Abstract
The zoonotic Plasmodium knowlesi parasite is the most common cause of human malaria in Malaysia. Genetic analysis has shown that the parasites are divided into three subpopulations according to their geographic origin (Peninsular or Borneo) and, in Borneo, their macaque host (Macaca fascicularis or M. nemestrina). Whilst evidence suggests that genetic exchange events have occurred between the two Borneo subpopulations, the picture is unclear in less studied Peninsular strains. One difficulty is that P. knowlesi infected individuals tend to present with low parasitaemia leading to samples with insufficient DNA for whole genome sequencing. Here, using a parasite selective whole genome amplification approach on unprocessed blood samples, we were able to analyse recent genomes sourced from both Peninsular Malaysia and Borneo. The analysis provides evidence that recombination events are present in the Peninsular Malaysia parasite subpopulation, which have acquired fragments of the M. nemestrina associated subpopulation genotype, including the DBPβ and NBPXa erythrocyte invasion genes. The NBPXb invasion gene has also been exchanged within the macaque host-associated subpopulations of Malaysian Borneo. Our work provides strong evidence that exchange events are far more ubiquitous than expected and should be taken into consideration when studying the highly complex P. knowlesi population structure.
Similar content being viewed by others
Introduction
Plasmodium knowlesi, a common malaria parasite of long-tailed (Macaca fascicularis) and pig-tailed (M. nemestrina) macaques, is now recognized as a significant cause of human malaria, with cases reported across all countries of Southeast Asia1,2,3,4. P. knowlesi is the predominant cause of malaria in Malaysia1,2,3,4. The Plasmodium species can be transmitted through several vector species, including Anopheles latens and A. balbacensis in Malaysian Borneo5,6,7 and A. hackeri and A. cracens in Peninsular Malaysia8. Severe disease occurs in 6–9% of clinical presentations and fatalities have been described1,9,10. Rapid human population growth and deforestation, which can drive both encroachment on wild macaque habitats and vector distribution changes11, are thought to increase human-macaque contact, change transmission dynamics, and drive up the incidence of human P. knowlesi infections12,13. Regional elimination efforts have targeted P. falciparum and P. vivax transmission, with significant progress being demonstrated by the near-elimination of these Plasmodium species from areas such as Malaysian Borneo3,4,14. However, due to the difficulties in reducing the monkey parasite reservoir, it is unclear if similar control approaches are able to limit the risk of humans acquiring P. knowlesi malaria15,16.
Appropriate molecular tools and sampling are needed to assist surveillance of P. knowlesi by malaria control programs, and to understand its genetic diversity and transmission. P. knowlesi genomics could lead to biological insights that inform control measures. Advances in whole genome sequencing (WGS) technologies have led to the characterization of single nucleotide polymorphisms (SNPs) across P. falciparum and P. vivax, with an improved understanding of their population structure and diversity, as well as loci underpinning drug resistance (e.g.17,18,19,20,21,22). For P. knowlesi WGS studies, the number of high quality isolates analysed in each study has been small (n < 70)23,24,25. However, these studies have revealed that the P. knowlesi genome is more polymorphic than P. falciparum, and that three main subpopulations exist based on geographical source (Peninsular-Malaysia vs. Malaysian Borneo) and, within Malaysian Borneo, different hosts (M. nemestrina [Mn-Pk] and M. fascicularis [Mf-Pk] macaques, and humans)23,24,25. These studies have also provided evidence that P. knowlesi nuclear genomes are not genetically isolated, and there have been chromosomal-segment exchanges between subpopulations23,24,25. This observation points to subpopulations that have diverged in isolation and then re-connected, possibly due to deforestation and disruption of wild macaque habitats13. The resulting genetic mosaics reveal traits selected by host-vector-parasite interactions in a setting of ecological transition12,13. However, despite these insights, P. knowlesi isolates from both macaques and humans in Peninsular Malaysia are under-represented in analyses, and the genetic diversity in that geographical region is less clear.
One roadblock to large-scale genomic studies of clinical P. knowlesi parasites is that the majority of infections have a low parasitaemia, leading to samples with high levels of human compared to parasite DNA. Until now the WGS data for Plasmodium parasites has been obtained from venous blood of clinical cases that were filtered to remove human leukocytes, and therefore reduce human DNA “contamination”. However, this approach does not always yield sufficient parasite DNA for WGS. Recently, a selective whole genome amplification (SWGA) strategy has been used to sequence P. falciparum and P. vivax genomes from non-filtered blood and from dried blood spots of clinical samples26,27,28. The SWGA method uses oligonucleotides that preferentially bind with high frequency to the target DNA, but less frequently to the “contaminating” genome29. The high fidelity Phi29 polymerase is then used to amplify long segments of DNA. Here, we developed a SWGA approach for P. knowlesi, and sequenced 26 isolates across Malaysia, including from Peninsular and Borneo, revealing new insights into the population structure and evolution of this parasite.
Results
Selective whole genome amplification of P. knowlesi parasite DNA from clinical samples
We performed SWGA on P. knowlesi DNA samples obtained directly from human blood using six selected primers that specifically amplify the parasite genome and bind less frequently to the human genome (See Methods). The primer set had a mean binding frequency of at least once every 4,826 bp to the P. knowlesi genome, much higher than the once every 40,307 bp to the human genome. Binding sites that are sufficiently near each other, as obtained with the primer set for P. knowlesi, enable the branching and displacement actions of the Phi29 polymerase and increase the success of the genome amplification30.
