PhoP-activated gene C)20,21, sen (Shigella flexneri enterotoxin 2) (Z4326)22, efa1 (EHEC factor for adherence) (Z4332), and efa1 (Z4333)23. OI-122 marker genes encode proteins that play a role in immunomodulation, adhesion to host enterocytes and survival in the host23. Furthermore, various genes located on OI-43/48 carry virulence factors24,25: iha (IrgA homologue adhesin) encodes an adhesin24,25, while terC and ureC encode tellurite resistance and urease, respectively26. Tellurite resistance aids bacteria in general stress response within the host environment26, while urease facilitates infection by lowering the STEC infective dose and enhancing bacterial survival in the host27,28.
STEC secrete effector proteins that are carried on a number of pathogenicity islands (PAIs) other than the LEE. These proteins have been termed “non-LEE effector proteins” (nles genes) because they are not encoded on the LEE pathogenicity island29. Important non-LEE effectors (nles) are located on OI-122 (nleB, nleE and ent/espL2), OI-57 (nleG2-3, nleG6-2 and nleG5-2), OI-71 (nleA, nleF, nleG, nleH1-2, nleG2-1 and nleG9) and OI-36 (nleC, nleD, nleB2 and nleH1-1)19,30,31,32. Non-LEE effectors have been associated with different functions including immunosuppression, adherence, invasion, colonization of host enterocytes, disruption of tight junctions and protein trafficking in the host18,19,30,33.
Escherichia coli strains are considered indicators of antimicrobial resistance. E. coli strains including STEC have been used for monitoring and surveillance of antimicrobial resistance in animals, various environments and humans. A number of studies have documented antimicrobial resistance among STEC isolates from cattle34,35,36,37. The emergence and spread of antimicrobial resistant E. coli strains has become a public health concern world-wide, as antimicrobial resistant STEC may be transferred from cattle to humans along the food chain, through occupational exposure or manure runoff from cattle farms. Monitoring of antimicrobial resistance in STEC provides information on antimicrobial abuse and the dynamics of transmission and development of antimicrobial resistant pathogens.
The first association of STEC with human disease in South Africa was reported in 199038. Later on, in 1992 a large outbreak was documented in South Africa and neighboring Swaziland after affected humans had consumed water, which had been contaminated by dead cattle carcasses after a long drought39,40. However, information on virulence characteristics, antimicrobial resistance profiles and genotypes of cattle STEC isolates from South Africa remains scanty. The overall aim of this study was to characterize STEC serotypes of cattle origin and assess their virulence potential for humans. One hundred and forty STEC isolates belonging to serogroups O26, O45, O103, O121, O145 and O157 were screened for a number of virulence-associated genes, and antimicrobial resistance profiles. In addition, pulsed-field gel electrophoresis was used to subtype and determine relatedness/diversity among STEC isolates.
Material and Methods
Bacterial strains
One hundred and forty (N = 140) STEC isolates representing 33 O:H STEC serotypes, which had been previously recovered from cattle on five cow-calf operations in two provinces of South Africa were characterized in this study (Mainga et al., 2018). The collection included STEC O26 serotypes: O26:H2 (20), O26:H4 (1), O26:H7 (3), O26:H8 (8), O26:H11 (3), O26:H16 (2), O26:H19 (2), O26:H21 (7), O26:H28 (2), O26:H38 (2), O26:H45 (1) and O26:HNT (4); STEC O45 serotypes: O45:H2 (1), O45:H8 (3), O45:H11 (8), O45:H16 (3), O45:H19 (3), O45:H21 (2), O45:H28 (1), O45:H38 (5) and O45:HNT (12); STEC O103 serotypes: O103:H2 (1) and O103:H21(1); STEC O121 serotypes: O121:H8 (8), O121:H21 (1) and O121:HNT (1); STEC O145 serotypes: O145:H2 (1), O145:H7 (1), O145:H8 (1), O145:H11 (1), O145:H19 (13), O145:H28 (3) and O145:HNT (3); and STEC O157 serotypes: O157:H2 (1), O157:H7 (9), O157:H19 (1) and O157:H28 (1).
DNA extraction
Frozen STEC cultures (−80 °C) were propagated aerobically overnight at 37 °C on Luria Bertani (LB) agar (DifcoTM Becton and Dickson & Company). Bacterial DNA was extracted using the boiling method as described previously with slight modifications41. Briefly, a loopful of bacterial cells was suspended into 1000 µL of sterile FA buffer (BactoTM FA Buffer, Becton and Dickson Company) in a 1.5 mL Eppendorf tube, mixed by vortexing and centrifuged at 12,000 rpm for 5 min. The supernatant was discarded and bacterial cells were re-suspended in 1000 µL of sterile FA buffer and centrifuged two more times. After the third centrifugation cycle, the supernatant was discarded. The pellet was re-suspended in 500 µL of sterile distilled water and boiled (heating block) for 20 min and cooled on ice for 10 min. Finally, the suspension was centrifuged at 12,000 rpm for 5 min, and DNA was stored at −20 °C for further use in PCR reactions.
Detection of stx1, stx2, eaeA and ehxA genes by PCR
A multiplex polymerase chain reaction (mPCR) was carried out to detect the stx1, stx2, eaeA and ehxA genes using previously described primers (Table 1) and cycling conditions42. Briefly, each PCR reaction (25 µL) contained 2.5 μL of 10X Thermopol reaction buffer, 2.0 μl of 2.5 mM dNTPs, 0.25 μl of 100 mM MgCl2, 0.3 µM of each primer, 1U of Taq DNA Polymerase (New England BioLabs® Inc.) and 5 μl of DNA template. Sterile distilled water was used to top up the reaction volume to 25 µL. STEC O157:H7 EDL933 and sterile water were used as positive and negative controls, respectively. All PCR reagents were supplied by New England BioLabs (NEB, USA) except for primers, which were obtained from Inqaba Biotec (South Africa). PCR reactions were carried out in a C1000 TouchTM (Bio-Rad, USA) or Veriti 96-well Thermal Cycler (Applied Biosystems, USA). PCR amplicons were electrophoresed in 2% (w/v) agarose gels in TAE (Tris–acetate-ethylenediamine tetraacetic acid) buffer, stained with ethidium bromide (0.05 mg/μl) and visualized under ultraviolet (UV) light with a Gel Doc system (Bio-Rad, USA).
