Abstract
Delta-9-tetrahydrocannabinol (THC) is the primary psychoactive compound in Cannabis, which is studied extensively for its medicinal value. A central gap in the science is the underlying mechanisms surrounding THC’s therapeutic effects and the role of gut metabolite profiles. Using a mass-spectrometry based metabolomics, we show here that intraperitoneal injection of THC in C57BL/6 mice modulates metabolic profiles that have previously been identified as integral to health. Specifically, we investigated the effects of acute (single THC injection denoted here as ‘1X’) and short -term (five THC injections on alternate days denoted as ‘5X’) THC administration on fecal and intestinal tissue metabolite profiles. Results are consistent with the hypothesis that THC administration alters host metabolism by targeting two prominent lipid metabolism pathways: glycerophospholipid metabolism and fatty acid biosynthesis.
Similar content being viewed by others
Introduction
The medical use of Cannabis (commonly termed ‘marijuana’), a product from the plant, Cannabis sativa, is becoming increasingly popular worldwide for its medicinal value1,2,3. In the USA, 33 states and District of Columbia have already legalized the medical use of marijuana4. Delta-9-tetrahydrocannabinol (THC) that was first described in 1964 is the primary psychoactive compound in Cannabis and is known to display therapeutic potentials as an analgesic, antiemetic and appetite stimulant2,3,5,6,7,8. Additionally, THC can be used for the treatment of multiple acute and chronic health disorders9,10,11. These include treatment of nausea and vomiting associated with cancer chemotherapy, anorexia, and cachexia associated with HIV and AIDS patients, pain and muscle spasms in multiple sclerosis12. Their anti-inflammatory effects have been tested in experimental models for autoimmune disorders such as multiple sclerosis, rheumatoid arthritis, colitis, hepatitis and cancer9. Importantly, during the past several years, THC content in marijuana has been steadily increasing13,14; and recreational use has expanded15. Hence, the scientific premise for studying the mechanisms involved in the modulation of disease by THC is strong.
It is well established that THC can be useful in preventing and ameliorating the symptoms of intestinal inflammation such as abdominal pain, diarrhea and reduced appetite in patients suffering from inflammatory bowel disease (IBD)16,17,18,19. In animal models of colitis, THC demonstrated successful reduction of 2,4,6-Trinitrobenzene sulphonic acid-induced mucosal damage of the intestine, neutrophil infiltration and in vitro motility disturbances. It has been shown20,21,22 that the epithelial wound healing, inhibition of pro-inflammatory cytokine and chemokine release, immune cell recruitment, are mediated through activation of cannabinoid receptors (CB1 and CB2) of the endogenous cannabinoid system. Activation of these receptors is critical in the neuromodulatory actions on the sensory and autonomic nervous system connected with the pharmacological function of THC23.
Despite such well-documented evidence supporting THC’s therapeutic impact in safeguarding the healthy functioning of the intestine, the entire mechanism underlying the intestinal protection and other pharmacological effects of THC still remains unclear. This gap in understanding is at least in part due to the absence of knowledge on whether THC influences host metabolism and how. The knowledge of the key metabolic targets of THC is critical for appropriate therapeutic applications of THC. Hence the main objective of this study was to investigate the influence of acute and short-term THC administration on fecal and intestinal tissue metabolome of wild-type C57BL/6 mice. A comparative untargeted metabolome profiling of fecal and intestinal tissue samples obtained from THC administered mice and control mice that received vehicle instead, was conducted using an ultra-performance liquid chromatography-time of flight-mass spectrometry (UPLC-TOF-MS). Presented in this report are the results of this investigation.
Materials and Methods
Experimental animals, diet and THC administration
Female C57BL/6 (BL6) mice, aged 8–10 weeks, obtained from Jackson Laboratories were used for this study. All mice were housed in pathogen-free conditions and allowed ad libitum access to filtered water and Teklad rodent diet 8604 (regular chow) at the Animal Research Facility located at the University of South Carolina School of Medicine. To understand the effect of THC on gut metabolites we have followed our previously published protocols of THC administration24,25,26,27,28, whereby we injected 20 mg/Kg THC intraperitoneally. The experimental group (n = 5) received THC every 48 h. THC was dissolved in a vehicle of 100% ethanol, and both treatment and vehicle were administered in 100 µL of a combination of ethanol, Tween-80, and saline, at a ratio of 2:1:17. Animals were regularly monitored during the period of the experiment for any body weight changes, signs of toxicity and mortality. Fecal samples analyzed for comparative metabolomics were collected 24 h after the first administration (denoted here as 1X) and 24 h after the 5th administration (denoted here as 5X). The 1X samples and 5X samples were used to study respectively, the acute and short-term effects of THC administration.
The rationale for using an intraperitoneal mode of exposure was the fast bioavailability of THC in the bloodstream, due to which we have consistently used this model25,26,27 as the closest intravenous self-administration paradigm because THC is unable to sustain in rodents upon intravenous administration29. As in our previous studies25,26,27, we reasoned here as well, that since i.p. administration would allow the THC to directly diffuse across the peritoneal membrane to the blood vessels of the abdominal viscera, musculature and mesentery, it would help in avoiding any possible artifacts resulting differential nutrient absorption rates caused by oral administration of THC.
The dose of THC in this study and our previous studies was determined according to body surface area normalization based calculations described earlier by Reagan-Shaw et al.30. Based on these calculations, our applied 20 mg/kg THC dose translates to 60 mg/m2 in humans, which is well within the maximum human recommended dose (MHRD) of synthetic THC of 90 mg/m2/day. We point out here that THC (also termed Dronabinol) has consistently been used for clinical use to reduce neuropathic pain in multiple sclerosis patients31,32,33. No teratogenic effects was reported in mice administered THC at up to 30 times the MRHD and up to 5 times the MRHD for patients with AIDS and cancer, respectively (see FDA data34). We have chosen mice of a single gender for our experiments to rule out any previously reported sex difference effects on THC metabolism35,36. Given that the prevalence of multiple sclerosis is about two to three times higher in women than men37, we have chosen female mice for this study.
