Abstract
Bee pollinators are exposed to multiple natural and anthropogenic stressors. Understanding the effects of a single stressor in the complex environmental context of antagonistic/synergistic interactions is critical to pollinator monitoring and may serve as early warning system before a pollination crisis. This study aimed to methodically improve the diagnosis of bee stressors using a simultaneous untargeted and targeted metabolomics-based approach. Analysis of 84 Bombus terrestris hemolymph samples found 8 metabolites retained as potential biomarkers that showed excellent discrimination for nutritional stress. In parallel, 8 significantly altered metabolites, as revealed by targeted profiling, were also assigned as candidate biomarkers. Furthermore, machine learning algorithms were applied to the above-described two biomarker sets, whereby the untargeted eight components showed the best classification performance with sensitivity and specificity up to 99% and 100%, respectively. Based on pathway and biochemistry analysis, we propose that gluconeogenesis contributed significantly to blood sugar stability in bumblebees maintained on a low carbohydrate diet. Taken together, this study demonstrates that metabolomics-based biomarker discovery holds promising potential for improving bee health monitoring and to identify stressor related to energy intake and other environmental stressors.
Similar content being viewed by others
Introduction
Bees are perhaps the best known beneficial insects, performing ecosystem services which are vital to both food security and biodiversity1. More specifically, bumble bees are key pollinators in temperate climate regions2, and the economic value derived from their pollination services is worth billions of dollars annually3. However, bee diversity and their essential pollination services are threatened4,5. Understanding synergistic/antagonistic interactions between drivers of decline, to ultimately identify dangerous combined effects, will be critical to save bees6,7. Having robust tools to measure bee health would allow us to identify how health relates to biotic and abiotic stressors. Meanwhile, pollinators, especially honey bees, have been employed as biological indicators to monitor environmental pollution since 19628, as well as to identify diseases and parasites in relation to chemical and physical factors9,10. Although relations with environmental stressors can be drawn, no identification or quantification on the severity of a specific stressor on the bees can be inferred. Hence, given that the biomarker-approach is capable of linking the physiological status of bees to the severity of specific stressor, the development of an effective and practical approach, which can identify the diagnostic biomarkers tracking a single stressor and its interplay with others, is crucially needed.
In this context, metabolomics has been shown to have some advantages over other post-genomic technology. The metabolome is the final downstream product of gene transcription, and therefore, it is the closest to the phenotype of the biological system studied11. Additionally, unlike the transcriptome or the proteome which are diversified from species to species, the basic metabolic pathways and their metabolites among different species are the same, making metabolomic analysis much more universal12. Moreover, metabolomic approaches have been successfully developed for environmental relevant species for biomarker discovery and risk assessment of toxicant exposure, metabolic responses to environmental stressors, and disease diagnosis and monitoring13,14. Within the field of entomology, the metabolomic approach was first applied in 1990 with the evaluation of the metabolism of parasitized Manduca sexta larvae15. Since then, metabolomics has been applied to a wide range of insect research including honey bees, providing new insights into biological processes that could hardly be obtained using any other approach16,17. Hence, this study has the objective to perform mass spectrometry-based metabolomics to identify biomarkers that can discriminate bee health status with or without an introduced single stressor.
Bees depend entirely on nutrition obtained through floral pollen and nectar for growth, reproduction, and health18. Growing evidence has linked the interaction between malnutrition and other stressors with current bee population declines18,19,20. Although this is not yet fully understood, it seems highly likely that nutritional stress is significantly contributing to the synergistic effect of numerous other stressors18,20. For instance, poor nutrition due to loss of food sources could be synergistically acting with emerging pathogens to cause bee population decline20,21. Moreover, some pathogens are known to directly affect the energy metabolism in bees22,23, but not always24. Hence, tools to identify in which environments bees suffer nutritional stress are therefore key for good conservation planning. Following this, we setup this proof-of-concept study, with the focus on testing the ability of a metabolomics-based approach to classify nutritional status of bees, whereby a mimic of low carbohydrate food forage stress was imposed to Bombus terrestris, a key pollinator in temperate climate regions of Europe25.