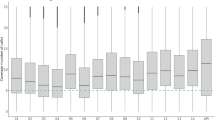
For ten samples, we performed WGS on both the non-amplified and the SWGA DNA. Both sets of samples (with and without amplification) were sequenced at the same theoretical depth, and we observed a significant increase (mean: 7.7-fold greater) in the proportion of reads that mapped to the P. knowlesi A1-H.1 reference genome after DNA amplification (Table 1, showing non-pooled sequencing results). As a result, amplified samples have higher genome coverage (mean: 6.8-fold greater) and a much greater number of callable SNPs (mean: 182-fold greater, with an average of 14,078 SNPs for no SWGA vs. 115,995 SNPs for SWGA) (Table 1). After amplification, higher coverage was observed in genes and intergenic regions (% of positions with a coverage >5-fold; within genes: no SWGA 5.3% vs. SWGA 43.3%; intergenic regions: no SWGA 4.1% vs. SWGA 30.4%) (Table 1). DNA from a further sixteen clinical isolates underwent SWGA and WGS. A trend towards improved coverage in samples with higher parasitaemia was observed (R2 = 0.6, Fig. 1), with superior results for the samples with ≥5,000 parasites/µl, consistent with data from P. vivax and P. falciparum isolates27,28. For the samples with <5,000 parasites/µl, results are more variable and do not correlate with an increase in parasitaemia. For these low-parasitaemia samples, the percentage of the genome coverage in excess of 5-fold ranged from 6% to 43% after amplification, and represent an average increase of 78% in coverage compared to non-amplified samples. This coverage difference led to an average of 66,143 callable SNPs post-amplification and 2,908 SNPs for non-amplified samples (Table 1). The average distribution of the genome coverage for the twenty high quality isolates undergoing SWGA, after applying quality filtering, is shown in Supplementary Fig. 1; a relatively uneven distribution of the coverage can be observed, which is similar to previous studies in other Plasmodium species27,28, but with no specific bias towards genic regions (Table 1). For samples with lower parasite densities, increased sequencing and merging of the resulting reads can lead to improved genome coverage, as shown for P. vivax28. Evidence of mixed infections (multiclonality) was detected in two SWGA samples, demonstrating that the method can amplify more than one clone present in an infection, as was observed for P. vivax amplified samples28.
P. knowlesi genetic variation and clustering of isolates
The sequence data for the 26 new P. knowlesi isolates and 156 previously sequenced samples (see Methods) were mapped to the A1-H.1 reference genome31. The new genomes include recently collected Peninsular Malaysian isolates (n = 5) and clinical isolates from Sabah, Malaysia Borneo (n = 21). From the resulting alignments, 1,741,056 high quality SNPs were identified across the 14 chromosomes. Isolates with high levels of multiplicity of infection (MOI) (>15% of genome with MOI > 1 or >0.0004% of SNPs with mixed calls, see Supplementary Fig. 2) were excluded. In addition, isolates with overall low genome coverage (<30%) were excluded. As a result of these filters, 103 isolates, including 20 of the 26 isolates undergoing SWGA, were carried forward for further analysis (see Supplementary Table 1). A neighbour-joining tree was constructed using the SNP data (Fig. 2) and revealed 3 predominant clusters, consistent with recent findings23,25. In particular, these clusters relate to the specific geographic Peninsular-Malaysia subpopulation (purple, π = 3.4 × 10−9), and Borneo macaque Mn-Pk (green, π = 2.23 × 10−9) and Mf-Pk (blue, π = 3.29 × 10−9) associated subpopulations. The genetic distances between the populations, estimated as the average FST32 for SNPs with minor allele frequencies >0.05, were 0.14 (Mf-Pk vs. Mn-Pk), 0.20 (Mf-Pk vs. Peninsular) and 0.28 (Mn-Pk vs. Peninsular). Furthermore, the tree showed a consistent positioning for SWGA isolates: 4 SWGA Peninsular isolates (red branches) clustered within the Peninsular Malaysia clade (purple), and of the 16 SWGA Sabah isolates, 2 and 14 clustered within the Mn-Pk (green) and Mf-Pk (blue) Borneo clades, respectively. This result demonstrates that the SWGA method can amplify all known sub-populations of P. knowlesi.
Neighbour-Joining tree for 103 P. knowlesi isolates shows three main clusters. The tree shows the expected split into three different clusters associated with: (i) Peninsular Malaysia (purple in tips), (ii) Malaysian Borneo Macaca nemestrina (Mn-Pk, green) and (iii) Malaysian Borneo M. fascicularis (Mf-Pk, blue). The tree also shows the correct positioning of the 4 newly generated Peninsular isolates (in red) within the Peninsular cluster, and the clustering of the 16 new Malaysian Borneo isolates from Sabah (in orange) within either of the Mf-Pk or Mn-Pk associated clusters. Bootstrapping was performed (n = 100) and all the nodes that split the relevant subpopulations presented with a value greater than 90.
