Abstract
The tricarboxylic acid cycle produces NADH for oxidative phosphorylation and fumarase [EC 4.2.1.2] is a critical enzyme in this cycle, catalysing the reversible conversion of fumarate and l-malate. Fumarase is applied to industrial l-malate production as a biocatalyst. l-malate is used in a wide range of industries such as food and beverage, pharmacy chemistry. Although the biochemical properties of fumarases have been studied in many organisms, they have not been investigated in cyanobacteria. In this study, the optimum pH and temperature of Synechocystis 6803 fumarase C (SyFumC) were 7.5 and 30 °C, respectively. The Km of SyFumC for l-malate was higher than for fumarate. Furthermore, SyFumC activity was strongly inhibited by citrate and succinate, consistent with fumarases in other organisms. Substitution of alanine by glutamate at position 314 of SyFumC changed the kcat for fumarate and l-malate. In addition, the inhibitory effects of citrate and succinate on SyFumC activity were alleviated. Phylogenetic analysis revealed cyanobacterial fumarase clades divided in non-nitrogen-fixing cyanobacteria and nitrogen-fixing cyanobacteria. SyFumC was thus biochemically characterised, including identification of an amino acid residue important for substrate affinity and enzymatic activity.
Similar content being viewed by others
Introduction
Fumarase, or fumarate hydratase [EC 4.2.1.2], is an enzyme in the tricarboxylic acid (TCA) cycle and is conserved in all organisms. Fumarase catalyses the reversible conversion of fumarate and l-malate. Two classes of fumarases are found in prokaryotes, namely Class I fumarases that are iron-dependent proteins, and Class II fumarases which resemble eukaryotic enzymes belonging to the aspartase/fumarase superfamily1. Escherichia coli possesses three genes encoding fumarases, fumA, fumB, and fumC, which are differently regulated by oxygen and growth conditions2,3. In E. coli, FumA and FumB are Class I enzymes and FumC is a Class II enzyme and no homology exists between the two classes of fumarases4. The FumC protein of E. coli resembles fumarases found in other bacteria such as Bacillus subtilis as well as in mammals5; it is an iron-independent enzyme and relatively heat-stable compared to FumA, and its biochemical properties are similar to those of mammalian fumarases6. The crystal structures of E. coli and Saccharomyces cerevisiae FumC have been resolved and their activity and regulatory domains determined7,8. Fumarase is thus a well-studied enzyme; however, recent studies have demonstrated that human fumarases are localized not only in mitochondria but also in the cytosol and function in response to DNA damage9,10. A Class II fumarase in B. subtilis also functions in response to DNA damage by producing l-malate, which regulates translation of RecN11. For the application, malate is used mainly in the food and beverage industry as an acidulant, flavor enhancer, food additive and a precursor for pharmaceutical chemicals12. Improvement of the thermostability of Corynebacterium glutamicum fumarase enhances l-malate production13 and an enzyme membrane reactor with immobilized fumarase is used to produce l-malate14. For the metabolic engineering, the production of l-malate is increased by expressing yeast fumarase in Aspergillus oryzae, suggesting that fumarase is a rate-limiting enzyme of malate production15. In this way, the importance of fumarases is conserved across the kingdoms and recognized in both basic and applied sciences.
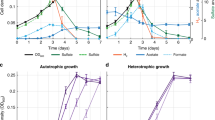
Cyanobacteria are photosynthetic prokaryotes and are known as a model for photosynthetic organisms. Cyanobacteria fix carbon dioxide and this activity influences the global carbon cycle, with cyanobacteria contributing nearly 30% of the global net primary production16. Moreover, engineered cyanobacteria can produce industrially relevant chemicals from fixed carbon dioxide and are suitable for biofuel and bulk chemical production17,18. Among cyanobacteria, Synechocystis sp. PCC 6803 (hereafter referred to as Synechocystis 6803) is a unicellular, non-nitrogen-fixing cyanobacterium, the genome of which was first sequenced in 199619. Substrains of this species are used for studies of photosynthesis and for metabolic engineering20,21,22. Transcriptomic analyses using microarrays can elucidate the mechanisms of environmental stresses and identify regulatory factors that transduce environmental signals23,24,25 and transcriptional regulators of Synechocystis 6803 carbon metabolism have been identified using these techniques26,27,28. Subsequently, metabolomic analyses of Synechocystis 6803 was performed and the carbon distribution and flux in response to environmental changes and genetic manipulation were demonstrated29,30,31,32. Fluxome analyses showed that TCA cycle and of acetyl-CoA metabolite pool sizes are smaller in Synechocystis 6803 than in E. coli30,32. Furthermore, fluxes in the TCA cycle remain low under all conditions tested30,33, indicating that the TCA cycle of unicellular cyanobacteria is unique among bacteria.
The cyanobacterial TCA cycle was thought to be incomplete because it lacked 2-oxoglutarate dehydrogenase, which generates succinyl-CoA from 2-oxoglutarate. Biochemical analysis based on Synechococcus sp. PCC 7002, however, has demonstrated that marine cyanobacteria possess the enzymes 2-oxoglutarate decarboxylase and succinic semialdehyde dehydrogenase, which generate succinate from 2-oxoglutarate by two enzymatic reactions34. In Synechocystis 6803, additional enzymatic reactions involving the γ-aminobutyric acid (GABA) shunt generate succinate from 2-oxoglutarate35. In this shunt, 2-oxoglutarate is converted to glutamate, followed by conversion to GABA35 which is then converted to succinyl-semialdehyde, and finally to succinate35. In addition to these bypasses, succinate is also produced in the reductive TCA cycle in Synechocystis 6803 under dark, anaerobic conditions36. Phosphoenolpyruvate carboxylase (PEPC), which generates oxaloacetate from phosphoenolpyruvate, is a rate-limiting enzyme for succinate production under these conditions36. Generally, PEPCs in bacteria and plants are inhibited by aspartate and malate; however, PEPC in Synechocystis 6803 is uniquely insensitive to these metabolites37. Substitution of the glutamate residue with lysine at position 954 restores these inhibitory effects, similar to that seen for PEPC in a nitrogen-fixing cyanobacterium37. These recent studies demonstrate that TCA cycle enzymes in cyanobacteria possess unique properties, making biochemical characterisation of these enzymes not only intriguing, but also indispensable for understanding their metabolism.