Results and Discussion
A schematic overview of the study design is presented in Fig. 1. A global data table characterizing each of the 84 samples by 2197 components was obtained and established as metabolomic fingerprints for B. terrestris hemolymph. PCA analysis revealed tight clustering of the QC samples (Fig. 2A), suggesting good instrumental stability during sample analysis. Datasets were validated by CV-ANOVA (P < 0.01) and permutation test. The OPLS-DA analysis showed a good discrimination in terms of nutritional stress (dataset T, Q2 = 0.616, R2Y = 0.981, Fig. 2B), even with variables from hierarchy and exposure time of the stressor. 6-day samples were also clearly distinct from 12-day samples (dataset T, Q2 = 0.754, R2Y = 0.958, Fig. 2C). By contrast, hierarchy effects on metabolic levels were not obvious (Table 1), although we observed dominance hierarchies which were established during the first days in queen-less conditions as previously described26. These results confirm that UHPLC-Orbitrap-MS-based metabolomics can successfully establish and distinguish the metabolic profiles of an individual B. terrestris using only microliter hemolymph samples.
Schematic diagram of the metabolomics-based bee health monitoring workflow. (A) Experimental setup in this prove-of-concept study with commercial bees and diluted sugar syrup diet. Five random Bombus terrestris callow workers were allocated to one microcolony. A total of 26 microcolonies were randomly assigned to the treatment (25% sugar syrup, 13 microcolonies) or control (50% sugar syrup, 13 microcolonies) groups. We sampled 2 specimens at day 6 (in 2 × 5 microcolonies) and 4 specimens at day 12 (in the other 2 × 8 microcolonies). In total for day 6 this is 10 dominant pseudo queens and 10 workers, for day 12 this is 16 dominant pseudo queens and 48 workers. Collected bee hemolymph has a variation in diet, age and hierarchy characteristics. (B) Metabolomics-based biomarker selection. Bee hemolymph extracts were processed by UHPLC-Orbitrap-MS for both untargeted and targeted metabolomics. In untargeted metabolomics, to assess the metabolic differences between the defined sample sets, PCA (principle component analysis) and OPLS-DA (orthogonal partial least square-discriminant analysis) were performed using Simca (soft independent modelling of class analogy)13 multivariate statistics software. Potential biomarkers annotation were consulted on online HMDB (Human Metabolome Database) and KEGG (Kyoto Encyclopedia of Genes and Genomes) database. In targeted metabolomics, Statistical analysis was performed with SPSS 22.0, applying two-way ANOVA and Tukey HSD test for post hoc comparisons to select significantly changed metabolites. To locate significantly expressed metabolites, the web-based platform MetaboAnalyst and KEGG database were used. Hereby, two candidate biomarker sets that could be associated with low-carbohydrate nutritional stress were selected from untargeted metabolomics (set 1) and targeted metabolomics (set 2), respectively. (C) Machine leaning algorithms are further implemented to build and evaluate the classification model. Classification sensitivity and specificity of the two selected biomarker sets (targeted and untargeted) and the combined set were tested using different machine learning algorithms. Well-validated classification models may be applied in real environments for bee health monitoring in terms of nutritional stress.
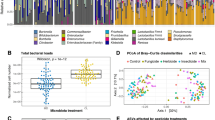
Plots from multivariate statistical analysis. (A) PCA-X score plots. Score plots for experimental (25% sugar syrup, n = 42), control (50% sugar syrup, n = 42), and quality control (QC) samples. (B) Low carbohydrate nutritional stress associated OPLS-DA score plot. (C) Exposure time associated OPLS-DA score plots. (D) Venn diagram of candidate biomarkers for nutritional stress from four datasets.
Seven sub-datasets were further created containing specimens of different hierarchy and exposure time of the stressor (see Table 1). First, the results showed that the introduced low-carbohydrate stress had a striking effect on the hemolymph metabolome in all four analyzed datasets (dataset 1, 2, 3 and T), suggesting poor food availability disturbs bee metabolic states. This contributes to our understanding that sufficient food availability is vital for bee health. For bees, sugars are almost exclusively the substrates used for flight27,28,29, and they store only limited amounts of glycogen in the fat body and flight muscle tissues30,31. Hence, hemolymph energy supplies are very important for most activities. Second, similar performance of exposure time was observed both in control (dataset 4) and stressed datasets (dataset 5), indicating that age itself already had a clear metabolic effect. This suggests that the intrinsic physiological changes on B. terrestris hemolymph did not depend solely on extrinsic stressors, which is consistent with the assessment of honey bee senescence32. Third, the impact of hierarchy on the metabolic fingerprint is not obvious here (dataset 6, 7 and T). A possible explanation is that the dominance phenotype was not so clear-cut as expected. In the wild queenright colonies, multiple worker bees may also start laying drone eggs once the queen has started to produce new queens and drones3. Likewise, in microcolonies it has been observed, aside from the dominant worker, that also other bees tend to develop their ovaries (data not shown).
