Abstract
Multiple lines of evidence have shown that systemic lupus erythematosus (SLE) is attributable to both genetic and environmental factors. The product of GRB2 is a key factor in the activation of B cells and has been reported to be significantly associated with SLE in European populations. In the study, we aimed to investigate the relationship between GRB2 and SLE. A total of 1,710 Han Chinese women comprising 567 SLE patients and 1,143 controls were recruited to genotype 20 selected tagging SNPs. We tested the potential association between 13 clinical variables of SLE and the significant polymorphisms related to SLE. The eQTL data were extracted from the GTEx database to examine the functional consequences of the targeted SNPs. A significant association signal was identified between rs36023980 and SLE in both genotypic and allelic analyses (OR = 0.61, P = 0.0003). Complement inhibition was shown to be significantly associated with the genotypes of SNP rs36023980 in SLE patients (Pgenotype = 0.003). Further stratification analyses showed that the genetic association signal of SNP rs36023980 on SLE could only be identified in cases with complement inhibition. SNP rs36023980 was also identified to be significantly associated with the expression of GRB2 in whole blood and sun-exposed skin. In conclusion, our findings confirm the results from the previous GWAS and are the first to report the association of GRB2 with SLE in Han Chinese population.
Similar content being viewed by others
Introduction
Systemic lupus erythematosus (SLE) is an autoimmune disease characterized by inflammation of connective tissues1. SLE could lower the life expectancy of patients by significantly increasing the risk of multiple clinical manifestations including cardiovascular disease2,3, nephritis4, anemia5 etc. There is no cure for SLE, and common treatments such as NASIDs and corticosteroids may only relieve SLE symptoms6. The incidence rate of SLE is approximately 20–70 per 100,000 people globally and is related to ethnicity, age and gender6. In general, women are 9 times more often than men to be affected by SLE6. Europeans are at lower risk compared to other ethnicities, including Africans and Asians6.
Multiple lines of evidence have shown that SLE is a disease attributable to both genetic and environmental factors4,7,8,9,10,11. Early twin studies have shown that higher concordance was observed in monozygous twins than in dizygous twins12, and the heritability was estimated to be approximately 0.6613. Further gene association mapping studies have identified multiple susceptible loci of SLE. Since 2007, a total of 33 genome-wide association studies have reported more than 700 association signals from hundreds of susceptible loci of SLE14. A recent GWAS published in 2018 by Julià et al. has reported SNP rs36023980 of GRB2 to be significantly associated with SLE disease status in European and Spanish populations15. In addition, Another bioinformatics study using genome molecular interaction networks have identified GRB2 as a potential candidate gene for multiple autoimmune disorders including SLE16. GRB2 encodes growth factor receptor-bound protein 2, which can form a complex with activated epidermal growth factor receptor (EGFR) and with the RAS-specific guanine nucleotide exchange factor SOS1, and thereby mediates the activation of RAS17. GRB2 is a key factor for the activation of B cell related signaling pathways18. Despite the recent significant findings reported by GWAS, further replication studies are still needed to determine the association between GRB2 and SLE in other populations to validate this previous GWAS hit. In addition, it is still not very clear of the functional significance for the SNP identified to be significantly associated with SLE disease status. It is more clinically valuable to examine the potential connections between polymorphisms of GRB2 and clinical variables of SLE.
In the present study, we aimed to investigate the genetic association between polymorphisms of GRB2 and SLE. A total of 1,710 Chinese women were recruited and 20 selected tagging SNPs were genotyped to examine the differences in distribution of genotypes in SLE patients and controls. We tested the potential association between 13 clinical variables of SLE and the significant polymorphism related to the disease status of SLE. eQTL analyses were also performed to explore the potential functional consequence of significant SNPs.
Methods
Study subjects
A total of 1,710 unrelated Han Chinese women comprising 567 SLE patients and 1,143 controls were recruited from the Second Affiliated Hospital of Xi’an Jiaotong University. All patients with SLE were diagnosed according to the 1997 American College of Rheumatology (ACR) classification criteria for SLE. Patients with other autoimmune diseases, cancer, systemic diseases, and other serious diseases were excluded from the study. Meanwhile, unrelated healthy controls were recruited from the same hospital. The inclusion criteria were as follows: (1) without autoimmune diseases, cancer, systemic diseases, and serious diseases and (2) without autoimmune diseases in members of their immediate family. All subjects were born in the local area. Subjects were randomly chosen, unrelated Han Chinese individuals without migration history, which ensures the genetic homogeneity in the study. Data on general characteristics and clinical information including malar rash, photosensitivity, leucopenia, anemia, complement depressed, renal disorder, neurologic disorder, arthritis, anti-dsDNA, anti-RNP, anti-Sm, anti-SSA and anti-SSB were obtained from medical records or questionnaire (Supplemental Table S1). No significant differences in age could be identified between cases and controls (P = 0.3903). Written informed consent was obtained from subjects.