In the present study, biochemical analysis of fumarase C from Synechocystis 6803 (SyFumC) revealed that the reversible reactions are regulated by TCA cycle metabolites and further showed the importance of the alanine residue at position 314 for substrate affinity.
Results
Affinity purification and biochemical characterisation of SyFumC
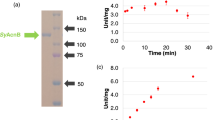
Genome sequence analysis indicated that Synechocystis 6803 does not possess a Class I fumarase but does express a Class II fumarase, SyFumC. To evaluate the biochemical properties of SyFumC, glutathione-S-transferase-tagged SyFumC (GST-SyFumC) was expressed in E. coli cells and purified from the soluble fraction by affinity chromatography (Figs 1a and S1). The enzymatic activity of SyFumC using fumarate as a substrate (hereafter referred to as “the enzymatic activity towards fumarate”) was highest at pH 7.5 (Fig. 1b); with l-malate as a substrate (hereafter referred to as “the enzymatic activity towards l-malate”), the enzymatic activity of SyFumC was also highest at pH 7.5, but was less affected by variation in pH. Enzymatic activities towards both fumarate and l-malate were highest at 30 °C and were inhibited at 60 °C (Fig. 1c). SyFumC activity towards l-malate was less sensitive to temperature variation than that toward fumarate (Fig. 1c). Therefore, for the ensuing experiments, SyFumC enzymatic assays were performed at pH 7.5 and 30 °C.
Determination of optimum conditions for Synechocystis sp. PCC 6803 fumarase C (SyFumC). (a) GST-tagged SyFumC after purification. Purified proteins were electrophoresed on 8% SDS-PAGE gel and stained using InstantBlue reagent. (b) Effect of pH on SyFumC activity. Data represent the means ± SD obtained from three independent experiments. For the enzyme assay, 15 pmol of SyFumC was used. One unit of SyFumC activity was defined as the consumption of 1 µmol fumarate or l-malate per min. Light blue circles and orange triangles represent activity specific for fumarate and l-malate, respectively. (c) Effect of temperature on SyFumC activity. Data represent the means ± SD obtained from three independent experiments. For the enzyme assay, 15 pmol of SyFumC was used. One unit of SyFumC activity was defined as the consumption of 1 µmol fumarate or l-malate per min. Light blue circles and orange triangles represent activity specific for fumarate and l-malate, respectively.
Substrate saturation curves were obtained using different concentrations of fumarate (Fig. 2a) and l-malate (Fig. 2b) as substrates. The Km of SyFumC for fumarate and l-malate was 0.244 ± 0.026 and 0.478 ± 0.112 mM, respectively; the kcat of SyFumC for fumarate and l-malate was 101.2 ± 6.7 and 42.3 ± 4.3 s−1, respectively; and the kcat/Km of SyFumC for fumarate and l-malate was 415.1 ± 17.0 and 90.3 ± 11.3 s−1 mM−1, respectively (Table 1).
In vitro enzyme assays for SyFumC using fumarate and l-malate as substrates. (a) Enzyme activity was measured by varying the fumarate concentration. The data represent the means ± SD obtained from three independent experiments. Light blue squares represent activity specific for fumarate. Yellow and green squares represent activity specific for fumarate with 4 mM citrate and 4 mM succinate, respectively. (b) Enzyme activity was measured by varying the l-malate concentration. The data represent the means ± SD obtained from three independent experiments. Orange triangles represent activity specific for l-malate. Navy and purple triangles represent activity specific for l-malate with 4 mM citrate and 4 mM succinate, respectively.
Identification of effectors altering SyFumC activity
Effectors of SyFumC activities were also evaluated. Citrate and succinate are competitive inhibitors of fumarases from pig heart, E. coli, and Pisum sativum38,39,40; therefore, the effect of citrate and succinate on SyFumC was examined at various concentrations (2–8 mM) under conditions of substrate saturation (Fig. 3a,b). SyFumC activity towards both fumarate and l-malate reactions decreased in a concentration-dependent manner in the presence of both citrate and succinate (Fig. 3a,b). The enzymatic activity of SyFumC towards fumarate and l-malate decreased to 45–58% of control in the presence of 4 mM succinate or 4 mM citrate (Fig. 3a,b). The Km and kcat for fumarate and l-malate in the presence of 4 mM succinate were 0.256 ± 0.017 mM and 79.5 ± 2.2 s−1; and 0.409 ± 0.034 mM and 65.4 ± 2.1 s−1, respectively (Table 1). The Km and kcat for fumarate and l-malate in the presence of 4 mM citrate were 0.616 ± 0.073 mM and 34.9 ± 1.8 s−1; and 0.921 ± 0.042 mM and 28.0 ± 0.4 s−1, respectively (Table 1). The kcat/Km for fumarate and l-malate in the presence of 4 mM succinate was 311.5 ± 12.4 and 57.0 ± 4.0, respectively; and in the presence of 4 mM citrate, it was 160.2 ± 8.8 and 30.4 ± 1.8 s−1 mM−1, respectively (Table 1).
Effect of citrate and succinate on SyFumC activity. Enzymatic activity was measured by varying the concentrations of citrate or succinate under conditions of substrate (fumarate and l-malate) saturation. (a) Yellow and green bars represent SyFumC enzymatic activity towards fumarate with citrate and succinate, respectively. (b) Navy and purple bars represent SyFumC enzymatic activity towards l-malate with citrate and succinate, respectively. Asterisk (*) indicates P < 0.05. The data represent the means ± SD from three independent experiments.
Additional effectors of SyFumC enzymatic activity were examined under conditions of substrate saturation. We selected 11 effectors which regulate the activities of enzymes in the TCA cycle and pyruvate metabolism in this cyanobacterium37,41. Among the 11 effectors tested, Co2+ and Zn2+ decreased SyFumC activity towards fumarate and l-malate (Fig. 4a,b). At 1 mM, Co2+ decreased SyFumC enzymatic activity towards fumarate and l-malate to 66.5% and 57.4% of control, respectively (Fig. 4a,b). The presence of 1 mM and 10 mM Zn2+ decreased SyFumC enzymatic activity towards fumarate to 0.99% and 1.99% of control, respectively (Fig. 4a,b); similarly, 1 mM and 10 mM Zn2+ decreased SyFumC enzymatic activity towards l-malate to 0.94% and 1.89% of control (Fig. 4a,b). At 10 mM, pyruvate and Mn2+ both decreased SyFumC enzymatic activity towards l-malate (Fig. 4a,b). SyFumC activity could not be measured in the presence of 10 mM Co2+, 2-oxoglutarate, and phosphoenolpyruvate due to interference of absorption by solvents (Fig. 4a,b).