With more and more genomes of different bee species being published33,34,35,36,37, there is a growing number of comparative genomics, transcriptomics, and proteomics studies regarding bee health38,39,40,41,42. For most of these “omics” studies, the analysis has focused on explaining individual impacts or eventually emphasizing the complex nature of this problem. However, they have overlooked the methodological potential to disentangle complex interacting drivers43, and thus we still lack effective bee health monitoring and risk assessment tools. With respect to the current study, three following features may be noted when making the comparison with previous studies on bee stressors.
Selection of diagnostic metabolic biomarkers for low-carbohydrate nutritional stress
A major feature of the current study is that we can promote biomarker discovery by selecting the candidate biomarkers not only through multiple validated OPLS-DA models but also by a dual targeted/untargeted data analysis strategy. As presented in Fig. 2D, with VIP > 1.0, a total of 44, 29, 59 and 45 components was assigned biomarker potential for nutritional stress in long-term stressed workers (dataset 1), long-term stressed bees (dataset 2), all stressed workers (dataset 3), and all stressed bees (total dataset), respectively. Furthermore, eight components were conserved in all four biomarkers sets and selected as the ideal untargeted biomarkers for nutritional stress. In parallel, a simultaneous targeted screening was performed. Among the nearly 300 metabolites included in our in-house library, 64 metabolites were identified and retained for semi-quantitative metabolic profiling. As summarized in Table 2, among these metabolites, there were 20% amino acids, 19% carbohydrates, 25% carboxylic acids and 36% of other chemical classes. Eight metabolites from sugar and amino acid metabolism were significantly changed during low-carbohydrate diet (Fig. 3) and selected as targeted biomarker set for nutritional stress. A total of 16 candidate biomarkers, including key metabolites of sugar and amino acid metabolism (set 2) and novel metabolites (set 1), are therefore ideal candidates to confirm our proof of concept.
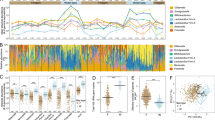
Comparison of nine key differential metabolites between diet (25% sugar syrup and 50% sugar syrup) and exposure time (day 6 and day 12). (A) main sugars. (B) main amino acids. The black cycles represent bees fed on 50% sugar syrup, and the green squares represent bees fed on 25% sugar syrup. Data were derived from targeted metabolomic assay data, and statistically significant results from two-way ANOVA comparing diet and age are indicated at the top left of the graphs. *P < 0.05, **P < 0.01, ***P < 0.001. NS = not significant.
Application of machine learning algorithms to metabolomic data for stressor classification
A second distinctive feature is the application of machine learning algorithms on two selected biomarker sets to improve the sensitivity and accuracy of stressor diagnosis44,45. Within two selected biomarker sets, the further challenge is to validate the diagnostic performance of these selected biomarker sets. The evaluation of several well-known machine learning algorithms were therefore applied to four datasets separately. Results were summarized in Table 3, where sensitivity (recall) and specificity (precision) are displayed. The receiver operating characteristic (ROC) areas indicate a numerical value between 0 and 1 that describes the relationship between sensitivity and specificity for a given diagnostic test46,47. For the eight targeted key metabolites, good classification performance was concluded with ROC areas ranging between 0.64 and 0.951, indicating that our methodology was working well. Notably, combing the biomarkers sets from the targeted and untargeted approach resulted in ROC areas up to 0.99–1.0. This excellent performance can be mainly attributed to the biomarkers from the untargeted approach. With the growing evidence of nutritional stress as one major factor contributing to the synergistic effect of many other stressors and current bee colony declines18,19,20,48, this is the most clear test of the biomarker strategy concerning nutritional stress to date. Our results also suggest that combining metabolomics with data-driven machine learning algorithms has promising potential in evaluating bee health status and early risk assessment.