This research was performed in accordance with the ethical guidelines of the Declaration of Helsinki (version 2002) and was approved by the Ethics Committee of Xi’an Jiaotong University.
SNP selection and genotyping
SNPs located within the GRB2 gene region, with a minor allele frequency (MAF) >0.05, were searched in the 1000-genomes CHB database. An r2 ≥0.8 was used as the cutoff criteria in pairwise tagging. Overall, 20 tagging SNPs were selected for further genotyping. The basic information for these 20 tagging SNPs is summarized in Supplemental Table S1. All of these 20 SNPs were intronic SNPs of GRB2. Genomic DNA was extracted from peripheral blood leukocytes according to the manufacturer’s protocol (Genomic DNA kit, Axygen Scientific Inc., CA, USA). The selected tagging SNPs were genotyped using the high-throughput Sequenom MassARRAY platform (Sequenom, San Diego, CA, USA) according to the manufacturer’s protocol. The results were processed using Sequenom Typer 4.0 software to generate genotypic data. For quality control, the disease state of the sample was unknown throughout the genotyping process. The final genotyping call rate for each SNP was greater than 99%, and the overall genotyping call rate was 99.9%. Subsequently, we randomly selected 5% of the samples for regenotyping, and the results were exactly the same as before.
Statistical analyses
Hardy-Weinberg equilibrium (HWE) tests were performed for each SNP in the control group (Supplemental Table S2). Logistic models were fitted for each SNP and age was included as a main covariate to adjust using Plink19. Linkage disequilibrium (LD) blocks were constructed by Haploview based on the algorithm proposed by Gabriel et al.20. Haplotype-based analyses were then performed for each LD block. Bonferroni corrections were applied to address multiple comparisons. For single-marker-based association analyses, the threshold used for significance of P values was 0.05/20 = 0.0025. In addition to genetic association analyses, we also examined the distributions of multiple clinical variables among different genotypic groups of SLE patients. Power analyses were performed by using GAS power calculator (http://csg.sph.umich.edu/abecasis/gas_power_calculator/index.html). For the present study, we could achieve 80% statistical power for a SNP with effect size equal or greater than 1.5 (Supplemental Figure S1).
Bioinformatics and eQTL analyses
To further investigate the functional consequences of SNPs significantly associated with SLE disease status, we extracted eQTL data of 47 human tissues from the GTEx database (https://gtexportal.org/home/)21. Differences in GRB2 gene expression levels among different genotypic groups of targeted SNPs were examined and significant eQTL signals in specific human tissues were reported.
Results
Association between SNP rs36023980 and SLE disease status
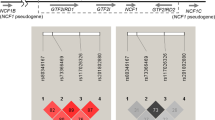
A significant association signal was identified between SNP rs36023980 and SLE disease status in both genotypic and allelic analyses (Table 1, Supplemental Table S3). The minor allele of rs36023980, T allele, could have a protective effect on SLE (OR = 0.61, P = 0.0003). Three LD blocks were constructed (Fig. 1). None of these LD blocks were significantly associated with SLE in the following haplotype-based analyses (Supplemental Table S4).
Association between SNP rs36023980 and SLE related clinical features
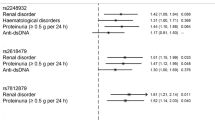
Complement inhibition was shown to be significantly associated with the genotypes of SNP rs36023980 in SLE patients (Padjusted = 0.003, Table 2). None of the other clinical variables were determined to be significantly associated with this SNP. The C allele of rs36023980 may significantly lower the rate for complement inhibition by approximately 50% (OR = 0.48). To further examine whether the association of the rs36023980 with SLE risk is independent of the complement status, we performed stratified logistic regression analyses. The results were summarized in Table 3. As we can see, the association signal of rs36023980 was mainly derived from SLE cases with complement depression (OR = 0.47,P = 1.44 × 10−5).
eQTL pattern of SNP rs36023980
Using the data set extracted from the GTEx database, we demonstrate that SNP rs36023980 is significantly associated with the gene expression levels of GRB2 in two types of human tissues: whole blood (P = 7.7 × 10−6) and sun-exposed skin (P = 0.0003) (Supplemental Table S5, Supplemental Figure S2). The C allele of rs36023980 can significantly increase the gene expression levels of GRB2 in both tissues.
