Abstract
Enterococci are opportunistic pathogens that can form biofilms during infections and many virulence determinants are involved in this process. Although the virulence factors are often analysed in Enterococcus spp. from humans and food animals, little is known about gut enterococcal isolates from wild birds. Therefore, the determination of virulence factors among enterococci isolated from wild birds may provide new information about a possible source of infection for humans and animals or vice versa via the environment. We analysed different phenotypic and genotypic traits in enterococci from wild birds related to potential virulence in humans and animals and to evaluate biofilm formation and its relationship to virulence genes. The E. faecalis isolates were characterised by greater frequency of biofilm formation in BHI than E. faecium. There was a correlation between hydrophobicity and biofilm formation in BHI broth in E. faecalis. None of the isolates was haemolytic. The presence of some adhesion and gelatinase genes was detected in biofilm-positive isolates. The enterococcal pathogenic factors (esp, hyl, and cyl operon genes) did not seem to be necessary or sufficient for production of biofilm by analysed bacteria. Enterococcus species isolated from wild birds should be considered as a possible source of some virulence determinants.
Similar content being viewed by others
Introduction
Biofilm is considered as an essential factor in the pathogenesis of various opportunistic bacteria, e.g. during enterococcal infections1. Moreover, many virulence determinants in Enterococcus spp. are involved in the pathological process and the ability to growth in biofilm2.
An important step in induction of infection and/or biofilm production is the adhesion of bacterial cells to host tissues. Surface proteins that bind to receptors on the surface of eukaryotic cells or extracellular matrix proteins are adhesion factors; in enterococcal species these factors mainly include enterococcal surface protein (encoded by esp), aggregation substance (encoded by agg or asa1), and collagen-binding protein (encoded by ace)1. Production of adhesin-like E. faecalis and E. faecium endocarditis antigen A (encoded by efaAfs and efaAfm, respectively) is also involved in enterococcal adhesion to biotic and abiotic surfaces. Moreover, expression of pili (encoded by ebpABC locus, srt, and pil) on the cell surface is considered as an important virulence factor facilitating cell adhesion and biofilm formation2.
Enterococcal cells are capable of communicating via peptide pheromones (e.g. encoded by cpd, cob, and ccf), which are secreted by recipient cells to induce the conjugative apparatus of donor cells. In this way they mediate the transfer of pheromone-responsive plasmids, which may carry virulence genes that promote biofilm formation or regulation. In response to increased cell population densities, important virulence factors in Enterococcus spp. are regulated by quorum sensing, including the fsr (faecal streptococci regulator) locus. Changes in the activity of particular genes lead to maturation of the biofilm and the appearance of appropriate phenotypic traits, depending on the conditions. In the case of Enterococcus spp., cytolysin, i.e. a haemolytic exotoxin (encoded by cyl), gelatinase (encoded by gelE), serine protease (encoded by sprE), and hyaluronidase (encoded by hyl) are the most frequent virulence factors affecting host cells by their toxic or destructive effects3.
Enterococci are Gram-positive elements of the gut microbiota and significant agents of opportunistic infections in animals and humans4,5. Enterococcus spp. from avian species associated with diseases include mainly Enterococcus faecalis, E. faecium, E. durans, E. hirae and E. cecorum6,7,8. The collection of Enterococcus isolates from wild birds analysed in this study, especially E. faecalis and E. faecium, are resistant to multiple antibiotics and represent different epidemic clones (e.g. CC17, CC81, CC116)9,10. The same clones have previously been found in hospitalised patients in Poland and other European countries, as well as globally in unhospitalised patients, livestock, pets, food and wastewater11,12,13. For bacteria to be pathogenic, in addition to antibiotic resistance they must also possess virulence factors14. In addition, enterococci are naturally able to acquire, accumulate, and transmit extrachromosomal elements encoding virulence traits15. Thus, wild birds may be considered a source of enterococci that are potentially pathogenic as well as resistant to antibiotics. Transmission of such enterococcal strains from wild birds to humans and animals is possible via environment, such as: water polluted with faeces, soil, dietary patterns in wild animals/birds, meat or direct contact during hunting in humans1. Taking into account migration of birds, the same etiological agent can occur in distant places.
Monitoring faecal Enterococcus spp. as indicator bacteria in different populations, including wild birds makes it feasible to compare the prevalence of virulence in determined STs/CC and to detect the transfer of potentially pathogenic strains from animals to humans and vice versa. However, a few reports of the prevalence of virulence factors in antibiotic resistant enterococci from wild animals have been published16,17,18,19.
The objective of this work is to analyse different phenotypic and genotypic traits in enterococcal isolates from the gut microbiota of different wild birds related to potential virulence in humans and animals and to evaluate in vitro biofilm formation and its relationship to presence of virulence genes.
