Abstract
Diffusional motion within the crowded environment of the cell is known to be crucial to cellular function as it drives the interactions of proteins. However, the relationships between protein diffusion, shape and interaction, and the evolutionary selection mechanisms that arise as a consequence, have not been investigated. Here, we study the dynamics of triaxial ellipsoids of equivalent steric volume to proteins at different aspect ratios and volume fractions using a combination of Brownian molecular dynamics and geometric packing. In general, proteins are found to have a shape, approximately Golden in aspect ratio, that give rise to the highest critical volume fraction resisting gelation, corresponding to the fastest long-time self-diffusion in the cell. The ellipsoidal shape also directs random collisions between proteins away from sites that would promote aggregation and loss of function to more rapidly evolving nonsticky regions on the surface, and further provides a greater tolerance to mutation.
Similar content being viewed by others
Introduction
Interactions between diffusing proteins are central to the function of the cell. The rate at which reactions of proteins occur in the cell is proportional to the product of their concentration and their rate of diffusion1. As protein concentration increases their translational diffusion \({D}_{t}^{0}\) slows due to interactions in the increasingly crowded environment2. For a system of hard spheres, which undergo a glass transition at a volume fraction ϕc ≈ 0.583, the translational diffusion constant falls with volume fraction ϕ approximately as \({D}_{t}={D}_{t}^{0}{(1-\varphi /{\varphi }_{\text{c}})}^{2}\)4. The product of diffusion constant and concentration is maximum when ϕ = 19%1,2, a value similar to that of proteins in the cell5,6,7.
The diffusion constant of a protein depends on its size and shape and on the shape of other macromolecules in its environment8. The translational diffusion constant of a spherical particle is proportional to its hydrodynamic radius, and the diffusion constant is smaller for an aspherical particle compared to the sphere of the same volume. Spherical proteins with their larger diffusion constants would, therefore, be expected to give rise to higher biochemical reaction rates than aspherical proteins of equivalent volume. Proteins are, however, generally not spherical9.
The glass transition point ϕc also depends on particle shape. Mode coupled theory predicts that ϕc is higher for spheroids (ellipsoids of revolution) than it is for spheres, suggesting spheroids prevent crystallization by raising the glass transition point10, and that ϕc of dumbbell-shaped particles increased with asymmetry up to a maximum when their length/diameter is around 1.411,12,13. Simulations of the liquid-to-solid phase transition also suggest that ϕc is higher for spheroids that it is for spheres14,15. Raising the value of ϕc for protein would increase the diffusion constant and biochemical reaction rate. Studying the relationship between particle shape and Dt and ϕc and hence the possible relationships between protein shape, their concentration and their reactions within a cell is the subject of this work.
Here, we first study the aspect ratios of globular proteins by taking both calliper measurements and determining triaxial ellipsoids of equivalent steric volume and diffusive properties. Using a combination of molecular dynamics and ellipsoidal packing and unpacking simulations we show how the critical volume fraction and translational diffusion of these triaxial ellipsoids depends on their shape and concentration. Assuming the diffusion of proteins follows that predicted for these ellipsoidal models these simulations suggest that proteins have evolved an ellipsoidal shape that limits aggregation and gelation in situations of high concentration, and to maximise diffusion-limited processes within the cell. We further analyse the collisions between diffusing ellipsoids and suggest that proteins have evolved binding sites on their surface at regions that limit the likelihood of aggregation during collisions.
Materials and Methods
47 677 structures from the Protein Data Bank clustered at 95% sequence identify using CD-HIT were downloaded from RCSB.org on 01/23/2018. The proteins in this set were taken as the single-chain, and multimerisation ignored. The 3D Complex16 (single chain proteins and protein dimers), PDB Select (asymmetric units)17, Dynameomics18, and species19 subsets were also studied.
Calliper measurements
To determine the extent of asphericity of proteins Feret diameters were measured by determining the largest and smallest differences between the maximum and minimum atomic coordinate values in the x, y, and z planes for all orientations of a protein around the x and y planes between 0 and π/2 in π/180 increments. The maximum error in this estimation is 1 − cos(π/180) = 0.02%.
Ellipsoid of equivalent steric volume
To calculate the principle radii of the ellipsoid of the equivalent steric volume to a protein we calculated the moment of inertia of a body representing the atoms20
where x, y, and z are the positions of the N atoms in the protein, and r is their vdW-radii (C 1.7 Å; N 1.55 Å; O 1.52 Å; S 1.8 Å; H 1.2 Å; other 1.7 Å). This tensor is diagonalized to give the three eigenvectors of the principle axes and their eigenvalues λ1 ≥ λ2 ≥ λ3. The lengths a, b, and c of the semi-axes of the ellipsoid of equivalent steric volume are then found as
where \(M={\sum }_{i=1}^{N}\,\frac{4\pi }{3}{r}_{i}^{3}\). Here, we express the ratios of these axes lengths by the parameters α = a/c, the aspect ratio of the ellipsoid, and αβ = b/c, which describes the shape from prolate (β = 0) to oblate (β = 1) spheroid. Code for this was written in C using diagonalization routines of Kopp21 (http://arXiv:physics/0610206).
Brownian and Langevin dynamics
HOOMD-blue (v2.1.1-92)22,23 was used to simulate the diffusive motion of soft spheroids at volume fractions between 0.001% and 55%. An initial starting configuration was generated by packing 300 randomly oriented spheroids to a volume fraction of 30% using PackLSD (see below). The position and orientation of the packed spheroids were then used to generate 8100 Gay-Berne particles by replicating in a 3 × 3 × 3 array. The Gay-Berne anisotropic pair potential used in HOOMD-blue is24
with \({\ell }_{||}\) and \({\ell }_{\perp }\) set such that V(2a) = V(2c) = 1
Dynamics of unit-volume spheroids was performed for 10 000 steps with kBT = 1, translational and rotational friction factors γ = 1, timestep δt = 0.0001 whilst the size of the periodic box was changed to reach the specified volume fraction, the system equilibrated for a further 10 000 steps and then particle diffusion monitored over 250 000 steps (t = 25). Translational diffusion was determined from the mean-squared displacement d of the particles Dt = <d2>/6t. Eq. (14) was fitted to the simulated diffusion data to determine ϕc using OriginPro (OriginLab Corporation, Northampton, MA 01060).
