Abstract
Telomerase is expressed in adult mouse, but not in most human, tissues and mouse telomeres are much longer than those in humans. This interspecies difference of telomere homeostasis poses a challenge in modeling human diseases using laboratory mice. Using chromatinized bacterial artificial chromosome reporters, we discovered that the 5′ intergenic region, introns 2 and 6 of human telomerase gene (hTERT) were critical for regulating its promoter in somatic cells. Accordingly, we engineered a humanized gene, hmTert, by knocking-in a 47-kilobase hybrid fragment containing these human non-coding sequences into the mTert locus in mouse embryonic stem cells (mESCs). The hmTert gene, encoding the wildtype mTert protein, was fully functional, as a mESC line with homozygous hmTert alleles proliferated for over 400 population doublings without exhibiting chromosomal abnormalities. Like human ESCs, the engineered mESCs contained high telomerase activity, which was repressed upon their differentiation into fibroblast-like cells in a histone deacetylase-dependent manner. Fibroblast-like cells differentiated from these mESCs contained little telomerase activity. Thus, telomerase in mESCs with the hmTert alleles was subjected to human-like regulation. Our study revealed a novel approach to engineer a humanized telomerase gene in mice, achieving a milestone in creating a mouse model with humanized telomere homeostasis.
Similar content being viewed by others
Introduction
Telomeres are specialized nucleoprotein structures at chromosomal ends and play key roles in human aging and tumorigenesis. Telomeric DNAs are replenished by telomerase, a ribonucleoprotein complex containing two essential components, a catalytic reverse transcriptase (TERT) and a template RNA (TERC), along with accessory proteins1,2. In humans, telomerase activity is low or undetectable in most adult tissues. Consequently, telomeres shorten upon successive cell divisions, thereby functioning as a biological clock. In contrast, telomerase activity is readily detected in adult mouse tissues and laboratory mice have very long telomeres (50–100 kb)3. Consequently, telomerase-null mice survive up to six generations with no discernible phenotypes in early generations4,5,6.
Studies have indicated that hTERT transcription is the primary step of telomerase regulation7. Whereas TERC RNA and telomerase-associated proteins are expressed in most somatic tissues, hTERT expression is highly regulated and correlated with telomerase activity in many adult cell types8. hTERT transcriptional regulation is a complex process that involves both the binding of transcription factors at its promoter and epigenetic repression that regulate chromatin environment of the hTERT locus9. Our previous studies revealed that the hTERT gene was embedded in a condensed chromatin domain in somatic cells and inhibition of histone deacetylation led to an opening of this domain and activated hTERT transcription10. We have recently shown that repression of the hTERT promoter was dictated by distal elements and required a human specific genomic context11,12.
The differences in telomerase regulation and telomere length among different species likely contribute to important differences in aging processes and cancer development between humans and mice. Indeed, many human somatic cells, but not mouse cells, undergo replicative senescence mediated by progressive telomere shortening13. Mouse models often give rise to tumors of different spectra from the types of human cancers they are intended to mimic. For example, p53-null mice of several inbred strains typically developed soft tissue sarcomas and lymphomas, the types of cancer mostly occurring in human childhood14. Conversely, p53 deficiency, along with shortened telomeres in late generations of telomerase-null mice resulted a shift of tumor spectrum to epithelial carcinomas of skin, breast, and colon origins, the most common adult human cancers15. Thus, telomere length and telomerase regulation contribute to the different tumor development in mouse and human cancers. Nonetheless, telomerase-null mice are not the best models for human cancers because their telomerase cannot be reactivated as it occurs in majority of human cancers. An ideal model would regulate telomerase in mice the same way as in humans. Here, as the first step towards the generation of a mouse model with humanized telomerase regulation and telomere homeostasis, we determined the cis regulatory elements involved in hTERT repression and revealed that human genomic sequences within the 5′ intergenic region (5IR), introns 2 and 6 (I2 and I6) contained elements that confer a human-like regulation in a mouse genomic context in telomerase-positive and telomerase-negative fibroblast lines. These elements also repressed the hTERT promoter during mouse ESC differentiation. We thus engineered an mTert locus by replacing its 5IR, I2, and I6 with corresponding human genomic sequences in mouse ESCs. Our results showed that mouse ESCs with the chimeric locus, hmTert, expressed a high level of telomerase activity and this activity was strongly repressed during in vitro differentiation, similar to human ESCs.
