Abstract
The design of microfluidic Lab on a Chip (LoC) systems is an onerous task requiring specialized skills in fluid dynamics, mechanical design drafting, and manufacturing. Engineers face significant challenges during the labor-intensive process of designing microfluidic devices, with very few specialized tools that help automate the process. Typical design iterations require the engineer to research the architecture, manually draft the device layout, optimize for manufacturing processes, and manually calculate and program the valve sequences that operate the microfluidic device. The problem compounds when engineers not only have to test the functionality of the chip but are also expected to optimize them for the robust execution of biological assays. In this paper, we present an interactive tool for designing continuous flow microfluidic devices. 3DμF is the first completely open source interactive microfluidic system designer that readily supports state of the art design automation algorithms. Through various case studies, we show 3DμF can be used to reproduce designs from literature, provide metrics for evaluating microfluidic design complexity and showcase how 3DμF is a platform for integrating a wide assortment of engineering techniques used in the design of microfluidic devices as a part of the standard design workflow.
Similar content being viewed by others
Introduction
The integration of microfluidics into experimental microbiology is limited to very few specific use cases in both academia and industry despite numerous advances in the technology1,2. Even in research areas where microfluidic devices have the potential to address numerous challenges in biological computation, primarily when used as the platform on which synthetic biological systems can be specified, designed, built, and tested3, the absence of widespread use of microfluidic devices is symptomatic of the problems faced in engineering and manufacturing them.
Currently, the standard engineering practices within research groups for microfluidics design fall within one of two distinct groups: (1) Component Designers (2) System Designers. Component Designers consist of researchers who engineer new components that take advantage of fluid mechanical phenomena. System Designers consist of researchers who design large scale systems with tens to thousands of geometric features. While many System Designers use Computer Aided Design (CAD) tools like Solidworks and AutoCAD, a sizable portion of researchers use vector design tools like Adobe Illustrator to draw the layouts of large multilayered designs manually. The designer’s need to organize geometries patterned at different depths drives the usage of these tools. For Component Designers, the ability to carefully draft complex parametric geometries and visualize 2D/3D projections of the device is essential. System Designers, on the other hand, value the ability to manage multilayered designs and perform rapid iterations efficiently. Despite being sophisticated software tools, both the vector editors and general purpose CAD tools place the burden of engineering the device for manufacturing and runtime design onto the user.
In electronics, Electronic Design Automation (EDA) tools take on the bulk of the labor-intensive design tasks. Without the availability of these tools, modern-day electronics would not exist. The formalization of the engineering process and the development of EDA tools have, in turn, affected the entire electronics and technology industry resulting in the significant proliferation of electronic devices. Hence, the potential benefits of introducing domain specific Computer Aided Design (CAD) tools have motivated researchers to develop design automation tools for microfluidic engineering. The survey4 published by Araci et al. provides an excellent overview of the various design automation tools developed in the field.
Additionally, current CAD tools and vector editors do not capture any information about the topology of the microfluidic device. The effects of this loss of information propagate throughout the design flow and deteriorate the researcher’s ability to cycle through design iterations rapidly. In light of the potential to automate microfluidic device designs, McDaniel et al. surveyed approximately 100 groups in academia and industry. This study, conducted as a part of an NSF I-Corps program on mVLSI (Microfluidics Very Large Scale Integration) software tools concluded that an interactive design tool which integrates design generation algorithms would be of high value to the microfluidics community5.
To that effect, we introduce 3DμF, an interactive design tool that can capture the architecture of the device during the design process. By incorporating the designs into a parametric microfluidic component library, 3DμF enables component designers to share their work with the rest of the community while simultaneously allowing system designers to quickly create large and complex microfluidic designs by reusing existing microfluidic technologies. This method increases standardization and helps mitigate a source of experimental non-reproducibility which plagues both microfluidics and the associated application fields. Various components that mix, distribute, and manipulate fluids, in addition to those intended for performing biological functions such as trapping and incubating cells, are a part of the microfluidic component library implemented by 3DμF. We envision that the microfluidic design automation (MDA) community would be able to extend the platform provided by 3DμF by leveraging the data models and visualization provided by the tool. The designs built in 3DμF would allow researchers to incorporate algorithms that could engineer the devices designed in 3DμF to become more fault-tolerant6, have a longer lifetime7, support automated testing8 and allow for the optimization of the control line layout9.