Detection of genes encoding Shiga toxin (stx) subtypes
To detect stx1c, stx1d, stx2a, stx2c, stx2d genes, single PCR assays were performed using primers (Table 1) and cycling conditions described elsewhere (Scheutz et al., 2012) (Table 1). Briefly, each PCR reaction (25 µL) contained 2.5 μL of 10X Thermopol reaction buffer, 2.0 μl of 2.5 mM dNTPs, 0.25 μl of 100 mM MgCl2, 0.3 µM final of each primer concentration, 1U of Taq DNA Polymerase (New England BioLabs® Inc.) and 5 μl of DNA template.
Detection of plasmid and pathogenicity island encoded genes
Primers (Table 1) and cycling conditions described in previous studies were used to amplify virulence-associated genes located on plasmids and pathogenicity islands. Amplification reactions for ehxA, saa, subA16,42,43, katP11, espP12, and etpD14 genes were conducted in singleplex PCR reactions. Amplification of OI-122 gene markers including pagC (Z4321), sen (Z4326), efa1 (Z4332 and Z4333) was carried out as previously described (Karmali et al., 2003). Amplification of non-LEE-encoded effector (nle) genes including nleA, nleB, nleB2, nleC, nleD, nleE, nleF, nleG, nleG2-1, nleG2-3, nleG5-2, nleG6-2, nleG9, nleH1, nleH2, and ent/espL2 were performed in singleplex PCR reactions according to Coombes et al.19. PCR reactions for OI-43/48 island markers, iha, ter-island and ureC, were also carried out according to previous studies26,27,44. STEC O157:H7 EDL933 and sterile distilled water were used as positive and negative controls, respectively.
Antimicrobial susceptibility testing
All the 140 STEC isolates were tested for resistance against a panel of 15 antimicrobials by the disk diffusion method as described by the Clinical and Laboratory Standards Institute (CLSI, 2014). Briefly, pure STEC colonies were inoculated on Mueller Hinton agar (MHA) (Oxoid, UK) and incubated overnight at 37 °C. Bacterial suspensions (0.5 McFarland) of overnight cultures were prepared in 0.85% physiological saline. A sterile cotton swab was used to inoculate MHA plates to achieve a confluent growth. Antimicrobial discs were placed on inoculated MHA plates by means of a BBL Sensi-disk or Oxoid disk dispenser and incubated aerobically at 37 °C ± 2 °C for 18 h. The panel of 15 antimicrobials consisted of amoxicillin-clavulanic acid (20 µg–10 µg), amikacin (30 µg), ampicillin (10 µg), ceftazidime (30 µg), cephalothin (30 µg), cefoperazone (75 µg), ceftriaxone (30 µg), chloramphenicol (30 µg), ciprofloxacin (5 µg), gentamicin (10 µg), kanamycin (30 µg), nalidixic acid (30 µg), trimethoprim-sulfamethoxazole (1.25 µg and 23.75 µg, respectively) and tetracycline (30 µg). Antimicrobial disks were obtained from Becton Dickinson (BD, USA) and Oxoid (Thermo Scientific, UK), respectively. Escherichia coli ATCC 25922 was used as the control strain. Isolates were classified as susceptible, intermediate or resistant to each antimicrobial agent according to the CLSI interpretative criteria. However, in the final analysis, intermediate readings were assigned to the resistant category.
Pulsed-field gel electrophoresis
To subtype STEC isolates, DNA was extracted, digested with the XbaI restriction enzyme and subjected to PFGE according to the CDC/PulseNet protocol (https://www.cdc.gov/pulsenet/pdf/ecoli-shigella-salmonella-pfge-protocol-508c.pdf). Salmonella enterica serotype Braenderup (strain H9812; American Type Culture Collection, BAA-664) DNA was used as the molecular weight marker. PFGE fingerprints were analyzed for similarity, and a dendrogram was generated by Bionumerics software, version 6.6 (Applied Maths, Sint Martens-Latem, Belgium) with the Dice similarity indices (complete linkage; optimization, 1.5%; position tolerance, 1.5%) and the unweighted-pair group method with arithmetic means (UPGMA).
Statistical analysis
Descriptive statistical analysis was performed using the statistical package for social sciences (SPSS) software version 21 (IBM® SPSS® Statistics 21). Fisher’s exact test was used to determine if there were statistically significant differences and associations between gene proportions. P values of < 0.05 were considered statistically significant.
Results
Virulence-associated genes
The distribution of stx-encoding virulence genes (N = 140) was as follows: 4.3% of isolates carried stx1 only, 34.3% carried stx2 only, and 61.4% carried both stx1 and stx2. Among the 92 stx1 positive isolates, 20.7% carried stx1c and 18.5% were stx1d positive; 6.5% possessed both stx1c + stx1d. The stx1c and stx1d subtypes were significantly (P < 0.05) detected in STEC O26 and STEC O45 serogroups. Of the 134 stx2 positive isolates, stx2 subtypes were distributed as follows: 91.8%, 97%, and 56% carried stx2a, stx2c, and stx2d, respectively (Table 2). The most common toxin gene combinations among stx2-positive isolates were stx2a + stx2c + stx2d, 37.1%, stx2c + stx2d, 35%, stx2c + stx2d, 5.7%, and stx1c + stx2a + stx2c + stx2d in 5% of isolates (Table 2). All isolates were negative for stx2e, stx2f and stx2g subtypes.
The eaeA gene was detected in only 12.1% of isolates. Among the 17 STEC isolates, which carried eaeA, nine possessed also stx2a + stx2c + stx2d and five had stx2a + stx2c concurrently (Table S1). However, all isolates that were eaeA positive, lacked saa, stx1c and stx1d. The eaeA gene was present in STEC O157:H7 (9 isolates), STEC O157:H28 (1 isolate), STEC O26:H2 (2 isolates), STEC O103:H2 (1 isolate), STEC O145:H28 (3 isolates) and STEC O145:HNT (1 isolate) (Table S1) isolates only.
The following rates were observed for plasmid-encoded genes (Table S2, Fig. 1): ehxA, 90.7%; subA, 85%; saa, 82.1%; espP, 79.3%; katP, 10% and etpD, 7.9% (Fig. 1). All the 14 katP positive isolates were also eaeA positive. The katP and etpD genes were significantly (P < 0.05) observed in eaeA positive isolates (P < 0.000). In addition to eaeA, all O157:H7 isolates possessed the full complement of plasmid markers, including ehxA, subA, katP, espP and etpD except saa. However, STEC O145:H28, O145: HNT and O157:H28 that were also eaeA positive, carried ehxA, subA, katP and espP but lacked etpD. The katP/eaeA genotype was observed in 13/17 isolates, including STEC O145:H28 (3), STEC O145:HNT (1), STEC O157:H7 (9) and STEC O157:H28 (1) whereas the etpD/eaeA genotype was present in 10/17 isolates, including STEC O103:H2 (1), and STEC O157:H7 (9) isolates (Table S1).