Results
Haemolytic and gelatinase activities
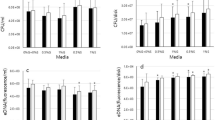
None of the isolates was haemolytic on 5% horse blood agar. Gelatinase was produced by the 27 isolates of E. faecalis (100%), one isolate of E. faecium (20%), and one isolate of E. hirae (33.3%) of the gelE-positive strains (Table S1).
Hydrophobicity and biofilm formation
The in vitro hydrophobicity test revealed twenty strains with hydrophobicity higher than 50%; eighteen of them represented E. faecalis and the other two strains belonged to the species E. faecium and E. casseliflavus. The E. faecalis strains showed significantly higher hydrophobicity than E. faecium (P < 0.001; Chi-square test), i.e. as many as 33.3% of the E. faecalis isolates displayed hydrophobicity at the level of 100% and another 33.3% exhibited %H in the range of 50–70%. Detailed data are shown in Table S1.
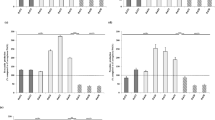
Biofilm formation by the tested E. faecalis and E. faecium detected with the microtitre plate method in BHI broth supplemented with 2% glucose was statistically significantly higher than in TSB broth supplemented with 1% glucose (P < 0.001; Chi-square test). Both species differed significantly in biofilm formation in TSB broth supplemented with 1% glucose (P = 0.00886; Chi-square test). However, the E. faecalis and E. faecium species did not differ in biofilm formation in BHI broth supplemented with 2% glucose (P = 0.59083; Fisher’s exact test) (Table S1). Among a total of isolates the ability to biofilm formation in BHI was observed in 87% (47 isolates); 70.4% (38 isolates) were classified as moderately or strongly adherent and 16.7% (9 isolates) were weakly adherent. Seven isolates (13%) showed no ability to form biofilm in BHI broth (Table 1).
In the E. faecalis strains, there was a link between biofilm formation in BHI broth supplemented with 2% glucose and hydrophobicity (P = 0.00718; Fisher’s exact test). However, there was no correlation between biofilm formation and hydrophobicity in the E. faecium strains in the same conditions (P = 0.83333; Fisher’s exact test).
Detection of virulence genes
The prevalence of virulence genes detected in all isolates is shown in Table 2. No esp, hyl, cylA, cylB, cylM, and cylL genes were detected in any of the tested isolates.
The screening results of pili components are summarised as follows: ebpA was present in E. faecalis (70.4%, 19 isolates), E. faecium (22.2%, 4 isolates) and E. hirae (20%, 1 isolate); ebpB was found only in E. faecalis (85.2%, 23 isolates); ebpC was present in E. faecalis (74.1%, 20 isolates), E. faecium (27.8%, 5 isolates), E. hirae (60%, 3 isolates), and E. casseliflavus (100%, 2 isolates); pil was detected in E. faecalis (70.4%, 19 isolates) and E. hirae (20%, 1 isolate), and srt was present in E. faecalis (96.3%, 26 isolates), E. faecium (33.3%, 6 isolates), and E. hirae (60%, 3 isolates). There was a correlation between biofilm formation in BHI broth supplemented with 2% glucose and the presence of the pil gene in the enterococcal bacteria (P = 0.03010; Chi-square test).
E. faecalis and E. faecium isolates differ in the number of ebpA, ebpB, ebpC and srt genes. The prevalence of ebpA (P = 0.00422; Chi-square test), ebpB (P = 0.0000; Chi-square test), ebpC (P = 0.00586; Chi-square test) and srt (P = 0.0000; Chi-square test) genes were significantly higher among E. faecalis isolates (19strains/70.4%, 23 strains/85.2%, 20 strains/74.1% and 26 strains/96.3%, respectively) than among E. faecium isolates (4 strains/22.2%, 0 strains/0%, 5 strains/27.8% and 6 strains/33.3%, respectively). The presence of pil gene between E. faecalis and E. faecium isolates was not found to be statistically significant (P = 0.0535; Fisher’s exact test). In addition, the pili genes were also found in E. hirae and E. casseliflavus isolates. The predominant gene was ebpC (E. hirae – 3 isolates/60% and E. casseliflavus – 2 isolates/100%). Moreover, the presence of ebpA, pil and srt was seen in only one E. faecalis isolate. Nine (33.3%) of E. faecalis and one (5.6%) E. faecium isolates co-harboured the genes agg and asa1. Additionally, the asa1 gene was found in one (5.6%) E. faecium and one (20%) E. hirae isolate. The E. faecalis and E. faecium isolates did not differ statistically for the presence of the asa1 gene (P = 0.08689; Fisher’s exact test); however, there was a difference in the presence of the agg gene (P = 0.02909; Fisher’s exact test).