Ethics statement
The mice employed in this study were housed at the American Association for the Accreditation of Laboratory Animal Care (AAALAC)-accredited Animal Resource Facility at the University of South Carolina, School of Medicine, Columbia, SC. All experimental procedures were performed according to National Institutes of Health (NIH) guidelines under protocols approved by the University of South Carolina Institutional Animal Care and Use Committee.
Sample preparation for metabolome analysis
Fecal samples were collected at the indicated time points by placing mice in individual cages with very little bedding but ad libitum access to food and water. Fecal pellets were immediately collected and placed on ice until a full sample was collected at which point the samples were immediately frozen in liquid nitrogen and transferred to −80. In order to ensure that only fresh feces were used for fecal metabolome analysis, the cages were changed every day. Fecal samples (0.1 g) from THC administered animals and controls were ultrasonically homogenized in 1 mL cold methanol/water (1:1) in for 30 mins followed by vortexing on cooled (4 °C) mixer for five mins. The homogenized samples were centrifuged for ten mins at 10000 × g at 4 °C. The supernatant (300 μL) was dried in a vacuum concentrator. The dry residue was re-dissolved in 150 μL methanol/water (1:1) before analysis. For intestinal tissues, the metabolites were extracted from 50 mg of lyophilized tissue samples with 800 µL of methanol. The samples were ground to fine powder using Grinding Mill at 65 Hz for 45 s. The ground samples were vortexed for 30 s, and centrifuged at 10,000 × g at 4 °C for 15 mins. Finally, 200 µL of supernatant was transferred to vial for LC-MS analysis.
UPLC-ESI-QTOF-MS profiling of fecal metabolites
Metabolites were separated from injected samples (5 μL aliquots) using Ultra Performance Liquid Chromatograph (1290 Infinity Binary LC System, Agilent Technologies, USA) and screened with ESI-MS (targeted MS/MS mode). The chromatograph system comprised of Waters ACQUITY UPLC HSS T3 (100 × 2.1 mm,1.8 um) with Phenomenex Security GuardTM ULTRA. The mobile phase consisted of 0.1% formic acid-water (solvent A) and 0.1% formic acid-acetonitrile (solvent B) with a gradient elution (0−1 min, 95% A; 1−6 min, 95−70% A; 6−20 min, 70−5% A). The flow rate of the mobile phase was set at 0.5 mL·min-1. The column temperature was maintained at 45 °C, and the sample manager temperature was set at 4 °C.
Mass spectrometry was performed on Quadrupole/Time-Of-Flight Mass Spectrometer (QTOF-MS; model G6540B Agilent Technologies, USA) using a Dual Agilent Jet Stream (AJS) ESI source. Spectra were recorded in the scanning mass-to-charge (m/z) range of 50 to 1500 with a scan rate of 1.00 spectra·sec-1. The capillary voltage was set to 4000 V, and 3500 V (positive and negative mode, respectively) and the fragmentor was set to 175 V. The pressure of the nebulizer was set at 35 psi, the gas temperature to 325 °C, and the continuous gas flow to 5 L·min-1. The instrument mode was set to an extended dynamic range. Quality control was maintained by injecting a control sample after analysis of every ten samples. The needle was washed (3X) with 50% methanol before every injection to avoid cross-contamination of samples. A volume of 20 μL of methanol was injected for rinsing.
UPLC-ESI-QTOF-MS profiling of intestinal tissue metabolites
Separation of the tissue metabolites was performed using a similar UPLC setup with the following modifications. The chromatography system comprised of an Agilent 959758-902 RRHD Eclipse Plus C18 (100 × 2.1 mm, 1.8 µm) with Phenomenex Security GuardTM ULTRA. The gradient elution was set to 0–1 min 95% A, 1–6 min 95–80% A, 6–9 min 80–50% A; 9–13 min 50–5% A; 13–15 min 5% A. The flow rate of the mobile phase was set at 0.35 mL·min−1. The column temperature was maintained at 40 °C, and the sample manager temperature is set at 4 °C. For mass spectrometry the spectra were recorded in the scanning mass-to-charge (m/z) range of 50 to 1000 with a inter scan time of 0.02 s. The capillary voltage was set to 4000 V and 3500 V (positive and negative mode, respectively) and the sampling cone was set to 35 kV and 50 kV (positive and negative mode, respectively). The cone gas flow was set to 50 L/h, the source temperature to 100 °C, and the extraction cone to 4 V. The desolvation temperature was set to 350 °C and 300 °C (positive and negative mode, respectively), and the desolvation gas flow is set to 600 L/h and 700 L/h (positive and negative mode, respectively).
Metabolite data processing and analysis
For metabolite data processing, the acquired raw data were aligned using Mass Hunter Workstation (B0.06.00, Agilent) based on the m/z value and the retention time of the ion signals. Ions from both ESI− and ESI+ were merged into the SIMCA-P program (version 14.1) for multivariate analysis. The data were stored in a table with one sample per row and one variable (bin/peak/metabolite) per column. The ion intensities for each peak detected were then normalized to the sum of the peak intensities in the sample (SI Fig. 1), finally rendering a multi-dimensional dataset, comprising of a peak number based on the m/z value and the retention time of the ion signals and ion intensities. This dataset was used for multivariate data analysis (MetaboAnalyst 3.0.), which included Univariate Analysis, Principal Component Analysis (PCA) and Partial Least Squares - Discriminant Analysis (PLS-DA). The univariate analysis was used for exploratory data analysis to determine Fold Change (FC) of metabolites between experimental and control groups and conduct t-tests. The unsupervised PCA was used to visualize the variance in a data set per group and the separation between the experimental and the control groups. The supervised PLS-DA was used to assign the class to the metabolites determine the difference between the groups for each class. Finally, the variable importance of projection values (the VIP values) was computed from the weighted sum of squares of the PLS loadings considering the amount of explained permuted class level variation in each dimension.