Effect of various metal ions and metabolites on SyFumC activity using fumarate and l-malate as substrates. Enzymatic activity was measured with various metabolites or metal ions under conditions of substrate (fumarate and l-malate) saturation. (a) The grey bars represent the control data. Red and blue bars represent concentrations of 1 mM and 10 mM metabolites or metal ions, respectively, using fumarate as a substrate. (b) The grey bar represents the control data. Green and dark blue bars represent, respectively, using l-malate as a substrate. NO, no enzymatic activity was detected; ND, enzymatic activity not determined in this study. The data represent the means ± SD from three independent experiments. Asterisk (*) indicates P < 0.05. Abbrevations; 2-OG, 2-oxoglutarate; PEP, phosphoenolpyruvate; Asp, aspartate; Pyr, pyruvate.
Amino acid substitution and substrate specificity of SyFumC
Amino acid sequences were compared by multiple sequence alignment analysis, using amino acid sequences of fumarases from other cyanobacteria, E. coli, Mycobacterium tuberculosis, and Arabidopsis thaliana (Fig. 5a). The amino acid residue at position 314 of SyFumC was found to be alanine; however, except for Synechococcus elongatus and Leptolyngbya sp., the equivalent residues of other fumarases are either glutamate or aspartate (Fig. 5a).
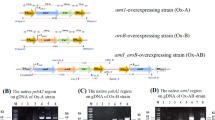
Multiple amino acid sequence alignment of Class II fumarases and determination of optimum conditions for Synechocystis sp. PCC 6803 fumarase C with glutamate substituted for alanine (SyFumC_A314E). (a) The multiple alignment of 13 amino acid sequences from Class II fumarases performed by CLC Sequence Viewer. The region containing the GSSxxPxKxN sequence, called an SS loop (indicated by a blue arrow), is shown. The green arrow indicates the alanine at position 314 in Synechocystis sp. PCC 6803. (b) GST-tagged SyFumC_A314E after purification. Purified proteins were electrophoresed on 8% SDS-PAGE gel and stained using InstantBlue reagent. (c) Effect of pH on SyFumC_A314E activity. Data represent the means ± SD obtained from three independent experiments. For the enzyme assay, 15 pmol of SyFumC_A314E was used. One unit of SyFumC activity was defined as the consumption of 1 µmol fumarate or l-malate per min. Light blue circles and orange triangles represent activity specific for fumarate and l-malate, respectively. (d) Effect of temperature on SyFumC_A314E. For the enzyme assay, 15 pmol of SyFumC_A314E was used. One unit of SyFumC activity was defined as the consumption of 1 µmol fumarate or l-malate per min. Light blue circles and orange triangles represent activity specific for fumarate and l-malate, respectively.
The impact of the alanine residue at position 314 of SyFumC was examined by substituting glutamate for alanine at position 314, and the recombinant protein was named SyFumC_A314E. The SyFumC_A314E protein was expressed in E. coli and purified by affinity chromatography (Figs 5b and S2). The optimal pH for SyFumC_A314E enzymatic activity towards fumarate was pH 7.3 and the optimal temperature was 30 °C (Fig. 5c). SyFumC_A314E was inactivated at 60 °C (Fig. 5d). Glutamate substitution had less impact at optimal pH and temperature using l-malate as a substrate (Fig. 5c,d). The maximum enzymatic activity of SyFumC_A314E for fumarate at the optimal pH and temperature decreased to 28.1% and 48.6% of SyFumC activity, respectively (Fig. 5c,d); the maximum enzymatic activity of SyFumC_A314E for l-malate at the optimal pH and temperature decreased to 41.5% and 41.6% of SyFumC activity, respectively (Fig. 5c,d).
The enzymatic activity of SyFumC_A314E was measured with various substrate concentrations to obtain saturation curves and calculate the kinetic parameters (Fig. 6a,b). The Km values for SyFumC_A314E towards fumarate and l-malate were 0.179 ± 0.036 and 0.442 ± 0.020 mM, respectively (Table 1), showing reductions to 73.4% and 92.5% of SyFumC values for fumarate and l-malate, respectively (Table 1). The kcat values for SyFumC_A314E towards fumarate and l-malate were 37.8 ± 3.0 and 21.6 ± 0.1 s−1, respectively (Table 1).
SyFumC_A314E enzyme assays using fumarate and l-malate as substrates. (a) Enzyme activity was measured by varying the fumarate concentration. The data represent the means ± SD obtained from three independent experiments. Light blue squares represent activity specific for fumarate. Yellow squares and green squares represent activity specific for fumarate with 4 mM citrate and with 4 mM succinate, respectively. (b) Enzyme activity was measured by varying the l-malate concentration. The data represent the means ± SD obtained from three independent experiments. Orange triangles represent activity specific for l-malate. Navy triangles and purple triangles represent activity specific for l-malate with 4 mM citrate and 4 mM succinate, respectively.
The Km and kcat for SyFumC_A314E were examined in the presence of 4 mM succinate or 4 mM citrate. The Km for SyFumC_A314E using fumarate and l-malate as substrates in the presence of 4 mM succinate was 0.214 ± 0.025 and 0.329 ± 0.030 mM, respectively (Table 1). The Km for SyFumC_A314E in the presence of 4 mM citrate, using fumarate and l-malate as substrates, was 0.391 ± 0.036 and 0.360 ± 0.020 mM, respectively (Table 1). The kcat values for SyFumC_A314E with fumarate and l-malate as substrates, in the presence of 4 mM succinate, were 44.0 ± 2.7 and 12.7 ± 0.7 s−1, respectively (Table 1). The kcat values for SyFumC_A314E with fumarate and l-malate as substrates, in the presence of 4 mM citrate, were 28.0 ± 0.4 and 10.4 ± 0.04 s−1, respectively (Table 1). The kcat/Km ratios for SyFumC_A314E with fumarate and l-malate as substrates were 206.3 ± 10.7 and 38.7 ± 1.6, respectively, in the presence of 4 mM succinate; and 121.7 ± 6.0 and 29.0 ± 1.7 s−1 mM−1, respectively, in the presence of 4 mM citrate (Table 1).