The efaAfs and efaAfm genes were found in all E. faecalis and E. faecium isolates, respectively. None of these genes was detected in E. hirae, E. casseliflavus, and E. durans.
The presence of the gelE and sprE genes associated with the fsrABC locus was detected in all 27 E. faecalis isolates, 5 E. faecium, and 1 E. hirae. Additionally, fsrA and fsrC were found in two gelE- and sprE-positive E. hirae isolates. Moreover, the ace gene was detected in the fsrABC-positive isolates, except one E. faecalis.
The sex pheromone determinants (cpd, cob, ccf) were widespread in the analysed E. faecalis isolates (100% strains). The ccf gene was also detected in all but one E. faecium. Moreover, the cpd and cob determinants were showed in E. faecium (five and two isolates, respectively) and E. hirae (tree and two isolates, respectively).
The genotypic patterns of the virulence factors detected in the isolates are shown in Table S1.
Discussion
The study was conducted to determine the prevalence of biofilm-forming ability among gut enterococci isolated from wild birds and its correlation with virulence genes.
There are no literature reports of biofilm formation and hydrophobicity in enterococci isolated from wild birds. In in vitro conditions, the microplate method was found to be the most common and effective approach for detection of biofilm production20. As was noted during our studies, the composition of the medium was important to show the ability of enterococci to in vitro biofilm formation. Other authors also pointed to similar dependencies20,21.
Moreover, we observed that the E. faecalis strains produced biofilm in BHI more often than E. faecium. Leuck et al.22 recently tested the biofilm-forming ability of E. faecalis isolates on polystyrene dishes. The authors reported that many of the clinical isolates showed low-level biofilm formation even when different types of media were used. However, they observed that clinical isolates were able to form biofilms on a relevant tissue surface.
Additionally, we found a correlation between hydrophobicity and biofilm formation in only E. faecalis. In addition, the E. faecalis isolates showed significantly higher hydrophobicity than E. faecium. It is known, that bacterial cell surface hydrophobicity is important for the interactions between the bacterium and host epithelial cells23. The hydrophobicity of the enterococcal cell surface is increased by the presence of aggregation substances. Strains possessing the agg or asa1 gene may form large cell aggregates during infection24. In the present study, the genetic determinants of aggregation substances were most frequently detected in E. faecalis, followed by E. faecium and E. hirae. The aggregation substance is a sex pheromone plasmid-encoded surface protein. Some of E. faecalis strains with all sex pheromone genes (cpd, cob, and ccf) exhibited the presence of the agg and asa1 genes as well. Similarly, Martin and coworkers25 noted the presence of the agg, cpd, and ccf genes in all E. faecalis isolates. Additionally, production of sex pheromones by E. faecalis may favour acquisition of antibiotic resistance and virulence from other enterococci, resulting in increased virulence. In the present study, a lower proportion of sex pheromone genes were observed in E. faecium in comparison with E. faecalis isolates. However, the sex pheromone genes in E. faecium may reflect sequence divergence, which may explain this result. Importantly, Eaton and Gasson26 showed that virulence determinants can be transferred from pathogenic strains to non-pathogenic strains (used as starters in food). The authors did not achieve transfer into E. faecium strains, although sex pheromone cross talk between E. faecium and E. faecalis has been demonstrated27.
Our finding corroborates the results reported by Martin and coworkers25, who demonstrated that all E. faecalis and E. faecium strains from Wood Pigeon (Columba palumbus) carried the efaAfs and efaAfm virulence genes, respectively, while the ace gene was more often found in E. faecalis strains from wild birds in Poland than from Partridges in Portugal16. Both efaAfs and ace genes play a role in the pathogenesis of endocarditis, whereas the role of efaAfm is yet unknown.
Some authors demonstrated that the presence of pili genes in enterococcal strains is required for the establishment of the first step of infection28. As shown in our study, most of the analysed Enterococcus spp. carried these genes; however, a correlation was only observed between biofilm formation in BHI with 2% glucose medium and the presence of the pil gene.
No esp, hyl, cylA, cylB, cylM, cylL genes were detected in any of the tested strains in our studies. The absence of the cyl operon was associated with negative β-haemolysis. Similarly, no β-haemolytic activity was found in enterococal strains from wild boars (Sus scrofa) in Spain, although the strains contained different combinations of cyl genes, in contrast to our results17. Olsen and coworkers29 showed that the cylA gene was detected more often in clinical than in commensal poultry isolates, where none of the isolates was haemolytic. As indicated previously, cytolysin activity requires the presence of the whole cyl operon (cyl LLLSABM)26,30. This was also confirmed in the study conducted by Silva and coworkers31. Cytolysin exerts activity against a broad spectrum of cell types including a wide range of Gram-positive bacteria, eukaryotic cells such as human, bovine, and horse erythrocytes, retinal cells, polymorphonuclear leukocytes, and human intestinal epithelial cells32.