Results
Regulatory genomic sequences required for hTERT repression
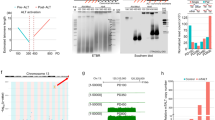
Histone deacetylase (HDAC)-dependent transcriptional repression is central to the regulation of hTERT gene in somatic cells16. To identify the genomic sequences responsible for the differential regulation of hTERT and mTert genes, we used the BAC reporters H(wt) and M(wt)17, which contained a 160-kb and a 135-kb genomic DNA encompassing the consecutive CRR9 (also called CLPTMIL), TERT, and XTRP2 (SLC6a18) loci of human and mouse genomic sequences, respectively (Fig. 1). Single-copy BACs were integrated into the same acceptor sites in Tel+ and Tel− human fibroblast lines by recombinase-mediated BAC targeting (RMBT)17, and thus had the same genomic environment. In this system, the activities of TERT promoters were measured as the ratios of Renilla luciferase (Rluc) activities from TERT promoters to Firefly luciferase (Fluc) activity from CRR9 promoters, because CRR9 promoters were constitutively active in most cells17. As shown in Fig. 1A, the hTERT promoter in H(wt) was 30–50 fold more active in Tel+ cells than that in Tel− cells and markedly activated in both cells by HDAC inhibitor trichostatin A (TSA), recapitulating the endogenous expression of hTERT gene17. While the mechanism of hTERT activation in Tel+ versus Tel− cells remained to be elucidated, it is likely that Tel+ cells contained different trans-acting factors, either increased activators or reduced repressors, that interact with the hTERT cis-regulatory elements. On the other hand, the mTERT promoter in M(wt) was moderately active in both Tel+ and Tel− cells. In our recent report12, the hTERT and mTert promoters, which were 472-bp and 474-bp genomic sequences upstream of the hTERT and mTert ATG codons, respectively, were swapped between H(wt) and M(wt), generating H(mPro) and M(hPro). Upon integration into Tel+ and Tel− cells by RMBT. The mTert promoter activity in the human genomic context, H(mPro), was higher in Tel+ cells than in Tel− cells and strongly activated by TSA in both cells (Fig. 1B). In contrast, the hTERT promoter in the mouse genomic context, M(hPro), was highly active in both Tel+ and Tel− cells, and TSA treatment had little or no effect on its activity. Thus, the human, but not the mouse, genomic sequence conferred a repressive chromatin environment for both TERT promoters12.
The involvement of distal genomic sequences in the repression of hTERT promoter in human fibroblasts. A schematic diagram of all BAC reporters is shown at the top. Shown below are illustrations of H(wt), M(wt), and chimera BACs. Black and light grey lines represent human and mouse genomic sequences, respectively. (A) BAC reporters made from human and mouse genomic sequences, H(wt) and M(wt), respectively. (B) H(mPro) and M(hPro), in which TERT promoter regions were swapped between H(wt) and M(wt). (C) M(h5IR), M(h5IRI2) and M(h5IRI2I6), 5IR, I2, and I6 represent 5′ intergenic region, introns 2 and 6 of the hTERT gene, replacing their corresponding mouse sequences in M(hPro). Bar graphs, activities of TERT promoters in chromatinized BAC reporters in Tel− (−) and Tel+ (+) human fibroblast cells. Luciferase activities were measured in 96-well plates, treated without (black bars) or with TSA (white bars). The TERT promoter activities were measured as the ratios of Firefly to Renilla luciferase activities. *p < 0.05; **p < 0.01; ***p < 0.001.
Whereas the coding regions are conserved between hTERT and mTert loci, non-coding regions are highly diverged both in sequences and in lengths. For example, 5IR of the hTERT gene is over 22 kb, yet its mouse counterpart is about 6 kb. The two largest introns, I2 and I6, are 10 kb and 5.5 kb in human, and 5.5 kb and 1.8 kb in mice, respectively. To determine if these regions are important for human-specific repression, we replaced the regions in M(hPro) with their human counterparts, generating M(h5IR), M(h5IRI2), and M(h5IRI2I6). As shown in Fig. 1C, 5IR replacement in M(hPro), i.e. M(h5IR), resulted in a 2-fold and a 20-fold HDAC-dependent repression of the hTERT promoter in Tel+ and Tel− cells, respectively. Inclusion of human I2 and I6, i.e. M(h5IRI2) and M(h5IRI2I6), led to more stringent repression of the hTERT promoter in these cells: 7 fold in Tel+ and 73 fold in Tel− cells for M(h5IRI2), and 15 fold in Tel+ and over 200 fold in Tel− cells for M(h5IRI2I6). Accordingly, the hTERT promoter in M(h5IRI2I6) was about 30 fold stronger in Tel+ cells than that in Tel− cells, similar to the endogenous promoter in host cells17. Therefore, all three non-coding regions of the hTERT gene contributed to an hTERT-like regulation in the context of chromatinized BAC reporters in human fibroblasts.
Regulatory functions of hTERT non-coding regions during ESC differentiation
Next, we determined whether the human genomic sequences conferred human-like TERT regulation in mouse cells. A subset of aforementioned BAC reporters were integrated into the same acceptor site in a mouse ESC line T2-5 by RMBT11. The TERT promoters in all the BAC reporters, M(wt), M(hPro), M(h5IR), M(h5IRI2), and M(h5IRI2I6), were highly active in ESCs as determined by ratios of Rluc/Fluc. To determine the regulation of TERT promoters in these chimera BACs during in vitro differentiation, ESCs were induced to differentiate in suspension cultures in the absence of leukemia inhibitory factor (LIF). Upon differentiation into embryoid bodies (EBs) for two weeks, the mTert and hTERT promoters in M(wt) and M(hPro) decreased by 40% and 55%, respectively, compared to their levels in undifferentiated ESCs (Fig. 2A). Addition of the 5IR in M(h5IR) led to a 70% decrease of hTERT promoter activity. In M(h5IRI2) and M(h5IRI2I6), the hTERT promoter was reduced by 85–90%, comparable to 90% of H(wt). These results indicated that human 5IR, I2, and I6 functioned to regulate hTERT promoter during mouse EB differentiation.
