Abstract
Dengue virus nonstructural protein 3 (NS3) fulfills multiple essential functions during the viral replication and constitutes a prominent drug target. NS3 is composed by a superfamily-2 RNA helicase domain joined to a serine protease domain. Quantitative fluorescence titrations employing a fluorescein-tagged RNA oligonucleotide were used to investigate the effect of salts on the interaction between NS3 and single stranded RNA (ssRNA). We found a strong dependence of the observed equilibrium binding constant, Kobs, with the salt concentration, decreasing at least 7-fold for a 1-fold increase on cation concentration. As a result of the effective neutralization of ~10 phosphate groups, binding of helicase domain of NS3 to ssRNA is accompanied by the release of 5 or 7 monovalent cations from an oligonucleotide or a polynucleotide, respectively and of 3 divalent cations from the same oligonucleotide. Such estimates are not affected by the type of cation, either monovalent (KCl, NaCl and RbCl) or divalent (MgCl2 and CaCl2), nor by the presence of the protease domain or the fluorescein label. Combined effect of mono and divalent cations was well described by a simple equilibrium binding model which allows to predict the values of Kobs at any concentration of cations.
Similar content being viewed by others
Introduction
Dengue virus (DENV) is an important human pathogen that belongs to the Flaviviridae family together with other emergent and re-emergent viruses such as Zika, yellow fever, West Nile and hepatitis C viruses1. Found in more than one hundred countries around the world, DENV causes over 390 million infections per year2 and puts about half of the world population at risk of infection3,4.
The Flaviviridae genome is a plus-stranded RNA molecule that is translated into a unique polyprotein which is subsequently processed by viral and host proteinases into 10 mature viral proteins: three structural (capsid, premembrane, and envelope proteins) and seven non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5)5,6,7.
DENV NS3 protein contains 618 amino acids with two well defined domains. At the N-terminal region, amino acid residues 1 to 170, together with part of the NS2B protein, form an active serine protease and at the C-terminal region, amino acid residues 171 to 618 folds into a tri-lobed domain with RNA-helicase, RNA 5’-triphosphatase (RTPase), nucleoside 5’-triphosphatase (NTPase), and RNA annealing activities8,9,10,11. In vitro, these two functional domains appear to work independently of each other. In this regard, it has been previously shown that the recombinant full-length NS3 protein (NS3f) including both the protease and helicase domains, and the NS3 helicase domain alone (NS3h) exhibit similar helicase and NTPase activities9,10,11,12. Nevertheless, the protease domain has been proposed to modulate the helicase/NTPase activities13. To further investigate this possibility, the NS3h and NS3f variants of the NS3 protein were included in this study.
RNA helicases are ubiquitous proteins that intervene in practically all pathways of RNA metabolism14. An extended property of these motor proteins is their ability to catalyze the hydrolysis of nucleotides (commonly ATP), which supplies the driving force for directional movement along RNA strands and for unwinding of double stranded RNA tracks.
The tertiary structure of DENV NS3h13 is shaped by an arrangement of two RecA-like folds, which includes the distinctive sequence motifs of superfamily 2 helicases15,16 and determines the core of many ATP-driven molecular motors14,17,18. The cleft between the RecA-like subdomains of NS3h (residues 171 to 481) and its C-terminal subdomain (residues 482 to 618) display numerous basic residues and is wide enough to harbor an ssRNA but not a duplex13,19.
Binding of a helicase to a polynucleotide track is the first necessary step of the mechanism of translocation and unwinding activities. Most in vitro studies have shown that RNA-helicases bind predominantly to single stranded RNA -and DNA in some cases- with a defined orientation with respect to the 3′-5′ nucleic acid polarity. Binding displays little or no specificity for particular sequences20,21 however Jankowsky & Harris22 reformulated the traditional specific/nonspecific in terms of affinity distribution along sequence space.
Several studies on nonspecific protein-nucleic acid interactions have provided a general understanding of the forces involved in these interactions as well as a theoretical framework to characterize them experimentally23. A general feature of this kind of interactions is their strong dependence on the salt concentration and type21,24. Such property has been successfully explained as the result of the net release of counterions from the nucleic acid and, in some cases, from the protein upon formation of the protein-nucleic acid complexes. Indeed, the free energy change proceeding from the release of cations from the nucleic acid to the bulk solution is generally recognized as a mayor contribution to the thermodynamic stability of protein-nucleic acid complexes24,25,26. From an experimental perspective the theory developed by Record et al. provides a means to determine the net number of ions released from the analysis of the dependence of the equilibrium binding constant for a given protein-nucleic acid interaction on salt concentration. In addition, this theory allows estimating the effective number of ionic pair interactions established between the protein and the nucleic acid phosphate groups. We hypothesize that this analysis provides information that is not only important to complete the thermodynamic characterization of the non-specific binding of NS3h to ssRNA, but also is relevant in the elucidation of the molecular mechanism of the NS3 helicase as a motor protein.
We have shown previously that DENV NS3h is a monomeric protein, binds nonspecifically to ssRNA with equal estimations for minimum and occluded binding site sizes (among 9 and 11 nucleotide residues), and that binding of several NS3h molecules to the same ssRNA track takes place with weak positive cooperativity27. In this study, we report an in vitro analysis to characterize the effects of concentration and type of monovalent and divalent cations on the equilibrium binding of DENV NS3h to ssRNA using quantitative fluorescence spectroscopy techniques.
Materials and Methods
General reagents
MgCl2, KOH, KCl and 3-(N-morpholino)propanesulfonic acid (MOPS) were obtained from Mallinckrodt Baker (ACS reagents). NaOH and HCl were from Carlo Erba. RbCl, CaCl2, adenosine 5′-triphosphate (ATP), 3-[(3-Cholamidopropyl) dimethylammonio]-1-propanesulfonate (CHAPS) and ethylenediaminetetraacetic acid (EDTA) were obtained from Sigma Aldrich. RNAse-free ultrapure water (Arium mini Ultrapure Lab Water, Sartorius) was used to prepare all stock solutions and buffers. Concentrations of CaCl2, MgCl2 and EDTA in stock solutions were determined by volumetric titrations using a Zn2+ standard solution and eriochrome black T indicator28.
Protein purification
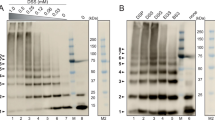
In the work presented here, we used two variants of the NS3 protein derived from the cDNA of an infectious clone of the DENV serotype 229. These proteins were expressed and purified in the same way that we explained in our previous works11,12. The NS3h variant is composed of residues 171 to 618 corresponding to the NTPase/helicase domain of the NS3 protein. The variant NS3f is an ensemble composed of the central region of the NS2B protein (residues 49 to 95) joined via a flexible linker peptide (GGGGSGGGG) to the full length NS3 protein (residues 1 to 618) so as to include both the protease domain and the NTPase/helicase domain. Additionally, the His51 residue corresponding to the catalytic triad of the protease domain was replaced by alanine to prevent auto-proteolytic degradation. Moreover, a hexa-histidine tag derived from the pET-28a plasmid was added to the N-terminal region of each protein construct to allow purification by affinity chromatography. The eluted fractions carrying the protein of interest were dialyzed twice at 4 °C in dialysis buffer (50 mM MOPS-KOH, pH 6.5 at 25 °C, 0.2 mM DTT, 200 mM KCl, 5% w/v glycerol). The protein samples were concentrated to 10 mg/ml using a centrifugal filter unit (Amicon Ultra, Millipore Corp.) and then centrifugated to remove any precipitated material. Supernatants were aliquoted and stored at −70 °C until use. Protein preparations were evaluated by SDS-PAGE and Coomassie staining to determine their homogeneity. Protein concentration was calculated by spectrophotometry according to the Edelhoch method30 using the empirical extinction coefficients obtained for NS3f and NS3h in our dialysis buffer (e280nm,25°C = 108.25 mM−1∙cm−1 and 73.13 mM−1∙cm−1, respectively)12.
RNA oligo and polynucleotides
Fluorescein-labeled and unlabeled RNA oligonucleotides were purchased from Midland CRC (Texas, USA). These oligonucleotides were purified by HPLC by the manufacturer. The lyophilized oligonucleotides were dissolved in water and their concentrations were calculated by spectrophotometry. The RNA oligonucleotides (both F-p-R10 and p-R10) have the same sequence 5′-AGUUGAGUUG-3′ and are phosphorylated at their 5′ ends, so both contain 10 phosphate groups. The F-p-R10 oligo differs from p-R10 in which incorporates a fluorescein moiety covalently attached to the 5′ terminal phosphate group through a phosphodiester bond.