As in our results, Silva and coworkers16 did not detect any esp genes in enterococci isolated from Partridge (Alectoris rufa), whereas this gene was described seven years later in six vanA-positive E. faecium isolated from the same bird species33. Additionally, the esp gene was found in two vanA/B2-positive E. faecalis strains34. Interestingly, the esp gene in E. faecalis is located on a large genetic component (150 kb) characterised by all features of the pathogenicity island, whose presence is characteristic for multiresistant isolates, including vancomycin-resistant35. Indeed, the esp gene is present predominantly in strains associated with infections and hospital outbreaks36,37.
In contrast to the isolates from the wild birds analysed in our study, the hyl gene was detected in two and one E. faecium strains isolated from Wild Boars and Partridges, respectively16,17. Moreover, the hyl gene was found in five vanA-positive E. faecium strains from wild partridges33 and three vanA/vanB2 E. faecalis strains from two Cattle Egrets and one Common Ringed Plover34. Gram-positive genera capable of elaborating hyaluronidase are able to cause infections initiated at a mucosal or skin surface of either humans or animals38.
It was reported in our study that all E. faecalis strains that exhibited gelatinase activity harboured the gelE, sprE, and fsrABC genes, which is in agreement with results reported by other authors17,18. Additionally, only one E. faecium and one E. hirae strains had gelatinase activity and harboured gelE, sprE, and all fsr operon genes. A discrepancy between the presence of gelE19 including sprE and fsrABC genes18 and production of gelatinase in enterococci was also observed which coincide with our results. Gelatinase (gelE) is co-transcribed with serine protease (sprE) and regulated by the quorum-sensing fsr locus. It can also cleave sex pheromones, which are known to be potent chemo-attractants39 and might therefore modulate the host response.
Many authors indicate the presence of numerous genes and virulence factors in both pathogenic and opportunistic bacteria28,40,41,42. However, they do not specify which of them may have of greatest importance for pathomechanism of infections, because it is a complex and multi-stage process and depends of many factors, including bacterial virulence as well as the conditions of the host and habitat and the presence of another components of microbiota. Similarly, it is difficult to do so in the case of enterococci isolated during our studies from the gut microbiota of wild birds. However, based on our findings, commensal enterococci from the wild birds had some virulence determinants and could be a source of potential pathogenic strains for humans and animals, especially that some of them were determined as antibiotic resistant epidemic clones. This hypothesis can be confirmed by the recent results of investigations of virulence factors in vancomycin-resistant enterococci from wild birds obtained by Ben Yahia and coworkers34, showing that vanA/vanB2 E. faecalis strains can also harbour the important virulence determinants.
Conclusion
In conclusion, the data presented in this study can help to elucidate the prevalence of virulence factors in enterococcal isolates from wild birds in Poland and indicate that Enterococcus species should be considered as a possible source for virulence determinants. None of the analysed genes should be considered definitive markers of pathogenicity in the tested bacteria. Moreover, the results of this study showed that the presence of pathogenic factors such as the esp, hyl, and cyl operon genes did not seem to be necessary or sufficient for the production of biofilm by enterococci in the analysed conditions. However, the presence of some adhesion and gelatinase genes has been detected in biofilm-positive isolates. It appears that many environmental conditions, e.g. the medium composition, and genetic factors may be associated with the pathomechanism and production of biofilm by enterococci. Therefore, the environment, e.g. organs outside the gastrointestinal tract where the bacteria live, affects their surface activity and intercellular interactions. In some cases, our strains also possessed silent virulence genes.
Methods
Collection of strains
The collection of 54 Enterococcus isolates (E. faecalis, 27 isolates; E. faecium, 18 isolates; E. hirae, 5 isolates; E. durans, 2 isolates; and E. casseliflavus, 2 isolates) from cloacal swabs of 52 free-living birds representing 25 species was studied (Table S1). The swabs for bacteriological analysis were collected from birds after their delivery to the Centre for Rehabilitation of Wild Birds, University of Life Sciences in Lublin, which receives injured or weak birds. The cloacal samples from the birds were collected by a veterinarian as part of his work and based on the authorization to collect biological material for research purposes by the Regional Directorate for Environmental Protection (WPN.6401.45.2015.MPR.1). The study was conducted in an ethical and responsible manner, in full compliance with all relevant codes of experimentation and legislation. Enterococci were isolated on Bile Esculin Azide Lab-Agar (Biocorp, Warsaw, Poland) at 37 °C for 24–48 h and identified to the species level by Ultraflextreme Matrix Assisted Laser Desorption Ionization Time of Flight Mass Spectrometry (MALDI-TOF MS) with MALDI-Biotyper 3.0 software (Bruker Daltonics, Bremen, Germany) and rpoA gene sequencing, as previously described43. In addition, antibiotic resistance and genetic diversity of analysedstrains were determined previously9,10. The isolates were stored at −80 °C in Brain Heart Infusion Broth (Oxoid, Basingstoke, Hampshire, UK) with 20% sterile glycerol for further analysis.