Identification of potential biomarkers and interpretation of metabolic signatures
The chemical structures of metabolites were identified according to online databases such as the Human Metabolome Database (www.hmdb.ca), Metlin (www.metlin.scripps.edu) and the Mass Bank (www.massbank.jp) using the data of accurate masses and MS/MS fragments. When necessary, further confirmation was acquired through comparisons with authentic standards, including retention times and MS/MS fragmentation patterns. To interpret the biological significance of the metabolic signatures, the metabolites that displayed significant differences between THC administered and control groups were imported to MBRole, a freely available web server for functional enrichment analysis on metabolic data from any organism38,39. Since a mouse model was used for this current study, the MBRole analysis was performed using the Mus musculus background.
Sample size determination and statistical analyses
We used power analysis to determine the ideal sample size for our experiments. With the assumption of a normal distribution, a 20% change in mean and 15% variation in THC effect on gut metabolome, we determined that a sample size ≥4 would be required per group to surpass 80% power for the study, given that we used concurrent controls for the study. Hence, have used 5 animals per group. Metabolites that showed significant differences between THC administered and control (vector administered) groups were identified by combining the results of students t-test (p < 0.05), fold change (FC > 2) and variable importance in projection values (VIP > 1). To address the cases when the quantified metabolites fail to satisfy the normality and equality of variance based on Kolmogorov-Smirnov test and Levene’s test respectively, we used the non-parametric Kruskal-Wallis test to determine metabolite differences between the THC-administered and the vector-control groups. The acquired p-values were corrected for multiple testing using Benjamini and Hochberg False Discovery Rate (FDR)40, a method applied previously for untargeted metabolomic analysis41.
Results
Analysis of fecal metabolite profile shifts upon acute THC administration
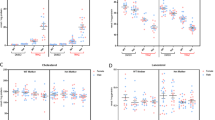
The experimental design of the experiment is illustrated in Fig. 1. To understand the acute effects of THC administration on gut metabolite profiles in C57BL/6 mice, we compared their fecal metabolite profiles 24 h after THC administration (sampling time point denoted here as ‘1X’). The results from our initial examination of base peak intensity chromatographs (SI Fig. 2) are shown as volcano plots in Fig. 2a, which indicated a shift in the profiles upon THC administration. An unsupervised evaluation of metabolic signatures was conducted using the indices PCA1 and PCA2 that were obtained upon PCA analysis by reduction of the multi-dimensional datasets to optimized and comparable datasets. As shown in Fig. 2b, the PCA scores scatter plot demonstrates a clear separation between the THC-treated and control fecal samples. Following PCA analysis, the identification of differential metabolites was performed using supervised partial least square- discriminant analysis (PLS-DA) on the MS data to predict the class membership and assess the significance of class discrimination. As shown in Fig. 2c, the PLS-DA scores scatter plot suggested that THC administration again demonstrates a significant class separation between the THC-treated and control fecal samples.
Study design. Fecal samples (n = 5) for analyses were collected at 2 time-points, 1X and 5X. The injections of THC and vehicle were administered intraperitoneally ~48 h apart (denoted dashed lines) Five administrations were conducted. Fecal and intestinal tissue samples were collected at time points 1X and 5X, which were 24 h after the first and the fifth administrations respectively.
Fecal metabolome changes in 1X samples. (a) Volcano plots enabling the visualization of metabolites showing differential abundance. These were selected based on fold change (X-axis) and p-value in (Y-axis). The m/z values (highlighted in pink) represent a fold change of ≥0.5 or ≤2.0 and p-value ≤ 0.05 in THC administered mice compared to the vehicle controls and were selected for further characterization. (b) PCA score scatter plots based on fecal metabolic profiling of THC (n = 5) and control (n = 5) mice c) PLS- DA score plots based on detected fecal metabolites from THC administered (n = 5) and control (n = 5) mice. T, THC administered mice, C, control mice.
Analysis of fecal metabolite profile shifts upon short-term THC administration
To understand the short-term effects of THC administration on gut metabolite profiles in C57BL/6 mice, we compared fecal metabolite profiles 24 h after five THC injections(sample denoted here as ‘5X’). The volcano plot (Fig. 3a) summarizing our initial examination of the fecal metabolites indicates a shift in fecal metabolite profiles with this multiple exposure protocol (base peak intensity chromatographs of fecal metabolites of 5x are shown in SI Fig. 2). The separation between the fecal metabolite profiles of the THC administered and control groups was further confirmed by the PCA scores scatter plot (Fig. 3b). Finally, the supervised PLS-DA scores obtained from the MS data was used to predict the class membership and demonstrate the significance of class discrimination (Fig. 3c).
Fecal metabolome changes in 5X samples. (a) Volcano plots showing the metabolites with differential abundance between THC treated mice and the controls. These were selected based on fold change (X-axis) and p-value in (Y-axis). The m/z values (highlighted in pink) represent a fold change of ≥0.5 or ≤2.0 and p-value ≤ 0.05 in THC administered mice compared to the vehicle controls and were selected for further characterization. (b) PCA score scatter plots based on fecal metabolic profiling of THC (n = 5) and control (n = 5) mice c) PLS- DA score plots based on detected fecal metabolites from THC administered (n = 5) and control (n = 5) mice. T, THC administered mice, C, control mice.