TERT promoter activities from chromatinized BAC reporter in mouse ESCs and their differentiated derivatives. (A) TERT promoter activities in differentiating EB cultures. ESCs containing BAC reporters were cultured in the absence of feeder cells and LIF to form EBs. Luciferase activities were measured in 3, 6, 10, and 14 days, and normalized to those of the undifferentiated ESCs. (B) TERT promoter activities in differentiated fibroblast-like cultures. The promoter activities were normalized to those of ESCs. *p < 0.05; ***p < 0.001, compared to their respective activities in ESCs.
Although EBs contained a variety of differentiated cells, there were also undifferentiated and partially differentiated cells with significant hTERT promoter activities in these structures. To study the repression of the hTERT promoter in differentiated cells, adherent cells from EBs were enriched by removing detached cells and cultured in osteogenic medium for an additional three weeks to allow their differentiation into fibroblast-like cells. During this process, the mTert and hTERT promoters in M(wt) and M(hPro) decreased to 16–18% of their activities in ESCs (Fig. 2B); the hTERT promoter was reduced to 12% in M(h5IR), and 4–6% in M(h5IRI2) and M(h5IRI2I6), compared to their respective activities in ESCs. The hTERT promoter activities in M(h5IRI2) and M(h5IRI2I6) was slightly higher than that of H(wt) at 2%. Thus, inclusion of the three non-coding regions of the hTERT locus, 5IR, I2 and I6, resulted in hTERT-like regulation in an otherwise mouse genomic context during mouse ESC differentiation.
Engineering a humanized mTERT locus (hmTERT)
To determine whether the non-coding genomic sequences confer human-like regulation at the endogenous mTert gene, we set out to engineer a humanized mTert gene, or hmTert, by knocking human 5IR, I2, and I6 into the mTert locus in ESCs via homologous recombination. Hence, a chimeric targeting BAC was constructed as illustrated in Fig. 3A. This construct contained a 47-kb hybrid region containing the human 5IR, I2, and I6, replacing the corresponding mouse sequences. The mTERT protein coding sequence was preserved except for the 9 silent nucleotide substitutions near the 3′ end of exon 2, allowing the design of PCR primers that distinguished mRNAs from the hmTert and wildtype mTert alleles. The targeting BAC was transfected into Tc1 ESCs together with two plasmids expressing sgRNAs targeting mouse 5IR and I6. The cells were selected sequentially for resistance to puromycin and ganciclovir (GCV), and the resulting clones were analyzed for the presence of the entire 47-kb genomic region resulting from homologous recombination by genomic PCR and Southern analyses as shown in Fig. S1A,B. From 800 puromycin-resistant clones, we identified one ESC clone that contained two hmTert alleles (hm/hm) and multiple clones with a single hmTert allele (m/hm) (Table S1). To avoid the potential effects of off target cleavage by Crispr/Cas9, we have analyzed multiple independent clones in subsequent studies.
Engineering of an hmTert allele in mouse ESCs. (A) A diagram of the targeting BAC. Black and grey portions of horizontal lines indicate human and mouse genomic sequences, respectively. Vertical bars show relative positions and sizes of exons of the CRR9 (left) and TERT (right) genes. TK and Puro are thymidine kinase and puromycin-resistant markers, respectively. The targeting BAC along with two pX458-sgRNAs targeting mouse sequences near the boundaries of homologous recombination (indicated by flash signs) were cotransfected into ESCs and puromycin/GCV-resistant colonies were selected. The Puro marker was subsequently removed following transient transfection of pCBM, a Cre-expressing plasmid. The knock-in of a 47-kb genomic region as the result of homologous recombination is indicated. A small white dot at the 3′ end of Tert exon 2 indicates the silent mutations that distinguish mRNAs from mTert and hmTert loci in RT-qPCR assays. (B) mRNA expression from the hmTert locus. Total RNAs from ESCs containing mTert and/or hmTert loci were subjected to RT-qPCR analysis using probes that distinguish mTert and hmTert mRNAs, and the data were normalized to 18S rRNA. a-d were four independent m/hm clones. (C) Telomerase activity in ESCs with mTert and/or hmTert alleles. Telomerase activities in extracts of 2000 or 20 cells were measured using TRAP assay. m/m, wildtype ESCs containing two mTert alleles; hm/hm, ESCs containing hmTert/hmTert alleles, m/hm, heterozygous ESCs with mTert/hmTert genes. MEF, wildtype mouse embryonic fibroblast cells. RNase A treated wildtype ESC extract was used as a negative control. MW, molecular weight control. IC, internal control. Underneath are relative intensities of all telomere bands in each lane, normalized to lysis buffer control (1).
