Abstract
The Poaceae constitute a taxon of flowering plants (grasses) that cover almost all Earth’s inhabitable range and comprises some of the genera most commonly used for human and animal nutrition. Many of these crops have been sequenced, like rice, Brachypodium, maize and, more recently, wheat. Some important members are still considered orphan crops, lacking a sequenced genome, but having important traits that make them attractive for sequencing. Among these traits is apomixis, clonal reproduction by seeds, present in some members of the Poaceae like Eragrostis curvula. A de novo, high-quality genome assembly and annotation for E. curvula have been obtained by sequencing 602 Mb of a diploid genotype using a strategy that combined long-read length sequencing with chromosome conformation capture. The scaffold N50 for this assembly was 43.41 Mb and the annotation yielded 56,469 genes. The availability of this genome assembly has allowed us to identify regions associated with forage quality and to develop strategies to sequence and assemble the complex tetraploid genotypes which harbor the apomixis control region(s). Understanding and subsequently manipulating the genetic drivers underlying apomixis could revolutionize agriculture.
Similar content being viewed by others
Introduction
Climate change modeling predicts sustained elevated temperatures in which C4 grasses will thrive1. E. curvula (Schrad.) Nees (weeping lovegrass) is a C4 perennial grass member of the Poaceae family, Chloridoideae subfamily. The E. curvula complex has a basic chromosome number of X = 10 and includes cytotypes with different ploidy levels (from 2X to 8X) that may undergo sexual reproduction and facultative or obligate apomixis2. Its drought tolerance and capacity to grow in sandy soils make it highly valued, especially for cattle feed in semiarid regions3. However, weeping lovegrass, like other C4 species, has lower nutritional quality compared to C3 species. Different molecular strategies have been developed in order to increase forage quality. Recently4, the genes for class I and class II caffeoyl shikimate esterase (CSE) have been discovered to be involved in lignin regulation being interesting targets to improve forage quality through genetic engineering5. In addition, E. curvula has been suggested as a potential biofuel crop6. In this context, the availability of a high-quality genome assembly for weeping lovegrass is essential to enable genetic improvement that aims to increase its digestibility and energy provision. Moreover, since E. curvula is a species adapted to high temperature, high radiation and drought, the characterization of the WRKY transcription factors could be central to understand the mechanism involved in resilience in case of environmental stresses.
Eragrostis is a poorly studied polyphyletic genus, with more than 400 species7, originating from Africa and now distributed in tropical and mid warm-season regions all over the world. Eragrostis tef, a cereal from Ethiopia, and E. curvula, a forage grass from the south of Africa, are the best-studied species of the genus. Setaria italica and Sorghum bicolor genomes were reported to be the closest relatives of E. tef8. However, comparisons of the E. curvula transcriptome with the E. tef genome and transcriptome sequences9 support the involvement of E. curvula in E. tef evolution. Like E. tef10, E. curvula is classified as an orphan, or underutilized, crop and despite its importance, very little research emphasis has been given to this species.
Until recently, the major limitation in any genome assembly project was given by the short length of the reads obtained11. The advent of new platforms for long molecule sequencing, such as the PacBio Sequel System and Oxford Nanopore Technologies systems, has greatly contributed in overcoming this limitation12. The former is ideal for sequencing large genomes and provides high-quality long reads that allow genome reconstruction with an accuracy of more than 99.99%13. High-quality assemblies based on PacBio sequences were recently published for Arabidopsis thaliana (135 Mb)13, Oropetium thomaeum (245 Mb)14, Utricularia gibba (82 Mb)15, Chenopodium quinoa (1500 Mb)16, Zea mays (2300 Mb)17 and Helianthus annuus (3000 Mb)18. However, to build chromosome-level genome assemblies, spanning long genomic distances and to order the contigs in the right orientation it is necessary to complement the PacBio system with other technologies. Chicago® and Dovetail™ Hi-C are two recently developed methodologies based on proximity DNA and chromatin ligation that complement the PacBio system and result in an extremely precise sequence assembly, orientating the contigs and increasing the N50 to values that can be as large as 30 Mb19. A good example of this integration of technologies is shown by the genome assembly of the tropical fruit, the durian, Durio zibethinus, where the N50 was 22.7 Mb20.
The availability of a high-quality diploid genome assembly of E. curvula will contribute to establishing its evolutionary relationships with other members of the Poaceae family, unraveling the taxonomy of the E. curvula complex, looking for new strategies to improve forage quality and directing the assembly of more complex and heterozygous tetraploid genomes harboring the apomixis control region(s). The elucidation of this region(s) and the associated apomixis genes could lead to revolutionary developments in terms of crop improvement.
To obtain a high-quality genome assembly of the diploid E. curvula genotype a combination of PacBio long read sequencing with Chicago and Hi-C technology was employed. The final genome assembly had 603 Mb distributed in 1,143 scaffolds, an N50 of 43.4 Mb with 28% of the bases corresponding to repetitive elements and was validated using DArT-seq and Simple sequence repeats (SSR) markers.
Results
Sequencing and Assembly
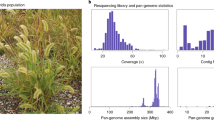
The PacBio Sequel platform was employed to sequence two E. curvula libraries of 10 and 20 kb resulting in 6,223,627 and 3,309,811 raw reads respectively. The average read length size was 5,296 for the 10 kb and 7,018 for the 20 kb libraries, covering 90.32X of the estimated genome size. The best assembly was obtained using the FALCON software with the following settings: i) a length cut-off of 7,500 bp, ii) overlap filtering parameters of minimum and maximum coverage of 5 and 120X, respectively and iii) a maximum difference of coverage of 120X between the 5′ and 3′ ends. The de novo assembly consisted of 3,118 contigs with an N50 of 378,697 bp, representing 97% of the genome length (Table 1). This assembly was later polished using the software Arrow, improving the accuracy per base content and increasing the percentage of complete BUSCO genes from 88.4% to 96% (Table 1). In this assembly 98.88% of the bases were found with a quality score more than 30, this means that 98.88% of the polished genome has an error rate less than 0.001.
The next step was to sequence the Chicago library, achieving 432 million 2 × 150 paired-end raw reads. The combination of these reads with the primary contigs of the FALCON assembly using the Hi-Rise assembler increased the N50 from 0.378 Mb to 0.791 Mb, decreasing the number of sequences to 1,884 (Table 1). Although this strategy vastly improved the contiguity of the assembly, another improvement involved the preparation of a Hi-C library, in which 333 million 2 × 150 bp paired-end raw reads were achieved. Using the Hi-C library as input within the Chicago assembly the N50 increased to 43.41 Mb and the number of scaffolds decreased to 1,143. The final BUSCO results were 96.4%, 80.3% of them were single copy while 16.1% were duplicated.
The pipeline used to sequence and assembly the genome, and the experimental steps in its analysis, are shown in Supplementary Fig. S1.