Identification of potential biomarkers
From the comparisons of fecal metabolite profiles between THC administered and the control groups, a group of ‘significant metabolites’ was identified using the criteria of VIP value > 1. The higher VIP values were indicative of a higher contribution from these metabolites toward the differential profiles between the THC and the control groups. A list of these metabolites from the comparisons of 1x with their controls and the 5x with their controls is provided in Table 1 a and b. Along with the VIP values for each metabolite, Table 1 also indicates the fold change of the increase or decrease in the metabolite concentration upon THC administration. The list of identified significant metabolites that were identified differentially upon THC administration was entirely different between samples 1X and 5X, suggesting that acute and short-term administrations have different functional impacts on the mouse gut.
Interpretation of metabolic signatures
To investigate the latent relationships of the differential metabolites listed in Table 1 and gain insights into metabolite enrichment representing specific metabolic pathway and molecular network perturbations induced by THC exposure, the information of the significant metabolites was imported to the MBRole platform. The enrichment analysis of the metabolic signatures was performed on Mus musculus background using annotations from the KEGG42 and HMDB databases43. While the former database primarily annotates metabolites with their associated pathways and enzymes, the later annotates metabolites with diseases, pathways, tissues, biofluids and the cellular localization. Figure 4 summarizes our observations from this enrichment analysis and indicates the potential metabolic processes that are impacted by THC administration, which in turn, could help explain its therapeutic impact on many diseases. Specifically, acute administration of THC demonstrated a positive correlation with the metabolic intermediates of Glycerolipid metabolism, PI3K/AKT/mTOR and lysophopholipid signaling, opioid peptide biosynthesis and endocannabinoid signaling. Also, the acute administration was negatively correlated with the metabolite intermediates of nicotinate degradation. Nicotinate being the precursor for the generation of nicotinamide adenine nucleotide (NAD+)44, our study suggests a positive effect of THC on cellular NAD+ levels and upregulation of systemic NAD+ is demonstrated to have profound health beneficial effects45,46,47. Short-term administration of THC was positively correlated with the fatty acid degradation pathway, sphingosine metabolism, caffeine metabolism as well as endocannabinoid signaling. Additionally, the short-term THC administration was negatively correlated with xenobiotic metabolism. These processes have significant cross-talks with several important metabolic processes such as amino acid, carbohydrate, and nucleotide metabolism. Specifically, the glutathione metabolism and branched chain amino acid (BCAA) pathways are highly enriched. Interestingly, dysregulation of these metabolic pathways are known to be associated with the etiology of diabetes, obesity, cancer, and neurodegeneration48,49,50. Our future work will investigate the specific metabolite changes in various preclinical models such as mice models for acute and short-term intestinal inflammation. Those studies will further elucidate the mechanistic details of the THC-mediated health benefits.
Functional pathway analysis showing the major pathways that are upregulated or downregulated upon administration of THC. Biological relationships of the pathways were adapted from KEGG, Lipid Maps and HMDB database based on the identification of the metabolite markers corresponding to the pathways. Based on number of identified metabolites with >2 fold increase/decrease, we indicate the processes that are upregulated with green circles and the processes that are downregulated with red circles. The green highlighted zone show the metabolic pathways for which metabolite enrichment was observed both in fecal and intestinal tissue metabolite profiling.
Metabolite profiling of intestinal tissues: validation of fecal metabolite profiles
Finally, to determine whether our metabolome analysis of fecal samples represent changes in host metabolism induced by THC administration, we performed similar untargeted metabolite profiling intestinal tissues obtained from 1X and 5X THC administered mice and the corresponding vector controls. The list of ‘significant metabolites’ identified from these comparisons using the criteria of FC > 2, VIP > 1 and adjusted P-value < 0.05 are shown in Table 2; associated volcano plots, PCA plots and the PLS-DA plots are included as supplementary Information (see SI Fig. 3). Also outlined in Table 2 are the potential metabolic processes that the identified metabolites associate with. Lipid metabolism, especially glycerophospholipid metabolism and fatty acid biosynthesis, emerged as the global metabolic process that was most significantly influenced by THC administration. As shown in Fig. 4, lipid metabolism is a core host metabolic process within the global host metabolic network. Results are consistent with the hypothesis that the altered fecal metabolite profiles seen upon 1X and 5X THC administrations, at least in part, were reflective of the THC-mediated alteration in host metabolism.
Discussion
This study highlights the metabolic changes induced by acute and short-term administration of THC in the gut of a murine model that has historically been used to demonstrate the positive health impacts of THC. To study these metabolic changes, comparative metabonomic profiling of fecal samples of THC-administered mice, and vector-administered mice were performed using a highly sensitive, accurate, and precise UPLC-ESI-QTOF-MS-based approach that has broad applications in metabonomic studies51,52.
With this, we have shown here that lipid metabolism, especially glycerophospholipid metabolism and fatty acid biosynthesis, is a key metabolic pathway targeted by THC following i.p. administration. Importantly, this pathway is intricately connected with several health disorders that are protected by THC; examples include Parkinson disease53, schizophrenia54, brain ischemia55, multiple sclerosis56 and cancer development57,58. Glycerophospholipids are precursors for several lipid mediators that, in collaboration with sphingolipids, participate in major signal transduction processes (see review by Farooqui et al.59) and along with sphingolipid metabolism, are functionally linked with several physiological and pathophysiological conditions that include but are not limited to pain, inflammation, metabolic syndrome, fibrosis, fertility, cancer and autoimmune and neurodegenerative disorders60. Others have also shown that glycerophospholipid and sphingolipid metabolism are the most significantly impaired pathways associated with the atherosclerosis progression61,62. Much of the protective role of cannabinoids on atherosclerotic coronary heart disease involves 15-lipoxygenase inhibitory activity, which in turn prevent lipid peroxidation, oxidative stress and atherosclerosis63. Based on our findings it is reasonable that THC-mediated protection against atherosclerosis and cardiovascular disorders can be linked to its regulatory effects on of glycerophospholipid and sphingolipid metabolism. We have conducted this study using adolescent mice to keep experimental consistency with our previous reports. While this age may seem irrelevant for some of the neurological disorders discussed above, we point out here that both young and adult mice have been used to understand the therapeutic impacts of THC on neuroinflammation and the associated health disorders such as autoimmune encephalitis64, Alzheimer’s disease65 and Parkinson’s disease66. Interestingly the increase in anti-inflammatory cytokine release in the brain of young mice can be mimicked by peripheral immune cells67.