In-house code was used to study the location of collisions between ellipsoids in Brownian motion described by Eq. (5) and Eq. (6). Here, a point x, y, z on the surface of an ellipsoid in its body frame of reference (i.e. axis a aligns with x, b with y and c with z) is defined by two angles θ and φ, where x = t cos(θ), y = t sin θ cos(φ), z = t sin(θ) sin (φ) with t given by
Packing and unpacking
Maximally random jammed packings of ellipsoids were generated using the neighbour list collision-driven molecular dynamics algorithm PackLSD of Donev25. Unpackings were performed by running PackLSD on an ordered array of ellipsoids, as described in ref.26, to decompress to a final volume fraction of 35%.
Estimation of the critical volume fraction ϕ c for ellipsoids
To estimate the critical volume fraction ϕc we studied the liquid-to-solid phase transition of triaxial ellipsoids. For a system of hard spheres the phase diagram of pressure as a function of volume fraction exhibits a freeze point ϕF ≈ 0.494, below which the suspension is a liquid27. Between ϕF and ϕM ≈ 0.545, the melt point, for a system in equilibrium solid and liquid coexist and above which the system is a crystal. Forcing a system above ϕF quickly enough to preclude equilibration sees the system enter a supercooled state of liquid until the glass transition ϕG ≈ 0.58 is reached, and between this and the maximally random jammed stated ϕMRJ ≈ 0.64 the system behaves as a glass14. There is some debate as which value is best for the glass transition: the value ϕG ≈ 0.58 or the maximally jammed state value ϕMRJ ≈ 0.64. Most experiments point to ϕG ≈ 0.58, for example eye-lens spherical α-crystallin multimers (ϕG = 0.579 ± 0.004)28,29, although some suggest the value of 0.64 is the true value30,31. Santamaria-Holek and Mendoza used ϕc = ϕMRJ when predicting the relative viscosity of ellipsoids32. Here, we assumed ϕc = ϕG, regardless of its value.
To estimate ϕc for triaxial ellipsoids we followed the method of Donev where a dense crystal arrangement of packed ellipsoids is unpacked and their order parameter and the pressure of the system monitored26. As the system was unpacked, the point at which order is lost precipitously was taken as the freeze point, ϕF, which also corresponds to an increase in pressure15,25. We took the melt point, ϕM, as the volume fraction of the unpacked ellipsoids that exerted the same pressure as this raised value of the freeze point. The value of ϕMRJ was determined by packing the ellipsoids from the random (liquid) state obtained at the end of the unpacking. We then studied how ellipsoid diffusion varied when taking the values found for ϕF, ϕM and ϕMRJ as ϕc.
Mapping
Maps of the location of contacts between ellipsoids and of residue type were generated by representing each θφ contact point as a Gaussian spot. To calculate the Gaussian, geodesic distances on the surface of the ellipsoid between two integer θφ points were estimated by considering the map as an image of 180 × 360 pixels, where each represents the integer degree value of the angles. All pixel values were initially set to zero. Starting at the pixel corresponding to the first point, the cartesian distance to the centres of its nearest unvisited neighbouring pixels was calculated at the closest found. The closest pixel was then given the value of the sum of its current value and this distance. Next, the pixel with the lowest value and with unvisited neighbours was selected. This process was iterated until the value of the pixel corresponding to the second point was set, which was the approximation of the geodesic distance from the first.
Positional evolutionary rates
The rates of residue evolution were calculated for the 382 orthologous sequences taken from the genomes of Saccharomyces cerevisiae and nine closely related species: Saccharomyces paradoxus, Saccharomyces mikatae, Saccharomyces bayanus, Candida glabrata, Saccharomyces castellii, Saccharomyces kluyveri, Kluyveromyces lactis, Kluyveromyces waltii, and Ashbya gossypii using Rate4Site33 as described in ref.34. As in that work, evolutionary rates are normalised to the average of all positions in all proteins in the set, and presented as \({\mathrm{log}}_{2}[{\rm{normalisedrate}}]\).
Equations of diffusion
Rotational and diffusional constants of triaxial ellipsoids
Many methods have been derived to predict the translational and rotational diffusion of proteins35,36,37,38,39. Here, using the semi-axis lengths of the ellipsoid of equivalent steric volume we calculated the translational \({D}_{{\rm{t}}a,b,c}^{0}\) and rotational \({D}_{{\rm{r}}a,b,c}^{0}\) diffusion coefficients along and around each semi-axis as (in the example of a)40
respectively, for a protein where a′, b′ and c′ are the semi-axis lengths of the equivalent ellipsoid increased by an amount δH to reflect the width of a stationary hydration layer surrounding the protein, and RF and RD are the Carlson symmetric elliptic integrals of the first RF(x, y, z), and second RD(x, y, z) kind
The algorithms to solve these were taken from Numerical Recipes41. The long-time translational and rotational diffusion constants are the arithmetic mean of the values for each axis
In terms of α′ = (a + δH)/(c + δH), α′β′ = (b + δH)/(c + δH) and \(r^{\prime} ={(a^{\prime} b^{\prime} c^{\prime} )}^{\frac{1}{3}}\)
Diffusion at finite concentration
Interactions between diffusing bodies leads to diffusive motion that is dependent on timescale and concentration42. There are several descriptions of the correlation between intrinsic viscosity or long-time diffusion of spheres and volume fraction, all sharing a critical volume fraction at the divergent point43,44,45. Tokuyama et al. described the short-time self-diffusion of spheres by4
where ϕ is the volume fraction of the particles in the suspension and L(ϕ) defined as
with B = (9ϕ/8)1/2 and C = 11ϕ/16. At longer times, a cage-effect of proteins surrounding others decreases diffusion further with proteins needing to transit from cage to cage. The long-time self-diffusion coefficient of a soft particle is described well by the expression
where ϕc is a singular point of the cessation of long-time self-diffusion; the critical volume fraction. Eqs (12) and (14) with a value κ = 2 have been shown to describe the volume fraction dependence of translational diffusion for a number of proteins, where the function has been fitted to the experimental data to determine the value of ϕc44,46,47,48.