Biochemistry of low-carbohydrate nutritional stress
The third distinct feature is that we were able to investigate the underlying biochemistry mechanisms of low-carbohydrate stress, which can help to correlate overall results with the metabolites supervising both local and systemic response of bees to stressors49. Knowledge of the underlying biochemical pathway can help to identify additional markers perhaps not detected in the current approach. Under the introduced low-carbohydrate stress, a very characteristic decrease in metabolic profile regarding carbohydrates, and more abundant profiles of some compensating fatty acids and amino acids were observed. As could be expected, sucrose and fructose, two major sugars present in the supplied sugar syrup, were significantly decreased in experimental bees, although it was somewhat surprising that the blood sugar trehalose was extremely stable under low-carbohydrate stress (Fig. 3A). This decreased levels of carbohydrates is also observed in honeybees with energetic and nutritional stress imposed by Nosema ceranae infection17,22. Interestingly, the stable levels of trehalose was prolonged in control starved bees opposed to the infected ones22. Furthermore, the metabolism of several important amino acids, including histidine, arginine, asparagine, L-glutamine, acetylcarnitine, and homoserine, were positively impacted in bees fed with low-carbohydrate diet (Fig. 3B). Pathway analysis showed that several pathways were significantly disturbed including metabolism of amino acids, sugars, and nitrogen (Fig. 4A). Five pathways (aminoacyl-tRNA biosynthesis, alanine, aspartate and glutamate metabolism, arginine and proline metabolism, D-glutamine and D-glutamate metabolism, and starch and sucrose metabolism) with P value < 0.05 were considered to be significantly associated with low carbohydrate diet stress-induced metabolic changes. These down-regulated metabolites were only located into starch and sucrose metabolism, whereas, up-regulated metabolites were found in the other four metabolic pathways, including aminoacyl-tRNA biosynthesis, alanine, aspartate and glutamate metabolism, arginine and proline metabolism and D-glutamine and D-glutamate metabolism. To our knowledge, the obligatory precursor of protein synthesis is aminoacyl-tRNA (AA-tRNA)50, whereby aminoacyl-tRNA biosynthesis was the most disturbed pathway, suggesting an increased protein catabolism (denoted by a significant increase in glutamine) in those nutritionally stressed bees. These results provide evidence that nutritionally stressed bumblebees probably respond with increased protein catabolism, which has also been reported in Diporeia51.
Pathway analysis of altered metabolites. (A) Summary of pathway analysis with MetaboAnalyst 3.0. Larger circles, higher and closer to the Y-axis, show a higher impact of the concerned pathway on the organism. (B) Scheme summarizing the proposed mechanisms underlying the response of bumblebee to low-carbohydrate food stress.
These results encourage us to further explore the pathways linked to trehalose under nutritional stress in terms of metabolic homeostasis. Trehalose has been reported as the main energy source in insect hemolymph and as a stress protectant during extreme environmental conditions52. Since glutamine, the substrate of gluconeogenesis, was significantly increased (P < 0.0001) in hemolymph after feeding with low carbohydrate diet, we propose that gluconeogenesis may be significantly contributing to trehalose steady state in bumblebees with nutritional stress as shown in Fig. 4B. It is easy to understand that amino acids, the major fraction of the pollen, play a more important role in promoting responses to nutritional stress53. The supporting argument is that the endogenous amino acids are the main source utilized through gluconeogenesis, and the freshly ingested amino acids from pollen are to promote protein synthesis.
In conclusion, in this proof-of-concept study, we demonstrated that metabolomics-based methods, coupled with machine learning algorithms, represent valuable tools for the analysis of single bee stressor. In addition, this technique also shows power and potential as an assessment tool of bee health status in the real environment. The next stage should involve the identification of the untargeted biomarkers and development of a large cohort of wild sampling sites with various factors influencing bee health to test the categorizing accuracy of this approach for discovering biomarkers in multiple stressor risk assessment.
Materials and Methods
Study design
In this study, we aimed at retrieving and validating biomarkers for the introduced nutritional stress. It was considered important to have some physiological differences within our specimens, for which we had: (i) bees with different social hierarchy and age within the nest; (ii) bees facing different levels of malnutrition stress. The study was organized into three discrete phases as presented in Fig. 1: Phase A, the experiment setup, encompassing 2 × 8 microcolonies (control vs malnutrition), and bees were sampled at day 12. This experiment is regarded as “longer” exposure to stress. In each microcolony we sampled 3 workers and 1 dominant pseudo queen. Additionally, from 2 × 5 microcolonies we sampled one worker and one pseudo queen at day 6, where the stressor had less time to manifest. This experimental setup allowed us to create 8 datasets to investigate the effect of exposure time of the stressor, bee hierarchy, and their interaction, offering the ability to detect suitable metabolic markers related to nutritional stress. Table 1 represents the details of which groups of specimens were joined to create a specific dataset. Phase B, global hemolymph metabolic fingerprinting and profiling of samples from phase A, yielding two sets (from untargeted and targeted metabolomics) of potential biomarkers. Phase C, evaluation of the performance of the selected potential biomarkers using machine learning algorithms.