Fecal metabolomics revealed an influence of THC on some additional major metabolic pathways which although connected with lipid metabolism, were not highlighted in our tissue metabolomic study. For example, a critical metabolite that feeds into Sphingolipid metabolism is L-serine, which is a metabolic output from the glycine, serine, and threonine metabolism68. The glycine, serine and threonine metabolic pathway feeds phosphatidylethanolamine to glycerophospholipid metabolism69. An upregulation of sphingolipid and glycerophospholipid metabolism, therefore, suggests an upregulation in Serine metabolism as well. Reduction of 2E-methyl glutaconic acid and tiglyglycine was observed upon 1X administration. These metabolites are often detected in human urine samples when the catabolism of branched-chain amino acids (BCAA) (especially isoleucine) is impaired70,71, suggesting that THC possibly influences BCAA catabolism. Emerging evidence supports the importance of BCAA catabolism in lowering the risk of type-2 diabetes72. While a previous study has shown that cannabidiol significantly reduces the incidence of diabetes in non-obese diabetic mice73, the relation between marijuana use and diabetes remains unclear. We also noted a significant reduction 2- methylene glutarate upon 1X administration suggesting downregulation of the metabolite flow from nicotinate degradation into pyruvate metabolism. This observation is in line with two recent reports that demonstrate a modulatory effect of cannabinoids and cannabinoid receptors on pyruvate (and energy) metabolism: (i) a report from Mendizabal-Zubiaga et al.74 which showed that expression of pyruvate metabolism genes increased in the striated muscle cells of CB1-knockout mice and, (ii) a report by Arrabal et al., which showed that pharmacological blockage CB1 was able to upregulate pyruvate metabolism enzymes75. It is hypothesized that such modulatory effects of THC and cannabinoids on energy metabolism may in part, contribute to their anti-tumor effects. Finally, an increased occurrence of two endogenous peptides upon 1X administration suggesting an activation of the endogenous opioid system. These peptides have receptors widely distributed in the central and peripheral nervous system and play key roles in immunity76, pain modulation64, emotion and stress response65, gut functioning66, neuroprotection with important implications in Parkinson’s disease67.
We point out here that, our study being untargeted in nature had three limitations that are typical for untargeted metabolomics: (1) a bias toward high-abundant metabolites (typical for LC-MS/MS), (2) the influence from exogenous metabolites such as those from gut microbiota (a common issue in fecal metabolome analysis) and (3) high-throughput analysis of samples without authentic standards, which although gives the advantage of the absence of a priori decisions, may lead to quantitative inaccuracy and in some cases compromise metabolite identity. The very high fold changes of enriched metabolites in fecal metabolite profiling could be either reflective of the influence of gut microbial metabolites while the differential abundance of certain metabolites only at one time point may indicate a bias toward high abundant metabolites. Regardless of these limitations, the strength of our study was our ability to conduct a comparative metabolomic examination of the fecal and intestinal tissue matrices (THC treated versus non-treated animals) in a holistic unbiased manner, which was helpful to test our central hypothesis and obtain a global understanding of how THC influences the host metabolic network. This provides us a scientific premise for developing new hypotheses for our future targeted metabolomic studies with diseased models. Such studies will focus on the cause-effect nature of the relationship between THC and the metabolic pathways identified in this study, under different pathophysiological conditions.
References
Hill, K. P. Medical marijuana for treatment of chronic pain and other medical and psychiatric problems: a clinical review. Jama 313, 2474–2483 (2015).
Koppel, B. S. et al. Systematic review: Efficacy and safety of medical marijuana in selected neurologic disorders Report of the Guideline Development Subcommittee of the American Academy of Neurology. Neurology 82, 1556–1563 (2014).
Whiting, P. F. et al. Cannabinoids for medical use: a systematic review and meta-analysis. Jama 313, 2456–2473 (2015).
ProCon. Should Marijuana Be a Medical Option?, https://medicalmarijuana.procon.org/ (2017).
Elikkottil, J., Gupta, P. & Gupta, K. The analgesic potential of cannabinoids. J Opioid Manag 5, 341–357 (2009).
Nahas, G., Harvey, D. J., Sutin, K., Turndorf, H. & Cancro, R. A molecular basis of the therapeutic and psychoactive properties of cannabis (delta9-tetrahydrocannabinol). Prog Neuropsychopharmacol Biol Psychiatry 26, 721–730 (2002).
Paris, M. Cannabis therapy. Ann Pharm Fr 60, 271–273 (2002).
Walsh, D., Nelson, K. A. & Mahmoud, F. A. Established and potential therapeutic applications of cannabinoids in oncology. Support Care Cancer 11, 137–143, https://doi.org/10.1007/s00520-002-0387-7 (2003).
Nagarkatti, P., Pandey, R., Rieder, S. A., Hegde, V. L. & Nagarkatti, M. Cannabinoids as novel anti-inflammatory drugs. Future Med Chem 1, 1333–1349, https://doi.org/10.4155/fmc.09.93 (2009).
Costa, B. On the pharmacological properties of Delta9-tetrahydrocannabinol (THC). Chemistry & biodiversity 4, 1664–1677, https://doi.org/10.1002/cbdv.200790146 (2007).