Genetic exchange events in P. knowlesi isolates from Peninsular Malaysia
It has been shown that the subpopulations of P. knowlesi in Malaysian Borneo, although presenting a strong genetic differentiation, are not genetically isolated. In particular, we have identified genetic exchanges predominantly between the Mf-Pk and Mn-Pk clusters23. We sought to investigate whether these events are also found in the clinical isolates from Peninsular Malaysia, by estimating SNP nucleotide diversity (SNP π) across the genome in sliding 50 kbp windows. In the two isolates where genome coverage was highest (P137 and P050), we identified several regions with an exceptional increase in similarity with the Mn-Pk cluster and reduced similarity with the Peninsular Malaysia cluster (Fig. 3). Analysis of the haplotypes for each individual isolate confirmed exchange events. The analysis of the individual sequence haplotypes of genes in the identified regions showed mis-clustering in a neighbour-joining tree when compared to the whole genome clustering patterns. These genes are represented in Supplementary Table 2. All the events observed were associated with genetic exchanges from Mn-Pk cluster haplotypes into the Peninsular Malaysia genomes and spanned mostly subtelomeric regions in chromosomes 1, 2, 7, 9, 10, 11, 12, 13 and 14. A high proportion of Plasmodium exported proteins with unknown function were found to be affected by the exchange, as well as tryptophan-rich antigens and lysophospholipases, genes associated with parasite invasion (Normocyte Binding Protein Xa (NBPXa), Duffy Binding Protein beta (DBPβ))33, and a cytoadherence linked asexual protein gene (PKNH_1401300). These results could indicate that the exchange events found in Peninsular Malaysia may affect vertebrate host-related factors in the erythrocytic stages of the parasite life cycle and could potentially impact the invasion of human red blood cells by P. knowlesi.
P. knowlesi isolates with the highest genomic coverage from Peninsular Malaysia (P137 and P050) present with genetic exchange events from the Mn-Pk sub-population. Peninsular isolate P137 was compared to DIM5 (top two panels) as a representative of the Mn-Pk cluster, and DIM6 (second row panels) of the Mf-Pk cluster; these isolates were selected based on sequencing quality and completeness, and an absence of evidence of multiplicity of infection. Isolate P050 was compared to the same isolates in the bottom 4 panels. On the top left panel each green dot represents a 50 kbp section of the DIM5 genome. Its position on the X-axis is defined by the average SNP π obtained by comparing its sequence in a pairwise manner to the same syntenic genome 50 kbp fragment in each isolate in the Mn-Pk cluster; in the Y-axis the average SNP π is compared to the same fragment of the Peninsular isolates. This average SNP π defines the similarity of each dot to the different clusters. The top most right panel represents the same data as the top left most panel with the P137 50 kbp fragments highlighted in purple for clarity. The same analysis was conducted in the second row of panels but using a Mf-Pk cluster isolate and the average SNP π to Mf-Pk as the X-axis. The dashed line represents the linear regression for the coloured dots in each plot, and the regions of interest were identified (in light green in the right panels) by finding the fragments of the Peninsular isolate genomes that presented low similarity to the Peninsular cluster and high similarity to either the Mn-Pk (green) or Mf-Pk (blue). This approach accounts for the highest residuals. After further filtering through the assessment of individual genes, we report the set of results in Supplementary Table 2.
Following the discovery of P. knowlesi invasion-related genes in the exchanged regions, we performed a comprehensive analysis of the genetic diversity harboured by the five reticulocyte binding like (RBL) and Duffy binding protein like (DBP) genes involved in erythrocyte invasion: DBPα (81 isolates, Fig. 4A), DBPβ (89 isolates, Fig. 4B), DBPγ (70 isolates, Fig. 4C), NBPXa (88 isolates, Fig. 5A), and NBPXb (90 isolates, Fig. 5B). For each of the high-quality isolates with coverage in excess of 30-fold across the different genes, we characterised their “invasion” haplotypes (Figs 4 and 5, left) and compared their resulting position on neighbour-joining trees to their expected clustering based on WGS data (Figs 4 and 5, right). For each of the 5 genes, there was evidence of strong genetic divergence of the sequences across the different clusters. Across all 5 genes, the Peninsular Malaysia cluster presented with marginally greater nucleotide diversity (Peninsular Malaysia: mean = 1.88 × 10−5; range = 1.63 × 10−5–2.17 × 10−5) when compared to the other two clusters (Mf-Pk: mean = 1.85 × 10−5, range = 1.15 × 10−5–2.71 × 10−5; Mn-Pk: mean = 1.01 × 10−5, range = 0.78 × 10−5–1.42 × 10−5).
Haplotype plot and neighbour-joining tree for three Duffy Binding Protein invasion genes ((A) DBPα, (B) DBPβ, (C) DBPγ) provides insights into the population dynamics of the different haplotypes. Only isolates with at least 30-fold coverage across the gene were used in each plot: 81 isolates for DBPα, 89 isolates for DBPβ and 70 isolates for DBPγ. A strong genetic divergence of the sequences from the different clusters was found for each of the 3 genes, and the Peninsular cluster had the highest diversity across all 3 genes (A–C). Red stars indicate isolates with differences in subpopulation clustering when compared to whole genome clustering, suggesting a genetic exchange. Bootstrapping was performed (n = 100) and all the nodes that split the relevant subpopulations and/or exchange events presented with a value greater than 84.