Results and Discussion
Proteins are naturally aspherical and have an aspect ratio around 1.6
The aspect ratio of proteins, calculated as the ratio of the largest-to-smallest calliper diameters, is broadly distributed around 1.6 and ranges from 1.2 to 18.6 (Fig. 1). The distribution of aspect ratio of the single chains of 47 677 PDB structures clustered at 95% sequence similarity is represented well by a log-lognormal with a modal value of 1.639 (±0.001). We divided the aspect ratio distributions into those of proteins that are generally prolate and of those generally oblate by determining the moment of inertia of the protein represented by its atoms as unit density van-der Waals radius spheres and determining the semi-axis lengths a ≥ b ≥ c of the ellipsoid with the same inertia; the so-called ellipsoid of equivalent steric volume20. We found the aspect ratio of the calliper measurements as longest-to-shortest of prolate and shortest-to-longest of oblate are distributed around 1.6 and 0.7 (\( \sim 1/1.6\)), respectively (Fig. 1 All). The modal aspect ratios are similar to that of the Golden ratio \({\rm{\Phi }}=\mathrm{(1}+\sqrt{5})/2=1.618\ldots \), and its reciprocal Φ−1. The distributions of aspect ratios of the 13 052 monomers (3D Monomers), 19 148 dimer components (3D Dimers A & B) and their 9574 dimeric complexes (3D Dimers AB) of the structures in 3D Complex database16, 3272 single chain asymmetric units in the PDBselect database17, 701 proteins in the Dynameomics database18, and 1243 proteins from different species (667 H. sapiens; 396 E. coli; 180 S. cerevisiae)19 are similarly shaped (Fig. 1). The modes of these distributions are given in Table 1.
The aspect ratio of proteins taken from calliper measurements ac and bc distribute around a modal value of 1.6. The ratio of the longest to shortest calliper measurements for 47 677 structures (single monomer chains) taken from the PDB database form a distribution around a modal value of 1.639. By determining the ellipsoid of equivalent steric volume the proteins can be separated into prolate and oblate-shaped structures. The distributions of aspect ratio of proteins in subsets of the PDB database (see text) are shown for oblate as bc/ac and prolate as ac/bc proteins.
This separation of the distribution shows that approximately 25% of proteins are oblate, a value similar to that found by Dima and Thirumalai who studied the proteins in the PDBselect17 subset of the Protein Data Bank9. As an unfolded chain is generally prolate we may question why some proteins fold to oblate structures as such would need to undergo significant changes in volume and/or surface area on folding49,50,51. This misconception arises from the approximation to purely oblate and prolate spheroids of revolution that suggests the change between the two requires a transition through a spherical state with corresponding differences in surface area or volume. The distribution of protein shape is better represented in two dimensions of αβ, where α = a/c and αβ = b/c. Figure 2 shows this 2D distribution for the 47 677 protein chains. The dashed line β = (ln(α + 1) − ln(2))/ln(α) for α > 1 shows the boundary between prolate (β = 0; lower) and oblate (β = 1; upper) proteins. We also plot (dotted) contours of the isoperimetric quotient (36πV2/A3) that represent ellipsoids of equivalent surface area and volume52. The αβ aspect ratios of proteins are broadly distributed around a modal value of α = 1.65 and β = 0.34. These values of aspect ratio correspond to the aspericity parameter Δ ≈ 0.1 reported in ref.9.
The distribution of the aspect ratios for the ellipsoids of equivalent steric volume of the 47 677 structures taken from the PDB database (single chains clustered at 95% sequence similarity). For each ellipsoid with semi-axis lengths a ≥ b ≥ c, α = a/c and αβ = b/c. The Gaussian-smoothed distribution has a maximum at α = 1.65, β = 0.34. The dashed line left-to-right for α > 1 shows the boundary between generally prolate (β = 0) and generally oblate (β = 1) structures. 75% of the structures are generally prolate. The dotted isoperimetric contour lines between values of β represent ellipsoids of equal volume and surface area.
The surface area A of an ellipsoid defined by α, β and c with volume \(V=\frac{4}{3}\pi {\alpha }^{(1+\beta )}{c}^{3}\) is approximately
with p = 1.6075, which for all values of β increases monotonically with α from the smallest (sphere) value at α = 1. For an ellipsoid of fixed volume and surface area there is a unique value of α for each value of β between 0 and 1. Notwithstanding constraints due to necessary rearrangements of the polypeptide chain a molten globule could morph from prolate to oblate maintaining constant volume and surface area (i.e. along an isoperimetric contour). This suggests, therefore, that the shape of the folded protein may not necessarily reflect the general shape of its denatured state. Interestingly, A has a minimum in β between 0 and 1 for values of α > 1, so the transition between prolate and oblate spheroids at constant volume and surface area requires a seemingly paradoxical increase in aspect ratio α. A value of α > 1, where the surface area of the protein is greater than the sphere of equivalent volume, affords a greater tolerance to mutation than a spherical protein would, as any mutations which cause a change in volume can be accommodated without a change in surface area, and vice versa53.
The volume of the ellipsoidal approximation of protein was found to scale with the number of residues N as V ≈ 203N Å3 (Fig. 3C), equal to an equivalent radius of 3.1 Å-per-residue at a packing density of 0.64, and the surface area of this ellipsoid of equivalent steric volume approximates to A ≈ 47Ns Å2 (equivalent radius of 3.7 Å-per-residue at the maximum disc packing density of 0.9), where Ns is the number of residues at the surface. These findings are in agreement with those of others9,54. As expected, the extent to which amino acids are buried was found to be correlated with their hydrophobicity and, in general, anticorrelated with their ‘stickiness’ as defined in ref.19 (Fig. 3A,B).
(A,B) The average fractional radial distance of the Cα of each type of residue from the core to the surface of a protein (A orange) is independent of protein shape and is ordered similarly to the frequency at which they are found at the surface (A magenta; B closed symbols), and opposite to their stickiness (A violet; B open symbols)19. (C) The volume of the ellipsoid of equivalent steric volume can be accurately predicted from the number of residues. (D) The number of residues at the surface of the protein can be predicted from the sum of each residue content and their surface propensities (partial least squares with one component, closed symbols; Bayesian Regularized Neutral Network with 20 neurons, open symbols).