Discussion
With the fast development and application of sequencing and genetic association analyses for studying genetic susceptibility of complex diseases, candidate gene-based association studies have successfully identified susceptibility loci for many complex diseases22,23,24,25,26,27,28,29,30,31,32,33,34,35,36. In this study, we have identified a significant signal association between GRB2 and disease status of SLE. Our findings are a successful replication of a recent GWAS meta-analysis conducted by Julia et al., in which SNP rs36023980 of GRB2 was found to be significantly associated with disease status of SLE in a combined dataset of European and Spanish populations. Our study and this GWAS agree to the direction of the effect of SNP rs36023980. Both studies identified the C allele of this SNP as a risk allele and the T allele as a protective allele. To the best of our knowledge, our study is the first to identify the genetic association between GRB2 and SLE in Chinese Han populations.
GRB2 encodes a receptor tyrosine-kinase (RTK) adaptor protein comprising one SH2 and two SH3 domains37. The RTK adaptor can bind to EGFR and can be involved with the activation of RAS16. An animal study using mice has shown that Grb2 could regulate B-cell maturation and memory responses17. The Grb2-dependent signaling pathways are crucial for control of secondary humoral immune responses17. Additional studies have shown that Grb2 is also very important for T Cell development and differentiation38.
SNP rs36023980 is located at the intronic region of GRB2 and therefore could not have significant functional consequences by altering the structure of the protein encoded by GRB2. Nevertheless, this SNP might play an important role in regulating the gene expression of GRB2. Using data extracted from the GTEx database, our eQTL analyses show that this SNP is significantly associated with the gene expression level of GRB2 in two types of human tissues: whole blood and exposed skin of the lower legs. This finding provides evidence for the functional consequences of SNP rs36023980 to GRB2. SNP rs36023980 located in a region enriched in CG and might involve a CpG site (change T/C). Therefore this SNP might affect the DNA methylation pattern in the region. However, these findings are insufficient to determine whether this signal is a direct or indirect association In this study, we genotyped only 20 intronic SNPs of GRB2. The genetic marker coverage in previous GWAS was even lower than the present study. Thus, rs36023980 may be just a surrogate of some underlying ungenotyped variants that are in strong LD with rs36023980. For future research, a sequencing-based study design would increase the coverage of genetic information and offer a more thorough genetic scan for associations between GRB2 and SLE.
In addition to the association analyses conducted for disease status, we examined the association between multiple clinical variables related to SLE and SNP rs36023980 in our patients. Among all of the 13 clinical variables, only complement inhibition was significantly associated with the genotype of rs36023980 after being corrected for multiple comparisons. The complement system, which is part of the innate immune system, enhances the ability of antibodies and promotes inflammation. Previous studies have indicated that complement system plays a major role in SLE39. Genetic studies have identified that the genetic deficiencies of many classical complement related pathway components are significantly associated with SLE status40. SLE is an autoimmune disorder characterized by widespread complement activation and deposition of complement fragments in the kidney1. In this sense, the direction of effect of the C allele of rs36023980 identified here was in accordance with its effect for SLE disease risk. The C allele was a risk allele of SLE disease status. Specifically, in SLE patients, the C allele of rs36023980 was related to no complement inhibition and therefore was related to increased complement activation. It seems that the genetic effect of SNP rs36023980 on SLE was mediated through complement inhibition based on the results from our stratification analyses. However, we need to be careful to interpret this result because our limited sample size (only 150 cases in the group of no complement inhibition). More research are needed in the future.
Our study suffers from several limitations. First, only women were enrolled in this study. Despite the sex ratio of 9:1 (female to male) for SLE, excluding male patients would harm the generalization of our study results. Second, population stratification might be a potential confounder for our significant hit. This might be even worse considering the fact that the Chinese Han population is highly genetically diverse. In addition, only SNPs located within the gene region of GRB2 were included in this study because of our limited funds. Multiple lines of evidence have shown that important regulatory regions are within 20 kb up- and downstream the gene region. In future studies, SNPs located within these important regulatory regions should be included.
In sum, we have identified a significant association between SNP rs36023980 of GBR2 and SLE disease status. This SNP was also identified to be significantly associated with clinical features of SLE patients. Our findings confirm the results from the previous GWAS and are the first to report the association of GRB2 with SLE in Han Chinese population.