Haemolysin and gelatinase activity screening
Haemolysis was evaluated by plating the strains on Columbia Agar Base (OXOID, Basingstoke, Hampshire, UK) supplemented with 5% defibrinated horse blood (Pro Animali Company, Wroclaw, Poland). The plates were incubated at 37 °C for 24 h in aerobic conditions. A positive result was indicated by the formation of haemolytic (clear) zones around the colonies. E. faecalis ATCC29212 (LGC Standards, Łomianki, Poland) was used as a positive control.
Gelatinase production was detected by inoculating the Enterococcus strains onto Trypticase Soy Agar (OXOID, Basingstoke, Hampshire, United Kingdom) containing 3% gelatine (Avantor Performance Materials, Gliwice, Poland). The appearance of a clear halo around the colonies after incubation at 37 °C for 24 h in aerobic conditions followed by refrigeration at 4 °C for 30 min was considered to be a positive indication of gelatinase production. E. faecalis ATCC29212 (LGC Standards, Łomianki, Poland) was used as a positive control.
Hydrophobicity and biofilm assays
Cell surface hydrophobicity was determined using the method developed by Dec and coworkers44. Each isolate was subcultured on Columbia Agar Base (OXOID, Hampshire, UK) supplemented with 5% defibrinated horse blood (Pro Animali Company, Wroclaw, Poland) at 37 °C. 24-h cultures were harvested and suspended in 5 ml of 0.9% NaCl to an optical density (OD600) of 0.8–1.0 (A0). Then, xylene (1.7 ml) was added to glass test tubes and the mixtures were vortexed vigorously for 90 s. After phase separation (ca. 15 min.), the optical density of the aqueous phase (A) was measured again and compared with the initial value. The percentage of cell surface hydrophobicity (%H) of the strain adhering to xylene was calculated using the equation: %H = [(A0 − A)/A0] × 100. Strains with hydrophobicity equal or higher than 50% were considered hydrophobic.
Biofilm assays were conducted based on a method described by Stepanović and coworkers20 using Tryptic Soy Broth (TSB) (Oxoid, Hampshire, UK) supplemented with 1% glucose and Brain Heart Infusion (BHI) (Oxoid, Hampshire, UK) supplemented with 2% glucose. Each isolate was subcultured on Columbia Agar Base (OXOID, Hampshire, UK) supplemented with 5% defibrinated horse blood (Pro Animali Company, Wroclaw, Poland) at 37 °C. After verification of the purity of the strain, a few colonies with identical morphology are suspended in physiological saline. Then, the turbidity of the bacterial suspension was adjusted to match turbidity comparable to that of the 0.5 McFarland standard (~108 CFU/ml) using a densitometer Biosan DEN-1 (Biogenet, Józefów-Otwock, Poland). Then, for each strain tested, 20 µl of bacterial suspensions were transferred to four wells of two separate sterile flat-bottomed 96-well polystyrene microtitre plates containing 180 µl of TSB supplemented with 1% glucose and 180 µl of BHI supplemented with 2% glucose, respectively. For the negative control, 200 µl of broths (TSB and BHI, both with glucose) were dispensed into eight vertical wells per plate. The plates were incubated under stationary aerobic conditions at 37 °C. After incubation for 24 hours, the broths were carefully removed. The wells were gently washed three times with phosphate-buffered saline (PBS, pH 7.2). Following every washing step, the wells were emptied by flicking the plates. Prior to biofilm staining, the plates were left at room temperature for drying in an inverted position overnight. The adherent biofilm layer formed in each microtitre plate well was stained with 200 µl of 0.1% crystal violet solution in water for 15 min at room temperature. After staining, the stain was aspirated with a pipette and excess stain was rinsed off by placing the microtitre plate under running tap water. Washing was continued until the washings were free of the stain. After the microplates were dried at room temperature, the dye bound to the cells was resolubilised with 200 µl of 96% ethanol per well for 30 min without shaking. The optical density (OD) of the resolubilised crystal violet was then measured at 570 nm (OD570) using a microplate reader (Bio-Rad, Model 680). Each assay was performed in quadruplicate on three occasions for 12 readings for each strain. Wells containing uninoculated medium served as negative controls to determine the background optical density. After subtracting the mean background OD570 readings, the 12 optical density readings per strain were averaged to obtain the mean OD570 reading for each strain. Based on the bacterial biofilm, the isolates were classified into four categories: non-biofilm producers, weak, moderate, or strong biofilm producer. The isolates were classified as follows: OD < ODc = non-biofilm producers (category I); ODc < OD < 2ODc = weak biofilm producers (category II); 2ODc < OD < 4ODc = moderate biofilm producers (category III); and OD > 4ODc = strong biofilm producers (category IV). The cut-off OD (ODc) was defined as three standard deviations above the mean OD of the negative control.