Through a series of case studies that include nine devices from literature and devices that capture microbiological protocols, we demonstrate how the tool can be used to design microfluidic technologies useful for Lab on a Chip (LoC) applications. We then introduce metrics that capture the effort required to design devices and show how 3DμF reduces the amount of effort to create them. We then show a collection of software examples, that demonstrate 3DμF’s potential as an extensible software platform by demonstrating various examples of how data captured by this tool can be used to integrate techniques that Specify, Design, Simulate, Build and Execute microfluidic devices. Through component level standardization and the facilitation of a systematic design process, we believe that 3DμF paves the first step in creating a standardized and automated engineering process for microfluidics similar to what is available to electronics designers.
Designing with 3DμF
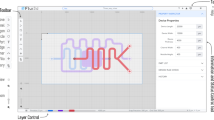
3DμF is a web-based design tool which allows System Designers to engineer their microfluidic designs. Available at http://3duf.org, the design environment is built entirely in Javascript and runs within the browser without the need for any complex cloud infrastructure. When loaded, the tool provides the user with a simple user interface as seen in Fig. 1J. which consists of a Design Canvas where the user can interactively construct their microfluidic design and a Primary Toolbar where they can select the different tools necessary to generate the design.
Sample steps to create a microfluidic device using 3DμF. The interface provides the user with a design canvas onto which they can place parametric components, and various tools in the toolbar that allows them to customize the placed components, select whether to place the component in the FLOW or the CONTROL layer and to choose how to export the designs. Users can utilize the design functionality provided by the tool via the component context menus and the primary toolbar as seen above. The video demonstration of this example is available in the SI.
Figure 1 shows how the Daridon Single Cell Trap10 is constructed using 3DμF. In Fig. 1A, the first step illustrates how the user can place components on the canvas by activating its corresponding tool on the primary toolbar Fig. 1J. Since the design requires a series of uniformly spaced PORTs, replicas of this component with the same parameters can be generated using the Generate Array tool as seen in step Fig. 1B. We then repeat the same process for the PUMP components to generate an array of actuators that pull liquid through each respective port. In step Fig. 1D, we manually create all connections between the ports and components using the CONNECTION tool (automated routing will be available in future versions based on Hightower et al.11).
Since the device we intend to replicate employs a custom cell trap10, we create a custom component in 3DμF by importing the design from an external CAD tool, as seen in Fig. 1E, in the form of a. DXF file. 3DμF then converts the imported geometry into a component object by inferring all the closed shapes described in the. DXF file. We complete the FLOW layer design by adding the remaining Reaction Chambers and creating the corresponding interconnections in Fig. 1F. Upon completing the FLOW layer, the user can switch to the CONTROL layer to add Valves (using the VALVE tool) and the corresponding control infrastructure. The tool then automatically aligns the valve and redraws the connection in the case of Valve3D12. Finally, the user then creates the corresponding control infrastructure by placing Ports and Connections on the control layer in step Fig. 1G, by repeating the steps A and D. Using 3DμF, this design with 47 components and 103 channel elements was constructed in less than 20 minutes and is available in the Supplementary Information (SI).