Finally, 11 effectors (metabolites and metal ions) were tested for their impact on SyFumC_A314E activity. At 1 mM, Zn2+ eliminated SyFumC_A314E enzymatic activity towards fumarate and l-malate (Fig. 7a,b). Also, at 1 mM, Co2+ decreased SyFumC_A314E enzymatic activity towards fumarate and l-malate to 54.6% and 47.6% of control, respectively (Fig. 7a,b). SyFumC_A314E enzymatic activity towards fumarate and l-malate was abolished with 10 mM Mn2+ (Fig. 7a,b). SyFumC_A314E enzymatic activity towards fumarate was marginally upregulated in the presence of 1 mM phosphoenolpyruvate (Fig. 7a); and at 10 mM, pyruvate promoted SyFumC_A314E enzymatic activity towards fumarate while enzymatic activity towards l-malate was suppressed (Fig. 7a,b).
Effect of various metal ions and metabolites on SyFumC_A314E activity towards fumarate and l-malate. Enzymatic activity was measured with various metabolites or metal ions under conditions of substrate (fumarate and l-malate) saturation. (a) The grey bar represents the control data. Red and blue bars represent concentrations of 1 mM and 10 mM metabolites or metal ions, respectively, using fumarate as a substrate. (b) The grey bar represents the control data. Green and dark blue bars represent concentrations of 1 mM and 10 mM metabolites or metal ions, respectively, using l-malate as a substrate. NO, no enzymatic activity was detected; ND, enzymatic activity not determined in this study. The data represent the means ± SD from three independent experiments. Asterisk (*) indicates P < 0.05.
Discussion
In this study, we performed the first biochemical analysis of class II fumarase in cyanobacteria, revealing that the modification of a residue at position 314 relieved the inhibition by citrate and succinate and reduced kcat of SyFumC. Previous studies have shown that the optimal pHs for fumarases from the thermophilic archaebacterium Sulfolobus solfataricus and from marine microorganisms are 8.0 and 8.5, respectively42,43. SyFumC is more active at a lower pH than these fumarases (Fig. 1b). The optimal pHs for fumarases from the eukaryotes Saccharomyces cerevisiae and Rhizopus oryzae are pH 7.5 and 7.244,45, respectively, and are thus similar to that for SyFumC (Fig. 1b). The growth pH ranges for S. solfataricus and S. cerevisiae are pH 1.0–5.8 and pH 4.0–4.5, respectively46,47, values inconsistent with the optimum pH for fumarases. Synechocystis 6803 shows optimal biomass production at pH 7.548, a pH for growth consistent with optimal enzymatic activity.
The optimal temperature for SyFumC activity was 30 °C, which is similar to that of fumarases from R. oryzae (30 °C) and the mesophilic Streptomyces coelicolor (30 °C), but lower than for bacterial and archaeal fumarases in Streptomyces lividans (45 °C), thermophilic Streptomyces thermovulgaris (50 °C) and Thermus thermophilus (85 °C), S. solfataricus (85 °C), and marine microorganisms (55 °C)42,43,45,49,50,51. S. thermovulgaris, T. thermophilus, and S. solfataricus are thermophilic organisms that thrive in hot springs, and higher optimal temperatures are not unexpected. The optimal temperature for Synechocystis 6803 growth is 30–32 °C52, which is close to the optimal temperature for SyFumC activity. The optimal temperatures for two other Synechocystis enzymes, SyPEPC and d-lactate dehydrogenase, are 30 °C and 30–40 °C, respectively37,53. The optimal temperature for SyFumC enzymatic activity is thus similar to that of other Synechocystis enzymes involved in primary carbon metabolism.
The Km for SyFumC enzymatic activity towards l-malate was higher than that for fumarate (Table 1). Fumarases from other organisms like E. coli54, Corynebacterium glutamicum55, S. solfataricus42, and marine microorganisms43 are similar; the Km ranges for fumarate and l-malate are 0.12–0.48 and 0.30–3.15 mM, respectively. Intracellular metabolite analysis using 13C-labeled glucose Synechocystis 6803 demonstrated that the number of 13C-atoms in malate was higher than in fumarate by approximately 5-fold at 37 °C in Synechocystis 680356. This 13C flux analysis indicates that the conversion reaction from fumarate to l-malate is stronger than that from l-malate to fumarate under aerobic conditions. Previous reports showed that the absolute concentration of l-malate (µmol/g dry cell weight) is higher than that of fumarate in Synechocystis 680357, indicating that SyFumC preferentially catalyses the reaction generating l-malate from fumarate. A different study shows that intracellular fumarate and malate pool sizes peak at 24 h under dark, anaerobic conditions, and then decrease during cultivation under the same conditions36. Carbon flow in the TCA cycle changes from using fumarate to using l-malate during dark, anaerobic incubation36. Nevertheless, intracellular l-malate concentrations (µmol/g dry cell weight) are still higher than fumarate concentrations36. The substrate specificity of SyFumC for fumarate well explains these results (Table 1). The Km of a malate dehydrogenase from Synechocystis 6803 (SyMDH) that catalyses the reversible conversion of l-malate and oxaloacetate is approximately 210-fold higher for the oxidative than for the reductive reaction and the catalytic efficiency of SyMDH is higher for the reductive than for the oxidation reaction58. Thus, both SyMDH and SyFumC preferentially catalyse reactions that produce l-malate. A previous study suggested that organic acids in the TCA cycle, including l-malate, play critical roles in the storage of carbon sources under nitrogen starvation31. SyFumC and SyMDH may preferentially convert metabolites to l-malate to promote this process.