The hmTERT gene maintained telomeres in ESCs
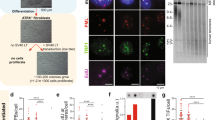
To determine the functionality of hmTert gene, ESC clones with hmTert alleles were further characterized. The hm/hm clone and two m/hm clones proliferated similarly to the wildtype m/m ESCs and had a correct number of 40 chromosomes (Fig. S2). The levels of Tert mRNA in a wildtype (m/m), an hm/hm, and four m/hm clones were determined by RT-qPCR analysis (Fig. 3B). Tert mRNA from the hmTert alleles was moderately higher (up to 2-fold) than those of the mTert alleles in both hm/hm and m/hm ESCs. Telomerase activities in hm/hm and m/hm ESCs were also similar to that in wildtype ESCs (m/m), as measured in a semi-quantitative TRAP assay (Fig. 3C). To assess the function of hmTert locus, ESCs were passaged for over 100 times (400 population doublings). Wildtype (m/m), heterozygous (m/hm), and homozygous (hm/hm) ESCs all proliferated similarly, whereas the proliferation of an ESC clone with homozygous mTert deletion (mTert −/−) slowed down significantly in late passages (Fig. 4A). The lengths of telomere in ESCs at different passages were measured by telomere fluorescence in situ hybridization (FISH) analysis and telomere restriction fragment analysis. Whereas telomere length varied during passages in both m/m and hm/hm clones, telomeres became significantly shorter in mTert −/− cells after 100 passages (Fig. 4B,C). Only 0.3 telomere signal-free chromosomal ends per metaphase were found in hm/hm cells after 110 passages, similar to m/m cells (Fig. 4D and Table 1). In contrast, about ten signal-free ends per metaphase were detected in mTert −/− cells at passage 100. Finally, while an average of five chromosomal end-end fusions were detected in mTert −/− cells after 100 passages, no such fusion was detected in hm/hm cells (Fig. 4E and Table 1). Taken together, these data indicated that hmTert loci encoded a functional telomerase gene and were able to maintain telomere stability in long-term cultures of mouse ESCs.
Proliferation and chromosome stabilities of ESCs with mTert and hmTert alleles. (A) Long-term proliferation of ESCs. ESCs with mTert/mTert (m/m), mTert/hmTert (m/hm), hmTert/hmTert (hm/hm), or mTert homozygous knockout (−/−) were passaged every 2 days for 110 times (m/m, m/hm, & hm/hm) or 100 times (−/−). (B) Telomere length in ESCs. Telomere lengths (Cy3) were measured by quantitative FISH (qFISH) analysis and the data were analyzed using the Metasystems software package. Mean telomere lengths are indicated by vertical dash lines and numbers in parentheses (mean ± s.d.). (C) Telomere restriction fragment length analysis. Genomic DNAs were extracted from ESCs at various passages and digested with HinfI and RsaI, followed by pulsed-field gel electrophoresis and Southern blotting. MW, molecular weight. Genotypes and passage numbers are indicated above each lane. (D) Images of telomeres (Cy3) in ESCs. (E) Telomere and chromosome integrity in ESCs. Red, Cy3 for telomere; green, FAM for centromere; blue, DAPI for DNA. Arrows point to telomere signal-less chromosomal ends and stars indicate chromosomal end-end fusion.
The hmTERT loci were repressed during ESC differentiation
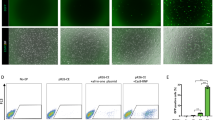
To determine the regulation of hmTert gene, Tert mRNA in differentiating EBs derived from ESCs with hmTert alleles were analyzed by RT-qPCR. As shown in Fig. 5A, the Tert mRNA levels were dramatically down-regulated during EB differentiation for 2 weeks in hm/hm cells, but not in wildtype m/m cells. Similarly in three clones of heterozygous m/hm cells, mRNA from the hmTert alleles decreased 8 and 15 folds as differentiation marker α-fetoprotein (AFP) increased during the differentiation process, whereas the down-regulation of expression from the mTert alleles was much less than the hmTert alleles in the same cells (Figs 5A and S3). Fibroblast-like cells from EBs of an m/hm clone were collected and passaged (Fig. S4). mRNA from both mTert and hmTert alleles progressively decreased upon passaging (Fig. 5B). The presence of incompletely differentiated cells might have contributed to the higher Tert mRNA levels in early passages of the cultures because fibroblast markers, type I collagen (Col1a1) and vimentin, progressively increased during passaging (Fig. S5). However, mRNA from the hmTert gene decreased much faster than that of mTert. The hmTert mRNA was about 100-fold lower than mTert mRNA after 11 passages. TSA treatment markedly induced hmTert mRNA, but had little effect on the level of mTert mRNA. Furthermore, telomerase activities were measured in ESCs and differentiated derivatives by TRAP assay. Whereas fibroblast-like cells derived from m/m ESCs contained a lower level of telomerase activity compared to that in undifferentiated ESCs, it was not detected in differentiated hm/hm cells (Fig. 5C), mimicking telomerase expression in human fibroblasts.