PCR
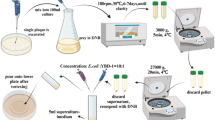
DNA was extracted using a commercial GeneMATRIX Bacterial & Yeast Genomic DNA Purification Kit (Eurx, Poland). To improve the nucleic acid extraction efficiency, lysozyme was used in the enzymatic lysis step. The presence of genes encoding putatitve virulence factors of the Enterococcus strains was evaluated using PCR with specific primers that encode the endocarditis- and biofilm-associated pili genetic locus (ebpABC) and an adjacent downstream sortase-encoding gene (srt), pili (pil), aggregation substance (agg and asa1), collagen binding protein (ace), enterococcal surface protein (esp), enterococcal endocarditis antigen (efaAfs for E. faecalis and efaAfm for E. faecium), gelatinase (gelE), serine protease (sprE), hyaluronidase (hyl), cytolysin (cylA, cylB, cylM and cylLL), the quorum sensing locus fsr (fsrA, fsrB, fsrC), and sex pheromones (cpd, cob and ccf). These virulence genes were chosen, because they are detected most frequently in clinical isolates of enterococci. All primer sequences are listed in Table S2. PCR was performed on a T100 thermal cycler (Bio-Rad Laboratories Inc., Hercules, CA, USA) in a final volume of 25 µl containing 1 µl (~20 ng) of DNA as template; 2.5 µl of reaction buffer (10x); 0.2 µl of Taq DNA polymerase (5U/1 µl); 1 µl of each of the two primers (10 pmol/1 µl, Genomed, Warsaw, Poland), 0.2 µl of 25 mM dNTPs MIX; and PCR pure water to a final volume. The reagents used in the PCR mixtures were purchased from AmpliKIT Allegro Taq (Novazym, Poznań, Poland).
The initial 5-min denaturation step at 95 °C was followed by 35 cycles of 1-min denaturation at 95 °C, annealing at the different temperatures shown in Table 1 for 1 min, and an extension at 72 °C for 1 min, followed by final extension at 72 °C for 8 min. The PCR products were analysed in 1.5% agarose–Tris–borate–EDTA gel containing 0.5 mg of ethidium bromide per ml and then visualised with a gel imaging analysis system with Quantity One software (Bio-Rad; Hercules, CA, USA). E. faecalis ATCC29212 (ace, asa1, gelE, efaAfs, cpd, cob, ccf, cylA), E. faecalis P33 (cylA, cylB, cylM, cylLL, esp), E. faecalis ATCC27285 (agg), E. faecalis OG1RF (ace, fsrA, fsrB, fsrC, gelE, sprE, ebpA, ebpB, ebpC, pil, srt), E. faecium ATCC19434 (efaAfm) were used as positive controls. E. faecalis P33 (sequence type 16) came from our collection and was isolated from one-day old broiler chickens with yolk sac infection. Nuclease-free water was used as a negative control.
Statistical analysis
All statistical analyses were performed using STATISTICA program version 13.1 (StatSoft Inc., 2014, Tulsa, OK, USA). A Chi-square independence test was performed to analyze the data. Additionally, to test the prevalence of a particular virulence factor among E. faecalis and E. faecium the Chi-square independence test with Yates correction or Fisher’s exact test for small samples (≤5) were used. P ≤ 0.05 was considered statistically significant.
References
Mohamed, J. & Huang, D. B. Biofilm formation by enterococci. J Med Microbiol 56, 1581–1588 (2007).
Hashem, Y. A., Amin, H. M., Essam, T. M. & Aziz, R. K. Biofilm formation in enterococci: genotype-phenotype correlations and inhibition by vancomycin. Sci Rep 7, 5733 (2017).
Semedo, T. et al. Comparative study using type strains and clinical and food isolates to examine hemolytic activity and occurrence of the cyl operon in enterococci. J Clin Microbiol 41, 2569–76 (2003).
Sieńko, A. et al. Comparison of biofilm-producing Enterococcus faecalis, Enterococcus faecium, and unusual Enterococcus strains. Eur J Biol Res 7, 291–298 (2017).
Pillay, S., Zishiri, O. T. & Adeleke, M. A. Prevalence of virulence genes in Enterococcus species isolated from companion animals and livestock. Onderstepoort J Vet Res. 85, e1–e8 (2018).
Szeleszczuk, P., Dolka, B., Żbikowski, A., Dolka, I. & Peryga, M. First case of enterococcal spondylitis in broiler chickens in Poland. Med Weter. 69, 298–303 (2013).