Haplotype plots and neighbour-joining trees for two Normocyte Binding Protein invasion genes ((A) NBPXa, (B) NBPXb) provides insights into the population dynamics of the gene haplotypes. Only isolates with at least 30-fold coverage across the gene were used in each plot: 88 isolates for NBPXa and 90 isolates for NBPXb. Red stars indicate isolates with differences in subpopulation clustering when compared to whole genome clustering, suggesting a genetic exchange. For the NBPXa gene (A), evidence of strong genetic divergence of the sequences from the different clusters was found. However, the NBPXb gene (B, right) presented a fairly distinct pattern of diversity. The clusters have small genetic distances between themselves, making the separation between them less obvious. Bootstrapping was performed (n = 100) and all the nodes that split the relevant subpopulations and/or exchange events presented with a value greater than 82.
For all the isolates, DBPα and DBPγ gene sequence-based clusters matched those from whole genome data. For DBPβ, the overall clustering pattern was still present, but one isolate from Peninsular Malaysia (P050; Fig. 4B, right; red star) had strong evidence of genetic exchanges with the Mn-Pk cluster. The longer branch length of the P050 isolate in the tree, where the genetic exchange is found, reveals a stronger genetic difference compared to the other Mn-Pk DBPβ haplotypes, thereby indicating that this exchange event could be non-recent. Analysis of the neighbouring genes revealed that this exchange spanned from approximately 31 kb to 96 kb in chromosome 14, affecting 14 genes. This observation is confirmed by the partial similarity of the P050 haplotype with the Mn-Pk haplotypic patterns (Fig. 4B, left). There was also evidence of genetic exchange events in the NBPXa, which had the lowest overall genetic diversity of all RBL/DBP genes (mean = 1.20 × 10−5). For example, the Peninsular Malaysia isolate P137 appears to have incorporated a haplotype from the Mn-Pk cluster, and its partial similarity to the current NBPXa haplotypes of the Mn-Pk’s clade suggests it could also be a non-recent event (Fig. 5A, right). An analysis of neighbouring genes revealed that the exchange event spanned from approximately 3,157 kb to 3,204 kb in chromosome 14, affecting 8 genes. Finally, NBPXb gene had low diversity, with intra-cluster genetic distances being smaller than those found in the DBP genes. Several genetic exchange events were identified, where 9 out of 33 (27%) of the Mn-Pk cluster isolates presented with Mf-Pk type haplotypes (Fig. 5B, right; red stars). Most of these isolates with genetic exchange evidence clustered together and were separated from the Mf-Pk isolates, which could be a reflection of a unique old genetic exchange event. This observation is consistent with the NBPXb gene, and not its neighbouring loci, being exchanged. The isolate KT233 positioned in the “Mn-Pk” group using all SNPs, has a much similar NBPXb haplotype to those found in the Mf-Pk samples, reflecting a more recent exchange event.
Discussion
The SWGA approach implemented led to reliable sequence data being generated for parasite isolates obtained from unprocessed human blood, and belonging to the three currently known subpopulations of P. knowlesi. This method is cost-effective, does not require sample processing at the time of collection, requires low quantities of input DNA, and is easy to implement. Importantly, the approach permits the genomic analysis of isolates that would otherwise be very difficult to investigate, as demonstrated by the poor WGS results of the non-SWGA DNA when compared with their respective SWGA samples. A neighbour-joining tree based on the SNP data generated revealed 3 predominant clusters, consistent with recent findings, and the positioning of the SWGA isolates confirmed their origin. The four recent Peninsular Malaysia isolates clustered closely with the long-term maintained samples originating from different regions in Peninsular Malaysia and the Philippines. Of the sixteen isolates originating from Sabah, two belonged to the Mn-Pk associated cluster, and fourteen belonged to the Mf-Pk associated cluster. This finding is consistent with the higher proportions of samples circulating in humans belonging to the Mf-Pk cluster34 and confirms the presence of both Borneo macaque host-related subpopulations in Sabah.
Previous population genetics studies on P. knowlesi subpopulations among human and macaque isolates from Malaysian Borneo provided evidence that chromosomal-segment exchanges between subpopulations have occurred recently23. This observation could be indicative of subpopulations that diverged in isolation and have re-connected, possibly due to deforestation and disruption of wild macaque habitats. Up until now these genetic exchanges had only been observed in parasites from Malaysian Borneo10, but the inclusion of recent Peninsular Malaysia isolates allowed us to scan for more widespread events. We identified regions that presented evidence of genetic exchange, in particular, with exceptionally high average SNP diversity when compared to the Peninsular isolates and exceptionally low diversity when compared to Mn-Pk or Mf-Pk isolates. This approach highlighted the presence of such events in two Peninsular samples, where the identified regions were found to be genetic exchanges with Mn-Pk type haplotypes and present across multiple chromosomes. This work shows that genetic exchange events are more widespread than previously thought, and they also affect the geographic subpopulation of Peninsular Malaysia. This finding is consistent with microsatellite analyses that have identified traces of Borneo-associated clusters in regions of Peninsular Malaysia34,35.