Bumblebee microcolonies and mimic of malnutrition: low carbohydrate food forage
All experiments were performed using commercial B. terrestris callow workers obtained from Biobest (Westerlo, Belgium). The callow or newborn workers were randomly collected from small queenright colonies at their initial phase of start-up. Each worker originated from a different queen. Five random callow workers were distributed as one microcolony and all the microcolonies were randomly assigned to the experimental or control groups, a total of 26 microcolonies with 130 bumblebees was used in this study. These microcolonies were placed in an incubator at 30 °C, 60% relative humidity, continuous darkness, and were all fed with gamma-irradiated pollen (Apihurdes, Pinofranqueado, Spain). Control groups (13 microcolonies) received a standardized sugar syrup (50 w/v%, BIOGLUC, Biobest) consisting of sucrose, fructose, dextrose and maltose, while experimental groups (13 microcolonies) received a 25% sugar syrup to mimic low carbohydrate nutritional stress (diluted in distilled water).
Hemolymph collection
Bee hemolymph was collected by making a small incision in the dorsal thorax and extracted for a total 10 μL per bee using Wiretrol II Capillary micropipettes (VWR) in phenylthiourea (PTU)-treated tubes to prevent melanization. The hemolymph sample was collected on ice and immediately put on dry ice afterwards. All hemolymph collection was performed under binocular microscope and three rules were strictly followed to guarantee the quality of sampling: i) the hemolymph should be pure and transparent; ii) no other tissues were perforated; iii) sampling time (incision and extraction) per bee is less than 35 seconds. All 84 samples were stored at −80 °C until chemical analysis.
Generic extraction of polar metabolites from bee hemolymph
Since there are no protocols available for bumblebee hemolymph extraction of polar metabolites, two different solvent systems were tested in a preliminary experiment, i.e. methanol and methanol-ethyl acetate (v/v, 1/1), whereby the latter proved more efficient in achieving high metabolome coverage (35% higher with methanol-ethyl acetate). As such, 40 μL of methanol-ethyl acetate mixture was used for the extraction of polar metabolites. To remove proteins, all hemolymph samples were precipitated with extraction solvent, and 5 μL internal standard valine-d8 (ISTD, 25 ng/μL) was pre-added. Subsequently, samples were incubated at 4 °C for 30 min to enhance protein precipitation and centrifugated at 15,000 g for 15 min at 4 °C to remove the resulting precipitate. Ultimately, the supernatant was transferred to a 1.5 mL microfuge tube, and consecutively dried using the Speed-Vac. All dried samples were suspended in 100 μL ultrapure water and transferred to an LC-MS vial with glass insert. Solvents used for extraction of hemolymph metabolites were of LC-MS grade, and obtained from Fisher Scientific (Loughborough, UK) and VWR International (Merck, Darmstadt, Germany).
UHPLC-Q-Orbitrap-HRMS analysis
The UHPLC-Orbitrap-MS method that was used in this study was adopted from54 as previously optimized55. An external standard mixture containing ca. 300 metabolites (including amino acids, monocarboxylic acids, phenols, multi-carboxylic acids, amines, carbohydrates, polyols, short chain fatty acids, inorganic acids, bile salts, and N-compounds) was used to assess instrumental performance and execute targeted profiling. A pool of all extracts (n = 84) was used to make quality control (QC) samples for instrument conditioning (external QC samples) and data normalization (internal QC samples). The Q-ExactiveTM Orbitrap mass analyzer (Thermo Fisher Scientific, San Jose, CA) was equipped with a heated electrospray ionization (HESI II) operating in polarity switching mode. An Acquity UPLC HSS T3 column (1.8 μm, 150 mm × 2.1 mm) (Waters, Zellik, Belgium) was used, whereby a binary solvent system using ultrapure water (A) and acetonitrile (B), both acidified with 0.1% formic acid, was applied at a constant flow rate of 0.4 mL min−1. All Solvents used were of LC-MS grade. Experimental samples were run in a randomized order (except for quality control samples, which were analyzed in duplicate after every nine experimental samples).