The distribution of OI-122 markers was as follows: pagC, 53.6%; sen, 34.3%; Z4332, 10.7%; and Z4333, 28.6% (Table S2, Fig. 2). All OI-122 (full complement OI-22 genes) markers were observed in only 7.1% of isolates, which belonged two only serotypes, including O157:H7 (9 isolates) and O103:H2 (1 isolate). An incomplete OI-122 was observed in 60% of isolates and 32.9% carried none of OI-122 markers. OI-43/48 encoded genes were distributed as follows: iha, 93.6%; terC, 80%; and ureC, 55.7% (Table S2, Fig. 2). All OI-43/48 markers were detected in 52.9% of isolates. Both terC (P = 0.032) and ureC (P < 0.000) were significantly (P < 0.05) prevalent among eaeA-positive STEC isolates. However, 2.1% (3/140) of isolates were negative for all OI-43/48 markers (Table S2).
The following proportions were observed for non-LEE effector (nle) genes: ent/espL2, 34.3%; nleB, 32.9%; nleE, 28.6%; nleG2-3, 30.7%; nleG6-2, 33.6%; nleG5-2, 19.3%; nleH1-2, 27.1%; nleG9, 16.4%: nleG2-1,14.3%; nleA, 12.1%; nleF, 5.0%; nleG, 2.1%; nleH1-1, 27.9% (39/140); nleD, 20.7%; nleB2,15.7%; and nleC, 12.1% (Table S2 and Fig. 2). More than 10 nle genes were observed in 12.9% of the isolates, which were mainly eaeA positive, eight to nine nle genes were present in 6.4%, and one to seven nle genes were detected in 30.7% of the isolates. The remaining 50% of isolates did not carry a non-LEE effector gene.
Overall, the highest number of virulence genes (more than 30 genes) was detected in STEC O157:H7 isolate. STEC O145:H28/HN isolate had 25–30 genes, a number of STEC O45 isolates (H21, H11, H2, H16, HNT) and 2 STEC O26:H2/H21 carried 20–30 virulence-associated genes.
Antimicrobial resistance
Of the 140 STEC isolates, 97.9% were susceptible to all the 15 antimicrobials. Only 2.1% of STEC isolates were antimicrobial resistant, including one STEC O26:H11 isolate which was resistant to tetracycline, one STEC O26:H4 which was resistant to ampicillin and tetracycline and one STEC O45:H21 isolate which was resistant ampicillin, tetracycline and cephalothin.
Pulsed field gel electrophoresis
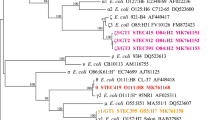
PFGE was conducted to investigate genetic relatedness among the STEC isolates. Six dendograms (Figs 3–6) that displayed relationships among individual serogroups were generated. All the 140 isolates yielded 101 distinct pulsotypes, including 43 for STEC O26, 27 for STEC O45, 2 for STEC O103, 6 for STEC O121, 7 for STEC O157, and 16 for O145 suggesting a high diversity (Dice similarity index < 70%) among STEC isolates in different serogroups. Most of the pulsotypes represented single isolates. The 39 isolates which shared identical PFGE profiles (100% similarity) in different serogroups either belonged to the same serotype or were recovered from the same animal or farm.
Discussion
STEC are frequently implicated in mild to severe human disease and outbreaks (EFSA, 2013). Since the first report on foodborne STEC in humans nearly 40 years ago, a number of studies have been published on virulence, antimicrobial resistance and molecular epidemiology of STEC around the world. However, most reports on STEC are based on data from high-income industrialised nations, while current studies on virulence, antimicrobial resistance and molecular epidemiology of STEC isolates from African countries including South Africa remain scanty. Cattle are a major reservoir of STEC. Molecular risk assessment studies on STEC isolates from cattle have contributed to a better understanding of the virulence potential cattle STEC present to humans and made it possible to differentiate low virulence from highly virulent STEC isolates. In this study, 140 STEC isolates from apparently healthy cattle on five cow-calf operations in South Africa were characterized for a panel 38 virulence-associated genes, antimicrobial resistance and PFGE profiles. The collection of isolates under study was a subset of a larger collection of STEC strains belonging 33 serotypes associated with serogroups O157, O45, O103, O121, O26 and O145.
The majority of STEC isolates carried both stx1 and stx2 concurrently. Almost all stx2-positive isolates (95.7%) harbored stx2a, stx2c and stx2d. Only a small fraction of isolates carried stx1 only. The stx2d subtype identified in this study was the stx2d-activatable variant45. The widespread distribution of stx2a, stx2c and stx2d subtypes in cattle isolates is in agreement with previous studies which have reported high rates of stx2a, stx2c and stx2d subtypes among STEC isolates from cattle in comparison to different stx1 subtypes46,47,48,49,50. STEC isolates that carry stx2 are more virulent compared to strains that possess stx1 alone or both stx1 and stx2 concurrently, and are frequently incriminated in outbreaks and severe human disease manifestations such as HC and HUS45,51,52. Furthermore, some studies have shown that Stx2, Stx2c and Stx2d subtypes are more potent than Stx145,53. In addition, Rasooly and Do54 reported that Stx2 was heat stable and not inactivated at currently approved pasteurization temperatures, making Stx2-producing isolates more likely to be implicated in human STEC disease outbreaks involving pasteurized cattle dairy products.
Less than 50% of isolates carried stx1c and stx1d subtypes. The stx1c subtype was significantly more frequent than stx1d, in agreement with a number of studies which have reported that STEC isolates of cattle origin are mainly stx1c positive47,49. STEC isolates that possess stx1c and/or stx1d subtypes have been mostly implicated in asymptomatic or mild diarrhea in humans52,55,56. However, some reports have implicated stx1c positive isolates in cases of human disease showing bloody diarrhea56,57,58. Interestingly, STEC O45:H2 and STEC O45:H11 isolates possessed stx1c, stx1d, stx2, stx2c, and stx2d concurrently. While STEC O45:H2 is a recognized enterohemorrhagic E. coli59, so far, there are no reports that have associated STEC O45:H11 isolates with severe human disease. Although STEC O45:H11 is not a recognised enterohemorrhagic E. coli, it is possible that the presence of numerous stx variants in this STEC serotype may be indicative of high virulence, assuming that all toxin encoding genes are maximally expressed during STEC infection in humans.