Polyadenylic acid (poly(A), potassium salt, Catalog number P-3001) and polyuridylic acid (poly(U), potassium salt, Catalog number P-3004) were from Midland Certified Reagent Company. Both nucleic acids were dissolved in 10 mM MOPS/KOH buffer (pH 6.5 at 25 °C, 2.0 mM K+) and stored at −80 °C. Their concentrations in mononucleotide units were determined spectrophotometrically31 employing the following molar extinctions coefficients: 10.3 mM−1∙cm−1 at 260 nm for poly(A) and 9.2 mM−1∙cm−1 at 260 nm for poly(U)32. According with specification of supplier, an analytical polyacrylamide gel electrophoresis showed nearly all of the polymer of poly(A) to be greater than 250 nucleotides in length and nearly all of the polymer of poly(U) to be less than 250 nucleotides in length.
Equilibrium fluorescence titrations
Steady-state fluorescence measurements were performed as described in our previous study27. Binding of NS3 variants to the fluorescent probeF-p-R10 was monitored in a Jasco FP-6500 spectrofluorometer equipped with a Peltier thermostat by following the fluorescence emitted between 510 and 640 nm upon excitation at 495 nm. Emission and excitation slits were set at 3 nm bandwidth and all measurements were done in a 3 mm-pathlength quartz cuvette. A linear correlation between the fluorescence intensity and the F-p-R10 concentration was checked in our experimental conditions. Due to the very low absorbance of the samples at the excitation wavelength of 495 nm (less than 0.005 for all the reaction mixtures), the inner filter effects were negligible (less than 1% of the observed signal) and therefore it was not necessary to apply corrections33. Fluorescence spectra of different mixtures of NS3 and ssRNA were measured repeatedly until no further change occurred between measurements. Approximately 30 minutes of incubation were enough to achieve constancy to all conditions tested. The mixtures were allowed to stand in reaction tubes (Greiner Bio-One) at 25 °C in a thermomixer prior to recording the fluorescence spectra27. In this way, concentrations of all the species present in the reaction mixtures correspond to those of the equilibrium27. Relative fluorescence change was defined as:
where F is the fluorescence intensity at a given NS3 protein variant concentration and F0 denotes the fluorescence intensity of the free F-p-R10.
Fluorescence titrations of F-p-R10 and competition titrations of p-R10 with NS3 protein
Titration curves were analyzed using a single-site binding isotherm:
where Kobs is the macroscopic association constant describing the binding of NS3 to the oligonucleotide and ΔFmax is the maximum relative fluorescence increase. Note that Eq. 2 is a function of the concentration of free NS3, which is unknown, and must be solved in terms of the total concentrations of both NS3 and RNA to fit this equation to the experimental titration curves:
Competition titration experiments were achieved according to the procedure defined by Jezewska et al.34 Briefly, if the fluorescent (F-RNA) and non-fluorescent (RNA) macromolecules compete for their binding to the ligand (NS3), mass balance on the ligand should read as:
where the binding density, ν, for labeled and unlabeled RNA are identified by subscripts f and u, respectively. Regardless of the total concentration of unlabeled macromolecule, a given value of ΔFrel would be reached at a unique value of free ligand concentration and thus of νu34.
Reaction media
Titration assays were performed at 25 °C in buffer BK (25.0 mM MOPS/KOH, 0.50 mM EDTA and 0.50 mM CHAPS) or BNa (25.0 mM MOPS/NaOH, 0.50 mM EDTA and 0.50 mM CHAPS), pH 6.5 (25 °C), containing variable amounts of monovalent and divalent salts (KCl, NaCl or RbCl and MgCl2 or CaCl2, respectively). As reported in a previous work, CHAPS detergent was added to avoid problems likely due to sticking of the protein to the reaction tubes and/or tips walls27. In the range of protein and salt concentrations employed we did not detect the formation of protein aggregates. An interesting observation in this regard is that, while the formation of aggregates is dependent on the total concentration of monovalent salts (data not shown) and they are rapidly formed at KCl concentrations below 20 mM at low concentrations of MgCl227, aggregation was not observed in mixtures containing low concentrations (~10 mM) of KCl in the presence of MgCl2 concentrations above 10 mM. This observation suggests that the formation of aggregates is prevented by the increase in the ionic strength.
The amount of monovalent cation proceeding from different sources was taken into account to compute total monovalent cation concentrations in the reaction media. The sources of monovalent cation were: the corresponding chloride salt (MCl) solutions, the titration of the MOPS buffer with KOH or NaOH solutions to the desired pH and the stock solutions of EDTA, protein and nucleic acid. The value of Mg-EDTA dissociation constant used to calculate the concentration of free Mg2+ in equilibrium was taken from the literature35.
Competition titrations with homopolyribonucleotides poly(U) and poly(A)
Mixtures of NS3 and the F-p-R10 oligonucleotide were titrated with poly(U) and poly(A). Fluorescence measurements were done as described previously but carried out in a 1 cm-pathlength cuvette equipped with a magnetic stirrer. Titrant solutions of polynucleotide contained the same concentrations of NS3 and F-p-R10 as the solution in the cuvette, so that corrections for dilution were not required. The initial volume in the cell was 1600 or 1800 μl and aliquots from 20 μl to 300 μl of poly(U) or poly(A)were added. Fluorescence emission spectra (495 nm → 510 to 640 nm) were recorded ~5 to 15 min after the addition of each aliquot.
The binding of NS3 to poly(U) and poly(A) was analysed with McGhee-von Hippel formalism for non-specific interactions of large ligands to a one-dimensional lattice of infinite length27,36. The parameters of the binding to F-p-R10 were set to those obtained in independent direct titrations of this oligonucleotide.
The mass balance equation for NS3 is:
where νf is the binding density for the labeled oligonucleotide and νpN is the binding density for poly(U) or poly(A),
where [NS3]pN is the concentration of NS3 bound to poly(U) or poly(A), and [poly(N)]total is the total concentration of poly(U) or poly(A) in nucleotide residues. According to the formalism of McGhee and von Hippel36 the generalized equation for the binding density is27:
where r is given by
KpN, m and ωpN denote the intrinsic association constant, the occluded site size and the cooperativity factor, respectively, characterizing the binding of NS3 to the corresponding homopolyribonucleotide. The occluded site size was set to 10 nucleotides according to our previous study27.
Data analysis
Model simulations and nonlinear regression analysis of the data were performed using Copasi version 4.13 (http://www.copasi.org/) and Libreoffice 4.2 spreadsheets (http://www.libreoffice.org/). Figures were edited in QtiPlot version 0.9.8.9 (http://www.qtiplot.com/).
Results
Previous studies from our laboratory have shown that NS3h is a monomeric protein with a ssRNA binding site size of 9 nucleotides27. Those experiments were performed at a fixed set of reaction conditions: pH 7.0, 30 °C, 1.9 mM free Mg2+ and 100 mM K+, with both cations added in the form of their chloride salts27. In the present work, we extend the analysis to characterize the effect of ions on the equilibrium constant for the binding of NS3 to ssRNA. To this aim, we performed fluorescence titration experiments using the signal proceeding from a fluorescein-labeled oligonucleotide (F-p-R10, see Materials and Methods) for which we have shown that NS3h binds with a 1:1 stoichiometry27.
Effect of monovalent cation concentration and type on the binding of NS3h to RNA oligonucleotide
Fluorescence titrations of the F-p-R10 oligonucleotide with the NS3h protein were carried out in buffer B (pH 6.5, 25 °C) in the presence of different concentrations of monovalent salts, KCl, NaCl and RbCl. As described in Materials and Methods, buffer B contains a basal amount of about 5 mM of monovalent cations K+ (buffer BK) or Na+ (buffer BNa) proceeding from the pH adjustment of the MOPS buffer and EDTA solution with KOH or NaOH, respectively. Additionally, a small amount of K+ (about 2 mM) is provided by the dialysis buffer in which the purified NS3h protein was stored, which was compensated with the variation of protein concentration along each titration curve. The rest of the monovalent cation to reach the total given concentration is added in the form of chloride salt.