Untargeted data analysis and metabolic fingerprinting construction
The LC–MS raw data were first exported using XcaliburTM 2.1 (Thermo Fisher Scientific, CA) and imported into Compound Discoverer 3.0 software (Thermo Fisher Scientific, CA) with the untargeted metabolomics workflow and differential analysis mode. As major parameters, a minimum peak intensity of 500,000 a.u., retention time width of 0.25 min, m/z scan range from 53.4–800 dalton, and m/z width of 6 ppm were applied for feature extraction. The final data matrix was composed of the peak intensities for the detected components (rows) and different samples (columns), and was exported to an excel file. The coefficient of variation (%CV) was calculated for each component in the collection of QC samples, and components with %CV lower than 30%, which is considered an acceptable value of repeatability in untargeted metabolomics56, were retained. Data normalization was then performed by dividing the peak intensity of each metabolite in every sample by its corresponding mean peak intensity, as determined based on the following two internal QC samples54. To assess the metabolic differences between the defined samples sets, PCA (principal component analysis) and OPLS-DA (orthogonal partial least square-discriminant analysis) were performed in SimcaTM 13 (Umetrics, Malmo, Sweden). S-plots were built using validated OPLS-DA models in order to select metabolites that are important for classifying treatment (full data set) and different levels under treatment (day 6 and day 12 data set). A variable importance in projection (VIP) plot was applied to evaluate the importance of a certain components with VIP-value > 1.0.
Targeted data analysis and putative metabolic mechanisms underlying low-carbohydrate stress
XcaliburTM 2.1 (Thermo Fisher Scientific, San Jose, USA) was used to process the data (peak area determination) from metabolites that were identified based on the m/z and retention time from those metabolites that were included in the in-house database (±300 compounds). Statistical analysis was performed using SPSS 22.0 software, applying two-way ANOVA and Tukey HSD test for post hoc comparisons, whereby a P < 0.05 was considered as statistically significant. To locate those significantly expressed metabolites, the metabolic pathways were drawn based on the knowledge of those metabolites and the web-based platform MetaboAnalyst (http://www.metaboanalyst.ca/). Investigation of hemolymph molecules by targeted analysis is envisaged to correlate overall results including those from the untargeted fingerprinting and provide more biochemical information, hence, the underlying metabolic mechanisms of bees’ response to nutritional stress was further explored.
Machine learning algorithms implementation
To provide a quantitative diagnostic approach for adaption to a field-based monitoring, we need tools capable of extending the utility of the selected biomarker sets from a complex multivariate analysis to an applicable binary or categorizable format. Within this context, machine learning methods along with several more specific classification algorithms were tested to reveal the categorical accuracy of the candidate biomarker signatures. Standard implementations of the classification algorithms were performed with the Waikato Environment for Knowledge Acquisition (WEKA, University of Waikato, New Zealand, https://www.cs.waikato.ac.nz/ml/weka/)57 with 10-fold cross-validation settings.
Data Availability
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Gallai, N., Salles, J. M., Settele, J. & Vaissière, B. E. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 68, 810–821 (2009).
Williams, P. H. An annotated checklist of bumble bees with an analysis of patterns of description (Hymenoptera: Apidae, Bombini). Bull. Br. Mus. Nat. Hist., Entomol. ser. 67, 79–152 (1998).
Goulson, D. Bumblebees: their behaviour and ecology. (Oxford University Press, USA, 2003).
Potts, S. G. et al. Global pollinator declines: trends, impacts and drivers. Trends Ecol. Evol. 25, 345–353 (2010).
Kearns, C. A., Inouye, D. W. & Waser, N. M. Endangered mutualisms: the conservation of plant-pollinator interactions. Annu. Rev. Ecol. Evol. Syst. 29, 83–112 (1998).
González Varo, J. P. et al. Combined effects of global change pressures on animal-mediated pollination. Trends Ecol. Evol. 28, 524–530 (2013).
Meeus, I., Pisman, M., Smagghe, G. & Piot, N. Interaction effects of different drivers of wild bee decline and their influence on host-pathogen dynamics. Curr. Opin. Insect Sci. 26, 136–141 (2018).
Celli, G. & Maccagnani, B. Honey bees as bioindicators of environmental pollution. BMC Genomics 56, 137–139 (2003).
Badiou Bénéteau, A. et al. Honeybee biomarkers as promising tools to monitor environmental quality. Environ. Int. 60, 31–41 (2013).
Bargańska, Ż., Ślebioda, M. & Namieśnik, J. Honey bees and their products: bioindicators of environmental contamination. Crit. Rev. Environ. Sci. Technol. 46, 235–248 (2016).
Urbanczyk Wochniak, E. et al. Parallel analysis of transcript and metabolic profiles: a new approach in systems biology. EMBO Rep. 4, 989–993 (2003).
Horgan, R. P., Clancy, O. H., Myers, J. E. & Baker, P. N. An overview of proteomic and metabolomic technologies and their application to pregnancy research. BJOG 116, 173–181 (2009).
Lin, C. Y., Viant, M. R. & Tjeerdema, R. S. Metabolomics: methodologies and applications in the environmental sciences. J. Pestic. Sci. 31, 245–251 (2006).