The kcat for SyFumC enzymatic activity towards l-malate was lower than those for fumarase C from E. coli54 and Class II fumarases from marine microorganisms and T. thermophilus43,51, and Homo sapiens59 (Table 2). Fluxome analysis indicates that TCA cycle fluxes in Synechocystis 6803 remain constant under dark, photoheterotrophic, photomixotrophic, and photoautotrophic conditions, suggesting that the cyanobacterial TCA cycle does not naturally produce energy sources33. Metabolic reaction catalysed by a fumarase is inclined to the flux from fumarate to malate in all condition33. In addition, molar-based, widely targeted metabolic profiling analysis reveals that concentrations of TCA cycle metabolites are lower than those of glycolysis metabolites57. We compared fumarases of organisms in different kingdoms (Table 2). The kcat of SyFumC is lower than kcat of other microorganisms fumarases. Km of SyFumC using fumarate as a substrate is lower than Km of other microorganisms fumarases using fumarate as a substrate except Km for fumarate of E. coli (Table 2). Since the specific activity of SyFumC is lower than those of other organisms, the turnover number limits SyFumC activity in vitro (Table 2). These results were consistent with a less active TCA cycle in cyanobacteria33. On the other hand, kcat/Km of SyFumC using l-malate was higher than its of C. glutamicum fumarase (Table 2). The result indicates the reductive TCA cycle is relatively efficient compared to other organisms, consistent with a previous study58.
Citrate is known as a competitive inhibitor of fumarases from pig heart38 and E. coli39, and citrate also inhibited SyFumC activity (Fig. 3, Table 1). Mitochondrial fumarase of a higher plant, P. sativum, is inhibited by 53% in the presence of 20 mM citrate, using l-malate as a substrate60. The activities of A. thaliana mitochondrial and cytosolic fumarases decrease to 15–37% in the presence of citrate61. Succinate is also known as an inhibitor of fumarases of pig and P. sativum60,62, and it also inhibited SyFumC (Fig. 3). Succinate at 100 mM inhibits P. sativum fumarase; the activity decreases to 55% using l-malate as a substrate60. The sensitivity of SyFumC to citrate and succinate was nearly identical with either fumarate and l-malate as substrates (Fig. 3a,b). Absolute quantification of metabolites (µmol/g dry cell weight) in Synechocystis 6803 shows that intercellular concentrations of citrate, succinate, fumarate, and malate were 2.16, 0.323, 0.163, and 0.182, respectively57. These concentrations suggest that citrate and succinate are inhibitory under typical physiological conditions and the reduced SyFumC activity may be one cause of the low fluxes through the TCA cycle observed in Synechocystis 6803.
Additionally, SyFumC activity was also inhibited by several metabolites and divalent cations (Fig. 4a,b). P. sativum mitochondrial fumarase activity towards l-malate was decreased to 14% at 10 mM pyruvate60. A. thaliana mitochondrial fumarase is activated to 178% by pyruvate using l-malate as a substrate61. At 10 mM, pyruvate marginally inhibited SyFumC activity towards l-malate (Fig. 4b), and the effect of pyruvate on fumarases is different among photosynthetic organisms. Zn2+ strongly inhibited SyFumC enzymatic activity using fumarate and l-malate as substrates (Fig. 4a,b). This inhibition is greater than reported for a fumarase from a marine microorganism (FumF), FumF enzymatic activity reduces to 45% by 5 mM Zn2+ 43. In contrast, a fumarase from R. oryzae is not inhibited by Zn2+ 45. The addition of 10 mM Ca2+ was less inhibitory by other divalent cations for SyFumC and SyFumC_A314E activity towards l-malate (Figs 4b and 7b), but the R. oryzae fumarase is slightly stimulated by 9.0 mM Ca2+ using l-malate as a substrate45. Compared with other organisms fumarases, SyFumC was differently inhibited by metabolites and metal ions. These results indicated that the regulation by metabolites and metals was not conserved among fumarases. Three TCA cycle enzymes (SyMDH, isocitrate dehydrogenase, and citrate synthase) in Synechocystis 6803 are regulated by divalent cations, particularly Mg2+ 41,58,63. SyFumC was severely inhibited by Zn2+, not by Mg2+, indicating the different regulatory manner of enzymatic activities in the TCA cycle in this cyanobacterium.
The alanine at position 314 is close to the sequence called the SS loop, a motif associated with substrate binding and catalytic activity1. The optimal pH and temperature for SyFumC_A314E were similar to the optima for SyFumC (Fig. 5c,d), but substrate affinity was enhanced by the A314E substitution (Fig. 6). The Km values for SyFumC_A314E activity towards fumarate and l-malate were not significantly different to those for SyFumC activity towards the same substrates (Table 1). Compared to SyFumC, the kcat for SyFumC_A314E activity towards fumarate and l-malate decreased to 0.37-fold and 0.51-fold, respectively (Table 1). The kcat for an E. coli recombinant FumC with substituting glutamine for glutamate at position 315 near the SS loop was decreased by 10-fold when using fumarate and l-malate as substrates, but no effect was observed for Km values54. This glutamate residue at position 315 in E. coli is correspond to a glutamate at position 316 in SyFumC (Fig. 5). These results indicate that amino acids near the SS-loop determine the maximum rate of reaction of bacterial fumarases. Other group demonstrate that the alanine at position 347 in S. coelicolor fumarase (correspond to the glutamine at position 354 in SyFumC) is important for thermostability49, and the improvement of the thermostability in SyFumC can be future theme for cyanobacterial fumarases.
Phylogenetic analysis revealed that SyFumC belongs to non-nitrogen-fixing cyanobacterial clade (except Trichodesmium erythraeum) being different from nitrogen-fixing cyanobacteria and Gram-negative bacteria (Fig. 8). Combined with previous data, the present results demonstrate that important amino acid residues are conserved between enteric bacteria and non-nitrogen-fixing cyanobacteria. However, succinate inhibited the SyFumC activity (Fig. 2a), while fumarase in C. glutamicum is not inhibited by succinate55, indicating the sensitivity of fumarases to the effetors are different among the clades. These differences were derived from several amino acid substitutions; the inhibitory effects of citrate and succinate on SyFumC activity were reduced by the alanine to glutamate amino acid substitution (Figs 2a and 6a). The effects of some metabolites were also changed after the amino acid substitution. Pyruvate at 10 mM and phosphoenolpyruvate at 1 mM activated enzymatic activity with fumarate as a substrate (Fig. 7a), and 10 mM pyruvate and 1 mM 2-oxoglutarate inhibited SyFumC_A314E activity. Additionally, 10 mM pyruvate elicited different effects depending on whether fumarate or l-malate were used as substrate (Fig. 7a,b). Pyruvate and PEP are compounds upstream of fumarase in the TCA cycle, and it is considered to be activated to acquire reducing power for fumarate.