Each experiment consisted of titrations of F-p-R10 with NS3 were done at two or three oligonucleotide concentrations. Figure 1 shows representative titrations in the presence of K+, Na+ or Rb+. At higher F-p-R10 concentration, more NS3 concentration is necessary to get the same value of ΔFrel. This is because at higher RNA concentrations more ligand is necessary to get identical degree of binding.
Representative fluorescence titration of F-p-R10 with NS3h. Dot symbols represent experimental values and solid lines are simulated titration curves based on Eq. 3 with parameter values obtained by non-linear regression analysis: KCl: Kobs = 3.6 × 107 M−1, ΔFmax = 1.6, RbCl: Kobs = 1.5 × 107 M−1, ΔFmax = 1.6 y NaCl: Kobs = 1.2 × 107 M−1, ΔFmax = 1.5. Binding of NS3h to F-p-R10 was monitored by fluorescence emission from 510 nm to 640 nm upon excitation at 495 nm. Titration experiments were performed in buffer BK or BNa supplemented with 130 mM NaCl, 120 mM RbCl and 105 mM KCl at 25.0 °C and pH 6.5.
Consistent with a 1:1 binding stoichiometry, each set of titration curves was well depicted by Eq. 3 with best-fitting values of the observed macroscopic association constant Kobs and the maximum relative fluorescence increase (ΔFmax) acquired from non-linear regression analysis of the results (solid lines in Fig. 1).
Logarithm of Kobs is plotted in Fig. 2 as a function of the logarithm of total monovalent cation concentration [M+] (Notice that, in order to apply logarithms, the quantities involved must of course be dimensionless. In accordance with usual practice this is implicitly done referring concentrations and equilibrium constant to 1 molar, M, standard states. That is, all logarithms are applied over the numerical values obtained when the quantities are expressed in molar units). It is observed that, within the experimental error, equal values of Kobs were obtained for the same values of [M+], regardless of the nature of the added cation, K+, Na+ or Rb+. In other words, the effect of monovalent salts on the binding of NS3h to the fluorescein-labeled oligonucleotide shows no specificity for the cation involved.
Effect of monovalent cations on Kobs for the interaction NS3h:F-p-R10. Experiments were performed with a series of salts differing in cation type: ( ) NaCl, (
) NaCl, ( ) KCl and (
) KCl and ( ) RbCl (plotted as total [M+]). Solid line is the representation of Eq. 9 with parameter values obtained from an overall fit by linear least-square analysis: a = 5.0 and b = 2.6. Experiments were carried out in buffer BK or BNa, at 25.0 °C and pH = 6.5.
) RbCl (plotted as total [M+]). Solid line is the representation of Eq. 9 with parameter values obtained from an overall fit by linear least-square analysis: a = 5.0 and b = 2.6. Experiments were carried out in buffer BK or BNa, at 25.0 °C and pH = 6.5.
Additionally, these results show a strong dependence of Kobs on the salt concentration, decreasing about 25-fold for a 1 fold increase on [M+], which is a common property of protein-nucleic acid interactions37.
In the range of salt concentration tested, 90 to 140 mM of M+, the logarithm of Kobs varies linearly with the logarithm of [M+] (Fig. 2) and this dependence can be described by the following equation:
where clearly −a is the slope (∂logKobs/∂log[M+])pH,T of the straight line defined by this equation, and b is the value of logKobs at the reference state [M+] = 1 M. Linear regression analysis of the results obtained in the presence of K+, Na+ and Rb+ provides the values of a and b shown in Table 1. These values were not significantly different for the three cations tested and simultaneous linear fitting to all the results provides the values (±standard error) a = 5.0 ± 0.2 and b = 2.6 ± 0.2.
The linear dependence of logKobs on log[M+] may be interpreted and analyzed by means of the theoretical formalism developed by Record et al. to analyze the linkage effects of multiple equilibria with small ligands on the binding between macromolecules24,38. This formalism indicates that, in the absence of preferential interactions of ions with the protein and of differential hydration effects, the slope of the dependence of logKobs on salt concentration, (∂logKobs/∂log[M+])pH,T, is dictated by the net number of cations released from the nucleic acid upon binding of the protein (a in the following reaction scheme)38. The intercept b is identified as the logarithm of the ‘thermodynamic’ equilibrium constant, logKT, of the binding reaction shown below, in which the release of cations is made explicit.
According with this interpretation, binding of DENV NS3h to the ssRNA oligonucleotide is accompanied by the release of 5.0 ± 0.2 monovalent cations M+. Using the linear fit value of b, one obtains KT = 392 ± 1 (referred to 1 M standard state for all reactants). Although KT, unlike Kobs, does not depend on [M+], still retains possible dependencies on other variables such as pH, buffer composition and temperature.
Notice that our analysis does not take into account potential effects of chloride ions39, whose concentration changes together with the concentration of cation. Thus, the meaning attributed to the value of the slope, -a, remains in a hypothetical ground and is open to further verification (see below on the discussion about the effect of divalent cations).
Regarding the spectroscopic parameter, ΔFmax, we observed variations between experiments. However, it did not appear to be dependent on the salt concentration (Fig. S1 in supplementary information) and take a mean value 2.50 ± 0.05.
Testing the impact of the fluorescein label: binding of NS3h to the unlabeled RNA oligonucleotide
In order to test whether the fluorescein label contributes to the salt dependence of the observed association constant, we performed competition titration experiments employing the unlabeled counterpart of the F-p-R10 oligonucleotide, p-R10. This unlabeled oligonucleotide, unlike those used in our previous work27, contains the same number of phosphate groups as the labeled one.
F-p-R10 at 30 or 50 nM concentration was titrated with NS3h in the presence of different concentrations of the p-R10 oligonucleotide, in buffer BK (pH 6.5, 25 °C) at different concentrations of KCl.
Each set of titration curves (at a given K+ concentration) was numerically analyzed employing a competition model in which labeled and unlabeled oligonucleotide compete with a 1:1 stoichiometry for their binding to NS3h (see Materials and Methods and Gebhard et al27.). Values of Kobs for the unlabeled oligonucleotide was determined by least-square fitting of the model, setting the value of Kobs for the labeled oligonucleotide to the value calculated from Eq. 9 using the parameters a and b obtained from the results shown in Fig. 2. The results were satisfactorily described by this competition model (Fig. 3).
Panel A. Effect of KCl on Kobs for the binding of NS3h to the unlabeled p-R10 oligonucleotide. Dot symbols represent logKobs values for the binding of NS3h to the unlabeled analogue of the F-p-R10 oligonucleotide and the blue solid line (−) represents graphically Eq. 9 with parameter values obtained by linear least-square fit: a = 4.9 and b = 2.1. Black solid line (−) is the representation of the same equation with the corresponding parameters for the binding of NS3h to F-p-R10. Experiments were carried out in buffer BK at 25.0 °C and pH 6.5. Panel B. Representative competition titration experiments of F-p-R10 and p-R10 with NS3h. Dots symbols represent experimental values and solid lines are simulated titration curves based on Eqs 3 and 4 with parameter values obtained by non-linear regression analysis: Kobs = 2.5 × 107 M−1 and ΔFmax = 1.8. Binding of NS3h to F-p-R10 (20 nM) was monitored by fluorescence emission from 510 nm to 640 nm upon excitation at 495 nm. Titration experiments were performed in buffer BK at 25 °C and pH 6.5 supplemented with 95 mM KCl and the indicated concentration of the p-R10 oligonucleotide.
Values of Kobs for the unlabeled p-R10 oligonucleotide were ~4-fold lower than for the F-p-R10 oligomer (Fig. 3) which is in agreement with the change in intrinsic affinity found in a previous work (;27 see Discussion). logKobs varied linearly with log[M+], with slope −a = −4.9 ± 0.2 and ordinate b = 2.1 ± 0.2. Thus, binding of NS3h to the unlabeled oligonucleotide is accompanied by a net release of 5 monovalent cations from the nucleic acid, in agreement with the value obtained for the labeled oligonucleotide.
Effect of monovalent cations on the binding of NS3h to polynucleotides
It has been previously reported that the number of ions released when a charged ligand binds to a polymeric nucleic acid is different from when it binds to short oligomers40,41,42. This difference was attributed to changes in the number of counterions thermodynamically associated per phosphate groups and explained by the contributions of coulombic end effect42. In order to test this phenomenon on the NS3h-RNA interaction we determined the association binding constant of NS3h to poly(A) and poly(U) in the presence of different concentrations of monovalent cations.