Bundy, J. G., Davey, M. P. & Viant, M. R. Environmental metabolomics: a critical review and future perspectives. Metabolomics 5, 3 (2009).
Thompson, S. N., Lee, R. W. K. & Beckage, N. E. Metabolism of parasitized Manduca sexta examined by nuclear magnetic resonance. Arch. Insect Biochem. Physiol. 13, 127–143 (1990).
Cox, J. E., Thummel, C. S. & Tennessen, J. M. Metabolomic studies in Drosophila. Genetics 206, 1169–1185 (2017).
Aliferis, K. A., Copley, T. & Jabaji, S. Gas chromatography–mass spectrometry metabolite profiling of worker honey bee (Apis mellifera L.) hemolymph for the study of Nosema ceranae infection. J. Insect Physiol. 58, 1349–1359 (2012).
Wright, G. A., Nicolson, S. W. & Shafir, S. Nutritional physiology and ecology of honey bees. Annu. Rev. Entomol. 63, 327–344 (2018).
Vaudo, A. D., Tooker, J. F., Grozinger, C. M. & Patch, H. M. Bee nutrition and floral resource restoration. Curr. Opin. Insect Sci. 10, 133–141 (2015).
Naug, D. Nutritional stress due to habitat loss may explain recent honeybee colony collapses. Biol. Conserv. 142, 2369–2372 (2009).
Vanbergen, A. J. Threats to an ecosystem service: pressures on pollinators. Front. Ecol. Environ. 11, 251–259 (2013).
Mayack, C. & Naug, D. Parasitic infection leads to decline in hemolymph sugar levels in honeybee foragers. J. Insect Physiol. 56, 1572–1575 (2010).
Vidau, C. et al. Differential proteomic analysis of midguts from Nosema ceranae-infected honeybees reveals manipulation of key host functions. J. Invertebr. Pathol. 121, 89–96 (2014).
Kurze, C. et al. Nosema spp. infections cause no energetic stress in tolerant honeybees. Parasitol Res. 115, 2381–2388 (2016).
Williams, P. H. & Osborne, J. L. Bumblebee vulnerability and conservation world-wide. Apidologie 40, 367–387 (2009).
Doorn, A. V. Factors influencing dominance behaviour in queenless bumblebee workers (Bombus terrestris). Physiol. Entomol. 14, 211–221 (1989).
Blatt, J. & Roces, F. Haemolymph sugar levels in foraging honeybees (Apis mellifera carnica): dependence on metabolic rate and in vivo measurement of maximal rates of trehalose synthesis. J. Exp. Biol. 204, 2709–2716 (2001).
Sacktor, B. Regulation of intermediary metabolism, with special reference to the control mechanisms in insect flight muscle. Adv. Insect Physiol. 7, 267–347 (1970).
Rothe, U. & Nachtigall, W. Flight of the honey bee. J. Comp. Physiol. B 158, 739–749 (1989).
Neukirch, A. Dependence of the life span of the honeybee (Apis mellifica) upon flight performance and energy consumption. J. Comp. Physiol. 146, 35–40 (1982).
Panzenböck, U. & Crailsheim, K. Glycogen in honeybee queens, workers and drones (Apis mellifera carnica Pollm.). J. Insect Physiol. 43, 155–165 (1997).
Remolina, S. C., Hafez, D. M., Robinson, G. E. & Hughes, K. A. Senescence in the worker honey bee Apis mellifera. J. Insect Physiol. 53, 1027–1033 (2007).
Consortium, H. G. S. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931 (2006).
Kapheim, K. M. et al. Genomic signatures of evolutionary transitions from solitary to group living. Science 348, 1139–1143 (2015).
Kocher, S. D. et al. The draft genome of a socially polymorphic halictid bee, Lasioglossum albipes. Genome Biol. 14, R142 (2013).
Park, D. et al. Uncovering the novel characteristics of Asian honey bee, Apis cerana, by whole genome sequencing. BMC Genomics 16, 1 (2015).
Sadd, B. M. et al. The genomes of two key bumblebee species with primitive eusocial organization. Genome Biol. 16, 76 (2015).
Grozinger, C. M. & Robinson, G. E. The power and promise of applying genomics to honey bee health. Curr. Opin. Insect Sci. 10, 124–132 (2015).
Johnson, R. M., Evans, J. D., Robinson, G. E. & Berenbaum, M. R. Changes in transcript abundance relating to colony collapse disorder in honey bees (Apis mellifera). Proc. Natl. Acad. Sci. USA 106, 14790–14795 (2009).