Regions identified with genetic exchange events were enriched with large multi-gene families coding for Plasmodium exported proteins and tryptophan-rich antigens, as well as loci associated with erythrocyte invasion by P. knowlesi. These findings contrast with our previous results found in isolates from Borneo23, where the genes involved in genetic exchange events were enriched by mosquito-stage related genes. This difference in gene ontology could suggest that there are different factors driving the exchanges between geographical regions. We found that all known RBL/DBP invasion genes (DBP α, β and γ; NBP Xa and Xb) are highly differentiated and cluster the isolates into three subpopulations. This clustering is consistent with vertebrate host-related factors being one of the main drivers for their genetic differentiation; although the Peninsular subpopulation is assumed to be a geographic subdivision rather than a macaque host-associated cluster. Across all five loci the Mf-Pk group is the most genetically diverse. The DBPα and DBPγ loci did not present with any genetic-exchange patterns. For the NBPXb gene, the three subpopulations were not as strongly differentiated as in other genes, and some of the Mn-Pk isolates (including from Sabah) presented with genetic exchanges with the Mf-Pk subpopulation. These genetic exchanges could be related to the adaptation of the parasites to different vertebrate hosts, especially as it has been shown that the two Borneo subpopulations can be found in both species of macaques24,25,34. Furthermore, the events observed here could involve yet another subpopulation of a Peninsular Mn-Pk type of P. knowlesi, especially as the level of sampling currently is very low and the haplotypes do show a degree of differentiation with the other Mn-Pk haplotypes. Other genes such as DBPβ and NBPXa presented genetic exchange events with Mn-Pk into the Peninsular subpopulation. NBPXa has the lowest level of genetic diversity, which suggests that this gene is highly conserved across subpopulations. This finding is important because NBPXa is required for the invasion of human red blood cells in vitro33. It has been shown that different haplotypes in the DBPα region II have differential binding affinities to the DARC receptor in human erythrocytes36. Therefore, these genetic exchanges affecting genes involved in invasion may reflect an adaptation to a new vertebrate host and may confer improved binding and increase invasion efficiency. It will be important to investigate whether changes in the haplotypes of other genes involved in erythrocyte invasion affect the ability of the parasite to invade and multiply in human cells. Genetic interactions between invasion genes may assist parasites with adapting more efficiently to humans and facilitate transmission, which could hamper malaria elimination efforts.
Overall, by establishing an effective SWGA strategy for P. knowlesi, it will be possible to perform much needed large-scale WGS studies of the parasite genomic diversity across Asia, as well as investigate important fundamental biology, such as the genetics underlying mechanisms for erythrocyte invasion.
Methods
Sample collection and preparation
For this project we use P. knowlesi DNA samples from Sabah in Malaysian Borneo (n = 21) (provided by the Menzies School of Health Research) and from Peninsular Malaysia (n = 5) (provided by the University of Malaya). Samples from Sabah were obtained from patients enrolled as part of clinical malaria studies conducted from 2010 to 201410,37. Ethical approval for these studies was obtained from the Ministry of Health, Malaysia, and Menzies School of Health Research, Australia. Samples from Peninsular Malaysia were collected from patients admitted to University Malaya Medical Centre (UMMC), Kuala Lumpur, from July 2008 to December 201438. Ethical approval was provided by the University of Malaya Medical Centre Medical Ethics Committee (MEC Ref. No: 817.18). Informed consent was obtained for study participation in both Sabah and Kuala Lumpur sites. All methods were performed in accordance with the relevant guidelines and regulations in both Sabah and Kuala Lumpur sites. All DNA samples were quantified using the Qubit Fluorometer using the dsDNA high sensitivity method (Invitrogen). All samples were screened by PCR targeting the genes encoding the Plasmodium 18S rRNA39. Confirmation of P. knowlesi mono-infection was performed using a heminested PCR assay based on a P. knowlesi specific conserved SICAvar region, to overcome possible cross reactivity between P. knowlesi and other Plasmodium species40,41. The relative amounts of parasite and human DNA in each sample was determined using a qPCR protocol using primers and probes specific for each species42,43,44. Pure human and P. knowlesi standard controls (range 0.0001–100 ng/ul concentrations) were included to determine the relative concentration (ng/ul) of each organism’s DNA in a sample.
Primer design for selective whole genome amplification
The swga program (www.github.com/eclarke/swga) was used to identify primer sets that preferentially amplify the P. knowlesi genome, providing as input the new A1-H.1 reference for the P. knowlesi human-adapted line A1-H.131 and the established human reference human_g1k_v37 (ftp://ftp.1000genomes.ebi.ac.uk). The resulting ten best sets consisted of combinations of four to six oligonucleotides each, with several overlapping primers, including two that were present in all sets. The set with the lowest Gini index and perfectly even binding across the genome consists of the following six primers: 5′-ATAATC*G*T-3′, 5′-ATTATC*G*T-3′, 5′-CGAAAT*A*G-3′, 5′-CGATAA*A*G-3′, 5′-GAATAA*C*G-3′ and 5′-TCGTAA*T*A-3′; where asterisks represent phosphorothioate bonds to prevent primer degradation by the exonuclease activity of the Phi29 polymerase
Selective whole genome amplification
Selective whole genome amplification (SWGA) was performed according to published protocols27. All SWGA reactions were carried out in the UV Cabinet for PCR Operations (UV-B-AR, Grant-Bio) to minimize contamination. SWGA reactions were performed containing a maximum of 50 ng of total input genomic DNA (and a minimum of 5 ng), 5 µl of 10 x Phi29 DNA Polymerase Reaction Buffer (New England BioLabs), 0.5 µl of Purified 100x BSA (New England BioLabs), 0.5 µl of 250 µM Primer mix of Pkset1, 5 µl 10 mM dNTP (Roche), 30 units Phi29 DNA Polymerase (New England BioLabs) and Nuclease-Free Water (Ambion, The RNA Company) to reach a final reaction volume of 50 µl. The reaction was carried out on a thermocycler with the following step-down program: 5 minutes at 35 °C, 10 minutes at 34 °C, 15 minutes at 33 °C, 20 minutes at 32 °C, 25 min 31 °C, 16 hours at 30 °C and 10 minutes at 65 °C. The SWGA samples were diluted 1:1 with EB buffer (Qiagen) and the reaction was purified using the AMPure XP beads (Beckman-Coulter), using a sample to beads ratio of 1:1 according to the protocol.