Results
Parametric Design Engineering with 3DμF
3DμF is a semi-automated design tool that employs design techniques typically seen in electronics CAD tools, such as PCB (printed circuit board) editors, in the design of microfluidic devices. 3DμF automatically captures the relationships between objects placed on the design canvas that would be necessary to understand the underlying architecture of a microfluidic device created within its design environment. While the provided user interface might seem like a simple environment lacking the advanced drafting and transformation features often seen in design tools such as SolidWorks and AutoCAD, the simple user interface abstracts much of the design automation that attempts to obviate the needs of the user to use the features as mentioned previously. The data captured using 3DμF drastically reduces the effort necessary to create larger designs with moderately high complexity. We compare the various features that are uniquely available within 3DμF against other tools typically used by researchers to design microfluidic devices, as seen in Fig. 2. While 3DμF may appear to fall short in Mechanical Design, the simple user interface is sufficient for visualizing the 2D profiles of the placed components. 3DμF excels in the areas of Design Techniques, Design Automation and Extensibility as it was designed explicitly for creating large microfluidic devices leveraging the various strategies used by researchers in microfluidic design automation.
This table compares the functionality of different design tools to design and optimize microfluidic systems. It also includes the concepts that enable 3DμF to provide the user with each of the features referenced in the Methods section: PCL - Parametric Component Library, FLD - Functional Layer Design, CCA - Component-Connection Architecture and DFM - Design for Manufacturing, Github - https://github.com/CIDARLAB/3DuF.
Since the effort required to design microfluidic devices can vary widely in each case, we developed metrics to measure the effort required to create each component in the 3DμF library. We can then compute the effort required to design the full device by computing the total effort required to create all the components. Since the process of designing a component consists of two stages, Design and Parameterization, the total effort required is expressed as EPrimitive = EDesign + EParameterization, where Effort E* is measured in actions. Figure 3 illustrates how the design and parameterization efforts can be captured as EDesign = CBasefProcedural(X) and EParameterization = CIdentification + NParams(CConstraint + CValue) respectively. Using these formulae, the Effort required to create a MIXER with 8 bends would be 88 actions (CBase = 6, CIdentification = 20, CValue = 4,fProcedural(X) = X|X = 8). Detailed explanation of each of the terms is available in SI, Section 8.1 and 8.2.
Effort can be quantitatively measured. The figure shows how the effort required create a component EPrimitive is composed of two expressions, Design (EDesign) and Parameterize (EParameterization). Both the Design and Parameterization involve two and three steps respectively and are represented by the different terms (CBasefProcedural(X), CIdentification, etc.) in the EDesign, EParameterization expressions. Additional details are available in the SI.
In addition to expanding the automation capabilities of the design, 3DμF helps reduce the debugging effort required to design devices. When designing complex microfluidic systems, one of the most time consuming and error-prone portions of the design drafting process is the parameterization of the various geometries constituting the device design. 3DμF inherently reduces a large amount of the effort that the designer has to undertake in assigning proper parametric geometry constraints.
Case Study 1 - Microfluidics from Literature
In practice, it is often difficult and sometimes impossible to recreate devices published in literature since the images and written specifications accompanying the manuscript are often insufficient to reproduce the device. Even in some cases, when the CAD designs are given, it is almost impossible to parameterize the design without substantial characterization work. Conventional CAD tools cannot capture the design as a composition of individual objects; they instead store the entire design as a large set of shapes. 3DμF’s capability to generate a structured device described as a network of connected components and connections partially addresses this fundamental problem in engineering communication. Further, since 3DμF encodes all of the designs in a superset of ParchMINT13 (an open microfluidic interchange format), anyone can write simple scripts to extract all the necessary design information. The effort estimations seen in Fig. 4 was computed by summing up the individual efforts required for designing each of the components in the design file using a python script(Available in SI, Section 9). In order to show the potential of 3DμF in designing devices with the standard component library, we designed nine microfluidic devices found in literature used for lab automation. The different devices designed as a part of the case study were selected for their popularity(citations) or as their relevance as scientific research platforms3. The designs seen in Fig. 4 are: (A) Quake Droplet Generator14 [Citations:1917] (B) Kobayashi Hydrogenation Chip15 [Citations:561] (C) Nozzle Droplet Generator Chip16 [Citations:298] (D) A single microchemostat unit17 [Citations:595] (E) Dynamic Signaling Chip18 [Citations:76] (F) Daridon Single Cell Trap10 [Citations:636] (G) Kinase Radioassay Chip19 [Citations:43] (H) Cell-Microenvironment Chip20 [Citations:58] (I) Oligonucleotide Synthesis Chip21 [Citations:85]. Each of these publications were only accompanied by pictures and limited textual descriptions of some design parameters making it virtually impossible to recreate the device from what is given in the paper. All of the devices in Fig. 4 were fabricated with the intention of demonstrating 3DμF’s capability for capturing the complexity of designs seen in literature and not to create functional prototypes. By fabricating the devices replicated in 3DμF, we were able to show that reasonably complex devices, as seen in literature, can be designed and fabricated using 3DμF. Further Fig. 4 1 and 2 shows how 3DμF reduces the amount of effort required by the system designer to create each of the designs. Additional information is available in the SI, Section 9. The nine designs are available at https://cidarlab.github.io/3DuF-Paper-Designs/.