Phylogenetic analysis of bacterial class II fumarases. Amino acid sequences of 31 bacterial fumarase obtained from GenBank were aligned using CLC Sequence Viewer ver. 8.0. A maximum-likehood tree based on 431 preserved amino acid residues was designed using PhyML online (http://www.atgc-montpellier.fr/phyml/). Bootstrap values were calculated by 500 replications.
The present results show that SyFumC has a higher affinity for fumarate than for l-malate. The enzymatic activity of SyFumC towards fumarate and l-malate were inhibited by citrate and succinate. Current study, a biochemistry of a fumarase in a new clade, contributes to understanding the diversity of the TCA cycle in bacteria, and potentially leads to the metabolic engineering using cyanobacteria.
Methods
Vector construction and expression of recombinant proteins
The genomic region of Synechocystis 6803 including the fumC (slr0018) open reading frame (ORF) with the BamHI-XhoI fragment was amplified by PCR using the KOD -plus- neo DNA polymerase (Toyobo, Osaka, Japan) and the following primer set: forward, 5′-GAAGGTCGTGGGATCATGGTAAATTCCCACCGC-3′ and reverse, 5′-GATGCGGCCGCTCGAGCTAGTCAGCAATCGGGG-3′; the BamHI and XhoI restriction enzymes were obtained from TakaraBio (Shiga, Japan). The resultant fragment was cloned into the BamHI-XhoI sites of pGEX5X -1 (GE Healthcare Japan, Tokyo, Japan). Amino acid substitution was performed commercially by TakaraBio (Shiga, Japan). For SyFumC_A314E, the region +940–942 from the start codon in the fumC ORF was changed from GCC to GAA.
These vectors were transformed into E. coli DH5α cells (TakaraBio) and 5 L of transformed E. coli were cultivated in LB medium at 30 °C with shaking (150 rpm); protein expression was induced overnight in the presence of 0.01 mM isopropyl β-D-1-thiogalactopyranoside (Wako Chemicals, Osaka, Japan).
Affinity purification of recombinant proteins
Affinity chromatography for protein purification was performed as previously described33. Harvested DH5α cells suspended in 40 mL PBST (1.37 M NaCl, 27 mM KCl, 81 mM Na2HPO4·12H2O, 14.7 mM KH2PO4, and 0.05% Tween 20) were disrupted by sonication (VC-750, EYELA, Tokyo, Japan) 10 times for 20 s at 20% intensity. The disrupted cells were removed by centrifugation at 5800 × g for 2 min at 4 °C. The supernatant was transferred to a 50-mL tube and placed on ice, and 560 μL of Glutathione Sepharose 4B resin (GE Healthcare Japan) was mixed into the supernatant, followed by gentle shaking for 30 min. After centrifugation (5800 × g for 2 min at 4 °C), the supernatant was removed and the resin was re-suspended in 700 μL of PBST. After washing five times, recombinant proteins were eluted five times with 700 μL of GST elution buffer [50 mM Tris-HCl (pH 8.0), 10 mM reduced glutathione]. Proteins were concentrated with a Vivaspin 500 MWCO 50000 device (Sartorius, Göttingen, Germany), and protein concentrations were analysed with a Pierce BCA Protein Assay Kit (Thermo Scientific, Rockford, IL). SDS-PAGE was performed to analyse protein purification with staining using InstantBlue (Expedion Protein Solutions, San Diego, CA).
Enzyme assay for SyFumC
SyFumC activity was measured using 15 pmol SyFumC mixed in 1 mL of an assay solution (100 mM Tris-HCl [pH 7.5] and 10 mM fumarate or 10 mM l-malate). Absorbance at 250 nm was monitored using a Shimadzu UV-1850 (Shimadzu, Kyoto, Japan). One unit of SyFumC activity was defined as the consumption or the generation of 1 µmol fumarate per min. Km and Vmax were calculated by curve fitting using KaleidaGraph v4.5 software, and kcat was calculated from the Vmax. Results were plotted as graphs of rate of reaction against substrate concentration.
Statistical analysis
The P-values were calculated using paired two-tailed Student’s t-tests with Microsoft Excel for Windows (Redmond, WA, USA). All results were obtained from three or four independent experiments.
References
Puthan, V. V., Fibriansah, G., Raj, H., Thunnissen, A. M. & Poelarends, G. J. Aspartase/fumarase superfamily: a common catalytic strategy involving general base-catalyzed formation of a highly stabilized aci-carboxylate intermediate. Biochemistry 51, 4237–4243 (2012).
Park, S. J. & Gunsalus, R. P. Oxygen, iron, carbon, and superoxide control of the fumarase fumA and fumC genes of Escherichia coli: role of the arcA, fnr, and soxR gene products. J. Bacteriol. 177, 6255–6262 (1995).
Tseng, C. P., Yu, C. C., Lin, H. H., Chang, C. Y. & Kuo, J. T. Oxygen- and growth rate-dependent regulation of Escherichia coli fumarase (FumA, FumB, and FumC) activity. J. Bacteriol. 183, 461–467 (2001).
Woods, S. A., Schwartzbach, S. D. & Guest, J. R. Two biochemically distinct classes of fumarase in Escherichia coli. Biochim. Biophys. Acta. 954, 14–26 (1988).
Woods, S. A., Miles, J. S., Roberts, R. E. & Guest, J. R. Structural and functional relationships between fumarase and aspartase. Nucleotide sequences of the fumarase (fumC) and aspartase (aspA) genes of Escherichia coli K12. Biochem. J. 237, 547–557 (1986).
Ueda, Y., Yumoto, N., Tokushige, M., Fukui, K. & Ohya-Nishiguchi, H. Purification and characterization of two types of fumarase from Escherichia coli. J. Biochem. 109, 728–733 (1991).
Weaver, T. M., Levitt, D. G., Donnelly, M. I., Stevens, P. P. & Banaszak, L. J. The multisubunit active site of fumarase C from Escherichia coli. Nat. Struct. Biol. 2, 654–662 (1995).
Weaver, T. et al. Crystal structures of native and recombinant yeast fumarase. J. Mol. Biol. 280, 431–442 (1998).
Yogev, O. et al. Fumarase: a mitochondrial metabolic enzyme and a cytosolic/nuclear component of the DNA damage response. PLoS Biol. 8, e1000328 (2010).