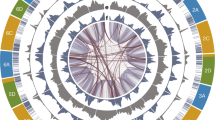
The E. curvula genome assembly presented here is one of the few published genomes (NCBI Bioproject PRJNA508722) with a scaffold N50 value greater than 10 Mb, having seven scaffolds with almost the size of complete chromosomes. A high level of contiguity was obtained, since 83.7% of the genome size was contained in the first 14 scaffolds (Fig. 1).
DArT-seq (Diversity Arrays Technology sequenciation) Analyses and Marker Mapping
DArT is a recently developed technology for SNP markers discovery based on the reduction of genome complexity. A total of 6,027 (95.5%) of the original 6,307 SNPs markers sequences present in cv. Victoria were successfully aligned to the genome assembly. The longest scaffolds present a proportionally higher number of markers than the shortest ones and the same tendency was appreciated in the gene number (Supplementary Table S1, Fig. 2).
Regression between number of gene models (light blue circles) and number of DArT markers (green circles) and the E. curvula scaffold length. Each circle represents a scaffold. The regression analysis shows that gene and DArT marker counts are directly proportional to the scaffold size, meaning that the largest scaffold, the higher the gene and DArT count.
DArT reads density in the chromosome-scale scaffolds was used to validate the assembly. DArT libraries are designed to avoid repetitive elements and target active regions covered predominantly by genes. A higher number of genes is expected in the distal arms regions than in the central regions. In fact, the seven longest scaffolds showed higher read density at the 3′ and 5′ ends than in the central region (Supplementary Fig. S2). Thus, contigs with the longest length, the greatest number of genes and DArT markers could be complete chromosomes.
Repetitive Elements and Gene Annotation
The analyses over the E. curvula genome assembly established that 28.7% is composed by repetitive elements, mainly Long Terminal Repeats retroelements (LTR-RT) (16.97%), followed by DNA and unclassified elements (Fig. 3 and Supplementary Table S2), as seen in most of the grasses21. Within the most representative LTR-RTs were the Gypsy and Copia superfamilies, accounting for 13.62% and 3.14% of the total, respectively, with the ratio between them of 4.3:1. This value is very close to the one found in E. tef (4.27:1)22 and higher than the corresponding one in S. italica (3.08:1)23, Z. mays (1.91:1)17 and S. bicolor (3.67:1)24 (Supplementary Table S3). Previously reported ESTs related to repetitive elements present in E. curvula´s floral and leaf libraries25 were mapped onto the genome assembly. Using this scheme a LTR structure of a Gypsy superfamily of retrotransposons identified from the EST EH191456 was validated, since it was present in at least 16 scaffolds (Supplementary Table S4).
Gene annotation was performed using an ab initio prediction algorithm combined with data from ESTs and RNA-seq databases from different tissues of E. curvula and from proteins of related species. After three iterations of the MAKER software, 56,469 gene models were obtained with an average size of 1,424 bp and 93.4% of the complete BUSCO genes (Supplementary Fig. S3 and Supplementary Table S5). These genes were classified into two main categories: High Confidence (HC) and Low Confidence (LC) genes, then divided into two and three subcategories, respectively. Using this strategy 13,376 HC genes and 20,330 LC1 genes were identified, representing approximately the number of genes expected for the species (Fig. 4). The protein domains were inferred using the InterProScan software (Supplementary Table S6), finding 35,713 matches in the Pfam domain database. Gene ontology analysis based on 56,469 genes classified 33,601 genes into biological processes, 17,710 into cellular components and 33,820 into molecular function, finding 29,462 genes with at least one GO annotation category (Supplementary Fig. S4).
Eragrostis curvula Genome Evolution Among the Poaceae Family
The duplication events of the Victoria cv. genome over the time has been evaluated through a paralogous genes analysis (Supplementary File 1). The Ks (synonymous substitutions rate) peaks show two paleopoliploidy events across the time, the ancestral paleoduplication event shared by all the grasses estimated that have occurred 80–90 Mya (Millions the years ago) and a recent (4–5 Mya) duplication before the in vitro culture diploidization of Victoria cv. (Fig. 5). Since the ancestral whole genome duplication a high contraction rate over the time was observed. In the BEP (Bambusoideae, Ehrhartoideae, and Pooideae) divergence from the PACCAD (Panicoideae, Arundinoideae, Chloridoideae, Centothecoideae, Aristidoideae, Danthonioideae) a 2.44 contraction/expansion rate was observed (Fig. 6). After that, the corresponding rate in the Panicoideae-Chloridoideae divergence was 6.2. Then, in the divergence between Eragrostis and Oropetium the rate increased up to 16.88. The contraction rate for O. thomaeum genome assembly was 7.14 being the expansion higher than the contraction in all the evolution history after the grasses whole genome duplication (gWGD). More recently, during the Eragrostis speciation the rate was stabilized in 1.04 and after the E. tef allotetraploidization (9–10 Mya) the rate increased again to 2.97. Correspondingly, E. curvula present an allotetraploidization event 4–5 Mya. However, it is not possible to calculate the gene loss after tetraplodization because one copy of the genome was lost during the in vitro diploidization event. The high proportion of expansions in E. curvula could be related to the low confidence genes models.
(a) Distribution of the estimated synonymous substitutions rate (Ks) between E. curvula and the selected Poaceae orthologous genes. Peaks represent the divergence between E. curvula and the selected species. (b) Distribution of the synonymous substitution rate within each selected genome paralogous genes. The whole genome duplication shared by all the grasses (gWGD) was estimated to occur 90 Mya. The peaks represent the tetraploidization events of E. tef (tWGD) E. curvula (cWGD) and Z. mays (mWGD).
Maximum likelihood phylogenetic tree of selected Poaceae species based on 1,185 orthogroups. The time scale above the tree indicates the evolution time in millions of years ago (Mya). The numbers in red in the nodes show the number of contracted gene families while the numbers in green refer to the number of expansions. The ancient WGD (red star) was calculated to happen around ~90 Mya. The Z. mays, E. tef and E. curvula tetraploidization events (blue stars) were placed at 24.1, 9.6 and 4.5 Mya, respectively. The yellow star represents the hexaploidization of T. aestivum. C4 grasses species are included in the red box and C3 in blue box.
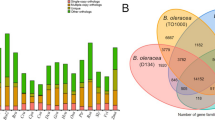
The gene models from selected monocots species, such as E. tef (A and B genome), Triticum aestivum (A, B and D genome), Oryza sativa, Z. mays, S. bicolor, S. italica, Panicum hallii, O. thomaeum, B. distachyon and Musa itinerans were grouped in orthogroups with the E. curvula genes models. The total number of defined orthogroups was 24,747, with 9,189 groups shared by the eleven species (Supplementary Fig. S5). To assess the evolutionary relationships within the subfamily we grouped them into Chloridoideae, Panicoideae, Pooideae and Ehrhartoideae. The number of gene families shared by the Chloridoideae subfamily was 12,940, while the number of genes families that are in common within the Panicoideae and Pooideae subfamilies were 13,621 and 15,532, respectively (Supplementary Fig. S6). Among the C4 grasses 11,570 common orthogroups were identified between both subfamilies, whereas 2,051 and 1,370 were found to be specific to the Panicoideae and the Chloridoideae subfamily, respectively. When an analogous analysis was performed with the C3 species, 14,203 shared orthogroups were identified, whereas 1,329 and 2,601 orthogroups were unique to the Pooideae and the Ehrhartoideae subfamily, respectively.