Whole-genome sequencing, bioinformatics analysis and population genetics
The SWGA products and unamplified DNA were sequenced on an Illumina MiSeq or HiSeq4000 platform. DNA and SWGA Libraries for MiSeq were prepared using the QIAseq FX DNA Library Kit (Qiagen) as per manufacturer’s instructions. A twenty-minute fragmentation step was optimized for Plasmodium samples. For the HiSeq runs, libraries were prepared using the NEB Next Ultra DNA Library Prep Kit for Illumina (from New England BioLabs Inc., E7370). All samples were run using 150 bp paired-end reads. The raw sequence data for the isolates was aligned against the new reference for the human-adapted line A1-H.1 (no regions were excluded for analysis)31 using bwa-mem software with default settings45. In order to establish the amount of human DNA in the isolate data, the sequence data was mapped to the GRCh37 human reference genome (NCBI; latest version, as accessed on 23/11/2018) using bwa-mem software with default parameters. WGS data from an extra 156 publicly available samples were also used for analysis (sourced from24,25,46, where sequencing accession numbers are listed). SNPs were called using the Samtools software suite47, and those of high quality were retained using previously described methods (phred score > 30, 1 error per 1 kbp)23. Samples with high levels of multiplicity of infection were detected using estMOI software48. For comparisons between populations, we applied principal component analysis (using R core functions dist and cmd.scale; results not presented) and neighbour-joining tree (using ape package in R49) approaches. These clustering approaches were implemented on a Manhattan distance matrix of pairwise identity by state values calculated from the SNP data. Nucleotide diversity (π) for pairwise isolate comparisons was calculated using in-house R scripts adapted from a previous study50. These scripts resulted in the number of SNP differences between the two sequences divided by the length of the DNA fragment. For population comparisons the function nuc.div from the pegas package in R was used51. R-base scripts were used to perform linear regression analyses for the SNP π plots. For the construction of the robust haplotypes of the invasion-related genes (DBPα, DBPβ, DBPγ, NBPXa and NBPXb) we used only reads mapped with high quality and discarded any SNPs with high levels of missing alleles or evidence of multiplicity of infection (MOI > 1). Genetic differentiation between subpopulations was calculated using the average FST32 value for each pairwise comparison, using only SNPs with minor allele frequencies in excess of 5%.
Data Availability
Previously published data can be found on the ENA using the Run accession codes in Supplementary Table 1. The newly generated data can be found in the ENA study accession number ERP110368.
References
Cox-Singh, J. et al. Plasmodium knowlesi Malaria in Humans Is Widely Distributed and Potentially Life Threatening. Clin. Infect. Dis. 46, 165–171 (2008).
Shearer, F. M. et al. Estimating Geographical Variation in the Risk of Zoonotic Plasmodium knowlesi Infection in Countries Eliminating Malaria. PLoS Negl. Trop. Dis. 10, e0004915 (2016).
William, T. et al. Changing epidemiology of malaria in Sabah, Malaysia: increasing incidence of Plasmodium knowlesi. Malar. J. 13, 390 (2014).
World Health Organization Malaria Policy Advisory Commitee. Outcomes from the Evidence Review Group on Plasmodium knowlesi (2017).
Vythilingam, I. et al. Natural transmission of Plasmodium knowlesi to humans by Anopheles latens in Sarawak, Malaysia. Trans. R. Soc. Trop. Med. Hyg. 100, 1087–1088 (2006).
Tan, C. H., Vythilingam, I., Matusop, A., Chan, S. T. & Singh, B. Bionomics of Anopheles latens in Kapit, Sarawak, Malaysian Borneo in relation to the transmission of zoonotic simian malaria parasite Plasmodium knowlesi. Malar. J. 7, 52 (2008).
Brant, H. L. et al. Vertical stratification of adult mosquitoes (Diptera: Culicidae) within a tropical rainforest in Sabah, Malaysia. Malar. J. 15, 370 (2016).
Vythilingam, I. et al. Plasmodium knowlesi in humans, macaques and mosquitoes in peninsular Malaysia. Parasit. Vectors 1, 26 (2008).
Daneshvar, C. et al. Clinical and Laboratory Features of Human Plasmodium knowlesi Infection. Clin. Infect. Dis. 49, 852–860 (2009).
Grigg, M. J. et al. Age-Related Clinical Spectrum of Plasmodium knowlesi Malaria and Predictors of Severity. Clin. Infect. Dis. 67, 350–359 (2018).
Overgaard, H. J., Ekbom, B., Suwonkerd, W. & Takagi, M. Effect of landscape structure on anopheline mosquito density and diversity in northern Thailand: Implications for malaria transmission and control. Landsc. Ecol. 18, 605 (2003).