Mixtures of 50 nM F-p-R10 and 0.20 or 0.60 µM NS3h were titrated with poly(A) or poly(U), in buffer BK (pH 6.5, 25 °C) at different concentrations of KCl. The addition of poly(A) or poly(U) caused a decrease in the value of ∆Frel as expected if both nucleic acids compete with F-p-R10 for the binding to NS3h (Fig. 4B, Panel B and Fig. S2 in Supplementary Information). Titration curves were analyzed employing a competition model in which F-p-R10 binds to NS3h following a 1:1 stoichiometry (Eq. 3) whereas binding to poly(A) or poly(U) is described using the McGhee-von Hippel formalism for the non-specific interaction of large ligands to a one-dimensional lattice of infinite length36,43 (see Materials and Methods). Binding constants for poly(A) and poly(U) at each salt concentration tested were acquired by non-linear regression analysis of the results using this competition model (see Materials and Methods).
Panel A: Effect of KCl on Kobs for the interactions of NS3 with polynucleotides. Dot symbols represent logKobs values for the binding of the following pairs: ( ) NS3h:poly(A), (
) NS3h:poly(A), ( ) NS3h:poly(U), (
) NS3h:poly(U), ( ) NS3f:poly(U). Solid lines are the representations of Eq. 9 with parameters values obtained by linear least-square fits: a = 6.0 and b = 1.1 for NS3h:poly(A), a = 7.7 and b = −2.8 for NS3h:poly(U) and a = 7.2 and b = -2.3 for NS3f:poly(U). Panel B: Representative competition titration experiments of NS3h:F-p-R10 with poly(A). Dot symbols represent experimental values and the solid lines are representation of Eq. 7 with parameter values obtained by non-linear regression analysis. Binding of NS3h to F-p-R10 was monitored by fluorescence emission from 510 nm to 640 nm upon excitation at 495 nm. Titration experiments were carried out in buffer BK at 25.0 °C and pH 6.5.
) NS3f:poly(U). Solid lines are the representations of Eq. 9 with parameters values obtained by linear least-square fits: a = 6.0 and b = 1.1 for NS3h:poly(A), a = 7.7 and b = −2.8 for NS3h:poly(U) and a = 7.2 and b = -2.3 for NS3f:poly(U). Panel B: Representative competition titration experiments of NS3h:F-p-R10 with poly(A). Dot symbols represent experimental values and the solid lines are representation of Eq. 7 with parameter values obtained by non-linear regression analysis. Binding of NS3h to F-p-R10 was monitored by fluorescence emission from 510 nm to 640 nm upon excitation at 495 nm. Titration experiments were carried out in buffer BK at 25.0 °C and pH 6.5.
In the range of salt concentrations tested, the values of Kobs for poly(U) were ~50–100 times lower than for poly(A) which is in accordance with the results obtained in a previous work27. Figure 4 shows that logKobs for poly(A) or poly(U) decreased linearly with log[K+]. Linear regression analysis of the results employing Eq. 9 provided the following best fitting values for a and b: 6.0 ± 0.2 and 1.1 ± 0.2 for poly(A) and 7.7 ± 0.5 and −2.8 ± 0.6 for poly(U). These results indicate that 6 monovalent cations are released when NS3h binds to poly(A), and 8 monovalent cations are released when it binds to poly(U).
Using the limiting-law predicted by the counterion condensation theory44,45, Record et al24,38. showed that, in the absence of preferential interaction of ions with the protein and differential hydration effects, the slope of the dependence of logKobs on log[M+] does not depend on M+ concentration and can be computed as the product of two constant factors:
where m′ is the number of ionic interactions between positive groups on the ligand (NS3h) and negatively charged phosphate group on the nucleic acid (ssRNA), and ψ is the fraction of a monovalent counterion thermodynamically associated per phosphate in the absence of protein and its value is a characteristic property of the nucleic acid24,45. In all, the empirical Eq. 9 is reformulated as:
This equation is of great practical use because by knowing the value of ψ, the estimation of the number of ionic interactions involved in a nucleic acid-protein system is straightforward. In this way, substituting in Eq. 11 the known value of ψ for poly(U) (ψ = 0.74 ± 0.04)46,47 and the value of slope obtained here, we estimate that NS3h establishes 10.4 ± 0.5 ionic interactions with poly(U). This estimate is coincident with both the occluded and minimum binding site sizes estimated for the NS3h protein, between 9 to 11 nucleotides12,27. We do not perform the same analysis with the results obtained from poly(A) because in this case the number of ion released does not depend linearly with the valence of oligocation48.
Testing the impact of the NS3 protease domain on the association between full-length NS3 and ssRNA
In this section we extended the study of the effect of monovalent cations on the NS3-ssRNA interaction using a full-length construct of the NS3 protein (NS3f) which includes its N-terminal protease domain (see Materials and Methods).
We conducted fluorescence titrations of the F-p-R10 oligonucleotide with the NS3f protein in buffer BK (pH 6.5, 25 °C) in the presence of different concentrations of monovalent cation K+. Titration curves were satisfactorily described by a 1:1 ligand binding model defined by Eq. 3 (Fig. 5B, Panel B) which was fitted to the experimental results to obtain values of Kobs and ΔFmax.
Panel A: Effect of KCl on Kobs for the interaction of NS3f with F-p-R10. Dot symbols represent logKobs values for the binding of full-length construct of NS3 to the oligonucleotide and blue solid line (−) is the representation of Eq. 9 with parameters values obtained by linear least-square fits: a = 5.0 and b = 2.6. Black solid line (−) is the representation of the same equation with the corresponding parameters for the binding of NS3h to F-p-R10. Experiments were carried out in buffer BK at 25.0 °C and pH 6.5. Panel B: Representative fluorescence titration of F-p-R10 with NS3f. Dot symbols represent experimental values and solid lines are simulated titration curves based on Eq. 3 with parameter values obtained by non-linear regression analysis: Kobs = 2.5 × 107 M−1 and ΔFmax = 1.8. Binding of NS3h to F-p-R10 was monitored by fluorescence emission from 510 nm to upon excitation at 495 nm. Titration experiments were performed in buffer BK supplemented with 100 mM KCl at 25 °C and pH 6.5.
In the range of K+ concentrations tested, the values of Kobs for NS3f were 1.4 to 1.6 times smaller than for the NS3h (Fig. 5). As with the NS3h construct, logKobs varied linearly with log[M+], with a = 4.8 ± 0.3 and b = 2.6 ± 0.3 (Eq. 9). Despite the differences in the absolute values of Kobs, these linear fit parameters are not significantly different from those obtained for the NS3h construct. According to the theory exposed in the preceding sections, this result indicates that binding of NS3f is accompanied by the release of ~5 monovalent cations from the ssRNA oligonucleotide.
Having characterized the salt effect on the interaction between NS3f and an oligonucleotide (F-p-R10), we next examined the effect of monovalent cation concentration on the binding to a polynucleotide (poly(U)). To this end, mixtures of 50 nM F-p-R10 and 400 nM NS3f were titrated with poly(U), in buffer BK (pH 6.5, 25 °C) at different concentrations of KCl, and the relative fluorescence increase ∆Frel was recorded (Fig. S2, Panel A). These results were analyzed as previously described in Material and Methods.
Values of Kobs for the binding of NS3h and NS3f to poly(U) were equal and ~2 orders of magnitude lower than for F-p-R10. Note that in the range of Kobs values obtained in these experiments, 2 pM−1, it is not possible to detect a change of 1.4–1.6 times, such as was observed for the binding of both constructs to F-p-R10.
Once again, logKobs varied linearly with log[M+] (Fig. 4), with a = 7.2 ± 0.1 and b = -2.3 ± 0.1 (Eq. 9). In accordance with the theory presented above, this result indicates that binding of NS3f is accompanied by the release of ~7 monovalent cations from the poly(U) that is practically equal to the results obtained with the helicase domain NS3h.
Effect of divalent cations concentration and type on the binding of NS3h to F-p-R10 oligonucleotide
Analogously to the experiments presented in the previous sections, we performed fluorescence titrations of the F-p-R10 oligonucleotide with the NS3h protein in buffer BK (pH 6.5, 25 °C, 6 mM basal K+) in the presence of different concentrations of divalent salts CaCl2 and MgCl2. Total K+ concentration was adjusted to 10 mM with KCl. Given that buffer BK contains 0.50 mM EDTA, the concentration of what we call ‘free divalent cation’ -not bound to EDTA- is not equal to total salt concentration and was instead computed taken into account the equilibrium of metal-EDTA complexation (See Material and Methods).