The average fractional radial extent of the alpha carbon of each residue toward the surface of the equivalent ellipsoid afforded an estimation of the likelihood that an amino acid of a type would be at the surface. By summing the product of these values by the number of each residue type in the protein we could estimate Ns, and hence surface area (Fig. 3D). This estimation could be improved by using the actual frequencies of each residue type being at the surface. It is possible, therefore, to estimate both volume and surface area of a protein’s equivalent ellipsoid, and therefore, from Eq. (15) its aspect ratio from knowledge of the amino acid content alone. Since the volume of a protein is related to the number of amino acids and its surface area related to the amino acid composition, Eq. (15) reveals that a protein aspect ratio α is dependent on the fractional content of surface-exposed residues. This indicates that for a chain of a given length there are far more different amino acid compositions that give rise to ellipsoids than give rise to spheres, and therefore suggest that proteins are naturally ellipsoidal irrespective of secondary structure. To support this we calculated the order parameter \(S= < \,1.5{\cos }^{2}\theta -0.5 > \) where θ is the angle between each secondary structural unit (helix or sheet) and the a axis of the protein (all, and separated into prolate and oblate structures) and found no correlation in alignment between secondary structure and the overall ellipsoidal shape of the protein (\({\bar{S}}_{{\rm{all}}}=\mathrm{0.159,}\,{\bar{S}}_{{\rm{prolate}}}=0.183,{\bar{S}}_{{\rm{oblate}}}=0.072\)) (Fig. 4).
The order parameter \(S= < \,1.5\,{\cos }^{2}\theta -0.5 > \) where θ is the angle between the vector defining the start (Cα) and end of each secondary structural unit (left sheet; middle helix; right both) in a protein and the a axis of its ellipsoid of equivalent steric volume was calculated (top oblate; bottom prolate). Average values are shown in 〈brackets〉.
By considering the volume of the protein is related to its chain length and its surface area related to its composition we suggest that the general ellipsoidal shape of a protein (and its molten globule) is inherent. The fact that proteins can fold successfully without the assistance of chaperones55, and can successfully refold following denaturation and hence not co-translationally proximal to the ribosomal vestibule56,57, suggests that their shape is not defined by the physical environment in which they fold. We estimated the extent of this ellipticity of proteins by considering the random sequence polypeptide chain as a binary sequence of either surface (polar) or buried (hydrophobic) residues, similar to a HP polymer model, which are also seen to collapse into ellipsoidal shapes58. The binomial theorem shows that the largest number of combinations of H and P is when they are in equal amounts. The dependence of aspect ratio α on chain length of this simple model when β = 0.5 is shown in Fig. 5A. For a chain of 400 residues, equal to the average length in the human proteome, the most frequent composition has 200 surface exposed residues giving rise to ellipsoid aspect ratios for varying β of αβ=0 = 1.57 to αβ=0.4 = 1.66 to αβ=1 = 1.52 (coincident with the contour line through the maximum of the distribution in Fig. 2). The ability of this model protein to accommodate mutations causing changes in volume and/or surface area is shown in Fig. 5B.
(A) Predicted aspect ratio αβ=0.5 for proteins where half of their residues are on the surface. (B) A 400 residue ellipsoidal protein can accommodate deletions(−)/insertions(+) of residues at the surface (triangles), deletions(−)/insertions(+) in the core (circles), and rearrangements from surface-to-core(−)/core-to-surface+ (diamonds) by changing aspect ratio.
Taking Ns/N to be 0.55, the value most found in the proteins and equal to the eleven out of twenty residues that have a surface preference value exceeding 0.5 (Fig. 3A), we find the length of protein where the predicted surface area is at least equal that of the sphere of equivalent volume is 36π(203)2/0.553473 = 270 (the value of N when in Eq. 1 A = 47(0.55N), V = 203N, and α = 1) residues. Proteins maintaining a 0.55/0.45 surface/core ratio of residues are predicted, therefore, to be at least approximately 270 residues in length. Larger proteins can be formed maintaining this surface/core ratio by becoming elliptical, but smaller proteins only formed through an increased prevalence of surface-preferred residues. Eukarya have median protein lengths longer at 361 residues, bacteria at 267 residues, and archaea are shorter at 247 residues59.
The ellipsoids of equivalent steric volume allow the accurate prediction of protein diffusion
Calculated values of the translational and rotational diffusion constants of the equivalent steric triaxial ellipsoid of proteins, using Eqs (5) and (6), respectively, matched the experimentally determined values after increasing the semi-axis lengths of the ellipsoid by 2.32 Å, for PDB structures without hydrogens, or 2.30 Å for those with (translation rRMSE = 4.7%, rotation rRMSE = 9.1%) (Fig. 6A)36,39. Thus, the ellipsoid of equivalent steric volume with a hydration layer of 2.3 Å, equivalent to a hydration level of 0.38 g/g (volume of equivalent ellipsoid with boundary layer ≈ 2.39 Mp, density of bound water 1.104 g/mL), facilitates the prediction of the translational and rotational diffusive properties of globular proteins, indicating the effective hydrodynamic radii of a protein for each of its semi-axes is equivalent to this ellipsoid of equivalent steric volume. The hydration value of 0.38 g/g is commensurate with a monolayer coverage of water. Taking the example of lysozyme with equivalent ellipsoid semi-axis lengths of a = 23.4 Å, b = 15.3 Å and c = 13.9 Å (α = 1.69, β = 0.19) we calculate a hydration shell volume of at most 9874 Å3 containing up to 365 water molecules. Microparticle dehydration studies estimate that lysozyme is hydrated by a stationary layer of ≈380 water molecules60.