The majority of STEC isolates lacked eaeA, consistent with previous reports on cattle STEC, which have shown that only a subset of cattle STEC are eaeA positive46,48,50. The eaeA gene was present in seropathotypes A and B STEC strains (STEC O157:H7, STEC O103:H2, STEC O26:H2, STEC O145:H28) that were also stx2c and/or stx2d positive but lacked stx1c and stx1d genes. Possession of stx2 and eaeA genes by a STEC strain is indicative of highly virulent STEC strains (EFSA, 2013). STEC seropathotypes A and B strains are highly pathogenic for humans, and commonly implicated in outbreaks and severe disease, including HC or HUS18,60 worldwide.
Plasmid-encoded virulence markers (ehxA, espP, and saa) were present in the majority of isolates. However, subA (37.9%), katP (10%) and etpD (7.9%) virulence markers were less frequent (<50%). Similar reports have documented high rates of ehxA, espP and saa and very low rates of katP and etpD in STEC isolates from cattle35,46,47,49,50,61. However, the rates of these genes in cattle STEC isolates are variable from country to country46,50,61,62,63. Furthermore, katP and etpD were exclusively detected in eaeA-positive STEC O157:H7, O103:H2 and O145:H28 that were also ehxA, espP positive. Possession of all the four plasmid-encoded genes (katP and etpD, ehxA, espP) concurrently is usually indicative of a complete plasmid (pO157 or its homologs). Carriage of a complete plasmid and eaeA is characteristic of highly virulent STEC isolates that are commonly incriminated in severe disease (HC and HUS) and outbreaks in humans64,65,66.
Plasmid-encoded genes saa and subA were exclusively detected among eaeA-negative STEC isolates only, consistent with other studies, which have reported the presence of saa and subA in eaeA-negative STEC isolates46,47,48,49,50. Both saa and subA genes are widespread among STEC serotypes that have been associated with uncomplicated diarrhea (O26:H2, O26:H8, O26:H21, O121:H8, O45:H2 and O145:H8), hemorrhagic colitis (O26:H7 and O145:H7) and hemolytic uremic syndrome (O26:H11) in humans43,67,68,69,70.
Karmali et al.18 suggested that possession of OI-122 marker genes pagC, sen, efa1 (Z4332), efa1 (Z4333) is indicative of a complete OI-122. A complete OI-122 was observed in only 7.1%. that were mostly stx2/eaeA-positive. Isolates which possessed the full complement of OI-122 marker genes belonged to serotypes which are commonly associated with STEC disease outbreaks, including HUS in humans (STEC O157:H7 and STEC O103:H2). Reports that have documented a complete OI-122 among clinically relevant isolates, including STEC O157:H7 and STEC O103:H2 have suggested that the presence of a complete OI-122 and stx2 in eaeA-positive strains is indicative of highly virulent STEC strains18,68,71,72.
Most STEC isolates that had an incomplete OI-122 were seropathotypes B and C STEC strains that are usually incriminated in mild or uncomplicated diarrhea (STEC O26:H2, O26:H8, O26:H21, O103:H21, O45:H2)18. However, isolates that were negative for all OI-122 markers were mainly seropathotypes D or E strains that are very rare in human disease or have never been incriminated in human disease18.
OI-43/48-encoded genes, including iha, terC and ureC are considered suitable markers of virulence in STEC serotypes which are implicated in severe human diseases and outbreaks27,73. OI-43/48 marker genes (iha, terC and ureC) were present in more than 80% of isolates in agreement with previous reports which have found that OI-43/48 marker genes (iha, terC and ureC) are widespread in cattle STEC46,74,75,76. However, the ureC gene was significantly detected in eaeA-positive STEC isolates (17/17) in comparison to eaeA-negative STEC (61/123)28,73,75,76. Furthermore, ureC positive STEC belonged to serotypes that have been incriminated in mild and severe STEC illness in humans, including STEC O26:H2, O26:H7, O26:H8, O26:H21, O45:H2, O103:H2, O145:H7, O145:H28 and O157:H7. The presence of urease genes in STEC has been associated with adherence and survival of bacteria within acidic environments in the host25,77 particularly in STEC serotypes that have been implicated in severe human disease27,73. The iha gene product is considered an additional adhesin in STEC strains24. Although the role of tellurite resistance genes in STEC virulence remains unclear, it has been hypothesized that tellurite resistance genes may promote adherence, STEC survival in the host, and resistance against pore-forming colicins and bacteriophage (T5) infection25.
A number of clinically relevant eaeA-positive STEC strains, including STEC O157:H7, STEC O145:H28, STEC O103:H2, STEC O26:H2 possessed the majority of nle genes. These isolates carried also the nle ‘virulence gene signature’, which includes of nleB, nleE, ent/espL2, nleG2-3, nleG5-2, nleG9, nleG2-1 and nleB2 concurrently18,19,64,65. However, clinically relevant eaeA-negative STEC serotypes, including O26:H2, O26:H21, O157:H19, O45:H11, O45:H16 and O45: HNT STEC isolates possessed 9 to 11/15 nle-encoding genes. While O26:H2 and O26:H21 strains have been previously implicated in mild diarrhea in humans59,60, STEC O157:H28 and O45:H11 have never been incriminated in human disease. The high proportion of nle-encoding genes in STEC O157:H19, O45:H11, O45:H16 and O45:HNT that have never been implicated in human disease may be an indication of emerging virulent cattle STEC strains that should be closely monitored in this part of the world as they may be high risk STEC serotypes with potential to cause severe disease in humans.
Fifty percent (50%) of isolates did not carry any nle-encoding genes. Isolates which did not posess any nle-encoding genes belonged to serotypes that have been incriminated in mild or uncomplicated diarrhea (STEC O26:H2, STEC O26:H8, STEC O26:H21, STEC O121:H8), hemorrhagic colitis (STEC O26:H7, STEC O145:H7) and HUS (STEC O26:H11) in humans59,60, and serotypes that have never been associated with human illness2,78,79. The lack of nle-encoding genes in STEC serotypes that have been previously implicated in mild to severe disease in humans, suggests that the capacity of these strains to cause disease in humans may not be dependent upon currently known non-LEE effectors. However, the absence of known non-LEE effector genes in STEC isolates that have never been implicated in human disease may also explain why these isolates have never been incriminated in human disease.