When the assembled genomes sharing orthogroups were analyzed, we found that the second most abundant group of species sharing orthogroups was constituted by a combination of two species (3,991 orthogroups). The combination of E. tef and E. curvula contributed to this group with 616 orthogroups, representing 15.5% of the total (Supplementary Fig. S5). This means that 616 orthogroups were shared by these two species exclusively being E. tef the most closely related species to E. curvula. This was confirmed by the construction of a phylogenetic tree with the eleven species, in which the divergence between E. curvula and E. tef was calculated as occurring 14.5 Mya and, as was expected, E. curvula was located close to the other C4 grasses (Fig. 6).
Using SyMAP to plot the syntenic regions between E. curvula and the other monocots species it was found that 79% of the E. curvula genome assembly length is covered by the Z. mays and S. bicolor syntenic blocks (Fig. 7, Supplementary Table S7). These analyses also revealed the presence of 182 reverse blocks between E. curvula and Z. mays and 149 between E. curvula and S. bicolor, representing genome rearrangements that occurred during the evolution of these grasses. Despite the divergence, 98% of the O. sativa assembly was covered by E. curvula scaffolds, sharing 262 syntenic blocks. Interestingly, O. sativa chromosome 3 is fully covered in the same orientation by the E. curvula Contig 3 (Fig. 7), indicating the close conservation of this chromosome among these grasses. The syntenic analysis over the O. thomaeum genome assembly revealed that 96% of the genome was covered by the E. curvula scaffolds (Supplementary Table S7). However, different patterns were observed between the E. curvula scaffolds and the O. thomaeum chromosomes (Fig. 8). For example, E. curvula Contig 3, that completely covers O. sativa chromosome 3, also covers the entire O. thomaeum chromosome 4, whereas O. thomaeum chromosome 3 is completely covered by E. curvula Contigs 25 and 8, suggesting that these two E. curvula scaffolds constitute one single chromosome. Other O. thomaeum chromosomes, such as 1, 2, 4, 5, 7, 8 and 10, are fully covered by Contigs 10, 38, 28, 6, 12 and 1, respectively, even when several rearrangements were detected.
Synteny between E. curvula and: (a) Zea mays; (b) S. bicolor; (c) O. thomaeum and (d) O. sativa genome assemblies. In E. curvula the fourteen longest scaffolds are represented. Seventy nine percent of the E. curvula genome was covered by the Z. mays and S. bicolor, 85% by O. thomaeum and 84% by O. sativa genomes, respectively.
Syntenic dotplot between E. curvula scaffolds and O. thomaeum (a) and E. curvula scaffolds and E. tef (b) genomes. The black dots indicate syntenic genes between the species. Blue shapes represent syntenic blocks between the species. Contigs 10, 12, 38, 6 and 28 constitute complete chromosomes in both species but the number of chromosome rearrangements is lower between E. curvula and E. tef.
The latest E. tef genome assembly version26 ratifies the chromosome scale of the Contigs 10, 12, 38, 6, 3, 1, 28 and provides evidences that the pairs of Contigs 8–25, 50–4, and 30–44 correspond to the E. tef chromosomes 3, 8 and 7 respectively, thus, assuming that there are not changes in the chromosome structure the ten E. curvula chromosomes seem to be present in the assembly with a high level of contiguity.
Genetic Relationships Among E. curvula Genotypes Assessed by SSR Analyses
SSR specific primers previously designed from transcriptomic sequences9 were mapped onto the E. curvula genome assembly. Twenty eight out of the 35 reported E. curvula SSRs primers (Supplementary Table S8) gave 100% identity and 100% of coverage with the E. curvula genome assembly. Additional SSR primers were designed based on the cv. Victoria assembly. Regarding the newly designed primers over the Victoria genome 14 out of 15 amplified with the expected amplicons size (Supplementary Fig. S7).
A phylogenetic tree was constructed using the Jaccard distance matrix calculated from the SSR markers (Fig. 9). This tree grouped the heptaploid cultivars Don Luis and Don Pablo together, showing the similarity between each other and the divergence from the other cultivars. Cultivar Victoria was located close to the diploid PI299920 and in the same branch as the tetraploid cultivars Tanganyika INTA and Tanganyika USDA. Tanganyika INTA is an apomictic tetraploid cultivar that gave rise to cv. Victoria through chromosome reduction by in vitro inflorescence culture27. Due to its similarity with cv. Victoria, it is a candidate for sequencing to identify the genomic region controlling apomixis.
Phylogenetic tree of different cultivars of E. curvula constructed using the Jaccard distance matrix calculated from SSR markers. Cultivar Victoria was placed next to the diploid cultivar PI299920 and the tetraploid Tanganyika INTA. The diploidization of this cultivar through in vitro inflorescence culture gave rise to the Victoria cv.
Lignin Pathway
Several reports have established that digestibility in forage grasses can be improved through downregulation of genes involved in the lignin pathway by genetic engineering28,29,30. Therefore, sequence information of lignin biosynthetic genes and their controlling elements is crucial to manipulate their expression. Using the KEGG31 database 16 gene models (Supplementary Fig. S8) were found to be involved in the E. curvula lignin biosynthesis pathway, from the first gene, corresponding to phenylalanine ammonia-lyase, to the final products, guaiacyl (G), p-hydroxyphenyl (H) and syringyl lignin (S) monolignols.
From this pathway we focused on the enzyme caffeoyl shikimate esterase (CSE) because of its recently identified role in lignin biosynthesis32,33. This enzyme affects the production of caffeoyl-coenzyme A 3-O-methyltransferase, previously studied by our group in E. curvula34 which could be used together with CSE to improve forage quality. Genes encoding class II enzymes are widespread in the plant kingdom, while genes encoding class I CSE enzymes are not present in all species. Aligning class I CSE from O. sativa and class II from S. bicolor against the E. curvula genome assembly we could identify both gene CSE classes. CSE genes were amplified from E. curvula using specifically designed PCR primers (Supplementary Table S9), and the resulting amplicons were cloned and sequenced, obtaining a perfect match with the sequences from the genome assembly. Moreover, our previous transcriptomic analysis9 showed the presence of mRNA from both genes, indicating that they are actively transcribed in this species.
The sequences of E. curvula class I and II genes were aligned to the corresponding sequences from other members of the Poaceae family (Supplementary Table S10). The two classes were found in O. thomaeum, P. hallii and O. sativa and only class II was present in Z. mays, S. bicolor, S. italic, B. distachyon and T. aestivum. We found single equal-sized amplicons in E. tef and E. curvula for class I CSE (Supplementary Fig. S9). Cloning these amplicons confirmed the existence of CSE class I in both species (Supplementary Fig. S10). Looking into the evolution of the grasses (Fig. 6) it is possible to deduce that the loss of the CSE class I occurred at different times during the divergence of these species, since the enzyme is absent from the Pooideae subfamily, present in the Chloridoideae and appears only in some members of the Panicoideae.