Protein diffusion is dependent on its ellipsoidal shape. (A) The translational diffusion constant \({D}_{{\rm{t}}}^{0}\) (solid) and rotational diffusion constant \({D}_{{\rm{r}}}^{0}\) (open) of 42 proteins is accurately predicted from their ellipsoids of equivalent steric volume dilated by 2.3Å. (B) Langevin and Brownian dynamics simulations of spheroids of varying aspect ratio reveal the reduction of translation diffusion with increasing volume fraction to the critical value ϕc. (C) The value of ϕc taken from fits to the Langevin (open circles) and Brownian (closed circles) dynamics reveal the dependence on aspect ratio which rises from 0.56 for spheres to 0.64 for prolate spheroids of aspect ratio 1.6, which then falls with increasing asphericity. The calculated freeze ϕF (triangles), melt ϕM (diamonds), and maximum jammed ϕMRJ (hexagons) volume fractions for prolate spheroids (β = 0) are also shown. (D) The product of the volume fraction and the predicted diffusion constant for varying aspect ratios using Eq. (14) with ϕc values taken from a polynomial fit to the Langevin and Brownian dynamics values shown in (C) shows a maximum for spheroids with aspect ratio of 1.58 at a volume fraction of 0.18.
By noting that the volume of the equivalent hydrated ellipsoid \(\frac{4\pi }{3}{r^{\prime} }^{3}\approx 2.39\,{M}_{{\rm{p}}}\), and that α = 1.65 and β = 0.34, we can approximate the translational and rotational diffusion constants from protein length N or weight Mp as (20 °C, ρ = 1.0016 mPa s)
The numerator in this approximation of the translational diffusion coefficient lies between the value of 244 given by Young-Carroad-Bell35 and 285 given by Polson61. It was noted by Hem and Neimeyer38 that the equation derived by Tyn and Gusek36 for the approximation of diffusion based on a protein’s radius of gyration implies a spheroidal geometry with an aspect ratio of 1.4 if prolate and 0.66 (1/1.5) if oblate.
Diffusion within the crowded cell is greater for ellipsoidal proteins
The Brownian and Langevin dynamics simulations of the diffusion of soft spheroidal particles, represented by the Gay-Berne potential, at various volume fractions confirmed the aspect ratio dependence of the diffusion of spheroids. Figure 6B is a plot of the normalised diffusion rate of Gay-Berne prolate spheroids of 1.0, 1.3, 1.6 and 2.0 aspect ratio at volume fractions of up to 55% determined from Brownian (solid symbols) and Langevin (open symbols) dynamics simulations, with fits of Eq. (14) with κ, a scaling parameter for L(ϕ), and ϕc fitted parameters. The extrapolated values of the critical volume fraction ϕc from both simulation methods show an identical dependence on shape (Fig. 6C), starting at a value of 0.565 for α = 1.0 (spheres) increasing to a maximum of 0.64 at α = 1.64 and then declining.
The crystal unpacking simulations gave values for ϕF, ϕM and ϕMRJ for spheres as 0.49, 0.56 and 0.68, respectively (Fig. 7). The value of ϕG ≈ 0.58 lies between ϕM and ϕMRJ. ϕM follows a similar trend as ϕc obtained from the Brownian and Langevin dynamics simulations (Fig. 6C), with both starting at 0.56 for α = 1.0 and exhibiting a maximum near αβ=0 = 1.6. The ϕMRJ for ellipsoids 1 ≤ α ≤ 3 and 0 ≤ β ≤ 1 has a single maximum value ϕMRJ ≈ 0.73 at α = 1.72, β = 0.5. The value of ϕF is maximal at 0.64 at α = 1.71, β = 0.4. The value of ϕM has three maxima in this αβ profile, with the highest of ϕM = 0.68 at α = 1.72, β = 0.36. As an extrapolation, however, we expect a degree of variability in the determination of ϕM. Ellipsoids with shape α ≈ 1.7, β ≈ 0.4 are expected, therefore, to have the highest value of ϕc, which suggests that the translational diffusion proteins of a similar shape is least retarded by crowding. Proteins of aspect ratio of α ≈ 1.7 and β ≈ 0.4 have both optimal diffusion and limited propensity to form a glass when at high concentration. Such heterogenous crowding in the cell can turn diffusion from normal to anomalous, where mean-squared displacement is no-longer linear in time62. A more complete model of the dependence of diffusion-limited reactions on cellular concentration that incorporates the subdiffusion due to crowding is required63,64, although we expect the dependence on aspect ratio of the diffusing particles to be similar to that found here.
Estimation of the critical freeze ϕF, melt ϕM, and maximum jammed ϕMRJ volume fractions. The ordered crystal state for each ellipsoid defined by αβ (top) is unpacked during which both the pressure of the system (blue) and order (black) are recorded (left). At the freeze point there is a slight increase in pressure as order is lost precipitously. The melt point is taken as the more packed state that exerted the same pressure as the raised value at the freeze point. After unpacking, the system is then packed to determine the maximum random jammed state (left, red).
Proteins are ellipsoidal to maximise their translational diffusion in the crowded cell
A biomolecular reaction limited by diffusion has a rate proportional to the product of the relative diffusion constant (i.e. the sum of the diffusion constants of the reactants) and the reactant concentrations. By combining Eqs (11), (12) and (14) we found the product \(\varphi {D}_{S}^{L}(\varphi ,{\varphi }_{{\rm{c}}})\) is maximum at αβ=0 = 1.58 and ϕ = 0.19 for prolate spheroids, and is maximum when α = 1.70, β = 0.5, ϕ = 0.18 using ϕc = ϕM, and maximum when α = 1.60, β = 0.5, ϕ = 0.20 for ϕc = ϕMRJ (Fig. 8). Thus, using ϕc of an ellipsoid suspension as either ϕM or ϕMRJ and assuming the diffusive behaviour of proteins at finite concentration remains equal to the ellipsoids of equivalent steric volume, we recover the cell volume fraction of 19% and additionally the optimum protein aspect ratio of a/c ≈ 1.7 and b/c ≈ 1.3, similar to that measued of proteins that form dimers (Table 1).
Using the critical volume fraction ϕc as the freeze point ϕF (left, blue), melt point ϕM (middle, red), and jammed point ϕMRJ (right, green) the maximum of the product (bottom row) of the predicted diffusion rate (middle row) and volume fraction (top row) for each ellipsoid αβ is found. The maximum of the product is shown as the yellow star and can be compared to the modal value found for the 47677 protein set (blue star).