Reproducing Designs from Literature (A-I). Graph 1 characterizes the designs using the number of components and connection elements present in the design. The radius of the markers shows that the total effort (\({E}_{Total}=\sum {E}_{Primitive}\)) required to draw and parameterize the design using a CAD tool eclipses the effort required to create the design using 3DμF. Graph 2 shows the breakdown of the effort required in Drawing and Parameterization for each of the design with different design tools. It also shows how the parameterization process requires more EParameterization since the manual addition of geometric constraints would typically require the user to continuously modify the constraints while incrementally building the design to prevent overconstraining. Graph 3 further shows the distribution by which different components compose the designs from literature, and it also shows how that even though the complexity of the design might scale in size, the relative distribution of different kinds of components is similar in all of the designs.
Case Study 2 - Modular System Design
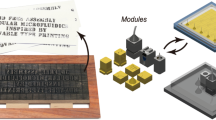
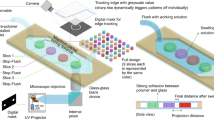
When designing large microfluidic systems to perform complex assays, using a modular design approach is advantageous. Each modular subsystem within a library is independently designed and tested to be functional. Therefore when building a complex system, each validated modular subsystem is already functional and “only” requires the user to create connections between them. Figure 5 represents a conceptual microfluidic system designed to perform cloning reactions illustrates an example of a system that performs complex assays. Cloning, a common molecular biology technique used heavily in synthetic biology, often uses restriction enzymes to digest DNA of interest into fragments with compatible ends which undergo ligation. This system, as shown in Fig. 5 reuses four different designs from the MARS repository (http://2017.igem.org/Team:BostonU_HW) which were each individually designed using 3DμF. Because of the component connection abstraction internally used by 3DμF, designers do not encounter problems where the geometric constraints of different geometries become entangled requiring them to spend a significant amount of time reassigning the geometric constraints when designs become ‘overdefined’(SolidWorks) or ‘overconstrained’(AutoCAD). Hence, users can safely import 3DμF designs requiring no additional user effort to prevent changes made to an individual component to propagate through the rest of the design — additional information on the four MARS designs is available in the SI, Section 10.
The figure shows the concept of a system that is intended to perform cloning. 3DμF was used to rapidly manufacture and iterate through each of the stages within four months. The system reuses existing 3DμF designs, allowing the user to design and test each subsystem separately before integrating them into a larger system with external purification steps22. In the Cell Lysis stage, the plasmids (genetic fragments) are released from cells and captured on magnets. In the DNA digestion stage, the plasmids are digested, using restriction enzymes, into compatible DNA fragments. In the Plasmid Ligation stage, the DNA fragments are ligated together to form a new, complete plasmid. Finally, in the Transformation stage, the new plasmid is transformed into E. coli cells for propagation and expression.
Platform Extensibility Demonstration
3DμF not only serves as a design tool but also as a framework for the integration of advanced engineering practices into the design workflow. In this case study, we show five examples of such methodologies demonstrated using 3DμF. They span five different stages of the device design lifecycle namely, Specify, Design, Simulate, Build, Execute. Extended summaries for each the demonstrations and links to the source code are available in the SI, Section 11.