Wilkinson, J. D. et al. Medicinal cannabis: is delta9-tetrahydrocannabinol necessary for all its effects? The Journal of pharmacy and pharmacology 55, 1687–1694, https://doi.org/10.1211/0022357022304 (2003).
Borgelt, L. M., Franson, K. L., Nussbaum, A. M. & Wang, G. S. The pharmacologic and clinical effects of medical cannabis. Pharmacotherapy 33, 195–209, https://doi.org/10.1002/phar.1187 (2013).
Cascini, F., Aiello, C. & Di Tanna, G. Increasing delta-9-tetrahydrocannabinol (Delta-9-THC) content in herbal cannabis over time: systematic review and meta-analysis. Current drug abuse reviews 5, 32–40 (2012).
Mehmedic, Z. et al. Potency trends of Delta9-THC and other cannabinoids in confiscated cannabis preparations from 1993 to 2008. Journal of forensic sciences 55, 1209–1217, https://doi.org/10.1111/j.1556-4029.2010.01441.x (2010).
Garcia-Planella, E. et al. Use of complementary and alternative medicine and drug abuse in patients with inflammatory bowel disease. Med Clin (Barc) 128, 45–48 (2007).
Lal, S. et al. Cannabis use amongst patients with inflammatory bowel disease. Eur J Gastroenterol Hepatol 23, 891–896, https://doi.org/10.1097/MEG.0b013e328349bb4c (2011).
Ravikoff Allegretti, J., Courtwright, A., Lucci, M., Korzenik, J. R. & Levine, J. Marijuana use patterns among patients with inflammatory bowel disease. Inflamm Bowel Dis 19, 2809–2814, https://doi.org/10.1097/01.MIB.0000435851.94391.37 (2013).
Storr, M., Devlin, S., Kaplan, G. G., Panaccione, R. & Andrews, C. N. Cannabis use provides symptom relief in patients with inflammatory bowel disease but is associated with worse disease prognosis in patients with Crohn’s disease. Inflamm Bowel Dis 20, 472–480, https://doi.org/10.1097/01.MIB.0000440982.79036.d6 (2014).
Ihenetu, K., Molleman, A., Parsons, M. E. & Whelan, C. J. Inhibition of interleukin-8 release in the human colonic epithelial cell line HT-29 by cannabinoids. Eur J Pharmacol 458, 207–215 (2003).
Storr, M. A. et al. Activation of the cannabinoid 2 receptor (CB2) protects against experimental colitis. Inflamm Bowel Dis 15, 1678–1685, https://doi.org/10.1002/ibd.20960 (2009).
Wright, K. et al. Differential expression of cannabinoid receptors in the human colon: cannabinoids promote epithelial wound healing. Gastroenterology 129, 437–453, https://doi.org/10.1016/j.gastro.2005.05.026 (2005).
Di Marzo, V., Bifulco, M. & De Petrocellis, L. The endocannabinoid system and its therapeutic exploitation. Nature reviews. Drug discovery 3, 771–784, https://doi.org/10.1038/nrd1495 (2004).
McKallip, R. J., Lombard, C., Martin, B. R., Nagarkatti, M. & Nagarkatti, P. S. Delta(9)-tetrahydrocannabinol-induced apoptosis in the thymus and spleen as a mechanism of immunosuppression in vitro and in vivo. J Pharmacol Exp Ther 302, 451–465, https://doi.org/10.1124/jpet.102.033506 (2002).
Sido, J. M., Jackson, A. R., Nagarkatti, P. S. & Nagarkatti, M. Marijuana-derived Delta-9-tetrahydrocannabinol suppresses Th1/Th17 cell-mediated delayed-type hypersensitivity through microRNA regulation. J Mol Med (Berl) 94, 1039–1051, https://doi.org/10.1007/s00109-016-1404-5 (2016).
Sido, J. M., Nagarkatti, P. S. & Nagarkatti, M. Delta(9)-Tetrahydrocannabinol attenuates allogeneic host-versus-graft response and delays skin graft rejection through activation of cannabinoid receptor 1 and induction of myeloid-derived suppressor cells. J Leukoc Biol 98, 435–447, https://doi.org/10.1189/jlb.3A0115-030RR (2015).
Sido, J. M., Yang, X., Nagarkatti, P. S. & Nagarkatti, M. Delta9-Tetrahydrocannabinol-mediated epigenetic modifications elicit myeloid-derived suppressor cell activation via STAT3/S100A8. J Leukoc Biol 97, 677–688, https://doi.org/10.1189/jlb.1A1014-479R (2015).
Yang, X., Bam, M., Nagarkatti, P. S. & Nagarkatti, M. RNA-seq Analysis of delta9-Tetrahydrocannabinol-treated T Cells Reveals Altered Gene Expression Profiles That Regulate Immune Response and Cell Proliferation. J Biol Chem 291, 15460–15472, https://doi.org/10.1074/jbc.M116.719179 (2016).
Lefever, T. W., Marusich, J. A., Antonazzo, K. R. & Wiley, J. L. Evaluation of WIN 55,212-2 self-administration in rats as a potential cannabinoid abuse liability model. Pharmacol Biochem Behav 118, 30–35, https://doi.org/10.1016/j.pbb.2014.01.002 (2014).
Reagan-Shaw, S., Nihal, M. & Ahmad, N. Dose translation from animal to human studies revisited. FASEB J 22, 659–661, https://doi.org/10.1096/fj.07-9574LSF (2008).
Fraguas-Sanchez, A. I. & Torres-Suarez, A. I. Medical Use of Cannabinoids. Drugs 78, 1665–1703, https://doi.org/10.1007/s40265-018-0996-1 (2018).
Pollmann, W. & Feneberg, W. Current management of pain associated with multiple sclerosis. CNS Drugs 22, 291–324, https://doi.org/10.2165/00023210-200822040-00003 (2008).