Antimicrobial resistance profiling showed that almost all (97.9%) the STEC isolates were susceptible to all the 15 tested antimicrobials except for three STEC isolates that were antimicrobial resistant. The three resistant isolates belonged to STEC O26:H11 (tetracycline), STEC O26:H4 (tetracycline and ampicillin) and STEC O45:H21 (amoxicillin-clavulanic acid and cephalothin). Similar findings were made by Dong et al.49, who also reported resistance to ampicillin and tetracycline among cattle STEC isolates. However, higher antimicrobial resistance levels to tetracycline, ampicillin, cephalothin and amoxicillin-clavulanic acid have been previously reported in a number of studies in STEC isolates35,36,80. The very low antimicrobial resistance rates observed in this study suggest that the selection pressure exerted on cattle farms from which the STEC isolates were recovered is negligible. Cattle on cow-calf operations in South Africa graze on pastureland all year round and are not supplemented with feed containing antimicrobial promoters that usually exert selective pressure on intestinal flora such as STEC and facilitate proliferation and development of antimicrobial resistant strains.
PFGE revealed that the STEC isolates under study were highly diverse and only a few isolates had identical fingerprints in individual serogroups. Isolates with identical fingerprints either belonged to the same serotype or were recovered from the same animal or farm. The high diversity observed among the STEC isolates under study is reflection of the high genetic flow occurring among STEC isolates through gene acquisition, shuffling and loss, particularly genes that are encoded on mobile genetic elements including plasmids, bacteriophages and pathogenicity islands.
In conclusion, the majority of STEC isolates were stx1, stx2a, stx2c and stx2d positive but lacked eaeA. Plasmid-encoded genes (hlyA, saa, subA and espP) were detected in most of the isolates but katP and etpD genes were only observed in a very small number of isolates that were also eaeA-positive. A complete plasmid, (ehxA, etpD, katP and espP) was observed in STEC O157:H7 isolates mainly. O island and nle marker genes were absent in most isolates, except for OI-43/48-associated genes (terC and iha), which were prevalent in more than 80% of isolates. STEC O157:H7, STEC O145:H28 and STEC O103:H2 and some STEC O26:H2 isolates possessed the highest number of virulence-associated genes. These serotypes which are frequently implicated in severe STEC disease in humans carried nle marker genes, such as nleB, nleE, ent/espL2, nleG2-3, nleG5-2, nleG9, nleG2-1 and nleB2, which are considered a “hallmark” of highly virulent STEC strains19.
To our knowledge, this is the first detailed characterization of a large number of cattle STEC isolates from South Africa. This study provides much needed data on the molecular characteristics of STEC serotypes from beef cattle in South Africa. Further studies using whole-genome sequencing (WGS) will be needed to fully assess the virulence potential of these cattle STEC isolates for humans.
References
Karmali, M. A. Infection by verocytotoxin-producing Escherichia coli. Clinical Microbiology Reviews 2, 15–38 (1989).
Hussein, H. S. & Bollinger, L. M. Prevalence of Shiga toxin-producing Escherichia coli in beef cattle. Journal of Food Protection 68, 2224–2241 (2005).
Johnson, K. E., Thorpe, C. M. & Sears, C. L. The emerging clinical importance of non-O157 Shiga toxin-producing Escherichia coli. Clinical Infectious Diseases 43, 1587–1595, https://doi.org/10.1086/509573 (2006).
Scotland, S. M., Smith, H. R., Willshaw, G. A. & Rowe, B. Vero cytotoxin production in strain of Escherichia coli is determined by genes carried on bacteriophage. Lancet 2, 216 (1983).
Scheutz, F. et al. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. Journal of Clinical Microbiology 50, 2951–2963 (2012).
Patel, I. R. et al. FDA Escherichia coli Identification (FDA-ECID) Microarray: a pangenome molecular toolbox for serotyping, virulence profiling, molecular epidemiology and phylogeny. Applied and Environmental Microbiology 82, 3384–3394, https://doi.org/10.1128/aem.04077-15 (2016).
Bai, X. et al. Identification and pathogenomic analysis of an Escherichia coli strain producing a novel Shiga toxin 2 subtype. Scientific Reports 8, 6756, https://doi.org/10.1038/s41598-018-25233-x (2018).
Yu, J. & Kaper, J. B. Cloning and characterization of the eae gene of enterohaemorrhagic Escherichia coli O157:H7. Molecular Microbiology 6, 411–417 (1992).
Schmidt, H., Beutin, L. & Karch, H. Molecular analysis of the plasmid-encoded hemolysin of Escherichia coli O157:H7 strain EDL 933. Infection and Immunity 63, 1055–1061 (1995).
Brunder, W., Schmidt, H. & Karch, H. KatP, a novel catalase-peroxidase encoded by the large plasmid of enterohaemorrhagic Escherichia coli O157:H7. Microbiology 142(Pt 11), 3305–3315, https://doi.org/10.1099/13500872-142-11-3305 (1996).
Brunder, W., Schmidt, H., Frosch, M. & Karch, H. The large plasmids of Shiga-toxin-producing Escherichia coli (STEC) are highly variable genetic elements. Microbiology 145(Pt 5), 1005–1014, https://doi.org/10.1099/13500872-145-5-1005 (1999).
Brunder, W., Schmidt, H. & Karch, H. EspP, a novel extracellular serine protease of enterohaemorrhagic Escherichia coli O157:H7 cleaves human coagulation factor V. Molecular Microbiology 24, 767–778 (1997).
Schmidt, H. & Karch, H. Enterohemolytic phenotypes and genotypes of shiga toxin-producing Escherichia coli O111 strains from patients with diarrhea and hemolytic-uremic syndrome. Journal of Clinical Microbiology 34, 2364–2367 (1996).
Schmidt, H., Henkel, B. & Karch, H. A gene cluster closely related to type II secretion pathway operons of gram-negative bacteria is located on the large plasmid of enterohemorrhagic Escherichia coli O157 strains. FEMS Microbiology Letters 148, 265–272, https://doi.org/10.1111/j.1574-6968.1997.tb10299.x (1997).
Hui, W., Paton, J. C. & Paton, A. W. Pathologic changes in mice induced by subtilase cytotoxin, a potent new Escherichia coli subAB toxin that targets the endoplasmic reticulum. The Journal of Infectious Diseases 196, 1093–1101 (2007).