References
Fortuna, G. & Brennan, M. T. Systemic lupus erythematosus: epidemiology, pathophysiology, manifestations, and management. Dental clinics of North America 57, 631–655, https://doi.org/10.1016/j.cden.2013.06.003 (2013).
Manzi, S. et al. Age-specific incidence rates of myocardial infarction and angina in women with systemic lupus erythematosus: comparison with the Framingham Study. American journal of epidemiology 145, 408–415 (1997).
Miner, J. J. & Kim, A. H. Cardiac manifestations of systemic lupus erythematosus. Rheumatic diseases clinics of North America 40, 51–60, https://doi.org/10.1016/j.rdc.2013.10.003 (2014).
Mohan, C. & Putterman, C. Genetics and pathogenesis of systemic lupus erythematosus and lupus nephritis. Nat Rev Nephrol. 11(6), 329–41, https://doi.org/10.1038/nrneph.2015.33 (2015).
Giannouli, S., Voulgarelis, M., Ziakas, P. D. & Tzioufas, A. G. Anaemia in systemic lupus erythematosus: from pathophysiology to clinical assessment. Ann Rheum Dis. 65(2), 144–8 (2006).
Danchenko, N., Satia, J. A. & Anthony, M. S. Epidemiology of systemic lupus erythematosus: a comparison of worldwide disease burden. Lupus 15, 308–318, https://doi.org/10.1191/0961203306lu2305xx (2006).
Lisnevskaia, L., Murphy, G. & Isenberg, D. Systemic lupus erythematosus. Lancet 384, 1878–1888, https://doi.org/10.1016/S0140-6736(14)60128-8 (2014).
Relle, M., Weinmann-Menke, J., Scorletti, E., Cavagna, L. & Schwarting, A. Genetics and novel aspects of therapies in systemic lupus erythematosus. Autoimmun Rev 14, 1005–1018, https://doi.org/10.1016/j.autrev.2015.07.003 (2015).
Teruel, M. & Alarcón-Riquelme, M. E. The genetic basis of systemic lupus erythematosus: What are the risk factors and what have we learned. Journal of Autoimmunity 74, 161–175 (2016).
Ceccarelli, F. et al. Genetic Factors in Systemic Lupus Erythematosus: Contribution to Disease Phenotype. Journal of immunology research 2015, 745647, https://doi.org/10.1155/2015/745647 (2015).
Lewis, M. J. & Jawad, A. S. The effect of ethnicity and genetic ancestry on the epidemiology, clinical features and outcome of systemic lupus erythematosus. Rheumatology 56, i67–i77, https://doi.org/10.1093/rheumatology/kew399 (2017).
Block, S. R., Winfield, J. B., Lockshin, M. D., D’Angelo, W. A. & Christian, C. L. Studies of twins with systemic lupus erythematosus. A review of the literature and presentation of 12 additional sets. The American journal of medicine 59, 533–552 (1975).
Lawrence, J. S., Martins, C. L. & Drake, G. L. A family survey of lupus erythematosus. 1. Heritability. The Journal of rheumatology 14, 913–921 (1987).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res 45, D896–D901, https://doi.org/10.1093/nar/gkw1133 (2017).
Julia, A. et al. Genome-wide association study meta-analysis identifies five new loci for systemic lupus erythematosus. Arthritis research & therapy 20, 100, https://doi.org/10.1186/s13075-018-1604-1 (2018).
Kara, S., Pirela-Morillo, G. A. & Gilliam, C. T. Wilson GD.Identification of novel susceptibility genes associated with seven autoimmune disorders using whole genome molecular interaction networks. J Autoimmun. 97, 48–58 (2019).
Lowenstein, E. J. et al. The SH2 and SH3 domain-containing protein GRB2 links receptor tyrosine kinases to ras signaling. Cell 70, 431–442 (1992).
Ackermann, J. A., Radtke, D., Maurberger, A., Winkler, T. H. & Nitschke, L. Grb2 regulates B-cell maturation, B-cell memory responses and inhibits B-cell Ca2+ signalling. The EMBO journal 30, 1621–1633, https://doi.org/10.1038/emboj.2011.74 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics 81, 559–575 (2007).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Carithers, L. J. & Moore, H. M. The Genotype-Tissue Expression (GTEx) Project. Biopreservation and biobanking 13, 307–308, https://doi.org/10.1089/bio.2015.29031.hmm (2015).