Stępień-Pyśniak, D. et al. Prevalence and antibiotic resistance of Enterococcus strains isolated from poultry. Acta Vet Hung 64, 148–163 (2016).
Dolka, B. et al. Occurrence of Enterococcus spp. in poultry in Poland based on 2014–2015 data. Med Weter. 73, 220–224 (2017).
Stępień-Pyśniak, D., Hauschild, T., Nowaczek, A., Marek, A. & Dec, M. Wild birds as a potential source of known and novel multilocus sequence types of antibiotic-resistant Enterococcus faecalis. J Wildl Dis 54, 219–228 (2018).
Stępień-Pyśniak, D., Hauschild, T., Dec, M., Marek, A. & Urban-Chmiel, R. Clonal structure and antibiotic resistance of Enterococcus spp. from wild birds in Poland. Microb Drug Resist. https://doi.org/10.1089/mdr.2018.0461 (2019).
PubMLST. 2019a. Public databases for molecular typing and microbial genome diversity. Enterococcus faecalis MLST databases. https://pubmlst.org/efaecalis/. Accessed March 2019.
PubMLST. 2019b. Public databases for molecular typing and microbial genome diversity. Enterococcus faecium MLST databases. https://pubmlst.org/efaecium/. Accessed March 2019.
Hammerum, A. M. Enterococci of animal origin and their significance for public health. Clin Microbiol Infect 18, 619–25 (2012).
Heidari, H., Emaneini, M., Dabiri, H. & Jabalameli, F. Virulence factors, antimicrobial resistance pattern and molecular analysis of enterococcal strains isolated from burn patients. Microb Pathog. 90, 93–97 (2016).
López, M., Tenorio, C. & Torres, C. Study of vancomycin resistance in faecal enterococci from healthy humans and dogs in Spain a decade after the avoparcin ban in Europe. Zoonoses Public Health 60, 160–167 (2013).
Silva, N. et al. Virulence factors in enterococci from partridges (Alectoris rufa) representing a food safety problem. Foodborne Pathog Dis 8, 831–833 (2011).
Poeta, P. et al. Virulence factors and bacteriocins in faecal enterococci of wild boars. J Basic Microbiol 48, 385–392 (2008).
Poeta, P., Costa, D., Klibi, N., Rodrigues, J. & Torres, C. Phenotypic and genotypic study of gelatinase and β-haemolysis activities in faecal enterococci of poultry in Portugal. J Vet Med B Infect Dis Vet Public Health 53, 203–208 (2006).
Radhouani, H. et al. Clonal lineages, antibiotic resistance and virulence factors in vancomycin-resistant enterococci isolated from fecal samples of red foxes (Vulpes vulpes). J Wildl Dis 247, 769–773 (2011).
Stepanović, S. et al. Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS. 15, 891–899 (2007).
Knobloch, J. K., Horstkotte, M. A., Rohde, H. & Mack, D. Evaluation of different detection methods of biofilm formation in Staphylococcus aureus. Med Microbiol Immunol 191, 101–106 (2002).
Leuck, A. M., Johnson, J. R. & Dunny, G. M. A widely used in vitro biofilm assay has questionable clinical significance for enterococcal endocarditis. PLoS One. 9, e107282 (2014).
Krasowska, A. & Sigler, K. How microorganisms use hydrophobicity and what does this mean for human needs? Front Cell Infect Microbiol 4, 112 (2014).
Kreft, B., Marre, R., Schramm, U. & Wirth, R. Aggregation substance of Enterococcus faecalis mediates adhesion to cultured renal tubular cells. Infect Immun. 60, 25–30 (1992).
Martín, M. et al. Genes encoding bacteriocins and their expression and potential virulence factors of enterococci isolated from wood pigeons (Columba palumbus). J Food Protect 69, 520–531 (2006).
Eaton, T. J. & Gasson, M. J. Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl Environ Microbiol 67, 1628–1635 (2001).
Heaton, M. P., Discotto, L. F., Pucci, M. J. & Handwerger, S. Mobilization of vancomycin resistance by transposon-mediated fusion of a vanA plasmid with an Enterococcus faecium sex pheromone-response plasmid. Gene. 171, 9–17 (1996).
Jarzembowski, T., Naumiuk, Ł. & Pałubicka, A. Prevalance of genes involved in pili and biofilm formation and in vitro adherence properties of medical and fecal strains of Enterococcus faecalis isolated in Gdańsk. Microb Ecol Health Dis 21, 100–103 (2009).
Olsen, R. H., Schønheyder, H. C., Christensen, H. & Bisgaard, M. Enterococcus faecalis of human and poultry origin share virulence genes supporting the zoonotic potential of E. faecalis. Zoonoses Public Health 59, 256–263 (2012).