WRKY Transcription Factor Family
The WRKY transcription factor family is one of the most studied gene families associated with biotic and abiotic stresses in plants. Since E. curvula is a species adapted to high temperature, high radiation and drought stress, classification of its WRKY transcription factors is central to understand the mechanism(s) involved in its tolerance to these conditions.
From the Pfam annotated gene models, 74 genes with WRKY and zinc finger motifs were found (Supplementary Fig. S11). Seven out of the 74 were classified as group I, 32 as group II and 35 as group III. Group II was divided into five subclasses according to the remaining sequence motifs and the phylogenetic distribution (Fig. 10). The EcWRKY family has 39 of its members clustered into 13 genomic regions of less than 100 kb while other members are isolated (Supplementary Table S11). This spatial distribution agrees with previously reported data for other grass species35,36,37.
Discussion
Here we present the first high-quality genome assembly of a diploid genotype of E. curvula. This diploid assembly is a starting point for the genome assembly of the most complex polyploids of the same genus, which harbor the region(s) that controls diplosporous apomixis, and may allow us to assess the complex relationship between apomixis and ploidy.
The final FALCON assembly, after polishing, rendered an N50 of 0.380 Mb, 3,118 contigs and 96% of complete BUSCO genes. At this point, due to the size, the complexity and repetitiveness of the E. curvula genome, we could not achieve the assembly metrics reached by other plant genomes based on PacBio assembly alone. However, the N50 of our assembly was higher than the ones obtained for other grasses using other sequencing technologies, as was the case for Aegilops tauschii (4.3 Gb)38, Triticum urartu (4.94 Gb)39 and S. Italica (490 Mb)23 with N50 values of 0.207 Mb, 0.064 Mb and 0.254 Mb, respectively.
The promising results obtained with the FALCON assembly encouraged us to look for a scaffolding technology to increase genome assembly contiguity. One of the latest methodologies to obtain chromosome-scale scaffolds is the proximity ligation-based technology used by Chicago and Hi-C. The combination of these technologies with the FALCON assembly increased the N50 to 43.41 Mb and decreased the number of scaffolds to 1,143. The only currently available report about this combination of technologies is for D. zibethinus (738 Mb)20, in which an N50 of 22.7 Mb was achieved, half of the final E. curvula N50. Other assemblies using a different combination of technologies such as optical mapping and Hi-C, for Chenopodium quinoa (1.45 Gb)16, M. truncatula (465 Mb)40 and Arabis alpine (370 Mb)41, rendered N50 values of 3.84 Mb, 12.5 Mb and 31 Mb, respectively. A combination of optical mapping and Chicago led to an N50 of 95 kb in Hevea brasiliensis (2.15 Gb)42. In Manihot esculenta (1.23 Gb)43 Chicago only was used with a resulting N50 value of 27.7 kb. These results show that the combination of Hi-C and Chicago is very powerful, increasing the N50 more than other combinations of technologies.
Our final E. curvula genome assembly has 96.4% of complete BUSCO genes, covers 97% of the estimated genome length and contains 95.5% of the DArT markers, all this components suggest the near completeness of this assembly.
An important feature of grass genomes is the presence of repetitive elements that differ in number and complexity among species, being higher in more complex genomes44,45. In the E. curvula genome assembly we found 28.8% of repetitive elements, a similar value to the 27.46% reported for E. tef22 and 21.4% for B. distachyon46, but lower than the 46.44% found in S. italica23 and the 62% from S. bicolor24.
The number of gene models found in E. curvula was 56,469. The BUSCO analysis performed on these models resulted in 28.1% of duplicated BUSCO genes. This overestimation seems to occur in most species, for example, the recently published T. aestivum genome assembly47 has 259,979 gene models, a very high number of genes even for an hexaploid species. From the gene models, we assessed the evolution of E. curvula within the Poaceae family, positioning the species close to E. tef and finding 9,189 orthogroups shared by the selected Poaceae subfamilies.
Seventy-four WRKY transcription factors were identified in the E. curvula genome assembly, and classified into three main groups and five subgroups, based on the results obtained by other researchers48. The identification, classification and characterization of this gene family constitute a key step into the elucidation of the molecular basis of the drought tolerance of this species49 since it allows the design of further specific expression studies that would contribute to dissect their involvement in this important trait of E. curvula.
One of the main limiting factors of E. curvula as a forage source it is its low digestibility50 a trait that has the potential to be improved by transgenic modification30. Two classes of CSE, an enzyme involved in lignin reduction, have been described32,33, with class II present in all species examined, whereas class I is only present in some of them. Interestingly, class II was present in all the evaluated Chloridoideae species, such as O. thomaeum, E. tef and E. curvula, finding not previously reported (Supplementary Table S10).
In conclusion, using a combination of different technologies to assemble and validate the E. curvula genome a notable advance in the de novo assembly of non-model genomes was achieved. This assembled genome also provides an invaluable tool to find new targets for crop improvement regarding classical focused traits, such as drought tolerance and digestibility. Finally, the availability of this assembly provides the foundation for the assembly of more complex tetraploid apomictic E. curvula genomes, aiding in the study of the reproductive mode.
Methods
Plant Material
DNA for genome sequencing was extracted from E. curvula cv. Victoria, a sexual diploid (2n = 2x = 20) genotype obtained from in vitro culture of inflorescences of cv. Tanganyika27 and registered at the National Cultivar Register, Argentina (UNST1122, RC9192). This accession couldn’t be selfed to decrease the heterozygosity because its self-incompatibility. Leaf samples for DNA extraction were collected from a plant growing in the greenhouse at CERZOS, CCT – CONICET Bahía Blanca, Argentina. DNA for DArT markers was extracted from leaves of a mapping population consisting of 63 individuals derived from the cross between two tetraploid E. curvula cultivars, OTA x Don Walter, and from two samples of cv. Victoria. DNA for SSRs amplification was extracted from leaves of different E. curvula cultivars: USDA accessions PI208214 (2x), PI299919 (2x), PI299920 (2x), PI299928 (2x), PI574506 (OTA, 4x), PI234217 (Tanganyika, 4x), and cultivars from INTA germplasm collection Tanganyika (4x), Don Walter (4x), Don Pablo (7x) and Don Luis (7x) (Supplementary Table S12).
DNA Extraction
DNA samples for PacBio long reads and DArT sequencing were obtained from 80 mg of fresh leaf tissue using a CTAB-based method51. The protocol was adapted to obtain large DNA molecules (gDNA), taking care in the critical steps to avoid breaking, resulting in an average length of 41,800 bp and a concentration of 0.7880 ng/μL.
For the Chicago and Dovetail Hi-C proximity ligation libraries, fresh leaf tissue from the same plant was delivered to Dovetail Genomics (www.dovetailgenomics.com) for extraction and sequencing of pure DNA and endogenous chromatin using a proximity ligation-optimized Dovetail in-house protocol.