Yogev, O., Naamati, A. & Pines, O. Fumarase: a paradigm of dual targeting and dual localized functions. FEBS J. 278, 4230–4242 (2011).
Singer, E., Silas, Y. B., Ben-Yehuda, S. & Pines, O. Bacterial fumarase and L-malic acid are evolutionary ancient components of the DNA damage response. eLife 6, e30927 (2017).
Thakker, C., Martínez, I., Li, W., San, K. Y. & Bennett, G. Metabolic engineering of carbon and redox flow in the production of small organic acids. J. Ind. Microbiol. Biot. 42, 403–422 (2015).
Lin, L. et al. Enhancing the thermostability of fumarase C from Corynebacterium glutamicum via molecular modification. Enzyme Microb. Technol. 115, 45–51 (2018).
Giorno, L., Drioli, E., Carvoli, G., Cassano, A. & Donato, L. Study of an enzyme membrane reactor with immobilized fumarase for production of l-malic acid. Biotechnol. Bioeng. 72, 77–84 (2001).
Liu, J. et al. Metabolic engineering of Aspergillus oryzae for efficient production of l-malate directly from corn starch. J. Biotechnol. 262, 40–46 (2017).
Rae, B. D. et al. Cyanobacterial carboxysomes: microcompartments that facilitate CO2 fixation. J. Mol. Microbiol. Biotechnol. 23, 300–307 (2013).
Matson, M. M. & Atsumi, S. Photomixotrophic chemical production in cyanobacteria. Curr. Opin. Biotechnol. 50, 65–71 (2018).
Oliver, N. J. et al. Cyanobacterial metabolic engineering for biofuel and chemical production. Curr. Opin. Chem. Biol. 35, 43–50 (2016).
Kaneko, T. et al. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 30, 109–136 (1996).
Branco Dos Santos, F., Du, W. & Hellingwerf, K. J. Synechocystis: Not just a plug-bug for CO2, but a green E. coli. Front. Bioeng. Biotechnol. 2, 36 (2014).
Kanesaki, Y. et al. Identification of substrain-specific mutations by massively parallel whole-genome resequencing of Synechocystis sp. PCC 6803. DNA Res. 19, 67–79 (2012).
Williams, J. G. K. Construction of specific mutations in Photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol. 167, 766–778 (1988).
Kanesaki, Y., Suzuki, I., Allakhverdiev, S. I., Mikami, K. & Murata, N. Salt stress and hyperosmotic stress regulate the expression of different sets of genes in Synechocystis sp. PCC 6803. Biochem. Biophys. Res. Commun. 290, 339–348 (2002).
Mikami, K., Kanesaki, Y., Suzuki, I. & Murata, N. The histidine kinase Hik33 perceives osmotic stress and cold stress in Synechocystis sp PCC 6803. Mol. Microbiol. 46, 905–15 (2002).
Yamaguchi, K. et al. A two-component Mn2+ -sensing system negatively regulates expression of the mntCAB operon in Synechocystis. Mol. Microbiol. 14, 2901–2913 (2002).
Azuma, M., Osanai, T., Hirai, M. Y. & Tanaka, K. A response regulator Rre37 and an RNA polymerase sigma factor SigE represent two parallel pathways to activate sugar catabolism in a cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 52, 404–412 (2011).
Kaniya, Y. et al. Deletion of the transcriptional regulator cyAbrB2 deregulates primary carbon metabolism in Synechocystis sp. PCC 6803. Plant Physiol. 162, 1153–1163 (2013).
Osanai, T. et al. Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 sigma factor sigE. J. Biol. Chem. 286, 30653–30659 (2005).
Eisenhut, M. et al. Metabolome phenotyping of inorganic carbon limitation in cells of the wild type and photorespiratory mutants of the cyanobacterium Synechocystis sp. strain PCC 6803. Plant Physiol. 148, 2109–2020 (2008).
Nakajima, T. et al. Integrated metabolic flux and omics analysis of Synechocystis sp. PCC 6803 under mixotrophic and photoheterotrophic conditions. Plant Cell Physiol. 55, 1605–1612 (2014).
Osanai, T. et al. Capillary electrophoresis-mass spectrometry reveals the distribution of carbon metabolites during nitrogen starvation in Synechocystis sp. PCC 6803. Environ. Microbiol. 16, 512–524 (2014).
Yoshikawa, K. et al. Integrated transcriptomic and metabolomic analysis of the central metabolism of Synechocystis sp. PCC 6803 under different trophic conditions. Biotechnol. J. 8, 571–580 (2013).
Wan, N. et al. Cyanobacterial carbon metabolism: Fluxome plasticity and oxygen dependence. Biotechnol. Bioeng. 114, 1593–1602 (2017).
Zhang, S. & Bryant, D. A. The tricarboxylic acid cycle in cyanobacteria. Science 334, 1551–1553 (2011).
Xiong, W., Brune, D. & Vermaas, W. F. The γ-aminobutyric acid shunt contributes to closing the tricarboxylic acid cycle in Synechocystis sp. PCC 6803. Mol. Microbiol. 93, 786–796 (2014).
Hasunuma, T., Matsuda, M. & Kondo, A. Improved sugar-free succinate production by Synechocystis sp. PCC 6803 following identification of the limiting steps in glycogen catabolism. Metab. Eng. Commun. 3, 130–141 (2016).
Takeya, M., Hirai, M. Y. & Osanai, T. Allosteric inhibition of phosphoenolpyruvate carboxylases is determined by a single amino acid residue in cyanobacteria. Sci. Rep. 7, 41080 (2017).
Teipel, J. W., Hass, G. M. & Hill, R. L. The substrate specificity of fumarase. J. Biol. Chem. 243, 5684–5694 (1968).
Weaver, T. & Banaszak, L. Crystallographic studies of the catalytic and a second site in fumarase C from Escherichia coli. Biochemistry. 35, 13955–13965 (1996).
Robert, H. B. & David, J. O. Biochemical and molecular characterization of fumarase from plants: Purification and characterization of the enzyme—cloning, sequencing, and expression of the gene. Arc. Biochem. Biophys. 348, 65–74 (1997).
Ito, S., Koyama, N. & Osanai, T. Citrate synthase from Synechocystis is a distinct class of bacterial citrate synthase. Sci. Rep. 9, 6038 (2019).