De Vuyst, L., Foulquié Moreno, M. R. & Revets, H. Screening for enterocins and detection of hemolysin and vancomycin resistance in enterococci of different origins. Int J Food Microbiol 84, 299–318 (2003).
Han, D. et al. The occurrence of virulence traits among high-level aminoglycosides resistant Enterococcus isolates obtained from feces of humans, animals, and birds in South Korea. Int J Food Microbiol 144, 387–392 (2011).
Coburn, P. S. & Gilmore, M. S. The Enterococcus faecalis cytolysin: a novel toxin active against eukaryotic and prokaryotic cells. Cell Microbiol. 5, 661–669 (2003).
Silva, V. et al. Genetic characterization of vanA-Enterococcus faecium isolates from Wild Red-Legged Partridges in Portugal. Microb Drug Resist 24, 89–94 (2018).
Ben Yahia, H. et al. Antimicrobial resistance and genetic lineages of faecal enterococci of wild birds: Emergence of vanA and vanB2 harboring Enterococcus faecalis. Int J Antimicrob Agents 52, 936–941 (2018).
McBride, S. M. et al. Genetic variation and evolution of the pathogenicity island of Enterococcus faecalis. J Bacteriol. 191, 3392–3402 (2009).
Willems, R. J. et al. Variant esp gene as a marker of a distinct genetic lineage of vancomycin-resistant Enterococcus faecium spreading in hospitals. Lancet. 57, 853–855 (2001).
Leavis, H. et al. A novel putative enterococcal pathogenicity island linked to the esp virulence gene of Enterococcus faecium and associated with epidemicity. J Bacteriol. 186, 672–682 (2004).
Hynes, W. L. & Walton, S. L. Hyaluronidases of Gram-positive bacteria. FEMS Microbiol Lett 183, 201–720 (2000).
Koch, S., Hufnagel, M., Theilacker, C. & Huebner, J. Enterococcal infections: host response, therapeutic, and prophylactic possibilities. Vaccine. 22, 822–830 (2004).
Dolka, B., Chrobak-Chmiel, D., Czopowicz, M. & Szeleszczuk, P. Characterization of pathogenic Enterococcus cecorum from different poultry groups: Broiler chickens, layers, turkeys, and waterfowl. PLoS One. 12, e0185199 (2017).
Jackson, C. R. et al. Antimicrobial resistance, virulence determinants and genetic profiles of clinical and nonclinical Enterococcus cecorum from poultry. Lett Appl Microbiol 60, 111–119 (2015).
Nowakiewicz, A. et al. Characterization of multidrug resistant E. faecalis strains from pigs of local origin by ADSRRS-fingerprinting and MALDI -TOF MS; evaluation of the compatibility of methods employed for multidrug resistance analysis. PLoS One. 12, e0171160 (2017).
Stępień-Pyśniak, D., Hauschild, T., Różański, P. & Marek, A. MALDI-TOF mass spectrometry as a useful tool for identification of Enterococcus spp. from wild birds and differentiation of closely related species. J Microbiol Biotechnol 27, 1128–1137 (2017).
Dec, M., Puchalski, A., Nowaczek, A. & Wernicki, A. Antimicrobial activity of Lactobacillus strains of chicken origin against bacterial pathogens. Int Microbiol. 19, 57–67 (2016).
Author information
Authors and Affiliations
Contributions
D.S.P. designed the study, analysed and interpreted the data, supervised all experiments, isolated enterococci, performed phenotypic analysis and virulence gene detection with PCR, and drafted the manuscript. T.H. was responsible for isolation of DNA from enterococci. U.K. analysed and interpreted the data of biofilms. M.D. analysed and interpreted the data of hydrophobicity. R.U.CH. revised the manuscript critically for important intellectual content. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stępień-Pyśniak, D., Hauschild, T., Kosikowska, U. et al. Biofilm formation capacity and presence of virulence factors among commensal Enterococcus spp. from wild birds. Sci Rep 9, 11204 (2019). https://doi.org/10.1038/s41598-019-47602-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47602-w
This article is cited by
-
Biofilm-specific determinants of enterococci pathogen
Archives of Microbiology (2024)
-
Characterization and anti-biofilm activity of bacteriophages against urinary tract Enterococcus faecalis isolates
Scientific Reports (2022)
-
Enterococci as Intestinal Microbiota: Investigation of Characteristics and Probiotic Potential in Isolates from Adults and Breast-Fed Infants
Probiotics and Antimicrobial Proteins (2022)
-
In vitro Biofilm Formation by Bioluminescent Bacteria Isolated from the Marine Fish Gut
Microbial Ecology (2021)
-
Improved detection of esp, hyl, asa1, gelE, cylA virulence genes among clinical isolates of Enterococci
BMC Research Notes (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.