Library Preparation and Sequencing
Long-read sequencing of cv. Victoria DNA was performed using Pacific Bioscience’s Single Molecule Real-Time (SMRT) chemistry through the Sequel platform (www.pacb.com) at the University of Liverpool Centre for Genomic Research (UK). Current PacBio systems generate reads with an average size of nearly 20 kb and a maximum length of over 60 kb. Two Libraries of 10 kb and 20 kb were prepared through the BluePippin (Sage Science) fragment selection method (http://www.sagescience.com/products/bluepippin/). After repairing the ends and an adapter ligation process the libraries were sequenced. The coverage of the estimated haplotype (620 Mb) was 90X, 10X greater than the one recommended for this technology for a genome with characteristics such as that of E. curvula (www.pacb.com/calculator-whole-genome-sequencing/).
To improve the assembly contiguity and to orientate the contigs, Chicago and Dovetail Hi-C libraries were sequenced starting from endogenous chromatin and high molecular weight DNA, respectively. The Chicago and Dovetail Hi-C libraries were sequenced through a 2 × 150 paired-end Illumina HiSeq2500 platform.
Genome Assembly
The PacBio raw reads were assembled with FALCON52 and Canu53 software exploring different parameters (Supplementary Table S13). The assembly quality was assessed by comparing numerous metrics (N50, assembly size, number of contigs and average contig length). The assembly was also evaluated using BUSCO v.354. This software uses a large selection of widespread orthologous single-copy genes as benchmarks to gauge the completeness of the novel assembled genome. The assembly with the highest N50, the least number of contigs, the highest average contig length and with more complete, less fragmented and/or missing BUSCO genes, was chosen.
After this procedure, the PacBio SMRT tool reference guide (https://www.pacb.com/wp-content/uploads/SMRT-Tools-Reference-Guide-v4.0.0.pdf) was followed to polish the draft genome assembly. The raw reads were aligned with the pbalign software against the assembly with the following criteria: minimum alignment length 50 bp; minimum similarity and minimum accuracy, 70%. The pbalign output was used as input for the Arrow software with the default parameters to polish the assembly by choosing the base with the highest coverage in each position.
The final assembly was obtained by scaffolding the polished genome assembly with the data obtained through the Chicago and Dovetail Hi-C libraries. The Dovetail Hi-Rise scaffolding software55 was used to integrate the data obtained from the Chicago library with the PacBio polished assembly. Finally, the Chicago assembly was combined with the Dovetail Hi-C files through Dovetail Hi-Rise to obtain scaffolds in ranges up to the size of whole chromosomes.
Diversity Arrays Technology (DArT) Validation
The entire DArT preparation procedure, including SNP calling, was provided by the Genetic Analysis Service for Agriculture Laboratory (SAGA, CIMMyT, México) using an in-house protocol. DArT technology uses the combination of PstI and MseI restriction enzymes in order to reduce the genome complexity. The process separate low copy sequences from the repetitive fraction of the genome (https://www.diversityarrays.com). The fragments obtained from the enzyme digestion were sequenced using the Illumina platform and the reads were used as input by the service provider protocol to de novo obtain the markers of 69 bp containing the SNP. The reads and markers are available under the NCBI bioproject PRJNA508722 (https://www.ncbi.nlm.nih.gov/bioproject/) with the SNP position and polymorphism present in the header of each marker sequence. The SNP markers were mapped onto the whole genome assembly using the Bowtie software56 with the end-to-end and -k 10 (up to 10 distinct valid alignments for each read) parameter to validate the final assembly with data from another source. Since the DArT libraries are designed to target active regions of the genome the 492,378 DArT reads were mapped with Bowtie onto all the scaffolds using the –k 10 and end-to-end parameter and the hits were plotted with a 500 kb window size.
Repetitive Sequence Assessment
Repetitive sequences were assessed through three different approaches. The first one was based on the generation of a de novo library by the RepeatModeler57 software, that uses a modeling package to de novo identify repeat families. The second approach uses the TransposonPSI software58 that identifies repetitive elements based on the homology to protein or nucleic acid sequences to proteins encoded by diverse families of transposable elements. The result of each program is merged with USERCH v759 taking only a single record when the repetition is included in both programs. The merged file is then classified with the RepeatClassifier script (included in the RepeatModeler package) according to the structure and the type of element present in the sequence in one of the main classes of repetitions (ALU, LINE, LTR, DNA elements and Unknown elements). The sequences were classified to subclass and superfamily level depending if they were complete or not. The final approach consisted of finding homologous repetitive elements in related monocot species. RepBase23, a reference database of repetitive DNA sequences from different eukaryotic species60, was used to find homologous sequences. Here, consensus sequences of large families and subfamilies of repeats from Z. mays, S. bicolor and Oryza sativa were used.
The previously reported ESTs25 related to repetitive elements were aligned to the repetitive elements present in the assembled E. curvula genome in order to find the complete structure of these elements using the BLASTn algorithm with an e-value of 1.0 e−10.
Gene Annotation
In order to annotate the gene models present in the E. curvula genome assembly, the repetitive DNA was masked using the RepeatMasker software61 using the de novo and the homology-based fasta files with the library (−lib) parameter. The -s parameter was used to increase sensitivity in the masking process. To find homology, RMblast, a RepeatMasker-compatible version of the standard NCBI BLASTn program, was used. The main difference between these two programs is that RMblast is optimized to compare the RepeatMasker matrix.
After this procedure two different approaches were used to annotate the genes; the first consisted of the alignment of the genome scaffolds against the E. curvula floral transcriptome9 and ESTs62, and protein data from related species like Eragrostis tef 8, S. italica23, S. bicolor24 and Z. mays17. This analysis was performed using the Exonerate software63, a general tool for sequence comparisons, setting the minimum alignment coverage to 80% and the minimum identity to 85%. The second method included ab initio gene prediction, an intrinsic method based on gene content and signal detection in which the genomic DNA sequence alone is systematically searched for certain tell-tale signs of protein-coding genes. For this, we used AUGUSTUS64 an HMM-based (Hidden Markov Model) gene finder and SNAP software65. In ab initio prediction is necessary to train the programs in order to create the best model to precisely find the genes. For this purpose, the model used to find the complete genes was extracted from the output of the BUSCO software.
The annotation was performed through the MAKER software66, using as input the RepeatMasker output and the protein and RNA alignment obtained from Exonerate in a splice-aware fashion to accurately identify splicing sites. MAKER also uses the gene models predicted by AUGUSTUS and SNAP, compares all the predicted gene models to RNA and protein alignment evidence, and then revises the ab initio gene models in order to predict the most confident gene models.
To assess the annotation completeness, the classification strategy adopted for T. aestivum47 was followed. This strategy classified the genes in three main categories, high confidence (HC), low confidence (LC), and transposons (TREP), based on the completeness (start and stop codon) and on the homology (coverage ≥ 90%; e-value ≤ 10 e−10) to unipoa (Poaceae proteins, SwissProt and trEMBL), unimag (Magnoliophyta proteins, SwissProt) and TREP database67 (transposons database) (Supplementary Table S14).