Wallberg, A. et al. A worldwide survey of genome sequence variation provides insight into the evolutionary history of the honeybee Apis mellifera. Nat. Genet. 46, 1081 (2014).
Hu, H. et al. Proteome analysis of the hemolymph, mushroom body, and antenna provides novel insight into honeybee resistance against Varroa infestation. J. Proteome Res. 15, 2841–2854 (2016).
Han, B., Zhang, L., Feng, M., Fang, Y. & Li, J. An integrated proteomics reveals pathological mechanism of honeybee (Apis cerena) sacbrood disease. J. Proteome Res. 12, 1881–1897 (2013).
Authority, E. F. S. Towards an integrated environmental risk assessment of multiple stressors on bees: review of research projects in Europe, knowledge gaps and recommendations. EFSA J. 12, 3594 (2014).
Sajda, P. Machine learning for detection and diagnosis of disease. Annu. Rev. Biomed. Eng. 8, 537–565 (2006).
Swan, A. L., Mobasheri, A., Allaway, D., Liddell, S. & Bacardit, J. Application of machine learning to proteomics data: classification and biomarker identification in postgenomics biology. OMICS 17, 595–610 (2013).
Lasko, T. A., Bhagwat, J. G., Zou, K. H. & Ohno Machado, L. The use of receiver operating characteristic curves in biomedical informatics. J. Biomed. Inform. 38, 404–415 (2005).
Swets, J. A. Measuring the accuracy of diagnostic systems. Science 240, 1285–1293 (1988).
Trapp, J., McAfee, A. & Foster, L. J. Genomics, transcriptomics and proteomics: enabling insights into social evolution and disease challenges for managed and wild bees. Mol. Ecol. 26, 718–739 (2017).
Boccard, J. et al. Standard machine learning algorithms applied to UPLC-TOF/MS metabolic fingerprinting for the discovery of wound biomarkers in Arabidopsis thaliana. Chemom. Intell. Lab. Syst. 104, 20–27 (2010).
Ibba, M. & Söll, D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 69, 617–650 (2000).
Maity, S. et al. Starvation causes disturbance in amino acid and fatty acid metabolism in Diporeia. Comp. Biochem. Physiol. B 161, 348–355 (2012).
Liebl, M., Nelius, V., Kamp, G., Ando, O. & Wegener, G. Fate and effects of the trehalase inhibitor trehazolin in the migratory locust (Locusta migratoria). J. Insect Physiol. 56, 567–574 (2010).
Crailsheim, K. The protein balance of the honey bee worker. Apidologie 21, 417–429 (1990).
Vanden Bussche, J., Marzorati, M., Laukens, D. & Vanhaecke, L. Validated high resolution mass spectrometry-based approach for metabolomic fingerprinting of the human gut phenotype. Anal. Chem. 87, 10927–10934 (2015).
De Paepe, E. et al. A validated multi-matrix platform for metabolomic fingerprinting of human urine, feces and plasma using ultra-high performance liquid-chromatography coupled to hybrid orbitrap high-resolution mass spectrometry. Anal. Chim. Acta 1033, 108–118 (2018).
Michopoulos, F., Lai, L., Gika, H., Theodoridis, G. & Wilson, I. UPLC-MS-based analysis of human plasma for metabonomics using solvent precipitation or solid phase extraction. J. Proteome Res. 8, 2114–2121 (2009).
Frank, E., Hall, M., Trigg, L., Holmes, G. & Witten, I. H. Data mining in bioinformatics using Weka. Bioinformatics 20, 2479–2481 (2004).
Jung, J. Y. et al. 1H-NMR-based metabolomics study of cerebral infarction. Stroke 42, 1282–1288 (2011).
Acknowledgements
Luoluo Wang is the recipient of a doctoral grant from China Scholarship Council (CSC) and co-funded by Ghent University (BOF-UGent). This work is supported by the Research Foundation-Flanders (FWO-Flanders).
Author information
Authors and Affiliations
Contributions
L.W., I.M. and G.S. conceived the presented idea. L.W. carried out the experiment and wrote the manuscript with help from I.M., C.R., L.V.M. and L.V. helped with the method optimization, and data analysis. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, L., Meeus, I., Rombouts, C. et al. Metabolomics-based biomarker discovery for bee health monitoring: A proof of concept study concerning nutritional stress in Bombus terrestris. Sci Rep 9, 11423 (2019). https://doi.org/10.1038/s41598-019-47896-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47896-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.