The predicted translational diffusion of the protein at a volume fraction of 19% is 35% of its value at infinite dilution, giving a simple expression for the approximate translational diffusion [×10−7 cm2 s−1] of a protein in the cell at 37 °C, ρ = 0.6913 mPa s of
Ellipsoidal shape helps proteins avoid non-functional interactions
Figure 9B shows the normalized frequency distribution in θφ space of the location of collisions between ellispoids (α = 1.78, β = 0.44) undergoing Brownian dynamics. Each collision at θφ on the surface of the ellipsoid is represented by a Gaussian spot with variance 10° arc. The locations of the collisions are distributed unevenly across the surface with the fewest collisions made at the poles of the c axis.
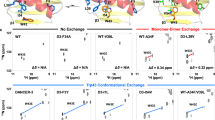
Proteins have surfaces that are nonsticky at the sites where they are most likely to make collisions during diffusion. (A) The effective translational diffusion over short time of points on the surface of a tumbling ellipsoid, defined by two angles θ from the a axis and φ from b, is lowest (light) at the poles of the c axis, the flattest face, and highest (dark) at the poles of the a axis. (B) Plot of the normalized distribution of collision frequency (low light to high dark) for each θφ point on the surface of a diffusing ellipsoid. (C) Plot of the frequency distribution of the location of points at the centre of the interface between monomers in a protein dimer (low light to high dark). (D) Plot of the average radial extent values (Fig. 3A) for the residues found on the surface of the monomers in protein dimers. (E) The most conserved residues on the surface of homologs are coincident with the centre of the pole at c where binding and active sites are common. (F) The a/c cross-section through the evolutionary rate data reveals that mutations of the faster-evolving residues on the surface away from the active site facilitate changes in the core.
The distribution is similar in form to that of the effective translational diffusion constants over short times (i.e. dominated by the rotation) Dtp(x, y, z) of points across the surface of the ellipsoid (Fig. 9A), calculated as
Points that are translating the quickest, i.e. the poles of the a axis, make more contacts than those which are translating the slowest, i.e. the poles of the c axis. Molecular dynamics simulations support this importance of rotational diffusion in directing collisions between proteins65.
We compared the collision probabilities to the points of the centres of protein binding patches measured from the 3D Complex database16. We calculated the ellipsoid of equivalent steric volume of each partner in a binary complex and determined their centre of contact by shrinking the ellipsoids until their surfaces just touched. Figure 9C shows the distribution of the θφ contact points of 9958 protein pairs, where each point is mapped as a Gaussian-spot with variance 10° arc across the surface. Figure 9D shows the map of the average extent to which the amino acid at the contact point is normally found at the surface (Fig. 3A), which reflects the patch stickiness as calculated by Levy et al.19. Protein-protein interaction sites were found predominately on the face of the poles of the c axis, at residues that are generally normally found more buried in a protein. Whilst the face at c presents the greatest surface area it is the area involved in the fewest collisions during diffusion. An ellipsoidal shape to protein directs random collisions between them to nonsticky regions of their surface and thereby limits the formation of non-functional interactions and aggregation.
A protein’s interaction or active site is expected to be the most conserved during evolution, since mutation of a residue at the site is more likely to lead to loss of function than a mutation elsewhere on the surface19,34. Therefore, we expected the residues at the poles of the c axis to be most conserved in homologs across species of known phylogeny. Following the method of Tóth-Petróczy and Tawfik34 we calculated the evolution rates-per-position of 382 protein domains of known structure in orthologs in 10 yeast species. In Fig. 9E we plot the average rates of the surface residues mapped onto θφ (the protein is rotated around a by 0° or 180° so the face at c with the slower rates is at 0 < φ < 180). The residues around the centre of the c face at θ = 90°, φ = 90°, coincident with the location of the active site most often, were found to have an evolution rate around half that of the other surface residues. A cross section taken through the data taken at φ = 90° highlights the association of evolutionary rates between the residues at the surface and those at the core. The evolutionary rates of residues decrease with their distance from the surface with core residues exchanging on average fourfold slower than those on the surface (away from the centre of c at θ = 90°, φ = 90°). The core region of the protein that is most conserved is offset from the centre of the protein towards the interface. This too is as expected following the surface–core association of evolutionary rates revealed by Tóth-Petróczy and Tawfik34. Mutations of the faster evolving residues on the surface away from the active site facilitate changes in the core. The rate of evolution decreases with distance from the surface, converging on a point that is offset the c axis (coincident with Φ). If core mutations were facilitating surface changes we would expect the most conserved residues to be found at the centre of the protein, which is not the case.
Summary
We have shown that proteins are generally aspherical with an aspect ratio distributed around 1.6; a value close to the Golden ratio. We have shown than proteins of random sequence greater than 270 residues in length are likely to be ellipsoidal irrespective of secondary structure and that a simple model of a protein of 400 residues has a shape similar to that most common in the PDB.
We have shown that the translational and rotational diffusion of proteins can be accurately modelled by considering the protein as a triaxial ellipsoid of equivalent steric volume. We have shown that the translational diffusion of such ellipsoids decreases with increasing concentration and that the critical volume fraction corresponding to the glass transition, where translational diffusion becomes negligible, is dependent on the shape of the ellipsoid. We found that the shape of ellipsoid that gives the highest predicted value for the critical volume fraction is coincident with the modal value found for proteins. We suggested, therefore, that proteins have a shape that maximizes their translational diffusion within the cell and limits the likelihood of gelation at high concentration.
We studied the location of contacts between diffusing ellipsoids and found a correlation between the location where collisions are least likely to occur and where the protein/protein interface is found in protein dimers. We found the same association between the type of residues found at these interaction sites as others19, and showed in a series of proteins that the residues that are found at these sites have evolved less quickly than those on other regions of the protein where random collisional contacts are more likely to occur. We suggested that proteins have evolved a shape which directs non-functional random collisions away from their sites of interaction to non-sticky residues that are least conserved.
References
Schmit, J. D., Kamber, E. & Kondev, J. Lattice model of diffusion-limited bimolecular chemical reactions in confined environments. Phys. Rev. Lett. 102, 218302 (2009).
Dill, K. A., Ghosh, K. & Schmit, J. D. Physical limits of cells and proteomes. Proc. Natl. Acad. Sci. USA 108, 17876–17882 (2011).