Zajicek, J. P. & Apostu, V. I. Role of cannabinoids in multiple sclerosis. CNS Drugs 25, 187–201, https://doi.org/10.2165/11539000-000000000-00000 (2011).
FDA. Marinol, https://www.accessdata.fda.gov/drugsatfda_docs/label/2005/018651s021lbl.pdf.
Watanabe, K., Matsunaga, T., Narimatsu, S., Yamamoto, I. & Yoshimura, H. Sex difference in hepatic microsomal aldehyde oxygenase activity in different strains of mice. Res Commun Chem Pathol Pharmacol 78, 373–376 (1992).
Wagner, E. J. Sex differences in cannabinoid-regulated biology: A focus on energy homeostasis. Front Neuroendocrinol 40, 101–109, https://doi.org/10.1016/j.yfrne.2016.01.003 (2016).
Wallin, M. T. et al. The prevalence of MS in the United States: A population-based estimate using health claims data. Neurology 92, e1029–e1040, https://doi.org/10.1212/WNL.0000000000007035 (2019).
Chagoyen, M. & Pazos, F. MBRole: enrichment analysis of metabolomic data. Bioinformatics 27, 730–731, https://doi.org/10.1093/bioinformatics/btr001 (2011).
Lopez-Ibanez, J., Pazos, F. & Chagoyen, M. MBROLE 2.0-functional enrichment of chemical compounds. Nucleic Acids Res 44, W201–204, https://doi.org/10.1093/nar/gkw253 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal statistical society: series B (Methodological) 57, 289–300 (1995).
Kim, H. H. et al. Metabolomic profiling of CSF in multiple sclerosis and neuromyelitis optica spectrum disorder by nuclear magnetic resonance. PLoS One 12, e0181758, https://doi.org/10.1371/journal.pone.0181758 (2017).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Wishart, D. S. et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res 46, D608–D617, https://doi.org/10.1093/nar/gkx1089 (2018).
Greenbaum, A. L. & Pinder, S. The pathway of biosynthesis of nicotinamide-adenine dinucleotide in rat mammary gland. Biochem J 107, 55–62 (1968).
Verdin, E. NAD(+) in aging, metabolism, and neurodegeneration. Science 350, 1208–1213, https://doi.org/10.1126/science.aac4854 (2015).
Tarrago, M. G. et al. A Potent and Specific CD38 Inhibitor Ameliorates Age-Related Metabolic Dysfunction by Reversing Tissue NAD(+) Decline. Cell Metab 27, 1081–1095 e1010, https://doi.org/10.1016/j.cmet.2018.03.016 (2018).
Zhang, H. et al. NAD(+) repletion improves mitochondrial and stem cell function and enhances life span in mice. Science 352, 1436–1443, https://doi.org/10.1126/science.aaf2693 (2016).
Schulz, J. B., Lindenau, J., Seyfried, J. & Dichgans, J. Glutathione, oxidative stress and neurodegeneration. Eur J Biochem 267, 4904–4911, https://doi.org/10.1046/j.1432-1327.2000.01595.x (2000).
Newgard, C. B. et al. A Branched-Chain Amino Acid-Related Metabolic Signature that Differentiates Obese and Lean Humans and Contributes to Insulin Resistance (vol 9, pg 311, 2009). Cell Metab 9, 565–566, https://doi.org/10.1016/j.cmet.2009.05.001 (2009).
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: Transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. P Natl Acad Sci USA 104, 19345–19350, https://doi.org/10.1073/pnas.0709747104 (2007).
Li, Y. et al. Analysis of 2-(2-Phenylethyl)chromones by UPLC-ESI-QTOF-MS and Multivariate Statistical Methods in Wild and Cultivated Agarwood. Int J Mol Sci 17, https://doi.org/10.3390/ijms17050771 (2016).
Zhang, X. et al. Metabolite profiling of plasma and urine from rats with TNBS-induced acute colitis using UPLC-ESI-QTOF-MS-based metabonomics–a pilot study. FEBS J 279, 2322–2338, https://doi.org/10.1111/j.1742-4658.2012.08612.x (2012).
Nakano, N. et al. PI3K/AKT signaling mediated by G proteincoupled receptors is involved in neurodegenerative Parkinson’s disease (Review). Int J Mol Med 39, 253–260, https://doi.org/10.3892/ijmm.2016.2833 (2017).
Norton, N. et al. Association analysis of AKT1 and schizophrenia in a UK case control sample. Schizophr Res 93, 58–65, https://doi.org/10.1016/j.schres.2007.02.006 (2007).
Kisoh, K. et al. Involvement of GSK-3beta Phosphorylation Through PI3-K/Akt in Cerebral Ischemia-Induced Neurogenesis in Rats. Mol Neurobiol 54, 7917–7927, https://doi.org/10.1007/s12035-016-0290-8 (2017).
Sattler, M. B. et al. Neuroprotective effects and intracellular signaling pathways of erythropoietin in a rat model of multiple sclerosis. Cell Death Differ 11(Suppl 2), S181–192, https://doi.org/10.1038/sj.cdd.4401504 (2004).
Bruhn, M. A., Pearson, R. B., Hannan, R. D. & Sheppard, K. E. AKT-independent PI3-K signaling in cancer - emerging role for SGK3. Cancer Manag Res 5, 281–292, https://doi.org/10.2147/CMAR.S35178 (2013).
Stegeman, H., Span, P. N., Kaanders, J. H. & Bussink, J. Improving chemoradiation efficacy by PI3-K/AKT inhibition. Cancer Treat Rev 40, 1182–1191, https://doi.org/10.1016/j.ctrv.2014.09.005 (2014).
Farooqui, A. A., Horrocks, L. A. & Farooqui, T. Interactions between neural membrane glycerophospholipid and sphingolipid mediators: a recipe for neural cell survival or suicide. Journal of neuroscience research 85, 1834–1850 (2007).