Finally, InterProScan version 568 was used to classify genes into families and predict the presence of domains and important sites. This software uses 14 different databases, retrieving information such as KEGG31 pathways, gene ontology and Pfam domains.
Synteny Analysis
SyMAP69 software was used to search for homologies among E. curvula and the genomic regions of other grasses. This tool generates whole genome synteny patterns plots between two organisms. The selected species for these comparisons were other C4 grasses, such as E. tef 26 S. bicolor24, Z. mays17, O. sativa70 and O. thomaeum14. SyMAP uses the alignment tool BLAT71, run with the following parameters: -minScore = 30, -minIdentity = 70, -tileSize = 10, -qMask = lower, and -maxIntron = 10000.
The paralogous and orthologous syntenic genes obtained were listed (Supplementary Files 1 and 2) and the Ks substitution rate was calculated for each pair of genes using the BioPerl package through the Nei–Gojobori method. To find WGD events the Ks rate was plotted and the peaks were corrected using the method proposed by Wang et al.72. Results were visualized in a circle plot showing the shared genomic regions with different colors and a table was constructed showing the number of anchors and blocks common to E. curvula, E. tef, S. bicolor, Z. mays, O sativa and O. thomaeum.
Analysis of the Evolutionary Relationships Among the Poaceae Family
The evolutionary relationships among E. curvula and monocots species like S. italica23, T. aestivum47, E. tef 8, O. sativa68, Z. mays17, S. bicolor24, B. distachyon46, O. thomaeum14, Musa itinerans73 and P. hallii74, were assessed by comparing the assembled genome annotations using the software Orthofinder75. This software groups in orthogroups genes originating from the same common ancestor and creates trees for each group and for all the species. The alignment of the orthogroups for all the species was used to create a maximum likelihood time phylogenetic tree using the Jones Taylor Thornton model with the software MEGAX76. The calibration was performed according to Prasad et al.77 considering the divergence time between O. sativa and Z. mays in approximately 70 Mya. M. itinierans was used as outgroup.
Gene expansion and contraction were assessed by the CAFEv3.078 software using the genes families obtained from Orthofinder. Multiple birth-death lambda (λ) was used in order to assess the different clade evolution rates.
Classification of the WRKY Transcription Factor Family
To identify the WRKYs from E. curvula the annotated genes with WRKY motifs were extracted from Pfam. The genes were filtered with the MEME software79, and those genes showing complete WRKY and zinc finger motifs were classified into three main groups (I, II and III) and into five subgroups (IIa, IIb, IIc, IId and IIe)48. Those genes with two WRKY motifs were classified in group I and those with only one into groups II and III. Then, if the terminal region of the zinc finger was H–X1–H, the gene was classified as group II and if the terminal region was H–X1–C, the gene was classified as group III. Based on the remaining motif of the sequences, group II was classified into five subgroups. Finally, to find the differences among the groups, a multiple sequence alignment was run with the MUSCLE software80, all the groups and subgroups were plotted in a phylogenetic tree constructed using MEGAX software76 with a maximum likelihood model
Analysis of Genes Involved in the Lignin Pathway
The KASS online tool81 assigns a biological role to new genes using the Ghostx aligner82, finding homology to known sequences in the KEGG31 database and assigning them a position in a pathway. Genes for class I and class II caffeoyl shikimate esterases (CSE) were targeted, since both have been recently mentioned as having important roles in the regulation of the lignin pathway32,33. For BLASTn, a class I O. sativa orthologous gene (accession XM_015768109.2) and a class II S. bicolor orthologous gene (accession XM_002462989.2) were used as queries. Specific primers for PCR amplification of these genes were designed based on the genome sequence, the PCR program consisted of an initial DNA denaturation at 94 °C for 2 min, followed by 38 cycles at 94 °C for 15 s, 50 °C for 20 s and 72 °C for 80 s and a final extension of 5 min at 72 °C. Amplicons were analyzed over 1.5% agarose gel. The amplicons were cloned into the pGEM-T Easy Vector (Promega), sequenced and BLAST searched were performed against the E. curvula transcripts, and the P halli, O. sativa, E. tef, O. thomaeum, Brachypodium distachyon, T. aestivum, Z. mays, S. bicolor and S italica genomes.
SSRs Discovery and Analysis of Genetic Relationships within the E. curvula Complex
SSRs discovery was conducted through SSR Locator software83 using the assembled genome sequence as input. Aiming to validate the genome assembly and to assess genetic relationships within the E. curvula complex, primers flanking each SSR were designed using the software Primer3 2.384. Fifteen randomly selected primer pairs (Supplementary Table S9) were synthesized and amplified in genomic DNA from 10 E. curvula genotypes (Supplementary Table S12). SSRs previously developed for E. curvula were also tested9 to increase the number of markers included in the phylogenetic tree, thus improving the accuracy of the results. PCR reactions used 1 μl of 10 mM dNTPs mix, 1× reaction buffer, 0.5 μl of each forward and reverse primer (100 pmol/μl), 1.5 U Taq DNA polymerase and 60 ng of template genomic DNA in 20 μl of reaction volume. The PCR program consisted of an initial DNA denaturation at 94 °C for 3 min, followed by 40 cycles at 94 °C for 30 s at the optimal annealing temperature for each primer pair and 72 °C for 30 s and a final extension of 5 min at 72 °C. The amplicons were validated in Victoria cultivar using 1.5% agarose gels and the presence/absence of the bands was assessed trough 6% polyacrylamide gels. To size the bands the ladder used was the Genebiotech 100 bp Plus DNA (L00307P).
The similarity of the cultivars and the relative position of cv. Victoria within the E. curvula complex were assessed regarding the presence or absence of individual bands in the polyacrylamide gels obtained from each primer pair in each individual tested. The pairwise distance of this binary (1 presence, 0 absence) data matrix was computed using the Jaccard method85 and the phylogenetic tree was created with hierarchical clustering and the ward D2 method in an R environment.
Data Access
Under the NCBI bioproject PRJNA508722 (https://www.ncbi.nlm.nih.gov/bioproject/) are available:
-genome sequences
-genome annotation
-DArT reads and markers sequences
References
Morgan, J. A. et al. C4 grasses prosper as carbon dioxide eliminates desiccation in warmed semi-arid grassland. Nature. 476, 202–205 (2011).
Voigt, P. W. & Bashaw, E. C. Facultative Apomixis in Eragrostis curvula. Crop Sci. 16, 803–806 (1976).
Farrington, P. The seasonal growth of lovegrass (Eragrostis curvula) on deep sandy soils in a semi-arid environment. Aust j exp agr. 13, 383–388 (1973).
Saleme, M. D. et al. Silencing CAFFEOYL SHIKIMATE ESTERASE affects lignification and improves saccharification. Plant physiol. 175, 1040–1057 (2017).
Spangenberg, G. et al. Breeding Forage Plants in the Genome Era. In Molecular Breeding of Forage Crops 1–39 (Lorne and Hamilton, 2001).