Specify - Support for High-Level Device Descriptions
Hardware Description Languages (HDL) have been critical in synthesizing digital electronic designs. They allow the engineer to specify the functionality of the device using programming language syntax, relying on automation algorithms to generate the actual circuit that performs the functionality. Figure 6A shows how MINT23 (a microfluidic HDL) can be compiled and imported into 3DμF. This demonstration shows how a platform like 3DμF can be used to integrate engineering workflows where designs are synthesized from high-level description languages.
3DμF is not just a design tool. The architectural topology captured by the tool allows 3DμF to be used as a platform for engineering microfluidic devices. A1, A2. (Specify) Shows how the design described in a hardware description language (MINT) can be imported into 3DμF and B1, B2 (Simulate) shows how the design files can be imported into external software that can analyze and estimate flow rates and pressure at different locations on the design.
Design - Support for Design of Experiment Techniques
Typically the design spaces for most microfluidic geometries are large, which can generate a wide range of performance. Researchers often utilize “Design of Experiments” methodologies to either simulate or rapid prototype24,25 orthogonal device designs that help them evaluate the performance of the device in the design space. Some of the methods often used in the generation of the orthogonal design of experiments methodologies are the Taguchi Methods26. System Designers can take advantage of such experimental methodologies and use 3DμF’s inference of the design to generate the orthogonal array automatically. A video of this demonstration is available in the SI.
Simulate - Support for Network Characterization
In order to circumvent the limitations of finite element simulation, researchers have devised strategies that rely on data-mining pre-simulated results for design exploration27 and the interaction of particles in fluid flow28. By leveraging the component connection abstraction, it is possible to apply techniques used in electronic circuit simulation29 on the microfluidic network by deriving electrical circuit equivalent models30. In order to illustrate how designs created using the Component-Connection Abstraction can be simulated, we demonstrate a simple tool that converts the design created in 3DμF into an equivalent electrical model to calculate the Pressure and Flow Rates at various points of interest as seen in Fig. 6.
Build - Support for Fabrication
The current release of 3DμF has built-in support for generating design files for CNC micro-milling, utilizing the 2.5D Manufacturing Layer Generation technique. Similarly, the 2.5D Manufacturing Layer Generation algorithm could be adapted to generate design files targeting other popular manufacturing processes like Soft Lithography31.
Execute - Support for Valve Control Programmability
A significant factor that inhibits the transfer of designs is the non-standard electronic infrastructure used to run microfluidic devices. By creating designs in 3DμF, designers can reuse the component names used in the design while describing the behavior in the form of valve actuation sequences(standardized). Additionally, the component-connection graph generated by 3DμF would allow researchers to design algorithms that can synthesize the valve sequences necessary to control the fluid flow when integrating with high-level languages such as BioStream32.
Discussion
3DμF’s capabilities as an open-source, extensible design editor enabled usage highlighted by Case Studies 1 and 2. In particular, the ability of the component library to be used to design the existing literature-grade microfluidic device was demonstrated by Case Study 1, covering a range of widely used device designs. 3DμF, as seen in Fig. 4 additionally reduces the effort to design devices by half. However, 3DμF has the potential to reduce the effort even further with its ability to allow for different steps in the design process to be automated. 3DμF’s use of standardization at multiple levels enables its interactions with existing microfluidic design tools and methodologies. Without any additional effort, each design in Case Study 1 could be imported into tools that integrate algorithmic advances in MDA6,7,9,33.
Though 3DμF supports the import of components using DXF, the size of the component library is a significant drawback with 3DμF since the geometries imported from DXF files lack the metadata required for fully integrating the design with any of its platform capabilities and since the user cannot create custom components using 3DμF. However, as the LoC community using 3DμF grows, the potential complexity of chips made by integrating existing work would only increase with every contribution. Every addition to the component library would then be able to leverage 3DμF’s capabilities and allow for large-scale standardization of microfluidic device design and their corresponding operation and manufacturing infrastructure.