Brock, P. M. et al. Plasmodium knowlesi transmission: integrating quantitative approaches from epidemiology and ecology to understand malaria as a zoonosis. Parasitology 143, 389–400 (2016).
Fornace, K. M. et al. Association between Landscape Factors and Spatial Patterns of Plasmodium knowlesi Infections in Sabah, Malaysia. Emerg. Infect. Dis. 22, 201–208 (2016).
Rajahram, G. S. et al. Falling Plasmodium knowlesi Malaria Death Rate among Adults despite Rising Incidence, Sabah, Malaysia, 2010-2014. Emerg. Infect. Dis. 22, 41–48 (2016).
Grigg, M. J. et al. Individual-level factors associated with the risk of acquiring human Plasmodium knowlesi malaria in Malaysia: a case-control study. Lancet. Planet. Heal. 1, e97–e104 (2017).
Imai, N., White, M. T., Ghani, A. C. & Drakeley, C. J. Transmission and Control of Plasmodium knowlesi: A Mathematical Modelling Study. PLoS Negl. Trop. Dis. 8, e2978 (2014).
Pearson, R. D. et al. Genomic analysis of local variation and recent evolution in Plasmodium vivax. Nat Genet in pess, 959–964 (2016).
Gomes, A. R. et al. Genetic diversity of next generation antimalarial targets: A baseline for drug resistance surveillance programmes. Int. J. Parasitol. Drugs Drug Resist. 7, 174–180 (2017).
Ravenhall, M. et al. Characterizing the impact of sustained sulfadoxine/pyrimethamine use upon the Plasmodium falciparum population in Malawi. Malar. J. 15, 575 (2016).
Diez Benavente, E. et al. Genomic variation in Plasmodium vivax malaria reveals regions under selective pressure. PLoS One 12, e0177134 (2017).
Miotto, O. et al. Genetic architecture of artemisinin-resistant Plasmodium falciparum. Nat. Genet. 47, 226–234 (2015).
Hupalo, D. N. et al. Population genomics studies identify signatures of global dispersal and drug resistance in Plasmodium vivax. Nat Genet 48, 953–958 (2016).
Benavente, E. D. et al. Analysis of nuclear and organellar genomes of Plasmodium knowlesi in humans reveals ancient population structure and recent recombination among host-specific subpopulations. PLoS Genet. 13, e1007008 (2017).
Pinheiro, M. M. et al. Plasmodium knowlesi Genome Sequences from Clinical Isolates Reveal Extensive Genomic Dimorphism. PLoS One 10, e0121303 (2015).
Assefa, S. et al. Population genomic structure and adaptation in the zoonotic malaria parasite Plasmodium knowlesi. Proc. Natl. Acad. Sci. USA 112, 13027–13032 (2015).
Sundararaman, S. A. et al. Genomes of cryptic chimpanzee Plasmodium species reveal key evolutionary events leading. Nat. Commun. 7, 1–14 (2016).
Oyola, S. O. et al. Whole genome sequencing of Plasmodium falciparum from dried blood spots using selective whole genome amplification. Malar. J. 15, 597 (2016).
Cowell, A. N. et al. Selective Whole-Genome Amplification Is a Robust Method That Enables Scalable Whole-Genome Sequencing of Plasmodium vivax from Unprocessed Clinical Samples. MBio 8 (2017).
Leichty, A. R. & Brisson, D. Selective whole genome amplification for resequencing target microbial species from complex natural samples. Genetics 198, 473–81 (2014).
Clarke, E. L. et al. Swga: a Primer Design Toolkit for Selective Whole Genome Amplification. Bioinformatics, 1–7, https://doi.org/10.1093/bioinformatics/btx118 (2017).
Benavente, E. D. et al. A reference genome and methylome for the Plasmodium knowlesi A1-H.1 line. Int. J. Parasitol, https://doi.org/10.1016/J.IJPARA.2017.09.008 (2017).
Holsinger, K. E. & Weir, B. S. Genetics in geographically structured populations: defining, estimating and interpreting FST. Nat Rev Genet 10, 639–650 (2009).
Moon, R. W. et al. Normocyte-binding protein required for human erythrocyte invasion by the zoonotic malaria parasite Plasmodium knowlesi. Proc. Natl. Acad. Sci. USA 113, 7231–7236 (2016).
Divis, P. C. S. et al. Admixture in Humans of Two Divergent Plasmodium knowlesi Populations Associated with Different Macaque Host Species. PLoS Pathog. 11, e1004888 (2015).
Divis, P. C. S. et al. Three Divergent Subpopulations of the Malaria Parasite Plasmodium knowlesi. Emerg. Infect. Dis. 23, 616–624 (2017).
Lim, K. L., Amir, A., Lau, Y. L. & Fong, M. Y. The Duffy binding protein (PkDBPαII) of Plasmodium knowlesi from Peninsular Malaysia and Malaysian Borneo show different binding activity level to human erythrocytes. Malar. J. 16, 331 (2017).
Barber, B. E. et al. A Prospective Comparative Study of Knowlesi, Falciparum, and Vivax Malaria in Sabah, Malaysia: High Proportion With Severe Disease From Plasmodium Knowlesi and Plasmodium Vivax But No Mortality With Early Referral and Artesunate Therapy. Clin. Infect. Dis. 56, 383–397 (2013).