Paton, A. W., Srimanote, P., Woodrow, M. C. & Paton, J. C. Characterization of Saa, a novel autoagglutinating adhesin produced by locus of enterocyte effacement-negative Shiga-toxigenic Escherichia coli strains that are virulent for humans. Infection and Immunity 69, 6999–7009, https://doi.org/10.1128/iai.69.11.6999-7009.2001 (2001).
Schmidt, H. & Hensel, M. Pathogenicity islands in bacterial pathogenesis. Clinical Microbiology Reviews 17(1), 14–56 (2004).
Karmali, M. A. et al. Association of genomic O island 122 of Escherichia coli EDL 933 with verocytotoxin-producing Escherichia coli seropathotypes that are linked to epidemic and/or serious disease. Journal of Clinical Microbiology 41, 4930–4940 (2003).
Coombes, B. K. et al. Molecular analysis as an aid to assess the public health risk of non-O157 Shiga toxin-producing Escherichia coli strains. Applied and Environmental Microbiology 74, 2153–2160, https://doi.org/10.1128/aem.02566-07 (2008).
Gunn, J. S., Alpuche-Aranda, C. M., Loomis, W. P., Belden, W. J. & Miller, S. I. Characterization of the Salmonella typhimurium pagC/pagD chromosomal region. J. Bacteriol. Journal of Bacteriology 177, 5040–5047 (1995).
Miller, S. I. & Mekalanos, J. J. Constitutive expression of the phoP regulon attenuates Salmonella virulence and survival within macrophages. Journal of Bacteriology, 172, 2485–2490 (1990).
Nataro, J. P. et al. Identification and cloning of a novel plasmid-encoded enterotoxin of enteroinvasive Escherichia coli and Shigella strains. Infection and Immunity 63, 4721–4728 (1995).
Nicholls, L., Grant, T. H. & Robins-Browne, R. M. Identification of a novel genetic locus that is required for in vitro adhesion of a clinical isolate of enterohaemorrhagic Escherichia coli to epithelial cells. Molecular Microbiology 35, 275–288 (2000).
Tarr, P. I. et al. Iha: a novel Escherichia coli O157:H7 adherence-conferring molecule encoded on a recently acquired chromosomal island of conserved structure. Infection and Immunity 68, 1400–1407 (2000).
Yin, X. et al. Contributions of O island 48 to adherence of enterohemorrhagic Escherichia coli O157:H7 to epithelial cells in vitro and in ligated pig ileal loops. Applied and Environmental Microbiology. 75, 5779 (2009).
Taylor, D. E. et al. Genomic variability of O islands encoding tellurite resistance in enterohemorrhagic Escherichia coli O157:H7 isolates. Journal of Bacteriology 184, 4690–4698 (2002).
Nakano, M. et al. Association of the urease gene with enterohemorrhagic Escherichia coli strains irrespective of their serogroups. Journal of Clinical Microbiology 39, 4541–4543, https://doi.org/10.1128/jcm.39.12.4541-4543.2001 (2001).
Friedrich, A. W. et al. Distribution of the urease gene cluster among and urease activities of enterohemorrhagic Escherichia coli O157 isolates from humans. Journal of Clinical Microbiology 43, 546–550 (2005).
Tobe, T. et al. An extensive repertoire of type III secretion effectors in Escherichia coli O157 and the role of lambdoid phages in their dissemination. Proceedings of the National Academy of Sciences 103, 14941–14946, https://doi.org/10.1073/pnas.0604891103 (2006).
Deng, W. et al. Dissecting virulence: systematic and functional analyses of a pathogenicity island. Proceedings of the National Academy of Sciences 101, 3597–3602, https://doi.org/10.1073/pnas.0400326101 (2004).
Gruenheid, S. et al. Identification and characterization of NleA, a non-LEE-encoded type III translocated virulence factor of enterohaemorrhagic Escherichia coli O157:H7. Molecular Microbiology 51, 1233–1249 (2004).
Wickham, M. E. et al. Bacterial genetic determinants of nonO157 STEC outbreaks and hemolytic uremic syndrome after infection. Journal of Infectious Diseases 194, 819–827 (2006).
Konczy, P. et al. Genomic O island 122, locus for enterocyte effacement, and the evolution of virulent verocytotoxin-producing Escherichia coli. Journal of Bacteriology 190, 5832–5840 (2008).
Mora, A. et al. Antimicrobial resistance of Shiga toxin (verotoxin)-producing Escherichia coli O157:H7 and non-O157 strains isolated from humans, cattle, sheep and food in Spain. Research in Microbiology 156, 793–806, https://doi.org/10.1016/j.resmic.2005.03.006 (2005).
Karama, M., Johnson, R. P., Holtslander, R., McEwen, S. A. & Gyles, C. L. Prevalence and characterization of verotoxin-producing Escherichia coli (VTEC) in cattle from an Ontario abattoir. Canadian Journal of Veterinary Research 72, 297–302 (2008).
Amézquita-López, B. A. et al. Antimicrobial resistance profiles of Shiga toxin-producing Escherichia coli O157 and non-O157 recovered from domestic farm animals in rural communities in Northwestern Mexico. Antimicrobial Resistance and Infection Control 5, 1–1, https://doi.org/10.1186/s13756-015-0100-5 (2016).
Mukherjee, S. et al. Antimicrobial drug-resistant shiga toxin-producing Escherichia coli Infections, Michigan, USA. Emerging Infectious Diseases 23, 1609–1611, https://doi.org/10.3201/eid2309.170523 (2017).
Browning, N. G., Botha, J. R., Sacho, H. & Moore, P. J. Escherichia coli O157:H7 haemorrhagic colitis. Report of the first South African case. South African Journal of Surgery 28, 28–29 (1990).
Isaacson, M. et al. Haemorrhagic colitis epidemic in Africa. Lancet 341, 961 (1993).
Effler, E. et al. Factors contributing to the emergence of Escherichia coli O157 in Africa. Emerging Infectious Diseases 7, 812–819, https://doi.org/10.3201/eid0705.017507 (2001).
Mainga, A. O. et al. Occurrence and characterization of seven major Shiga toxin-producing Escherichia coli serotypes from healthy cattle on cow-calf operations in South Africa. Zoonoses and Public Health 65(7), 777–789, https://doi.org/10.1111/zph.12491 (2018).
Paton, A. W. & Paton, J. C. Detection and characterization of Shiga toxigenic Escherichia coli by using multiplex PCR assays for stx1, stx2, eaeA, enterohemorrhagic E. coli hlyA, rfbO111, and rfbO157. Journal of Clinical Microbiology 36, 598–602 (1998).