Titration curves were satisfactorily described by the equations corresponding to a single-site binding model (Eqs 2 and 3). Figure 6 shows the dependence of the logarithm of best-fitting values of Kobs on the logarithm of free divalent cation concentration, log[Mg2+] and log[Ca2+].
Effect of divalent cations on Kobs for the interaction of NS3h with F-p-R10. Experiments were performed with a series of salts differing in cation type: ( ) CaCl2 and (
) CaCl2 and ( ) MgCl2 (plotted as total [M2+]). Solid line is the representation of Eq. 9 with parameters values obtained by linear least-square fits: a = 3.0, b = 2.3 and a = 2.9, b = 2.1 for the experiments performed with CaCl2 and MgCl2 respectively. Experiments were carried out in buffer BK containing 10 mM K+ at 25.0 °C and pH 6.5.
) MgCl2 (plotted as total [M2+]). Solid line is the representation of Eq. 9 with parameters values obtained by linear least-square fits: a = 3.0, b = 2.3 and a = 2.9, b = 2.1 for the experiments performed with CaCl2 and MgCl2 respectively. Experiments were carried out in buffer BK containing 10 mM K+ at 25.0 °C and pH 6.5.
Unlike what was observed for monovalent salts NaCl, KCl and RbCl, values of Kobs were different for the two divalent cations (Fig. 6), being about 2 fold greater in the presence of Ca2+ than of Mg2+. For both salts tested, logKobs varied linearly with the logarithm of divalent cation concentration and with practically equal values of slope (−2.9 ± 0.2 for MgCl2 and −3.0 ± 0.1 for CaCl2).
As will be shown in the following section, these slopes, obtained at 10 mM K+, are not expected to be significantly different from the values that would be observed in the absence of monovalent cation. This is because in the range of divalent cation concentrations tested, monovalent cations would be fully displaced from the oligonucleotide (see Discussion). Therefore, under the same assumptions as in the interpretation given above for the effect of monovalent salts, these results indicate that ~3 divalent cations are released upon binding of NS3h to the ssRNA oligonucleotide. Importantly, the ratio between the experimentally observed slopes for divalent and monovalent cations,
satisfies the expected relation for a charged ligand-ssRNA interaction in which the concentration of anions (chloride in this case) has no appreciable effects on the association constant Kobs, i.e.,
where ψ is the same as in Eq. 10 and σ denotes the number of divalent cations bound to the nucleic acid per phosphate residue and its value is a characteristic property of the nucleic acid47,49 (see Discussion).
Combined effects of monovalent and divalent cations
A complete characterization of the effects of monovalent and divalent cations on the interaction between NS3h and ssRNA requires an experimental examination of their combined effects. This is because both cations are counterions of the nucleic acid which compete with each other and with the protein for association to the nucleic acid24,38,39,49,50, and therefore their effects on the NS3h-ssRNA binding reaction are necessarily linked.
We performed fluorescence titrations of the F-p-R10 oligonucleotide with the NS3h protein in buffer BK (pH 6.5, 25 °C) in the presence of different concentrations of K+ and Mg2+. In Fig. 7, Panel A shows the dependence of the logarithm of best-fitting values of Kobs on log[K+] obtained at different magnesium concentrations whereas Panel B shows the dependence upon log[Mg2+] at different potassium concentrations. The range of salt concentration tested was set by the technical limits in the determination of Kobs. A limitation is given by the minimum concentration of the fluorescent oligonucleotide, F-p-R10, that can be used to obtain a detectable spectroscopic signal with adequate precision. At the other extreme, at high fluorescent oligonucleotide concentrations, loss of linearity or saturation of the spectroscopic signal occur. When increasing NS3 concentration, the difficulties derived from protein aggregation and self-assembly are also present. So that the range of values of Kobs that we can determine is approximately between 2 × 106 and 8 × 107 M−1. As Mg2+ concentration is increased, the plots of logKobs as a function of log[K+] becomes increasingly non-linear. The flattening of the plots at low K+ concentrations is an expected behavior due to the fact that Mg2+ displaces K+ from the RNA oligonucleotide and does so more readily at low concentrations of its competing ligand; therefore, as more Mg2+ is added, less K+ ions are released from the RNA upon binding of the NS3h protein38,39,47,49. The observation that the curves tend to converge as K+ concentration is increased is explained on the same grounds as the result of competition between both cations.
Combined effect of monovalent and divalent cations on Kobs for the interaction of NS3h with F-p-R10. Panel A: logKobs values are shown as a function of M+. Dots symbols represent data obtained from experiments performed in the presence of ( ) non MgCl2, (
) non MgCl2, ( ) 3,(
) 3,( ) 5, (
) 5, ( ) 13.5, (
) 13.5, ( ) 17, (
) 17, ( ) 19.5 and (
) 19.5 and ( ) 25 mM MgCl2. Solid lines are representations of Eq. 14 with parameter values obtained by non-linear regression analysis: KT,H = 170 M−1 and KT,Mg = 2.1 M−1. Panel B: logKobs values are shown as a function of M2+. Dot symbols (
) 25 mM MgCl2. Solid lines are representations of Eq. 14 with parameter values obtained by non-linear regression analysis: KT,H = 170 M−1 and KT,Mg = 2.1 M−1. Panel B: logKobs values are shown as a function of M2+. Dot symbols ( ), (
), ( ), (
), ( ) represent data obtained from experiments performed with MgCl2 at 10, 25 and 130 mM KCl respectively. Red dot (
) represent data obtained from experiments performed with MgCl2 at 10, 25 and 130 mM KCl respectively. Red dot ( ) indicated experiments carried out in the presence of CaCl2 at 10 mM KCl. Solid lines are representations of Eq. 14 with best parameter values obtained by non-linear regression analysis of all the data simultaneously. Black solid line simulates the performance of the model in the absence of M+. Titration experiments were performed in buffer BK at 25.0 °C and pH 6.5.
) indicated experiments carried out in the presence of CaCl2 at 10 mM KCl. Solid lines are representations of Eq. 14 with best parameter values obtained by non-linear regression analysis of all the data simultaneously. Black solid line simulates the performance of the model in the absence of M+. Titration experiments were performed in buffer BK at 25.0 °C and pH 6.5.
In order to provide a quantitative and unified description of the simultaneous dependence of Kobs on K+ and Mg2+ concentration we interpreted the effect of Mg2+ as the result of its competitive binding to the nucleic acid which reduces the effective concentration of protein binding sites, such as was done in previous works39,47,51. In this view, Mg2+ binding to the nucleic acid is assumed to take place through a definite number of binding sites -occupying a definite number of nucleotide residues- and thus it is described by the conventional binding isotherm for the nonspecific interaction of a large ligand to a one-dimensional lattice52.
Taking into account that the F-p-R10 oligonucleotide contains 10 phosphate groups and assuming that NS3h binding involves the displacement of cations from all of them (m′ = 10 from Gebhard et al27.) and that binding of Mg2+ to the oligonucleotide involves a ~3:1 phosphate:Mg2+ stoichiometry (see Supplementary Material Text S1), the simultaneous effect of K+ and Mg2+ concentrations on Kobs can be condensed in the following expression:
where [M+] is the total concentration of monovalent cation, KT and a are the same as defined above, KT,Mg is the intrinsic binding constant for Mg2+ ions in the presence of 1 M monovalent cation and aMg denotes the number of monovalent cations released from the nucleic acid upon binding of Mg2+. Finally, coef0,I denotes the combinatorial coefficient that gives the number of possible configurations for a complex with i bound Mg2+ ions (and without bound protein) deduced by the Epstein’s formalism for the nonspecific interaction of a large ligand to a one-dimensional lattice52 (see Supplementary Material Text S1). In the present case, for a 10-mer oligonucleotide and an occluded site size of 3 nucleotide residues, these coefficients take the values 8, 15 and 4 for i = 1, 2 and 3, respectively. As a consequence of the assumptions under which it was deduced, this expression only considers the possibility of binding of Mg2+ to the free RNA, not bound to the protein, but its extension to more general cases is straightforward.
Parameters appearing in Eq. 14, KT and a were determined from the results shown in the preceding sections, whereas aMg was computed assuming that each Mg2+ ion displaces an amount of monovalent cations proceeding from 3 phosphate residues, which for the F-p-R10 oligonucleotide corresponds to 1.5 monovalent cations (see Supplementary Material Text S1). Finally, the value of KT,Mg, was estimated by simultaneous least-square fitting of Eq. 14 to all the results shown in Fig. 7.