Puchegger, S., Redl, B. & Stoffler, G. Purification and properties of a thermostable fumarate hydratase from the archaeobacterium Sulfolobus solfataricus. J. Gen. Microbiol. 136, 1537–1541 (1990).
Jiang, C. et al. Identification and characterization of a novel fumarase gene by metagenome expression cloning from marine microorganisms. Microb. Cell Fact. 9, 91 (2010).
Keruchenko, J. S., Keruchenko, I. D., Gladilin, K. L., Zaitsev, V. N. & Chirgadze, N. Y. Purification, characterization and preliminary X-ray study of fumarase from Saccharomyces cerevisiae. BBA-Protein Struct. Mol. Enzymol. 1122, 85–92 (1992).
Song, P., Li, S., Ding, Y., Xu, Q. & Huang, H. Expression and characterization of fumarase (FUMR) from Rhizopus oryzae. Fungal Biol. 115, 49–53 (2011).
Brock, T. D., Brock, K. M., Belly, R. T. & Weiss, R. L. Sulfolobus: a new genus of sulfur-oxidizing bacteria living at low pH and high temperature. Arch. Microbiol. 84, 54–68 (1972).
Thomas, K. C., Hynes, S. H. & Ingledew, W. M. Influence of medium buffering capacity on inhibition of Saccharomyces cerevisiae growth by acetic and lactic acids. Appl. Environ. Microbiol. 4, 1616–1623 (2002).
Touloupakis, E., Cicchi, B., Benavides, A. M. S. & Torzillo, G. Effect of high pH on growth of Synechocystis sp. PCC 6803 cultures and their contamination by golden algae (Poterioochromonas sp.). Appl. Microbiol. Biot. 100, 1333–1341 (2016).
Lin, W., Chan, M., Goh, L. L. & Sim, T. S. Molecular basis for thermal properties of Streptomyces thermovulgaris fumarase C hinge at hydrophilic amino acids R163, E170 and S347. Appl. Microbiol. Biotechnol. 75, 329–335 (2007).
Su, R. R. et al. Identification of a novel fumarase C from Streptomyces lividans TK54 as a good candidate for l-malate production. Molecul. Biol. Rep. 41, 497–504 (2014).
Mizobata, T. et al. Purification and characterization of a thermostable class II fumarase from Thermus thermophilus. Arch. Biochem. Biophys. 355, 49–55 (1998).
Tasaka, Y. et al. Targeted mutagenesis of acyl-lipid desaturases in Synechocystis: evidence for the important roles of polyunsaturated membrane lipids in growth, respiration and photosynthesis. EMBO J. 15, 6416–6425 (1996).
Ito, S., Takeya, M. & Osanai, T. Substrate specificity and allosteric regulation of a d-lactate dehydrogenase from a unicellular cyanobacterium are altered by an amino acid substitution. Sci. Rep. 7, 15052 (2017).
Estevez, M., Skarda, J., Spencer, J., Banaszak, L. & Weaver, T. X-ray crystallographic and kinetic correlation of a clinically observed human fumarase mutation. Protein Sci. 11, 1552–1557 (2002).
Genda, T., Watabe, S. & Ozaki, H. Purification and characterization of fumarase from Corynebacterium glutamicum. Biosci. Biotechnol. Biochem. 70, 1102–1109 (2006).
Hasunuma, T., Matsuda, M., Kato, Y., Vavricka, J. C. & Kondo, A. Temperature enhanced succinate production concurrent with increased central metabolism turnover in the cyanobacterium Synechocystis sp. PCC 6803. Metab. Eng. 48, 109–120 (2018).
Dempo, Y., Ohta, E., Nakayama, Y., Bamba, T. & Fukusaki, E. Molar-based targeted metabolic profiling of cyanobacterial strains with potential for biological production. Metabolites. 4, 499–516 (2014).
Takeya M., Ito S., Sukigara H. & Osanai T. Purification and characterisation of malate dehydrogenase from Synechocystis sp. PCC 6803: Biochemical barrier of the oxidative tricarboxylic acid cycle. Front. Plant Sci. 10.3389 (2018).
Bulku, A., Weaver, T. M. & Berkmen, M. B. Biochemical characterization of two clinically-relevant human fumarase variants defective for oligomerization. Open Biochem. J. 12, 1–15 (2018).
Behal, R. H. & Oliver, D. J. Biochemical and molecular characterization of fumarase from plants: purification and characterization of the enzyme– cloning, sequencing, and expression of the gene. Arch. Biochem. Biophys. 348, 65–74 (1997).
Zubimendi, J. P. et al. The complex allosteric and redox regulation of the fumarate hydratase and malate dehydratase reactions of Arabidopsis thaliana Fumarase 1 and 2 gives clues for understanding the massive accumulation of fumarate. FEBS J. 285, 2205–2224 (2018).
Wigler, P. W. & Alberty, R. A. The pH dependence of the competitive inhibition of fumarase. J. Amer. Chem. Soc. 82, 5482 (1960).
Muro-Pastor, M. I. & Florencio, F. J. Purification and properties of NADP-Isocitrate dehydrogenase from the unicellular cyanobacterium Synechocystis sp. PCC 6803. Eur. J. Biochem. 203, 99–105 (1992).
Acknowledgements
This work was supported by the Ministry of Education, Culture, Sports, Science, and Technology, Japan, by a grant to T.O. from ALCA from the Japan Science and Technology Agency, Grant Number JPMJAL1306 and by JSPS KAKENHI Grant-in-Aid for Scientific Research on Innovative Areas, Grant Number 16H06559.
Author information
Authors and Affiliations
Contributions
N.K. designed the research, performed the experiments, analysed the data, and wrote the manuscript. M.T. analysed the data. T.O. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katayama, N., Takeya, M. & Osanai, T. Biochemical characterisation of fumarase C from a unicellular cyanobacterium demonstrating its substrate affinity, altered by an amino acid substitution. Sci Rep 9, 10629 (2019). https://doi.org/10.1038/s41598-019-47025-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47025-7
This article is cited by
-
Arginine inhibition of the argininosuccinate lyases is conserved among three orders in cyanobacteria
Plant Molecular Biology (2022)
-
Biochemical elucidation of citrate accumulation in Synechocystis sp. PCC 6803 via kinetic analysis of aconitase
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.