While Case Study 2 highlights 3DμF’s usefulness in enabling larger microfluidic system designs, it could take substantial effort to modify such a design based on changes to a single component between design iterations and different operating conditions. Future work could automate this process by incorporating algorithms that synthesize designs for these different scenarios. 3DμF could then be used to resize designs leveraging the underlying component-connection architecture. However, most of these parameters and rules for the synthesis of a microfluidic system would be context-, application-, flowrate-specific. Hence, we believe that the research community would leverage 3DμF and create the necessary extensions.
Methods
Parametric Component Library
The components that are a part of the component library are all procedurally generated by 3DμF. The components not only abstract microfluidic functionality but also lay the foundation to standardize the engineering flow. Figure 7 shows how using 3DμF can drastically reduce the effort required to design and parameterize different geometries. Figure 7 also shows how the benefits of having procedurally generated component libraries can drastically reduce the effort involved in generating components like MIXER and MUX which require effort that scales by linear and polynomial difficulty. Effort, a heuristic metric used throughout the paper (measured in actions) attempts to naively capture the number of actions required by the user in different design tools to draw and parameterize for creating each component as seen in the SI, Section 8.
The parametric component library significantly reduces the effort required to draw different microfluidic geometries. Graph A compares the effort taken to design microfluidic geometries using CAD design tools vs. the effort required to design using 3DμF. Graphs B and C show how the effort required to design the procedurally generated geometries scales with the number of outputs when created using CAD tools and how the effort required is nearly negligible when designing using 3DμF. The Procedure and Calculations for computing the Effort for all the components is available in the SI, Section 8.
Component Connection Architecture
The component connection architecture used by 3DμF to model the microfluidic design is the core enabling methodology that allows the tool to set itself apart from all the other available tools. It enables the designer to capture the architecture underlying the geometries that are drawn on the canvas and is the foundation for most of the 3DμF’s functionality. Rather than drawing polygons, manually setting geometric constraints, the component connection architecture allows the user to place predefined geometries in the form of components directly and then connect them using connections. Components allow the complexity of fluid mechanics to be abstracted, allowing engineers with little or no experience in the specific area of fluid mechanics to implement a diverse range of microfluidic functionality quickly.
Functional Layer Design
Figure 1 shows how a microfluidic device’s design abstracted into components and connections belonging to different layers, the Flow LAYER which consists of geometries that perform the primary operation of the microfluidic device and the Control LAYER which consists of the geometries that control the movements of the fluids in its corresponding Flow LAYER. 3DμF allows for multilayered designs in a manner that replicates the visualization in AutoCAD and Illustrator. For designs that have more than a single Flow and Control layer, 3DμF allows for the creation of multi-leveled designs that can be easily added/deleted using the ‘+’ and ‘delete’ buttons seen in Fig. 1H. 3DμF automatically creates a pair of corresponding Flow and Control layers for every new level that is added to design. The tool additionally updates the internal data model to store the relationship between the channels and the valves used in automatic control sequence generation algorithms34 as key-value pairs.
Design for Manufacturing
3DμF allows for the components placed in a single functional layer to have multiple heights. Typically during the fabrication, the designer would have to manually keep track of the features belonging to different depths (layers when using Adobe Illustrator or using a hierarchical coloring system when using AutoCAD35). 3DμF internally assigns each of the geometries a DFM (Design for Manufacturing) class, namely XY, XYZ, Z and EDGE that indicating the fabrication complexity of the geometry. By grouping all of the features with different heights in the XY class, 3DμF obviates the need for the user to follow any of the cumbersome schemes described earlier. In practice, the user would specify the ‘height’ parameter for each of the components placed and let 3DμF handle the process of generating the different “Manufacturing Layers”. The combination of the “DFM Classes”, “Manufacturing Layers” and “Functional Layers” captures a majority of the complexity faced when designing mLSI designs and allows for the fabrication support to be extended for different kinds of fabrication processes. Details of the DFM classes, manufacturing layer generation, and the fabrication protocol are available in the SI.