Eq. 14 provided a good description of the experimental data (solid lines in Fig. 7) with a best-fit value of KT,Mg = 2.1 ± 0.2. This result suggests that the effect of MgCl2 concentration on the interaction between DENV NS3h and ssRNA can be attributed to the binding of the Mg2+ to the nucleic acid, without the need of invoking an interaction of Mg2+ or Cl− ions with the protein.
Discussion
The general properties and interpretation of the salt effect on the nonspecific interaction between proteins and nucleic acids were extensively studied and characterized by Record, Lohman & De Haseth24,37,39,44,45,53. These studies developed the theoretical framework and the experimental basis to apply it to any particular protein-nucleic acid system providing thermodynamic information of great interest. In the present study we implemented this analysis as a tool to expand the thermodynamic characterization of the interaction between DENV NS3h and ssRNA.
Effect of monovalent cations
Fluorescence titration experiments were performed with the fluorescein-labeled RNA oligonucleotide F-p-R10 to determine the dependence of observed binding constant Kobs on the concentration of monovalent cations K+, Na+ and Rb+ added in the form of their chloride salts. It was observed that the binding affinity of NS3h for the ssRNA oligonucleotide -quantified as Kobs- was not dependent on the nature of the added cation (Fig. 2 and Table 1). This observation suggests that preferential cation interactions with RNA do not differ significantly K+, Na+ and Rb+ as was found in previous studies on the interaction of oligolysines with ssRNA47,54.
In all cases, log Kobs decreased linearly with log[M+] with a slope (∂logKobs/∂log[M+])pH,T of −5.0 ± 0.2. As was stated in Results, we interpreted this slope as the result of the preferential interaction of cations with the nucleic acid. It should be noted, however, that in the general case the value of (∂logKobs/∂log[M+])pH,T is governed by the preferential interaction coefficients of electrolyte ions (added cation M+ and its anion Cl−) and H20 with nucleic acid, protein and the complex protein-nucleic acid (cf. Eq. 20 in Record 199138).
With respect to the preferential interaction of water molecules, the net release or uptake of water molecules upon protein-nucleic acid binding can give rise to a non-linear dependence of logKobs on log[M+], which was not observed in our results (Fig. 2). However, theory24 and experiments39 show that the range of salt concentrations used in our work could be not wide and high enough for non-linearity to be observed, mainly because the sensitivity of slope (∂logKobs/∂log[M+])pH,T with respect to log[M+] increases exponentially with log[M+] (see Eq. 20 in Record 199138). Even in such case where nonlinearity cannot be observed, differential hydration could still affect the value of (∂logKobs/∂log[M+])pH,T through an additive term, which sign will be negative or positive if water is released or taken up in the binding reaction. In this sense, employing Eq. 2 from Ha et al55. we calculated that the release or uptake of about 250 water molecules is equivalent to the release of 1 monovalent cation in its contribution to the value of (∂logKobs/∂log[M+])pH,T in the range of salt concentration between 90 mM and 140 mM as employed in our experiments. Given the values observed in previous experimental studies of hydration effects on nonspecific protein-nucleic acid interactions (uptake 1.7 ± 0.356; none57; 3 ± 5255; none58), we think it is appropriate to neglect the potential effects of preferential water interaction on the analysis of the results presented in this work.
The potential effect of preferential interactions of anions -added as counterion of the salt- would proceed from differences in the interaction with free protein and protein bound to the nucleic acid24. Under conditions in which the degree of anion binding to the protein increases with salt concentration, slope (∂logKobs/∂log[M+])pH,T is also expected to increase with salt concentration. For that reason, as in the case of preferential interactions of water, anion effects can be evidenced by loss of linearity in the plot of log Kobs as a function log[M+], which was not observed in our results. Here again, linearity does not ensure that the observed value of slope (∂logKobs/∂log[M+])pH,T is not affected by anion release. However, the ratio between slope (∂logKobs/∂log[M2+])pH,T (obtained with MgCl2 and CaCl2) and (∂logKobs/∂log[M+])pH,T satisfies the expected value for the case in which anion and hydration effects are absent (see Eq. 13 and below). This fact provides support to the interpretation of −(∂logKobs/∂log[M+])pH,T as the net number of monovalent cations released in the NS3-RNA binding reaction24,39.
With these considerations in mind, our results indicate that binding of DENV NS3h to the F-p-R10 is accompanied by the release of ~5 monovalent cations.
Testing the dependence of the ion release with the nucleic acid length
We determined that binding of NS3h to poly(U) or poly(A) was more sensitive to KCl concentration than binding to the 10-mer oligonucleotide (Fig. 4). In terms of the interpretation of (∂logKobs/∂log[M+])pH,T given throughout this work, the results indicate that one and two/three more monovalent cations are released in the interaction of NS3h with poly(A) (a = 6.0 ± 0.3) and poly(U) (a = 7.7 ± 0.5), respectively, compared to its interaction with the 10-mer oligonucleotide (a ~ 5). This is an expected result. In previous experimental and theoretical studies of binding of oligolysines to single-stranded oligonucleotides and polynucleotides, it was found that (∂logKobs/∂log[M+])pH,T increases with nucleic acid length until it reaches a maximum value. This effect of nucleic acid length was interpreted as the result of an increase of the extent of counterion binding to longer nucleic acids due to a higher negative charge density within the nucleic acid strand (away from its 3′ and 5′ ends)41,42,59.
To distinguish the properties of the nucleic acid and of the protein contained in the observed values of a (ψ and m′ in Eq. 10, respectively) we employed as a reference the value of ψ for poly(U) obtained by Mascotti and Lohman46 from their study of the binding of oligolysines of different valence to the nucleic acid. Comparison of the values of (∂log Kobs/∂log[M+])pH,T for the F-p-R10 oligonucleotide and poly(U) allowed us to determine a value of ψ for the 10-mer oligonucleotide of about 0.50 units, 32% smaller than the known value for poly(U) (0.74). This represents a similar change as that observed for the binding of oligolysines to dT oligomers of increasing lengths, were (∂logKobs/∂log[M+])pH,T changed from 3.8 for a 10-mer to 6.4 for poly(dT)45.
Results shown in Fig. 3 provided us a value of m’ = 10 ± 1 (see Eq. 10) which, following Record and co-workers’ interpretation60, represents the effective charge on the binding surface of the NS3h implicated in the interaction with RNA. The magnitude of m’ is coincident with that of the minimum and occluded binding site size of NS3h obtained in previous works12,27. Although these two interaction parameters are expected to have some kind of relationship, given that both are related to the extension of the nucleic acid binding site on the protein, there is no reason to expect them to be equal. From a structural point of view, crystallographic data19 from DENV4 NS3h bound to a 12- or 13-mer RNA oligomer shows a segment of 7 nucleotide residues, with 6 phosphate groups, in the nucleic acid binding site. These structural data indicate that there are 6 positively charged residues (4 Arg and 2 Lys) and 2 negatively charged Asp residues within a radius of 8 Å from the phosphate groups of the ssRNA. The differences between the number of ionic interactions obtain in this work and that reported from crystallographic studies is not unusual. In this sense Record38 suggested that “neutralization of nucleic acid phosphates by proteins is in general not accomplished by classical salt bridges (ion pairs)” and Harrison and Aggarwal61 concluded that “hydrogen bonds from positively charged side chains to nucleic acid phosphates appear with only modest frequency. Coulombic interactions from less strongly anchored arginines and lysines do appear to be important, however, since in each structure a number of such residues lie near the DNA backbone”.
Testing the effect of the NS3 protease domain
Previous studies addressed the effect that the protease domain of NS3 exerts on the NTPase and helicase activities11,12,13,27. Dissimilar results were obtained from such studies which did not permit to reach a general conclusion on this issue. In this work we inquired the impact of the protease domain on the equilibrium interaction of DENV NS3 with ssRNA.
Experimental results shown in Fig. 5 indicate that Kobs for the binding of NS3f to F-p-R10 are 1.4–1.6 times smaller than that for NS3h while both constructs released ~5 monovalent cations upon binding to the oligonucleotide. This coincidence could be due to the fact that the number of cations released by each construct is limited by the total number of counterions thermodynamically bound to F-p-R10. To test this possibility, we determined the salt dependence of the NS3-poly(U) association constant since the length of this nucleic acid does not limit the number of counterions that could be released upon binding of the protein. (Fig. 4). We found that both, NS3h and NS3f, released the same number of monovalent cations. Taken together, our results indicate that the appending of the protease domain does not affect the value of the parameter m’ which characterizes the interaction of NS3 with ssRNA.