Weeks, E. R., Crocker, J. C., Levitt, A. C., Schofield, A. & Weitz, D. A. Three-dimensional direct imaging of structural relaxation near the colloidal glass transition. Science 287, 627–631 (2000).
Tokuyama, M., Moriki, T. & Kimura, Y. Self-diffusion of biomolecules in solution. Phys. Rev. E 83, 051402 (2011).
Fulton, A. B. How crowded in the cytoplasm? Cell 30, 345–347 (1982).
Zimmerman, S. B. & Trach, S. O. Estimation of macromolecular concentration and excluded-volume effects for the cytoplasm of Escherichia coli. J. Mol. Biol. 222, 599–620 (1991).
Ellis, R. J. Macromolecular crowding: Obvious but underappreciated. Trends Biochem. Sci. 26, 597–604 (2001).
Balbo, J., Mereghetti, P., Herten, D.-P. & Wade, R. C. The shape of protein crowders is a major determinant of protein diffusion. Biophys. J. 104, 1576–1584 (2013).
Dima, R. I. & Thirumalai, D. Asymmetry in the shapes of folded and denatured states of proteins. J. Phys. Chem. B 108, 6564–6570 (2004).
Pfleiderer, P., Milinkovic, K. & Schilling, T. Glassy dynamics in monodisperse hard ellipsoids. EPL 84, 16003 (2008).
Chong, S. H. & Gotze, W. Idealized glass transitions for a system of dumbbell molecules. Phys. Rev. E 65, 041503 (2002).
Chong, S. H. & Gotze, W. Structural relaxation in a system of dumbbell molecules. Phys. Rev. E 65, 051201 (2002).
Yatsenko, G. & Schweizer, K. S. Ideal glass transitions, shear modulus, activated dynamics, and yielding in fluids of nonspherical objects. J. Chem. Phys. 126, 014505 (2007).
Torquato, S. Random heterogeneous materials. In Interdisciplinary Applied Mathematics, vol. 16 (Springer-Verlag, New York, 2002).
Donev, A., Torquato, S. & Stillinger, F. H. Neighbor list collision-driven molecular dynamics simulation for nonspherical hard particles. II. Applications to ellipses and ellipsoids. J. Comput. Phys. 202, 765–793 (2005).
Levy, E. D., Pereira-Leal, J. B., Chothia, C. & Teichmann, S. A. 3D Complex: A structural classification of protein complexes. PLoS Comp. Biol. 2, e155 (2006).
Griep, S. & Hobohm, U. PDBselect 1992–2009 and PDBfilter-select. Nucleic Acids Res. 38, D318–D319 (2010).
van der Kamp, M.W. et al. Dynameomics: a comprehensive database of protein dynamics Structure 18, 423–435 (2010).
Levy, E. D., De, S. & Teichmann, S. A. Cellular crowding imposes global constraints on the chemistry and evolution of proteomes. Proc. Natl. Acad. Sci. USA 109, 20461–20466 (2012).
Fogolari, F., Esposito, G., Viglino, P. & Cattarinussi, S. Modeling of polypeptide chains as C-alpha chains, C-alpha chains with C-beta, and C-alpha chains with ellipsoidal lateral chains. Biophys. J. 70, 1183–1197 (1996).
Kopp, J. Efficient numerical diagonalization of hermitian 3x3 matrices. Int. J. Mod. Phys. C 9, 523–548 (2008).
Anderson, J. A., Lorenz, C. D. & Travesset, A. General purpose molecular dynamics simulations fully implemented on graphics processing units. J. Comput. Phys. 227, 5342–5359 (2008).
Glaser, J. et al. Strong scaling of general-purpose molecular dynamics simulations on gpus. Comput. Phys. Commun. 192, 97–107 (2015).
Allen, M. P. & Germano, G. Expressions for forces and torques in molecular simulations using rigid bodies. Mol. Phys. 104, 3225–3235 (2006).
Donev, A., Torquato, S. & Stillinger, F. H. Neighbor list collision-driven molecular dynamics simulation for nonspherical hard particles. I. Algorithmic details. J. Comput. Phys. 202, 737–764 (2005).
Donev, A., Stillinger, F. H., Chaikin, P. M. & Torquato, S. Unusually dense crystal packings of ellipsoids. Phys. Rev. Lett. 92, 255506 (2004).
Gokhale, S., Sood, A. K. & Ganapathy, R. Deconstructing the glass transition through critical experiments on colloids. Adv. Phys. 65, 363–452 (2016).
Foffi, G. et al. Hard sphere-like glass transition in eye lens alpha-crystallin solutions. Proc. Natl. Acad. Sci. USA 111, 16748–16753 (2014).
Vodnala, P. et al. Hard-sphere-like dynamics in highly concentrated alpha-crystallin suspensions. Phys. Rev. E 97, 020601 (2018).
Russel, W. B., Wagner, N. J. & Mewis, J. Divergence in the low shear viscosity for Brownian hard-sphere dispersions: At random close packing or the glass transition? J. Rheol. 57, 1555–1567 (2013).
Hunter, G. L. & Weeks, E. R. The physics of the colloidal glass transition. Rep. Prog. Phys. 75, 066501 (2012).
Santamaria-Holek, I. & Mendoza, C. I. The rheology of concentrated suspensions of arbitrarily-shaped particles. J. Colloid. Interface Sci. 346, 118–126 (2010).
Mayrose, I., Graur, D., Ben-Tal, N. & Pupko, T. Comparison of site-specific rate-inference methods: Bayesian methods are superior. Mol. Biol. Evol. 21, 1781–1791 (2004).
Tóth-Petróczy, A. & Tawfik, D. S. Slow protein evolutionary rates are dictated by surface-core association. Proc. Natl. Acad. Sci. USA 108, 11151–11156 (2011).
Young, M. E., Carroad, P. A. & Bell, R. L. Estimation of diffusion-coefficients of proteins. Biotechnol. Bioeng. 22, 947–955 (1980).
Tyn, M. T. & Gusek, T. W. Prediction of diffusion-coefficients of proteins. Biotechnol. Bioeng. 35, 327–338 (1990).