Lauriault, L. M. et al. A Screening for Biofuel Feedstock Quality of Perennial Warm-Season Grasses in Semiarid Subtropical Environments. Res Rep New Mexico Agric Exp Sta. 774, 1–8 (2012).
Roodt‐Wilding, R. & Spies, J. J. Phylogenetic relationships in southern African chloridoid grasses (Poaceae) based on nuclear and chloroplast sequence data. Syst Biodivers. 4, 401–415 (2006).
Cannarozzi, G. et al. Genome and transcriptome sequencing identifies breeding targets in the orphan crop tef (Eragrostis tef). BMC genomics. 15, 581, https://doi.org/10.1186/1471-2164-15-581 (2014).
Garbus, I. et al. De novo transcriptome sequencing and assembly from apomictic and sexual Eragrostis curvula genotypes. PloS one. 12, https://doi.org/10.1371/journal.pone.0185595 (2017).
Kamies, R., Farrant, J. M., Tadele, Z., Cannarozzi, G. & Rafudeen, M. S. A proteomic approach to investigate the drought response in the orphan crop Eragrostis tef. Proteomes. 15, 32, https://doi.org/10.3390/proteomes5040032 (2017).
Jiao, W. B. & Schneeberger, K. The impact of third generation genomic technologies on plant genome assembly. Curr Opin Plant Biol. 36, 64–70 (2017).
Nakano, K. et al. Advantages of genome sequencing by long-read sequencer using SMRT technology in medical area. Hum Cell. 30, 149–161 (2017).
Berlin, K. et al. Assembling large genomes with single-molecule sequencing and locality-sensitive hashing. Nat Biotechnol 33, 623–630 (2015).
VanBuren, R. et al. Single-molecule sequencing of the desiccation-tolerant grass Oropetium thomaeum. Nature. 527, 508–511 (2015).
Lan, T. et al. Long-read sequencing uncovers the adaptive topography of a carnivorous plant genome. Proc Natl Acad Sci USA 114, 4435–4441 (2017).
Jarvis, D. E. et al. The genome of Chenopodium quinoa. Nature. 542, 307–301 (2017).
Jiao, Y. et al. Improved maize reference genome with single-molecule technologies. Nature. 546, 524–527 (2017).
Badouin, H. et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature. 546, 148–152 (2017).
Putnam, N. H. et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res. 26, 342–350 (2016).
Teh, B. T. et al. The draft genome of tropical fruit durian (Durio zibethinus). Nat genet. 46, 1633–1641 (2017).
Estep, M. C., DeBarry, J. D. & Bennetzen, J. L. The dynamics of LTR retrotransposon accumulation across 25 million years of panicoid grass evolution. Heredity. 110, 194–204 (2013).
Gebre, Y. G., Bertolini, E., Pè, M. E. & Zuccolo, A. Identification and characterization of abundant repetitive sequences in Eragrostis tef cv. Enatite genome. BMC Plant Biol. 16, 39, https://doi.org/10.1186/s12870-016-0725-4 (2016).
Zhang, G. et al. Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol. 30, 549–554 (2012).
Paterson, A. H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature. 457, 551–556 (2009).
Romero, J., Selva, J. P., Pessino, S., Echenique, V. & Garbus, I. Repetitive sequences in Eragrostis curvula cDNA EST libraries obtained from genotypes with different ploidy. Biol Plant. 60, 55–67 (2016).
VanBuren, R. et al. Exceptional subgenome stability and functional divergence in allotetraploid teff, the primary cereal crop in Ethiopia. bioRxiv 580720, https://doi.org/10.1101/580720.
Cardone, S. et al. Novel genotypes of the subtropical grass Eragrostis curvula for the study of apomixis (diplospory). Euphytica. 151, 263–272 (2006).
Chen, L. et al. Improved forage digestibility of tall fescue (Festuca arundinacea) by transgenic down‐regulation of cinnamyl alcohol dehydrogenase. Plant Biotechnol J. 1, 437–49 (2003).
He, X., Hall, M. B., Gallo-Meagher, M. & Smith, R. L. Improvement of forage quality by downregulation of maize O-methyltransferase. Crop Sci. 43, 2240–2251 (2003).
Giordano, A. et al. Reduced lignin content and altered lignin composition in the warm season forage grass Paspalum dilatatum by down-regulation of a Cinnamoyl CoA reductase gene. Transgenic Res. 23, 503–517 (2014).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, 590–595 (2018).
Vanholme, R. et al. Caffeoyl shikimate esterase (CSE) is an enzyme in the lignin biosynthetic pathway in Arabidopsis. Science. 341, 1103–1106 (2013).
Ha, C. M. et al. An essential role of caffeoyl shikimate esterase in monolignol biosynthesis in Medicago truncatula. Plant J. 86, 363–375 (2016).
Díaz, M. L., Garbus, I. & Echenique, V. Allele-specific expression of a weeping lovegrass gene from the lignin biosynthetic pathway, caffeoyl-coenzyme A 3-O-methyltransferase. Mol Breed. 64, 627–637 (2010).
Muthamilarasan, M. et al. Global analysis of WRKY transcription factor superfamily in Setaria identifies potential candidates involved in abiotic stress signaling. Front Plant Sci. 26, 910, https://doi.org/10.3389/fpls.2015.00910 (2015).
Wei, K. F., Chen, J., Chen, Y. F., Wu, L. J. & Xie, D. X. Molecular phylogenetic and expression analysis of the complete WRKY transcription factor family in maize. DNA res. 19, 153–64 (2012).
Ning, P., Liu, C., Kang, J. & Lv, J. Genome-wide analysis of WRKY transcription factors in wheat (Triticum aestivum L.) and differential expression under water deficit condition. PeerJ. 5, 3232, https://doi.org/10.7717/peerj.3232 (2017).
Luo, M. C. et al. Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature. 551, 498–502 (2017).
Ling, H. Q. et al. Draft genome of the wheat A-genome progenitor Triticum urartu. Nature. 496, 87–90 (2013).
Moll, K. M. et al. Strategies for optimizing BioNano and Dovetail explored through a second reference quality assembly for the legume model, Medicago truncatula. BMC genomics. 18, 578, https://doi.org/10.1186/s12864-017-3971-4 (2017).
Jiao, W. B. et al. Improving and correcting the contiguity of long-read genome assemblies of three plant species using optical mapping and chromosome conformation capture data. Genome Res. 27, 778–786 (2017).
Pootakham, W. et al. De novo hybrid assembly of the rubber tree genome reveals evidence of paleotetraploidy in Hevea species. Sci Rep. 7, 41457, https://doi.org/10.1038/srep41457 (2017).
Bredeson, J. V. et al. Sequencing wild and cultivated cassava and related species reveals extensive interspecific hybridization and genetic diversity. Nat Biotechnol. 34, 562–270 (2016).
Dubin, M. J., Scheid, O. M. & Becker, C. Transposons: a blessing curse. Curr Opin Plant Biol. 42, 23–29 (2018).
Lee, S. I. & Kim, N. S. Transposable elements and genome size variations in plants. Genomics inform. 12, 87–97 (2014).
Vogel, J. P. et al. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature. 463, 763–768 (2010).