Paton, A. W., Srimanote, P., Talbot, U. M., Wang, H. & Paton, J. C. A new family of potent AB(5) cytotoxins produced by Shiga toxigenic Escherichia coli. The Journal of Experimental Medicine 200, 35–46, https://doi.org/10.1084/jem.20040392 (2004).
Janka, A. et al. Presence and characterization of a mosaic genomic island which distinguishes sorbitol-fermenting enterohemorrhagic Escherichia coli O157:H- from E. coli O157:H7. Applied and Environmental Microbiology 71, 4875–4878, https://doi.org/10.1128/AEM.71.8.4875-4878.2005 (2005).
Bielaszewska, M., Friedrich, A. W., Aldick, T., Schurk-Bulgrin, R. & Karch, H. Shiga toxin activatable by intestinal mucus in Escherichia coli isolated from humans: predictor for a severe clinical outcome. Clinical Infectious Diseases 43, 1160–1167, https://doi.org/10.1086/508195 (2006).
Monaghan, A. et al. Serotypes and virulence profiles of non-O157 Shiga toxin-producing Escherichia coli isolates from bovine farms. Applied and Environmental Microbiology 77, 8662–8668, https://doi.org/10.1128/aem.06190-11 (2011).
Bosilevac, J. M. & Koohmaraie, M. Prevalence and characterization of non-O157 shiga toxin-producing Escherichia coli isolates from commercial ground beef in the United States. Applied and Environmental Microbiology 77, 2103–2112, https://doi.org/10.1128/aem.02833-10 (2011).
Akiyama, Y. et al. Shiga toxin subtypes and virulence genes in Escherichia coli isolated from cattle. Japanese Journal of Infectious Diseases 70, 181–185, https://doi.org/10.7883/yoken.JJID.2016.100 (2017).
Dong, H. J. et al. Prevalence, virulence potential, and pulsed-field gel electrophoresis profiling of Shiga toxin-producing Escherichia coli strains from cattle. Gut Pathogens 9, 22, https://doi.org/10.1186/s13099-017-0169-x (2017).
Gonzalez, A. G. et al. Serotypes, virulence markers and cell invasion ability of Shiga toxin-producing Escherichia coli strains isolated from healthy dairy cattle. Journal of Applied Microbiology 121, 1130–1143, https://doi.org/10.1111/jam.13230 (2016).
Boerlin, P. et al. Associations between virulence factors of Shiga toxin-producing Escherichia coli and disease in humans. Journal of Clinical Microbiology 37, 497–503 (1999).
Friedrich, A. W. et al. Escherichia coli harboring Shiga toxin 2 gene variants: frequency and association with clinical symptoms. The Journal of Infectious Diseases 185, 74–84, https://doi.org/10.1086/338115 (2002).
Fuller, C. A., Pellino, C. A., Flagler, M. J., Strasser, J. E. & Weiss, A. A. Shiga toxin subtypes display dramatic differences in potency. Infection and Immunity 79, 1329–1337, https://doi.org/10.1128/iai.01182-10 (2011).
Rasooly, R. & Do, P. M. Shiga toxin Stx2 is heat-stable and not inactivated by pasteurization. International Journal of Food Microbiology 136, 290–294, https://doi.org/10.1016/j.ijfoodmicro.2009.10.005 (2010).
Eklund, M., Leino, K. & Siitonen, A. Clinical Escherichia coli strains carrying stx genes: stx variants and stx-positive virulence profiles. Journal of Clinical Microbiology 40, 4585–4593, https://doi.org/10.1128/jcm.40.12.4585-4593.2002 (2002).
Friedrich, A. W. et al. Shiga toxin 1c-producing Escherichia coli strains: phenotypic and genetic characterization and association with human disease. Journal of Clinical Microbiology 41, 2448–2453, https://doi.org/10.1128/JCM.41.6.2448-2453.2003 (2003).
Zhang, W., Bielaszewska, M., Kuczius, T. & Karch, H. Identification, characterization, and distribution of a Shiga toxin 1 gene variant (stx(1c)) in Escherichia coli strains isolated from humans. Journal of Clinical Microbiology 40, 1441–1446 (2002).
Matussek, A. et al. Genetic makeup of Shiga toxin-producing Escherichia coli in relation to clinical symptoms and duration of shedding: a microarray analysis of isolates from Swedish children. European Journal of Clinical Microbiology & Infectious Diseases 36, 1433–1441, https://doi.org/10.1007/s10096-017-2950-7 (2017).
Beutin, L. & Fach, P. Detection of Shiga toxin-producing Escherichia coli from nonhuman sources and strain typing. Microbiology Spectrum 2, https://doi.org/10.1128/microbiolspec.EHEC-0001-2013 (2014).
EFSA Panel on Biological Hazards (BIOHAZ). Scientific Opinion on VTEC-seropathotype and scientific criteria regarding pathogenicity assessment. EFSA Journal 11 (2013).
Wu, Y. et al. Distribution of virulence genes related to adhesins and toxins in shiga toxin-producing Escherichia coli strains isolated from healthy cattle and diarrheal patients in Japan. Journal of Veterinary Medical Science 72, 589–597 (2010).
Lucchesi, P. M., Krüger, A. & Parma, A. E. Distribution of saa gene variants in verocytotoxigenic Escherichia coli isolated from cattle and food. Research in Microbiology 157, 263–266 (2006).
Cadona, J. S., Bustamante, A. V., Gonzalez, J. & Sanso, A. M. Genetic relatedness and novel sequence types of non-O157 Shiga toxin-producing Escherichia coli strains Isolated in Argentina. Frontiers in Cellular and Infection Microbiology 6 (2016).
Bugarel, M., Beutin, L., Martin, A., Gill, A. & Fach, P. Micro-array for the identification of Shiga toxin-producing Escherichia coli (STEC) seropathotypes associated with hemorrhagic colitis and hemolytic uremic syndrome in humans. International Journal of Food Microbiology 142, 318–329, https://doi.org/10.1016/j.ijfoodmicro.2010.07.010 (2010).
Bugarel, M., Martin, A., Fach, P. & Beutin, L. Virulence gene profiling of enterohemorrhagic (EHEC) and enteropathogenic (EPEC) Escherichia coli strains: a basis for molecular risk assessment of typical and atypical EPEC strains. BMC Microbiology 11, 142, https://doi.org/10.1186/1471-2180-11-142 (2011).