Zhou, H. X. Calculation of translational friction and intrinsic viscosity. 2. Application to globular proteins. Biophys. J. 69, 2298–2303 (1995).
Hem, L. H. & Niemeyer, B. A novel correlation for protein diffusion coefficients based on molecular weight and radius of gyration. Biotechnol. Prog. 19, 544–548 (2003).
Halle, B. & Davidovic, M. Biomolecular hydration: From water dynamics to hydrodynamics. Proc. Natl. Acad. Sci. USA 100, 12135–12140 (2003).
Happel, J. & Brenner, R. Low Reynolds Number Hydrodynamics: With Special Applications to Particulate Media. (Kluwer Academic Publishers Group, The Hague, 1983).
Press, W. H., Teukolsky, S. A., Vetterling, W. T. & Flannery, B. P. Numerical Recipes 3rd Edition: The Art of Scientific Computing., 3 edn. (Cambridge University Press, New York, NY, USA, 2007).
Roosen-Runge, F. et al. Protein self-diffusion in crowded solutions. Proc. Natl. Acad. Sci. USA 108, 11815–11820 (2011).
Medina-Noyola, M. Long-time self-diffusion in concentrated colloid dispersions. Phys. Rev. Lett. 60, 2705–2708 (1988).
Giannopoulou, A., Aletras, A. J., Pharmakakis, N., Papatheodorou, G. N. & Yannopoulos, S. N. Dynamics of proteins: Light scattering study of dilute and dense colloidal suspensions of eye lens homogenates. J. Chem. Phys. 127, 205101 (2007).
Dörrr, A., Sadiki, A. & Mehdizadeh, A. A discrete model for the apparent viscosity of polydisperse suspensions including maximum packing fraction. J. Rheol. 57, 743–765 (2013).
Nesmelova, I. V. & Fedotov, V. D. Self-diffusion and self-association of lysozyme molecules in solution. Biochim. Biophys. Acta-Protein Struct. Molec. Enzym. 1383, 311–316 (1998).
Porcar, L. et al. Formation of the dynamic clusters in concentrated lysozyme protein solutions. J. Phys. Chem. Lett. 1, 126–129 (2010).
Doster, W. & Longeville, S. Microscopic diffusion and hydrodynamic interactions of hemoglobin in red blood cells. Biophys. J. 93, 1360–1368 (2007).
Aronovitz, J. A. & Nelson, D. R. Universal features of polymer shapes. J. Phys. (Paris) 47, 1445–1456 (1986).
Rudnick, J. & Gaspari, G. The shapes of random walks. Science 237, 384–389 (1987).
Blavatska, V., von Ferber, C. & Holovatch, Y. Universal features of polymer shapes in crowded environments. Phys. Lett. A 374, 2861–2864 (2010).
Damasceno, P. F., Engel, M. & Glotzer, S. C. Predictive self-assembly of polyhedra into complex structures. Science 337, 453–457 (2012).
Sasidharan, R. & Chothia, C. The selection of acceptable protein mutations. Proc. Natl. Acad. Sci. USA 104, 10080–10085 (2007).
Shen, M. Y., Davis, F. P. & Sali, A. The optimal size of a globular protein domain: A simple sphere-packing model. Chem. Phys. Lett. 405, 224–228 (2005).
Saibil, H. Chaperone machines for protein folding, unfolding and disaggregation. Nat. Rev. Mol. Cell. Biol. 14, 630–642 (2013).
Batey, S. & Clarke, J. Apparent cooperativity in the folding of multidomain proteins depends on the relative rates of folding of the constituent domains. Proc. Natl. Acad. Sci. USA 103, 18113–18118 (2006).
Borgia, A. et al. Transient misfolding dominates multidomain protein folding. Nat. Commun. 6, 8861 (2015).
Pham, T. T., Duenweg, B. & Prakash, J. R. Collapse Dynamics of copolymers in a poor solvent: Influence of hydrodynamic interactions and chain sequence. Macromolecules 43, 10084–10095 (2010).
Brocchieri, L. & Karlin, S. Protein length in eukaryotic and prokaryotic proteomes. Nucleic Acids. Res. 33, 3390–3400 (2005).
Rickard, D. L., Duncan, P. B. & Needham, D. Hydration potential of lysozyme: Protein dehydration using a single microparticle technique. Biophys. J. 98, 1075–1084 (2010).
Polson, A. & Vanderreyden, D. Relationship between diffusion constants and molecular weight. Biochim. Biophys. Acta. 5, 358–366 (1950).
Ghosh, S. K., Cherstvy, A. G., Grebenkov, D. S. & Metzler, R. Anomalous, non-gaussian tracer diffusion in crowded two-dimensional environments. New J. Phys. 18, 013027 (2016).
Weiss, M., Elsner, M., Kartberg, F. & Nilsson, T. Anomalous subdiffusion is a measure for cytoplasmic crowding in living cells. Biophys. J. 87, 3518–3524 (2004).
Matzler, R. & Klafter, J. The restaurant at the end of the random walk: Recent developments in the description of anomalous transport by fractional dynamics. J. Phys. A: Math. Gen. 37, R161–R208 (2004).
Nawrocki, G., Karaboga, A., Sugita, Y. & Feig, M. Effect of protein-protein interactions and solvent viscosity on the rotational diffusion of proteins in crowded environments. Phys. Chem. Chem. Phys. 21, 876–883 (2019).
Acknowledgements
This work was supported by the Biotechnology and Biological Sciences Research Council and the University of Nottingham via a PhD sponsorship for Gareth Shannon; and the Engineering and Physical Sciences Research Council and the University of Nottingham via PhD sponsorships for Callum Marples and Rudesh Toofanny. High-performance computing infrastructure and support were provided by the University of Nottingham.
Author information
Authors and Affiliations
Contributions
R.D.T. posed the question whether the Golden ratio is found in proteins; G.S. and P.M.W. designed the research; C.R.M., G.S. and P.M.W. performed the research; C.R.M., G.S. and P.M.W. analysed the data; and G.S. and P.M.W. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shannon, G., Marples, C.R., Toofanny, R.D. et al. Evolutionary drivers of protein shape. Sci Rep 9, 11873 (2019). https://doi.org/10.1038/s41598-019-47337-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47337-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.