Effect of divalent cations and combined effect of mono and divalent cations
Application of counterion condensation theory in the analysis of the results shown in Fig. 6 provides a means to test the validity of the assumption that preferential interactions with small anions and water molecules can be neglected. According to this theory, in the absence of anion effects the ratio between the slopes (∂logKobs/∂log[M2+])pH,T and (∂logKobs/∂log[M+])pH,T should be equal to σ/ψ, the ratio of the number of divalent and monovalent cations thermodynamically bound to the nucleic acid per phosphate residue49 (Eqs 12 and 13). This relationship with σ/ψ is strictly applicable for slopes obtained from experiments performed in the presence of only one kind of small cation, divalent or monovalent. However, the same relationship may hold if the slopes are obtained in a mixture where the concentration of one of the cations -the one kept constant- is sufficiently small (See Supplementary material, Text S2). This was the condition under which we performed the experiments to determine the value of (∂logKobs/∂log[M2+])pH,T (where M2+ was Mg2+ or Ca2+), where a basal amount of K+ (10 mM) was present along with the divalent salts tested (Fig. 7). In this sense, fitting of an equilibrium thermodynamic model that takes into account the simultaneous effect of K+ and Mg2+ on Kobs, to the set of experimental results obtained in the presence of both cations (Fig. 7) allowed us to evaluate to what extent the limiting value of (∂logKobs/∂log[M2+])pH,T would differ from the value obtained in the presence of 10 mM K+. According to our analysis (see Supplementary Information, Text S2) the value of (∂logKobs/∂log [M2+])pH,T at 10 mM K+ obtained in the range of Mg2+ and Ca2+ employed would be no more than 3% smaller than its limit value, a difference even lower than the experimental uncertainty.
Therefore, we can interpret the coincidence between the experimental value of the ratio (∂logKobs/∂log[M2+])pH,T / (∂logKobs/∂log[M+])pH,T (~0.58) and the expected value of σ/ψ (~0.60)49 as evidence that, if there are preferential interactions of anions and water molecules, they have a negligible effect on the dependence of Kobs on salt concentration.
As was mentioned above, we studied the combined effect of mono and divalent cations on Kobs. A simple model fitted to the results provided a means to predict the value of Kobs in any concentration of mono and divalent cations (see the model in Supplementary Information, Text S1). Importantly, the model allowed us to explain the effect of both cations solely as the results of the preferential interaction of the cations with the RNA, without the need to invoke interactions of cations and/or anions with the protein. Additionally, this working model indicates that the difference in Kobs values observed in the experiments performed in the presence of Ca2+ or Mg2+ (Fig. 6) can be explained by a difference in the binding affinity of these cations for RNA phosphate groups (See Supplementary Information, Text S2).
References
Gubler, D. J. Epidemic dengue/dengue hemorrhagic fever as a public health, social and economic problem in the 21st century. Trends Microbiol 10, 100–103 (2002).
Bhatt, S. et al. The global distribution and burden of dengue. Nature 496, 504–507 (2013).
WHO. Dengue: Guidelines for diagnosis, treatment, prevention, and control. (World Health Organization, Geneva, 2009).
Weaver, S. C. & Vasilakis, N. Molecular evolution of dengue viruses: contributions of phylogenetics to understanding the history and epidemiology of the preeminent arboviral disease. Infect Genet Evol 9, 523–40 (2009).
Chambers, T. J., Hahn, C. S., Galler, R. & Rice, C. M. Flavivirus genome organization, expression, and replication. Annu Rev Microbiol 44, 649–688 (1990).
Perera, R. & Kuhn, R. J. Structural proteomics of dengue virus. Curr Opin Microbiol 11, 369–377 (2008).
Bollati, M. et al. Structure and functionality in flavivirus NS-proteins: Perspectives for drug design. Antiviral Res 87, 125–148 (2010).
Cui, T. et al. Recombinant dengue virus type 1 NS3 protein exhibits specific viral RNA binding and NTPase activity regulated by the NS5 protein. Virology 246, 409–417 (1998).
Li, H., Clum, S., You, S., Ebner, K. E. & Padmanabhan, R. The serine protease and RNA-stimulated nucleoside triphosphatase and RNA helicase functional domains of dengue virus type 2 NS3 converge within a region of 20 amino acids. J. Virol. 73, 3108–3116 (1999).
Bartelma, G. & Padmanabhan, R. Expression, purification, and characterization of the RNA 5’-triphosphatase activity of dengue virus type 2 nonstructural protein 3. Virology 299, 122–132 (2002).
Gebhard, L. G., Kaufman, S. B. & Gamarnik, A. V. Novel ATP-independent RNA annealing activity of the dengue virus NS3 helicase. PLoS ONE 7(4), e36244, https://doi.org/10.1371/journal.pone.0036244 (2012).
Incicco, J. J., Gebhard, L. G., González-Lebrero, R. M., Gamarnik, A. V. & Kaufman, S. B. Steady-state NTPase activity of dengue virus NS3: Number of catalytic sites, nucleotide specificity and activation by ssRNA. PLoS One 8(3), e58508, https://doi.org/10.1371/journal.pone.0058508 (2013).
Xu, T. et al. Structure of the dengue virus helicase/nucleoside triphosphatase catalytic domain at a resolution of 2.4 Å. J Virol 79, 10278–88 (2005).
Jankowsky, E. & Fairman, M. E. RNA helicases - one fold for many functions. Curr Opin Struct Biol 17, 316–324 (2007).
Gorbalenya, A. E., Koonin, E. V., Donchenko, A. P. & Blinov, V. M. Two related superfamilies of putative helicases involved in replication, recombination, repair and expression of DNA and RNA genomes. Nucleic Acids Res 17, 4713–4730 (1989).
Cordin, O., Banroques, J., Tanner, N. & Linder, P. The DEAD-box protein family of RNA helicases. Gene 367, 17–37 (2006).
Wolfe, A. R. & Meehan, T. Use of binding site neighbor-effect parameters to evaluate the interactions between adjacent ligands on a linear lattice: Effects on ligand-lattice association. J Mol Biol 223, 1063–1087 (1992).
Liao, J. C. Mechanical transduction mechanisms of RecA-like molecular motors. J Biomol Struct Dyn 29, 497–507 (2011).
Luo, D. et al. Insights into RNA unwinding and ATP hydrolysis by the flavivirus NS3 protein. EMBO J 27, 3209–3219 (2008).
Tanner, N. K. & Linder, P. DExD/H box RNA helicases: from generic motors to specific dissociation functions. Mol Cell 8, 251–262 (2001).
von Hippel, P. H. F. “simple” DNA-protein interactions to the macromolecular machines of gene expression. Annu Rev Biophys Biomol Struct 36, 79–105 (2007).
Jankowsky, E. & Harris, M. E. Specificity and nonspecificity in RNA-protein interactions. Nat Rev Mol Cell Biol 16, 533–544 (2015).
Anderson, C. F. & Record, M. T. Jr. Salt-nucleic acid interactions. Annu Rev Phys Chem 46, 657–700 (1995).
Record, M. T. Jr., Anderson, C. F. & Lohman, T. M. Thermodynamic analysis of ion effects on the binding and conformational equilibria of proteins and nucleic acids: the roles of ion association or release, screening, and ion effects on water activity. Q Rev Biophys 11, 103–178 (1978).
Lohman, T. M., De Haseth, P. L. & Record, M. T. Jr. Analysis of ion concentration effects on the kinetics of protein-nucleic acid interactions: Application to lac repressor-operator interactions. Biophys Chem 8, 281–294 (1978).
Overman, L. B., Bujalowski, W. & Lohman, T. M. Equilibrium binding of Escherichia coli single-strand binding protein to single-stranded nucleic acids in the (SSB) 65 binding mode. Cation and anion effects and polynucleotide specificity. Biochemistry 27, 456–471 (1988).
Gebhard, L. G. et al. Monomeric nature of dengue virus NS3 helicase and thermodynamic analysis of the interaction with single-stranded RNA. Nucleic Acids Res 42, 11668–11686 (2014).