mRNA expression and telomerase activity from mTert and hmTert loci during in vitro ESC differentiation. (A) Tert mRNA expression in differentiating EB cultures. Tert mRNA were measured by RT-qPCR and normalized to 18S rRNA. Light gray, mTert; dark gray, hmTert. (B) mTert mRNA levels in differentiated fibroblast-like cells. 3-day old EBs (m/hm) were passaged in DMEM with 10% fetal bovine serum for 2, 5, and 11 times. Passage 11 cells were also treated for 24 h with 200 ng/ml TSA. mRNAs from mTert and hmTert loci were measured using specific primers spanning intron 2. (C) Telomerase activities in ESCs and differentiated cells. Telomerase activities in extracts of 2000 or 20 ESCs and fibroblast-like cells (passage 11) were measured using TRAP assay. Numbers below are relative intensities of all telomere bands in each lane, normalized to lysis buffer control (1). (D) Covalent histone modifications at genomic regions around the hmTert and mTert promoters in ESCs and differentiated cells. Chromatin fragments from ESCs (m/hm) and differentiated cells (DIFF) were precipitated using antibodies against specific histone marks, followed by qPCR analysis. H3Ac and H4Ac refer to actylated histones H3 and H4. H3K27Ac and H3K4me3 are acetylated K27 and trimethylated K4 residues of histone H3, respectively. Pro, hmTert or mTert promoters. Up5k, Up2k, and Dn2k are chromosomal sites 5-kb and 2-kb upstream and 2-kb downstream of the Tert promoters, respectively.
The epigenetic status of the hmTert locus in ESCs and their differentiated derivatives was also examined using chromatin immunoprecipitation. As shown in Fig. 5D, the region around the hmTert promoter was hyperacetylated at both histones H3 and H4 in ESCs and became deacetylated in differentiated cells. In contrast, the acetylation of histones at the mTert locus was not dramatically altered at the wildtype mTert locus during differentiation. As a control, the level of H3K4me3, a marker of active transcription, decreased significantly at the hmTert, but not mTert, locus. Altogether, these results indicated that the expression from the hmTert locus was tightly regulated and repressed by an HDAC-dependent mechanism in differentiated mouse cells, recapitulating the hTERT locus in human fibroblasts.
Alternative splicing of hmTERT mRNAs
The transcript from the hTERT gene is spliced into several forms, providing an additional regulation for its activity7. The major alternative splicing product is the minus beta form, which skips exons 7–8 and likely encodes an inactive form due to a downstream reading frame shift. It was reported that genomic sequence within human intron 6 was essential for this splicing event18. Thus, we examined splicing patterns around human I6 sequence in the hmTert locus by RT-PCR analysis using primers spanning exons 6 to 9. As shown in Fig. 6, whereas the mTert gene produced the full-length (FL) mRNA exclusively, the hmTert locus yielded an additional short form consistent with the skipping of exon 7 (∆E7), which was validated by sequencing the PCR fragment. To measure the levels of splicing variants quantitatively, RT-qPCR analysis was performed using primers specific for FL and ∆E7. The levels of FL form were similar in m/m and hm/hm ESCs, but hm/hm cells contained an additional 1/5 amount of ∆E7 form (Fig. 6C). Similarly, the endogenous hTERT gene in human ESCs also produced mostly the FL form, one quarter of ∆E7-8, and little ∆E7 (Fig. 6D). In addition, the ∆E7 mRNA was also induced by TSA treatment in differentiated cells (Fig. S6). Therefore, human I6 sequence facilitated the splicing of ∆E7-8 in the hTERT gene and ∆E7 in the hmTert locus in human and mouse ESCs, respectively. Although it was unlikely that this alternative splicing affected telomerase activity significantly in ESCs, the presence of I6 in the hmTert gene might help it to recapitulate the regulation of hTERT gene in somatic cells.
The impact of human intron 6 on hmTert splicing. (A) PCR analysis of mTert, hmTert, and hTERT mRNAs. Total RNAs from mouse and human ESCs were subjected to PCR analysis using primers spanning introns 6–8 and analyzed on an Agarose gel. Numbers on the left are sizes of molecular weight markers in bp. (B) Diagrams of the genomic regions spanning exons 6–9. Mouse exons and introns are designated by grey lines and boxes and human sequences are shown as black lines and boxes. Splicing patterns are indicated by thin lines below. Arrowheads show the positions of PCR primers used in panel (A,C). Alternative splicing of exon 7 at the mTert and hmTert loci. mTert mRNA levels in ESCs were determined by RT-qPCR analysis using primers spanning intron 6 (FL) or introns 6–7 (∆E7). (D) Splicing of exon 7 of the hTERT locus. hTERT mRNA levels in human H9 ESCs were measured by RT-qPCR analysis using primers spanning intron 6 (FL), introns 6–7 (∆E7), or introns 6–8 (∆E7-8).