Bielaszewska, M. et al. Enterohemorrhagic Escherichia coli O26:H11/H-: a new virulent clone emerges in Europe. Clinical Infectious Diseases 56, 1373–1381, https://doi.org/10.1093/cid/cit055 (2013).
Tozzoli, R. et al. Production of the subtilase AB5 cytotoxin by Shiga toxin-negative Escherichia coli. Journal of Clinical Microbiology 48, 178–183, https://doi.org/10.1128/JCM.01648-09 (2010).
Buvens, G. et al. Incidence and virulence determinants of verocytotoxin-producing Escherichia coli infections in the Brussels-Capital Region, Belgium, in 2008–2010. Journal of Clinical Microbiology 50, 1336–1345, https://doi.org/10.1128/jcm.05317-11 (2012).
Galli, L., Miliwebsky, E., Irino, K., Leotta, G. & Rivas, M. Virulence profile comparison between LEE-negative Shiga toxin-producing Escherichia coli (STEC) strains isolated from cattle and humans. Veterinary Microbiology 143, 307–313, https://doi.org/10.1016/j.vetmic.2009.11.028 (2010).
Michelacci, V. et al. A new pathogenicity island carrying an allelic variant of the Subtilase cytotoxin is common among Shiga toxin producing Escherichia coli of human and ovine origin. Clinical Microbiology and Infection 19, E149–156, https://doi.org/10.1111/1469-0691.12122 (2013).
Morabito, S., Tozzoli, R., Oswald, E. & Caprioli, A. A mosaic pathogenicity island made up of the locus of enterocyte effacement and a pathogenicity island of Escherichia coli O157:H7 is frequently present in attaching and effacing E. coli. Infection and Immunity 71, 3343–3348 (2003).
Wickham, M. E. et al. Bacterial genetic determinants of non-O157 STEC outbreaks and hemolytic-uremic syndrome after infection. The Journal of Infectious Diseases 194, 819–827, https://doi.org/10.1086/506620 (2006).
Friedrich, A. W. et al. Urease genes in non-O157 Shiga toxin-producing Escherichia coli: mostly silent but valuable markers for pathogenicity. Clinical Microbiology and Infection 12, 483–486, https://doi.org/10.1111/j.1469-0691.2006.01379.x (2006).
Karama, M., Johnson, R. P., Holtslander, R. & Gyles, C. L. Production of verotoxin and distribution of O islands 122 and 43/48 among verotoxin-producing Escherichia coli O103:H2 isolates from cattle and humans. Applied and Environmental Microbiology 75, 268–270, https://doi.org/10.1128/aem.01445-08 (2009).
Orth, D., Grif, K., Dierich, M. P. & Wurzner, R. Prevalence, structure and expression of urease genes in Shiga toxin-producing Escherichia coli from humans and the environment. International Journal of Hygiene and Environmental Health 209, 513–520, https://doi.org/10.1016/j.ijheh.2006.06.003 (2006).
Ju, W., Shen, J., Toro, M., Zhao, S. & Meng, J. Distribution of pathogenicity islands OI-122, OI-43/48, and OI-57 and a high-pathogenicity island in Shiga toxin-producing Escherichia coli. Applied and Environmental Microbiology 79, 3406–3412, https://doi.org/10.1128/AEM.03661-12 (2013).
Steyert, S. R. & Kaper, J. B. Contribution of urease to colonization by Shiga toxin-producing Escherichia coli. Infection and Immunity 80, 2589–2600, https://doi.org/10.1128/IAI.00210-12 (2012).
Hussein, H. S. & Sakuma, T. Prevalence of shiga toxin-producing Escherichia coli in dairy cattle and their products. Journal of Dairy Science 88, 450–465 (2005).
Hussein, H. S. & Sakuma, T. Shiga toxin-producing Escherichia coli: pre- and postharvest control measures to ensure safety of dairy cattle products. Journal of Food Protection 68, 199–207 (2005).
Iweriebor, B. C., Iwu, C. J., Obi, L. C., Nwodo, U. U. & Okoh, A. I. Multiple antibiotic resistances among Shiga toxin producing Escherichia coli O157 in feces of dairy cattle farms in Eastern Cape of South Africa. BMC Microbiology 15, 213, https://doi.org/10.1186/s12866-015-0553-y (2015).
Acknowledgements
This research was funded by the Gauteng Department of Agriculture and Rural Development (GDARD) (Grant No. FY 2013/14‐A0W907), the Global Disease Detection (GDD) Program of the Centers for Disease Control and Prevention (CDC) (Grant No. 1U2GGH001874‐01) and the National Research Foundation (NRF) of South Africa Thuthuka (TTK13062619943), Research Technology (RTF14012762427) Funds. This manuscript is part a dissertation submitted in the Veterinary Public Health section, Department of Paraclinical Sciences, University of Pretoria, in partial fulfilment of the requirements for the degree of Master of Science (Veterinary Science).
Author information
Authors and Affiliations
Contributions
M.K. and A.K. conceptualized and designed the study; A.M., B.T., M.M. and M.K. designed laboratory assays and generated data; A.M. and M.K. analysed data; A.M. and M.K. and S.E. drafted the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karama, M., Mainga, A.O., Cenci-Goga, B.T. et al. Molecular profiling and antimicrobial resistance of Shiga toxin-producing Escherichia coli O26, O45, O103, O121, O145 and O157 isolates from cattle on cow-calf operations in South Africa. Sci Rep 9, 11930 (2019). https://doi.org/10.1038/s41598-019-47948-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47948-1
This article is cited by
-
Molecular detection of Shiga toxin and extended-spectrum beta-lactamase (ESBL)-producing Escherichia coli isolates from sheep and goats
Molecular Biology Reports (2024)
-
Occurrence, serotypes and virulence characteristics of Shiga toxin-producing and Enteropathogenic Escherichia coli isolates from dairy cattle in South Africa
World Journal of Microbiology and Biotechnology (2024)
-
Antimicrobial growth promoters approved in food-producing animals in South Africa induce shiga toxin-converting bacteriophages from Escherichia coli O157:H7.
Gut Pathogens (2023)
-
Genetic and antimicrobial resistance profiles of non-O157 Shiga toxin-producing Escherichia coli from different sources in Egypt
BMC Microbiology (2021)
-
Prolonged carriage of ESBL-producing enterobacterales and potential cross-transmission among residents in geriatric long-term care facilities
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.