Kolthoff, I. M., Sandell, E. B., Meehan, E. J. & Bruckenstein, S. Quantitative Chemical Analysis (ed. Harris, D.) (Macmillan, 1969).
Kinney, R. M. et al. Construction of infectious cDNA clones for dengue 2 virus: strain 16681 and its attenuated vaccine derivative, strain PDK-53. Virology 230, 300–308 (1997).
Pace, C. N., Vajdos, F., Fee, L., Grimsley, G. & Gray, T. How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4, 2411–2423 (1995).
Cavaluzzi, M. J., Borer, P. N. & Revised, U. V. extinction coefficients for nucleoside-5’-monophosphates and unpaired DNA and RNA. Nucleic Acids Res 32(1), e13, https://doi.org/10.1093/nar/gnh015 (2004).
Kowalczykowski, S. C., Lonberg, N., Newport, J. W. & Von Hippel, P. H. Interactions of bacteriophage T4-coded gene 32 protein with nucleic acids: I. Characterization of the binding interactions. J Mol Biol 145, 75–104 (1981).
Lohman, T. M. & Mascotti, D. P. Nonspecific ligand-DNA equilibrium binding parameters determined by fluorescence methods. Methods Enzymol 212, 424–458 (1992).
Jezewska, M. J. & Bujalowski, W. A general method of analysis of ligand binding to competing macromolecules using the spectroscopic signal originating from a reference macromolecule. Application to Escherichia coli replicative helicase DnaB protein-nucleic acid interactions. Biochemistry 35, 2117–2128 (1996).
Kolthoff, I. M. et al. Análisis Químico Cuantitativo (ed. Harris, D.) (Macmillan, 1979).
McGhee, J. D. & von Hippel, P. H. Theoretical aspects of DNA-protein interactions: Co-operative and non-co-operative binding of large ligands to a one-dimensional homogeneous lattice. J Mol Biol 86, 469–489 (1974).
Record, M. T. Jr., Lohman, T. M. & De Haseth, P. Ion effects on ligand-nucleic acid interactions. J Mol Biol 107, 145–158 (1976).
Record, M. T. Jr., Ha, J. H. & Fisher, M. A. Analysis of equilibrium and kinetic measurements to determine thermodynamic origins of stability and specificity and mechanism of formation of site-specific complexes between proteins and helical DNA. Methods Enzymol 208, 291–343 (1991).
Record, M. T. Jr., De Haseth, P. L. & Lohman, T. M. Interpretation of monovalent and divalent cation effects on the lac repressor-operator interaction. Biochemistry 16, 4791–4796 (1977).
Zhang, W., Bond, J. P., Anderson, C. F., Lohman, T. M. & Record, M. T. Jr. Large electrostatic differences in the binding thermodynamics of a cationic peptide to oligomeric and polymeric DNA. PNAS 93, 2511–2516 (1996).
Zhang, W. et al. The importance of Coulombic end effects: experimental characterization of the effects of oligonucleotide flanking charges on the strength and salt dependence of oligocation (L8+) binding to single-stranded DNA oligomers. Biophys J 76, 1008–1017 (1999).
Ballin, J. D., Shkel, I. A. & Record, M. T. Jr. Interactions of the KWK6 cationic peptide with short nucleic acid oligomers: demonstration of large Coulombic end effects on binding at 0.1-0.2 M salt. Nucleic Acids Res 32, 3271–3281 (2004).
Bujalowski, W., Lohman, T. M. & Anderson, C. F. On the cooperative binding of large ligands to a one-dimensional homogeneous lattice: The generalized three-state lattice model. Biopolymers 28, 1637–1643 (1989).
Manning, G. S. Limiting laws and counterion condensation in polyelectrolyte solutions I. Colligative properties. J Chem Phys 51, 924–933 (1969).
Manning, G. S. The molecular theory of polyelectrolyte solutions with applications to the electrostatic properties of polynucleotides. Q Rev Biophys 11, 179–246 (1978).
Mascotti, D. P. & Lohman, T. M. Thermodynamic extent of counterion release upon binding oligolysines to single-stranded nucleic acids. PNAS 87, 3142–3146 (1990).
Mascotti, D. P. & Lohman, T. M. Thermodynamics of single-stranded RNA binding to oligolysines containing tryptophan. Biochemistry 31, 8932–8946 (1992).
Mascotti, D. P. & Lohman, T. M. Thermodynamics of single-stranded RNA and DNA interactions with oligolysines containing tryptophan. Effects of base composition. Biochemistry 32, 10568–10579 (1993).
Lohman, T. M. & Mascotti, D. P. Thermodynamics of ligand-nucleic acid interactions. Methods Enzymol 212, 400–424 (1992).
De Haseth, P. L., Lohman, T. M. & Record, M. T. Jr. Nonspecific interaction of lac repressor with DNA: an association reaction driven by counterion release. Biochemistry 16, 4783–4790 (1977).
Lohman, T. M., De Haseth, P. L. & Record, M. T. Jr. Pentalysine-deoxyribonucleic acid interactions: a model for the general effects of ion concentrations on the interactions of proteins with nucleic acids. Biochemistry 19, 3522–3530 (1980).
Epstein, I. R. Cooperative and non-cooperative binding of large ligands to a finite one-dimensional lattice. A model for ligand oligonucleotide interaction. Biophys Chem 8, 327–339 (1978).
Lohman, T. M. Kinetics of protein-nucleic acid interactions: use of salt effects to probe mechanisms of interaction. CRC Crit Rev Biochem 19, 191–245 (1986).
Mascotti, D. P. & Lohman, T. M. Thermodynamics of oligoarginines binding to RNA and DNA. Biochemistry 36, 7272–7279 (1997).
Ha, J. H., Capp, M. W., Hohenwalter, M. D., Baskerville, M. & Record, M. T. Jr. Thermodynamic stoichiometries of participation of water, cations and anions in specific and non-specific binding of lac repressor to DNA. J Mol Biol 228, 252–264 (1992).
Galletto, R., Jezewska, M. J. & Bujalowski, W. Multistep sequential mechanism of Escherichia coli helicase PriA protein-ssDNA interactions. Kinetics and energetics of the active ssDNA-searching site of the enzyme. Biochemistry 43, 11002–11016 (2004).
Garner, M. & Rau, D. Water release associated with specific binding of gal repressor. EMBO J 14, 1257–1263 (1995).
Sidorova, N. Y. & Rau, D. C. Differences in water release for the binding of EcoRI to specific and nonspecific DNA sequences. PNAS 93, 12272–12277 (1996).
Olmsted, M. C., Anderson, C. F. & Record, M. T. Jr. Monte Carlo description of oligoelectrolyte properties of DNA oligomers: range of the end effect and the approach of molecular and thermodynamic properties to the polyelectrolyte limits. PNAS 86, 7766–7770 (1989).
Shkel, I. A., Ballin, J. D. & Record, M. T. Jr. Interactions of cationic ligands and proteins with small nucleic acids: analytic treatment of the large coulombic end effect on binding free energy as a function of salt concentration. Biochemistry 45, 8411–8426 (2006).
Harrison, S. C. & Aggarwal, A. K. DNA recognition by proteins with the helix-turn-helix motif. Annu Rev Biochem 59, 933–969 (1990).
Acknowledgements
This work was supported by the Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, Argentina) [PICT-2015-2555 to A.V.G. and S.B.K., PICT-2016-2685 to S.B.K.]; R.M.G.L., E.A.R., L.G.G. and A.V.G. are members of the Concejo Nacional de Investigaciones Científicas y Técnicas (CONICET, Argentina). L.A.C. and J.J.I. were funded by doctoral and postdoctoral fellowships from CONICET, respectively. Funding for open access charge: Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, Argentina).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: L.A.C., J.J.I., A.V.G. and S.B.K. Performed the experiments: L.A.C., J.J.I., L.G.G. and E.A.R. Analyzed the data: L.A.C., J.J.I., R.M.G.L., E.A.R., A.V.G. and S.B.K. Wrote the paper: L.A.C., J.J.I., L.G.G., A.V.G. and S.B.K. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cababie, L.A., Incicco, J.J., González-Lebrero, R.M. et al. Thermodynamic study of the effect of ions on the interaction between dengue virus NS3 helicase and single stranded RNA. Sci Rep 9, 10569 (2019). https://doi.org/10.1038/s41598-019-46741-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46741-4
This article is cited by
-
Thermodynamic and mechanistic analysis of the functional properties of dengue virus NS3 helicase
Biophysical Reviews (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.