Discussion
A principal difference between mice and humans relates to a longtime observation that murine fibroblasts growing in culture undergo spontaneous immortalization at a high frequency, owing to their long telomeres and constitutive telomerase expression19. Data from multiple laboratories, including our own, have indicated that transcriptional regulation of the TERT genes is fundamental to this interspecies difference3,8,20. As we have shown in our previous studies10,21, a central difference between the regulation of hTERT and mTert genes is the stringent repression of hTERT promoter in somatic cells. In this report, we identified 5IR, I2, & I6 as genomic sequences that confer human-specific repression during differentiation. The chromatinized BAC reporter, M(h5IRI2I6), containing human 5IR, I2, & I6 replacing their mouse counterparts in M(wt), was much more active in Tel+ human fibroblasts than in Tel− cells, recapitulating the endogenous hTERT gene in host cells. This reporter was active in ESCs and repressed upon differentiation; hence, we knocked this entire 47-kb chimeric genomic sequence (from 5IR to I6) into the mTert locus in ESCs. The resulting ESC clone with homozygous hmTert loci contained a high level of telomerase and was able to proliferate for over 400 population doublings without detectable telomere deficiencies, indicating that the Tert protein expressed from this engineered gene was fully functional. Moreover, the regulation of hmTert gene recapitulated that of the hTERT gene during in vitro ESC differentiation.
Our ultimate goal is to engineer a mouse strain with humanized telomere homeostasis. Mouse models of human diseases have become a central part of biomedical research. Laboratory mice provide the most experimentally accessible mammalian models that bear close similarities of genes, organs, and systemic physiology with humans. However, species-specific telomerase regulation and telomere homeostasis remain to be a major challenge in using mouse models to study human diseases. This is because telomere homeostasis is involved in the pathogenic processes of many types of diseases, especially those related to aging and cancer. One approach of generating such a mouse model is to complement mTert −/− mice with a transgenic hTERT gene. However, such an approach is not possible because the hTERT protein does not function with mTerc RNA22,23. In contrast, the Tert protein encoded by the hmTert locus is identical to that of the wildtype mTert gene. Thus, humanization of the mTert locus, like the hmTert allele we engineered here, is a new and practical approach for the generation of such a mouse model.
It remains to be elucidated how distal elements within the human 5IR, I2, and I6 regions control the regulation of hTERT gene. It has been noted that these human genomic regions contain abundant repetitive sequences, much more than their mouse counterparts. These repetitive elements include Alu, LINE, endogenous retroviral elements, and variable number tandem repeats (VNTRs). The functions of these elements in regulating hTERT expression are a focus of ongoing studies. For example, I6 was found to be involved in alternative splicing of the hTERT transcript18. In particular, a 1-kb VNTR within this intron was able to alter hTERT mRNA splicing24, and it also affected mRNA splicing in the hmTert locus (Fig. 6). The presence of abundant repetitive elements at the hTERT locus also likely helps to establish a repressive chromatin environment and repress its promoter activity.
In short, the new hmTert locus encoded an mTert protein that was able to maintain telomere length during long-term ESC proliferation. Its expression was repressed in differentiated cells by a HDAC-dependent mechanism, similar to the native hTERT gene. This hmTert locus provided a foundation for our long-term goal of generating a novel mouse model with humanized telomere homeostasis.
Materials and Methods
Bacterial Artificial Chromosomes (BACs) and Plasmids
BAC reporters H(wt), M(wt), H(mPro) and M(hPro), were described previously11,12,17. Other chimeric BACs and the targeting BAC were constructed using a multi-step BAC recombineering strategy25,26. Briefly, 5′ intergenic (h5IR, 22-kb), introns 2 (hI2, 10-kb) and 6 (hI6, 5.5-kb) of the hTERT gene replaced the syntenic regions in M(wt) to generate M(h5IR), M(h5IRI2), and M(h5IRI2I6), respectively (Fig. 1). The targeting BAC for the knock-in experiment was constructed using M(h5IRI2I6) with the following modifications: (1) The Rluc cassette was removed and the mTert initiation codon was restored; (2) Nine silent nucleotide substitutions were introduced into the coding region near 3′ end of exon 2 (AAAGTAGAGGATTGCCACTGGCTCCGCAGCAGCCCGG to AAGGTCGAAGACTGCCACTGGCTGAGGTCCAGCCCGG); (3) A puromycin-resistant marker (Puro), surrounded by a pair of lox511 sites, was inserted behind the 3′ end of CRR9 gene; (4) A thymidine kinase (TK) cassette replaced exons 1–12 of the CRR9 gene; (5) A 300-bp sequence from the 5′ end of mTert intron 2 was inserted before the hTERT intron 2 to restore its efficient splicing; (6) Sequences downstream of mTert exon 11 were deleted. The resulting BAC had about 4- & 3-kb of 5′ and 3′ homology arms, respectively. Single guide RNAs (sgRNAs) targeting mouse introns 2 and 6 were cloned into pX45827.
Cell culture and ESCs differentiation
Telomerase-positive 3C167b3.1 (Tel+) and -negative GM847.7 (Tel−) human fibroblast lines and mouse ESCs Tc1 (129S6/SvEvTac) were cultured as previously described11,12. BAC reporters were integrated into Tel+, Tel−, and T2-5 (a clone of Tc1 cells), all containing a single BAC acceptor site, using RMBT12,17. ESC differentiation into embryoid bodies (EBs) and fibroblast-like cells were also described28. Individual EBs were collected for luciferase assay. 3d-old EBs were treated with 10−7 M Retinoic Acid (RA) (Sigma-Aldrich, US) for 2d and cultured for another 2d without RA. Adherent cells migrated out of EBs were collected and passaged. Fibroblast-like cells were characterized by the expression of vimentin11.