Gardell, S. E., Dubin, A. E. & Chun, J. Emerging medicinal roles for lysophospholipid signaling. Trends Mol Med 12, 65–75, https://doi.org/10.1016/j.molmed.2005.12.001 (2006).
Jiang, X. C. & Liu, J. Sphingolipid metabolism and atherosclerosis. Handb Exp Pharmacol, 133–146, https://doi.org/10.1007/978-3-7091-1511-4_7 (2013).
Dang, V. T., Huang, A., Zhong, L. H., Shi, Y. & Werstuck, G. H. Comprehensive Plasma Metabolomic Analyses of Atherosclerotic Progression Reveal Alterations in Glycerophospholipid and Sphingolipid Metabolism in Apolipoprotein E-deficient Mice. Sci Rep 6, 35037, https://doi.org/10.1038/srep35037 (2016).
Singla, S., Sachdeva, R. & Mehta, J. L. Cannabinoids and atherosclerotic coronary heart disease. Clin Cardiol 35, 329–335, https://doi.org/10.1002/clc.21962 (2012).
Mayer, E. & Saper, C. Pain modulation: expectation, opioid analgesia and virtual pain. The Biological Basis for Mind Body Interactions 122, 245 (2000).
Drolet, G. et al. Role of endogenous opioid system in the regulation of the stress response. Progress in Neuro-Psychopharmacology and Biological Psychiatry 25, 729–741 (2001).
Kromer, W. Endogenous opioids, the enteric nervous system and gut motility. Digestive Diseases 8, 361–373 (1990).
Benarroch, E. E. Endogenous opioid systems Current concepts and clinical correlations. Neurology 79, 807–814 (2012).
Momin, A. A. et al. A method for visualization of “omic” datasets for sphingolipid metabolism to predict potentially interesting differences. J Lipid Res 52, 1073–1083, https://doi.org/10.1194/jlr.M010454 (2011).
Zhang, H. et al. Metabolomic study of corticosterone-induced cytotoxicity in PC12 cells by ultra performance liquid chromatography-quadrupole/time-of-flight mass spectrometry. Molecular BioSystems 12, 902–913 (2016).
Duran, M. et al. The identification of (E)‐2‐methylglutaconic acid, a new isoleucine metabolite, in the urine of patients with β‐ketothiolase deficiency, propionic acidaemia and methylmalonic acidaemia. Biological Mass Spectrometry 9, 1–5 (1982).
Bennett, M. J., Powell, S., Swartling, D. J. & Gibson, K. M. Tiglylglycine excreted in urine in disorders of isoleucine metabolism and the respiratory chain measured by stable isotope dilution GC-MS. Clinical chemistry 40, 1879–1883 (1994).
Lotta, L. A. et al. Genetic Predisposition to an Impaired Metabolism of the Branched-Chain Amino Acids and Risk of Type 2 Diabetes: A Mendelian Randomisation Analysis. PLoS Med 13, e1002179, https://doi.org/10.1371/journal.pmed.1002179 (2016).
Weiss, L. et al. Cannabidiol lowers incidence of diabetes in non-obese diabetic mice. Autoimmunity 39, 143–151, https://doi.org/10.1080/08916930500356674 (2006).
Mendizabal-Zubiaga, J. et al. Cannabinoid CB1 Receptors Are Localized in Striated Muscle Mitochondria and Regulate Mitochondrial Respiration. Front Physiol 7, 476, https://doi.org/10.3389/fphys.2016.00476 (2016).
Arrabal, S. et al. Pharmacological blockade of cannabinoid CB1 receptors in diet-induced obesity regulates mitochondrial dihydrolipoamide dehydrogenase in muscle. PloS one 10, e0145244 (2015).
Carr, D. J. The role of endogenous opioids and their receptors in the immune system. Proceedings of the Society for Experimental Biology and Medicine 198, 710–720 (1991).
Cook, J. A. et al. Mass Spectrometry-Based Metabolomics Identifies Longitudinal Urinary Metabolite Profiles Predictive of Radiation-Induced Cancer. Cancer Res 76, 1569–1577, https://doi.org/10.1158/0008-5472.CAN-15-2416 (2016).
Acknowledgements
A pilot project award to A.C. from the Center of Biomedical Research Excellence (Center for Dietary Supplements and Inflammation) at University of South Carolina, Columbia, funded this study. The parent grant # 1P20GM103641 from the National Institutes of Health (NIH) and the National Institute of General Medical Sciences (NIGMS) to P.N. funded this pilot project. Creative Proteomics Inc. provided support in the form of salary for author L.C., but did not have any role in the research design, data collection and analysis, and preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.C., M.N. and P.N. conceptualized the project and designed the experiments. M.N. and P.N. provided the resources for the project. W.B. conducted the THC intervention treatment. M.O., T.D., P.M.G., M.H.O., L.C., C.M., R.J. and P.D.R.M. conducted the metabolite profiling experiments. L.C. and P.C. performed the statistical analysis of the metabolite profiling data. M.O., P.M.G., L.C., K.B., Q.W., M.S., L.J.H., P.D.R.M. and A.C. collaboratively conducted the interpretation of the metabolite profiles. A.C., P.M.G., P.C., M.S. and L.J.H. wrote the manuscript, and the final manuscript was reviewed and approved by all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oza, M., Becker, W., Gummadidala, P.M. et al. Acute and short-term administrations of delta-9-tetrahydrocannabinol modulate major gut metabolomic regulatory pathways in C57BL/6 mice. Sci Rep 9, 10520 (2019). https://doi.org/10.1038/s41598-019-46478-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46478-0
This article is cited by
-
Targeting the endocannabinoid system for the treatment of abdominal pain in irritable bowel syndrome
Nature Reviews Gastroenterology & Hepatology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.