Appels, R. et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. 361, 6403, https://doi.org/10.1126/science.aar7191 (2018).
Eulgem, T. & Somssich, I. E. Networks of WRKY transcription factors in defense signaling. Curr Opin Plant Biol. 10, 366–371 (2007).
Colom, M. R. & Vazzana, C. Water stress effects on three cultivars of Eragrostis curvula. Ital. j. agron. 6, 127–32 (2002).
Gargano, A. O., Adúriz, M. A., Arelovich, H. M. & Amela, M. I. Forage yield and nutritive value of Eragrostis curvula and Digitaria eriantha in central-south semi-arid Argentina. Trop. grassl. 35, 161–167 (2001).
Meier, M., Zappacosta, D., Selva, J. P., Pessino, S. & Echenique, V. Evaluation of different methods for assessing the reproductive mode of weeping lovegrass plants, Eragrostis curvula (Schrad.) Nees. Aust J Bot. 59, 253–61 (2011).
Chin, C. S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat Methods. 13, 1050–1054 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 31, 3210–3212 (2015).
Koch, L. Technique: Chicago HighRise for genome scaffolding. Nat Rev Genet. 17, 194, https://doi.org/10.1038/nrg.2016.23 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods. 9, 357–359 (2012).
Smit, A., Hubley, R. & Green, P. Open-1.0, http://www.repeatmasker.org (2008–2015).
Altschul, S. F. & Koonin, E. V. Iterated profile searches with PSI-BLAST—a tool for discovery in protein databases. Trends Biochem Sci. 23, 444–447 (1998).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 26, 2460–2461 (2010).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA. 6, 11, https://doi.org/10.1093/nar/gkp335 (2015).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0, http://www.repeatmasker.org (2013–2015).
Cervigni, G. D. et al. Expressed sequence tag analysis and development of gene associated markers in a near-isogenic plant system of Eragrostis curvula. Plant Mol Biol. 67, 1–10 (2008).
Slater, G. S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC bioinformatics. 6, 31, https://doi.org/10.1186/1471-2105-6-31 (2005).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, 309–312 (2004).
Johnson, A. D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics. 24, 2938–2939 (2008).
Campbell, M. S., Holt, C., Moore, B. & Yandell, M. Genome annotation and curation using MAKER and MAKER‐P. Curr Protoc Bioinformatics 48, 11–39 (2014).
Schulman, A. H. & Wicker, T. A Field Guide to Transposable Elements. In Plant Transposons and Genome Dynamics in Evolution (ed. Fedoroff, N. V.) 15–40 (John Wiley & Sons, Pennsylvania State, University, 2013).
Finn, R. D. et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 45, 190–199 (2016).
Soderlund, C., Bomhoff, M. & Nelson, W. SyMAP: A turnkey synteny system with application to plant genomes. Nucleic Acids Res. 39(10), e68, https://doi.org/10.1093/nar/gkr123 (2010).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice. 6, 4, https://doi.org/10.1186/1939-8433-6-4 (2013).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome res. 12, 656–664 (2002).
Wang, X. et al. Genome alignment spanning major Poaceae lineages reveals heterogeneous evolutionary rates and alters inferred dates for key evolutionary events. Mol Plant. 8, 885–98 (2015).
Wu, W. et al. Whole genome sequencing of a banana wild relative Musa itinerans provides insights into lineage-specific diversification of the Musa genus. Sci Rep 6, 31586 (2016).
Lovell, J. T. et al. The genomic landscape of molecular responses to natural drought stress in Panicum hallii. Nature com. 9, 5213 (2018).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157, https://doi.org/10.1186/s13059-015-0721-2 (2015).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol. 35, 1547–1549 (2018).
Prasad, V., Strömberg, C. A., Alimohammadian, H. & Sahni, A. Dinosaur coprolites and the early evolution of grasses and grazers. Science 310, 1177–1180 (2005).
Han, M. V., Thomas, G. W., Lugo-Martinez, J. & Hahn, M. W. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol Biol Evol. 30, 1987–1997 (2013).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nuc acids research. 37, 202–208 (2009).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, 182–185 (2007).
Suzuki, S., Kakuta, M., Ishida, T. & Akiyama, Y. GHOSTX: an improved sequence homology search algorithm using a query suffix array and a database suffix array. PloS one. 9, e103833, https://doi.org/10.1371/journal.pone.0103833 (2014).
Da Maia, L. C. et al. SSR locator: tool for simple sequence repeat discovery integrated with primer design and PCR simulation. Int J Plant Genomics. 2008, 412696, https://doi.org/10.1155/2008/412696 (2008).
Koressaar, T. & Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics. 23, 1289–1291 (2007).
Hamers, L. Similarity measures in scientometric research: The Jaccard index versus Salton’s cosine formula. Inf Process Manag. 3, 315–18 (1989).
Acknowledgements
This work was granted funded by the Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, PICT Raíces 2014–1243 and PICT Raíces 2017 - 0879), Universidad Nacional del Sur (PGI 24/A199), Consejo Nacional de Investigaciones Científicas y Técnologicas (CONICET) and the European Union’s Horizon 2020 Research and Innovation Programme under the Marie Skłodowska-Curie Grant Agreement No. 645674 (PROCROP).
Author information
Authors and Affiliations
Contributions
The genome assembly, the annotation and bioinformatics validation was made by Carballo J. with Santos B.A.C.M. and Gallo C.A. assistance. The SSR molecular validations were designed and performed by Zappacosta D. and Garbus I. All the DNA samples were extracted by Selva J.P. The forage quality genes molecular validations were carried out by Selva J.P. and Diaz A. The manuscript was written by Carballo J., Echenique V., Zappacosta D., Caccamo M. and Albertini E. Work coordination was made by Echenique V and Albertini E. Bioinformatics supervision was leader by Caccamo M. Supervision in terms of biological and genetic analyses and the overall work planning and coordination was made by the team leader Echenique V.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Carballo, J., Santos, B.A.C.M., Zappacosta, D. et al. A high-quality genome of Eragrostis curvula grass provides insights into Poaceae evolution and supports new strategies to enhance forage quality. Sci Rep 9, 10250 (2019). https://doi.org/10.1038/s41598-019-46610-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46610-0
This article is cited by
-
Chromosome-scale genome assembly and annotation of Paspalum notatum Flüggé var. saurae
Scientific Data (2024)
-
Effect of dietary inclusion of Pennisetum purpureum (Napier) grass on growth performance, rumen fermentation and meat quality of feedlot sussex red steers
Tropical Animal Health and Production (2024)
-
Differentially methylated genes involved in reproduction and ploidy levels in recent diploidized and tetraploidized Eragrostis curvula genotypes
Plant Reproduction (2024)
-
Identification of WRKYs using in silico tools for unraveling the modulation during abiotic stress response in Tef [Eragrostis tef (Zucc.) Trotter] a super grain
Genetic Resources and Crop Evolution (2023)
-
High-quality chromosome-scale de novo assembly of the Paspalum notatum ‘Flugge’ genome
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.