Fong, M. Y., Wong, S. S., De Silva, J. R. & Lau, Y. L. Genetic polymorphism in domain I of the apical membrane antigen-1 among Plasmodium knowlesi clinical isolates from Peninsular Malaysia. Acta Trop. 152, 145–150 (2015).
Singh, B. et al. A genus- and species-specific nested polymerase chain reaction malaria detection assay for epidemiologic studies. Am. J. Trop. Med. Hyg. 60, 687–692 (1999).
Lubis, I. N. D. et al. Contribution of Plasmodium knowlesi to Multispecies Human Malaria Infections in North Sumatera, Indonesia. J. Infect. Dis. 215, 1148–1155 (2017).
Imwong, M. et al. Spurious amplification of a Plasmodium vivax small-subunit RNA gene by use of primers currently used to detect P. knowlesi. J. Clin. Microbiol. 47, 4173–4175 (2009).
Auburn, S. et al. An Effective Method to Purify Plasmodium falciparum DNA Directly from Clinical Blood Samples for Whole Genome High-Throughput Sequencing. PLoS One 6, e22213 (2011).
Divis, P. C., Shokoples, S. E., Singh, B. & Yanow, S. K. A TaqMan real-time PCR assay for the detection and quantitation of Plasmodium knowlesi. Malar. J. 9, 344 (2010).
Reller, M. E., Chen, W. H., Dalton, J., Lichay, M. A. & Dumler, J. S. Multiplex 5′ nuclease quantitative real-time PCR for clinical diagnosis of malaria and species-level identification and epidemiologic evaluation of malaria-causing parasites, including Plasmodium knowlesi. J. Clin. Microbiol. 51, 2931–8 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Divis, P. C. S., Duffy, C. W., Kadir, K. A., Singh, B. & Conway, D. J. Genome-wide mosaicism in divergence between zoonotic malaria parasite subpopulations with separate sympatric transmission cycles. Mol. Ecol. 27, 860–870 (2018).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Assefa, S. A. et al. estMOI: estimating multiplicity of infection using parasite deep sequencing data. Bioinformatics 30, 1292–1294 (2014).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20, 289–290 (2004).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 76, 5269–5273 (1979).
Paradis, E. pegas: an R package for population genetics with an integrated–modular approach. Bioinformatics 26, 419–420 (2010).
Acknowledgements
We thank Don Van Schalkwyk for providing biological materials from the P. knowlesi reference strain. We thank the KAUST Bioscience Core Lab (BCL) facility for sequencing the unamplified isolates. AI is funded by a UK Medical Research Council LiD PhD studentship. RWM is supported by an MRC Career Development Award (MR/M021157/1) jointly funded by the UK Medical Research Council and UK Department for International Development. TGC is funded by the Medical Research Council UK (Grant No. MR/M01360X/1, MR/N010469/1, MR/R025576/1, and MR/R020973/1) and BBSRC (Grant No. BB/R013063/1). SC is funded by Medical Research Council UK grants (MR/M01360X/1, MR/R025576/1, and MR/R020973/1) and BBSRC (Grant no. BB/R013063/1). The Sabah studies were funded by the Malaysian Ministry of Health (grant number BP00500420) and the Australian National Health and Medical Research Council (grant numbers 1037304, 1132975 and 1045156) and fellowships (1042072 and 1135820 to NMA, 1088738 to BEB, and 1138860 to MJG). AP and AR are supported by a faculty baseline fund (BAS/1/1020-01-01) to AP. We gratefully acknowledge the Scientific Computing Group for data management and compute infrastructure at Genome Institute of Singapore for their help. The MRC eMedLab computing resource was used for bioinformatics and statistical analysis.
Author information
Authors and Affiliations
Contributions
S.C. and T.G.C. conceived and directed the project. L.Y.L., T.W. and N.M.A. coordinated sample collection. A.R.G., S.C., J.R.D.S., M.G., A.R., A.I., T.Y., S.A., A.P. and R.W.M. undertook sample collection, processing and DNA extraction. P.F.d.S., M.L.H., A.P. and S.C. coordinated sequencing. E.D.B. performed bioinformatic and statistical analyses under the supervision of T.G.C. and S.C. T.G.C., S.C. and E.D.B. interpreted results. E.D.B., T.G.C. and S.C. wrote the first draft of the manuscript. All authors commented and edited on various versions of the draft manuscript and approved the final manuscript. E.D.B., T.G.C. and S.C. compiled the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Benavente, E.D., Gomes, A.R., De Silva, J.R. et al. Whole genome sequencing of amplified Plasmodium knowlesi DNA from unprocessed blood reveals genetic exchange events between Malaysian Peninsular and Borneo subpopulations. Sci Rep 9, 9873 (2019). https://doi.org/10.1038/s41598-019-46398-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46398-z
This article is cited by
-
Rapid profiling of Plasmodium parasites from genome sequences to assist malaria control
Genome Medicine (2023)
-
Population genetic analysis of Plasmodium knowlesi reveals differential selection and exchange events between Borneo and Peninsular sub-populations
Scientific Reports (2023)
-
A systematic review of asymptomatic Plasmodium knowlesi infection: an emerging challenge involving an emerging infectious disease
Malaria Journal (2022)
-
Plasmodium knowlesi: the game changer for malaria eradication
Malaria Journal (2022)
-
Optimization of whole-genome sequencing of Plasmodium falciparum from low-density dried blood spot samples
Malaria Journal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.