Genomic Knock-In
Knock-in experiments were performed by cotransfection of the targeting BAC and two pX458-sgRNAs (targeting the 5′ intergenic region and intron 6 of the mTert locus) into Tc1 cells29 (Fig. 3A). Following transfection, puromycin and GCV-resistant colonies are isolated. Genomic PCR and Southern analyses were performed to identify clones with correct knock-in in all three regions, 5IR, I2, and I6 (Fig. S1A,B)8,10. Probes are listed in Supplemental Table S1 and results are described in Table S2. Finally, the puromycin marker was removed following transient transfection of pCBM, a Cre-expressing plasmid17. As a control, ESCs with mTert homozygous knockout were also isolated following transfection of the abovementioned pX458-sgRNA plasmids.
Gene expression analyses
Luciferase activities were measured using Dual-Luciferase Reporter Assay System (Promega, Madison, WI). Firefly luciferase (Fluc) from the CRR9 genes was constitutively expressed in ESCs and differentiated cells. Renilla luciferase (Rluc) activities from TERT promoters were normalized to Fluc activity17 and the data were verified by normalizing Rluc activity to the number of cells, as determined by thiazolyl blue tetrazolium bromide (MTT) assay (data not shown). Quantitative RT-PCR (RT-qPCR) assays were conducted as previously described11.
Telomerase activity and telomere length analyses
Telomerase activities were determined using a modified telomeric repeat amplification protocol (TRAP) assay30,31. Telomere length was measured by quantitative fluorescence in situ hybridization (qFISH). Metaphases were prepared and qFISH was performed as described previously32,33 using peptide nucleic acid (PNA) probes according to manufacturer’s instruction (PNA Bio Inc.). PNA probes were TelC-Cy3 (CCCTAA repeats) for telomere and CENPB-FAM (ATTCGTTGGAAACGGGA) for centromere. Fluorescence images were captured using Axioscope Z2 (Zeiss, Germany). The fluorescence signals were integrated from at least 15 metaphases. Quantitative image analyses of telomere were done following the procedure as described by Perner et al.34 with Isis software (MetaSystems, Altlussheim, Germany).
Telomere length in mESCs was also assessed by telomeric restriction fragment analysis. Genomic DNAs were digested with Hinf I and Rsa I, and subjected to pulsed-field gel electrophoresis using CHEF-DR III Pulsed field Electrophoresis Systems (Bio-Rad), followed by Southern blotting. Telomere restriction fragments were hybridized to a (TTAGGG)3-Biotin probe and detected North2South™ Chemiluminescent Hybridization and Detection Kit (Thermo Scientific).
Statistical analysis
Two tails Student’s t test was used to compare relative gene expression by RT-qPCR.
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Change history
21 January 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Venteicher, A. S. et al. A human telomerase holoenzyme protein required for Cajal body localization and telomere synthesis. Science 323, 644–648 (2009).
Cohen, S. B. et al. Protein composition of catalytically active human telomerase from immortal cells. Science 315, 1850–1853 (2007).
Gomes, N. M. et al. Comparative biology of mammalian telomeres: hypotheses on ancestral states and the roles of telomeres in longevity determination. Aging Cell 10, 761–768, https://doi.org/10.1111/j.1474-9726.2011.00718.x (2011).
Blasco, M. A. et al. Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell 91, 25–34 (1997).
Erdmann, N., Liu, Y. & Harrington, L. Distinct dosage requirements for the maintenance of long and short telomeres in mTert heterozygous mice. Proc Natl Acad Sci USA 101, 6080–6085, https://doi.org/10.1073/pnas.0401580101 (2004).
Chiang, Y. J. et al. Expression of telomerase RNA template, but not telomerase reverse transcriptase, is limiting for telomere length maintenance in vivo. Molecular & Cellular Biology 24, 7024–7031 (2004).
Aisner, D. L., Wright, W. E. & Shay, J. W. Telomerase regulation: not just flipping the switch. Curr Opin Genet Dev 12, 80–85 (2002).
Jia, W. et al. A BAC transgenic reporter recapitulates in vivo regulation of human telomerase reverse transcriptase in development and tumorigenesis. FASEB J 25, 979–989, https://doi.org/10.1096/fj.10-173989 (2011).
Zhu, J., Zhao, Y. & Wang, S. Chromatin and epigenetic regulation of the telomerase reverse transcriptase gene. Protein & Cell 1, 22–32 (2010).
Wang, S. & Zhu, J. The hTERT gene is embedded in a nuclease-resistant chromatin domain. J Biol Chem 279, 55401–55410 (2004).
Cheng, D. et al. Regulation of human and mouse telomerase genes by genomic contexts and transcription factors during embryonic stem cell differentiation. Sci Rep 7, 16444, https://doi.org/10.1038/s41598-017-16764-w (2017).