Guan, F. et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr Res. 152, 97–104 (2014).
Chen, G., Guan, F., Lin, H., Li, L. & Fu, D. Genetic analysis of common variants in the HDAC2 gene with schizophrenia susceptibility in Han Chinese. Journal of human genetics. 60, 479–484 (2015).
Guan, F. et al. Evaluation of genetic susceptibility of common variants in CACNA1D with schizophrenia in Han Chinese. Scientific reports. 5, 12935 (2015).
Guan, F. et al. Evaluation of association of common variants in HTR1A and HTR5A with schizophrenia and executive function. Scientific reports. 6, 38048 (2016).
Guan, F. et al. Evaluation of voltage-dependent calcium channel γ gene families identified several novel potential susceptible genes to schizophrenia. Scientific reports. 6, 24914 (2016).
Guan, F. et al. A population-based association study of 2q32.3 and 8q21.3 loci with schizophrenia in Han Chinese. Journal of psychiatric research. 47, 712–717 (2013).
Yang, H. et al. 4q22.1 contributes to bone mineral density and osteoporosis susceptibility in postmenopausal women of Chinese Han population. PloS one. 8, e80165 (2013).
Guan F, et al. Two-stage association study to identify the genetic susceptibility of a novel common variant of rs2075290 in ZPR1 to type 2diabetes. Scientific reports. 6, 29586 (2016).
Jia, X. et al. Two-stage additional evidence support association of common variants in the HDAC3 with the increasing risk of schizophrenia susceptibility. American journal of medical genetics. Part B, Neuropsychiatric genetics. 171, 1105–1111 (2016).
Liu, X. et al. Dopamine D3 Receptor‐Regulated NR2B Subunits of N‐Methyl‐d‐Aspartate Receptors in the Nucleus Accumbens Involves in Morphine‐Induced Locomotor Activity. CNS neuroscience & therapeutics. 20(9), 823–829 (2014).
Zhu, L. et al. mRNA changes in nucleus accumbens related to methamphetamine addiction in mice. Scientific reports. 6, 36993 (2016).
Zhang, T. et al. Voltage-gated calcium channel activity and complex related genes and schizophrenia: A systematic investigation based on Han Chinese population. Journal of psychiatric research. 106, 99–105 (2018).
Li, J. et al. Relationship between schizophrenia and changes in the expression of the long non-coding RNAs Meg3, Miat, Neat1 and Neat2. Journal of psychiatric research. 106, 22–30 (2018).
Han, W. et al. Relationship of common variants in CHRNA5 with early-onset schizophrenia and executive function. Schizophr Res. 206, 407–412 (2019).
Han, W. et al. Combining information from multiple bone turnover markers as diagnostic indices for osteoporosis using support vector machines. Biomarkers. 24(2), 120–126 (2019).
Feng, G. S. et al. Grap is a novel SH3-SH2-SH3 adaptor protein that couples tyrosine kinases to the Ras pathway. Journal of Biological Chemistry 271, 12129 (1996).
Radtke, D. et al. Grb2 Is Important for T Cell Development, Th Cell Differentiation, and Induction of Experimental Autoimmune Encephalomyelitis. Journal of immunology 196, 2995 (2016).
Leffler, J. & Bengtsson, A. A. Blom AM.The complement system in systemic lupus erythematosus: an update. Ann Rheum Dis. 73(9), 1601–6 (2014).
Truedsson, L., Bengtsson, A. A. & Sturfelt, G. Complement deficiencies and systemic lupus erythematosus. Autoimmunity. 40, 560–6 (2007).
Acknowledgements
This research was supported by the general project of key research and development program in Shaanxi Province (No. 2018SF-008). The funding sources had no role in the design of this study, the collection, analysis and interpretation of data, the writing of the report, or the decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
Authors Xu M.F., Du S.Y. and Xiao S.X. conceived and designed the study. Xu M.F. and Du S.Y. carried out candidate SNPs selection and statistical analyses. Xu M.F., Liu Y., Liu Y.L., Li X.L. and Cheng C.T. conducted subject screening. Xu M.F., Liu Y., Liu Y.L., Li X.L. and Dong W. contributed to the collection and preparation of control DNA samples. Xu M.F. wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, M., Liu, Y., Li, X. et al. Evaluation of genetic susceptibility between systemic lupus erythematosus and GRB2 gene. Sci Rep 9, 10335 (2019). https://doi.org/10.1038/s41598-019-46827-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46827-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.