Cheng, D. et al. Repression of telomerase gene promoter requires human-specific genomic context and is mediated by multiple HDAC1-containing corepressor complexes. FASEB J 31, 1165–1178 (2017).
Rangarajan, A., Hong, S. J., Gifford, A. & Weinberg, R. A. Species- and cell type-specific requirements for cellular transformation. Cancer Cell 6, 171–183 (2004).
Donehower, L. A. et al. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356, 215–221 (1992).
Artandi, S. E. et al. Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice. Nature 406, 641–645 (2000).
Zhang, F., Cheng, D., Wang, S. & Zhu, J. Human Specific Regulation of the Telomerase Reverse Transcriptase Gene. Genes (Basel) 7, https://doi.org/10.3390/genes7070030 (2016).
Wang, S., Zhao, Y., Leiby, M. A. & Zhu, J. Studying human telomerase gene transcription by a chromatinized reporter generated by recombinase-mediated targeting of a bacterial artificial chromosome. Nucleic Acids Res 37, e111 (2009).
Wong, M. S. et al. Regulation of telomerase alternative splicing: a target for chemotherapy. Cell Rep 3, 1028–1035, https://doi.org/10.1016/j.celrep.2013.03.011 (2013).
Rangarajan, A. & Weinberg, R. A. Opinion: Comparative biology of mouse versus human cells: modelling human cancer in mice. Nature Reviews. Cancer 3, 952–959 (2003).
Horikawa, I. et al. Differential cis-regulation of human versus mouse TERT gene expression in vivo: identification of a human-specific repressive element. Proc Natl Acad Sci USA 102, 18437–18442 (2005).
Wang, S., Zhao, Y., Hu, C. & Zhu, J. Differential repression of human and mouse TERT genes during cell differentiation. Nucleic Acids Res 37, 2618–2629 (2009).
Beattie, T. L., Zhou, W., Robinson, M. O. & Harrington, L. Reconstitution of human telomerase activity in vitro. Current Biology 8, 177–180 (1998).
Chen, J. L. & Greider, C. W. Determinants in mammalian telomerase RNA that mediate enzyme processivity and cross-species incompatibility. Embo J 22, 304–314 (2003).
Wong, M. S., Shay, J. W. & Wright, W. E. Regulation of human telomerase splicing by RNA:RNA pairing. Nat Commun 5, 3306, https://doi.org/10.1038/ncomms4306 (2014).
Wang, S., Zhao, Y., Leiby, M. & Zhu, J. A new positive/negative selection scheme for precise BAC recombineering. Mol Biotechnol 42, 110–116 (2009).
Zhao, Y., Wang, S. & Zhu, J. A multi-step strategy for BAC recombineering of large DNA fragments. Int J Biochem Mol Biol 2, 199–206 (2011).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8, 2281–2308, https://doi.org/10.1038/nprot.2013.143 (2013).
Wang, S., Hu, C. & Zhu, J. Transcriptional silencing of a novel hTERT reporter locus during in vitro differentiation of mouse embryonic stem cells. Mol Biol Cell 18, 669–677 (2007).
Leder, A., Daugherty, C., Whitney, B. & Leder, P. Mouse zeta- and alpha-globin genes: embryonic survival, alpha-thalassemia, and genetic background effects. Blood 90, 1275–1282 (1997).
Krupp, G. et al. Molecular basis of artifacts in the detection of telomerase activity and a modified primer for a more robust ‘TRAP’ assay. Nucleic Acids Research 25, 919–921 (1997).
Wang, S. & Zhu, J. Evidence for a relief of repression mechanism for activation of the human telomerase reverse transcriptase promoter. J Biol Chem 278, 18842–18850 (2003).
Zhu, J., Wang, H., Bishop, J. M. & Blackburn, E. H. Telomerase extends the lifespan of virus-transformed human cells without net telomere lengthening. Proc Natl Acad Sci USA 96, 3723–3728 (1999).
Canela, A., Vera, E., Klatt, P. & Blasco, M. A. High-throughput telomere length quantification by FISH and its application to human population studies. Proc Natl Acad Sci USA 104, 5300–5305, https://doi.org/10.1073/pnas.0609367104 (2007).
Perner, S. et al. Quantifying telomere lengths of human individual chromosome arms by centromere-calibrated fluorescence in situ hybridization and digital imaging. Am J Pathol 163, 1751–1756, https://doi.org/10.1016/S0002-9440(10)63534-1 (2003).
Acknowledgements
We thank Chloe Dugger for her technical support and Dr. Gang Chen for the discussion of this manuscript. We also thank the core facilities at WSU Spokane campus. The authors declare that they have no professional or financial affiliations that could be perceived to bias the presentation of this manuscript.
Author information
Authors and Affiliations
Contributions
S. Wang and J. Zhu designed research; D. Cheng, Y. Zhao, F. Zhang, and J. Zhang performed experiments; D. Cheng, S. Wang and J. Zhu analyzed data and prepared the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheng, D., Zhao, Y., Zhang, F. et al. Engineering a humanized telomerase reverse transcriptase gene in mouse embryonic stem cells. Sci Rep 9, 9683 (2019). https://doi.org/10.1038/s41598-019-46160-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46160-5
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.