Conclusion
The design of microfluidic devices for the biological sciences is currently limited for numerous reasons. These include the lack of domain-specific computer-aided design environments. This paper presents a lightweight, web-based, open source tool (3DμF) that addresses many of the current software shortcomings while providing several innovative features. We illustrate both how existing, popular microfluidics designs can be created as well as how to quantify the effort associated with creating microfluidic designs in-silico. Our approach builds on techniques used in the EDA community and provides a solid foundation that can be expanded on by individual, motivated software engineers. We have provided numerous design examples, videos, and demonstration software that complement the design tool its capabilities.
In electronics, the availability of free PCB design tools such as KiCAD (kicad-pcb.org/) (open source) and Eagle(autodesk.com/products/eagle/) helped reduce the barrier of entry and push the envelope of what can be designed by individuals, thus increasing new market opportunities. Similarly, 3DμF is poised to play an essential role in the growing microfluidic design ecosystem that includes Metafluidics36, AutoPAD37 and other approaches to democratize microfluidics38,39.
References
Whitesides, G. M. The origins and the future of microfluidics. Nature 442, 368–373, https://doi.org/10.1038/nature05058 (2006).
Sackmann, E. K., Fulton, A. L. & Beebe, D. J. The present and future role of microfluidics in biomedical research. Nature 507, 181–189, https://doi.org/10.1038/nature13118 (2014).
Huang, H. & Densmore, D. Integration of microfluidics into the synthetic biology design flow. Lab on a Chip 14, 3459–3474, https://doi.org/10.1039/C4LC00509K (2014).
Araci, I. E., Pop, P. & Chakrabarty, K. Microfluidic very large-scale integration for biochips: Technology, testing and fault-tolerant design. In 2015 20th IEEE European Test Symposium (ETS), 1–8, https://doi.org/10.1109/ETS.2015.7138736 (2015).
McDaniel, J., Grover, W. H. & Brisk, P. The case for semi-automated design of microfluidic very large scale integration (mVLSI) chips. In Design, Automation Test in Europe Conference Exhibition (DATE), 2017, 1793–1798, https://doi.org/10.23919/DATE.2017.7927283 (2017).
Huang, W. L., Gupta, A., Roy, S., Ho, T. Y. & Pop, P. Fast architecture-level synthesis of fault-tolerant flow-based microfluidic biochips. In Design, Automation Test in Europe Conference Exhibition (DATE), 2017, 1667–1672, https://doi.org/10.23919/DATE.2017.7927262 (2017).
Tseng, K.-H., You, S.-C., Minhass, W. H., Ho, T. Y. & Pop, P. A network-flow based valve-switching aware binding algorithm for flow-based microfluidic biochips. In 2013 18th Asia and South Pacific Design Automation Conference (ASP-DAC), 213–218, https://doi.org/10.1109/ASPDAC.2013.6509598 (2013).
Hu, K., Ho, T. Y. & Chakrabarty, K. Test generation and design-for-testability for flow-based mVLSI microfluidic biochips. In 2014 IEEE 32nd VLSI Test Symposium (VTS), 1–6, https://doi.org/10.1109/VTS.2014.6818760 (2014).
Yao, H., Ho, T.-Y. & Cai, Y. PACOR: Practical Control-layer Routing Flow with Length-matching Constraint for Flow-based Microfluidic Biochips. In Proceedings of the 52Nd Annual Design Automation Conference, DAC’15, 142:1–142:6, https://doi.org/10.1145/2744769.2744887 (ACM, New York, NY, USA, 2015).
Wheeler, A. R. et al. Microfluidic Device for Single-Cell Analysis. Analytical Chemistry 75, 3581–3586, https://doi.org/10.1021/ac0340758 (2003).
Hightower, D. W. A Solution to Line-routing Problems on the Continuous Plane. In Proceedings of the 6th Annual Design Automation Conference, DAC’69, 1–24, https://doi.org/10.1145/800260.809014 (ACM, New York, NY, USA, 1969).
Grover, W. H., Skelley, A. M., Liu, C. N., Lagally, E. T. & Mathies, R. A. Monolithic membrane valves and diaphragm pumps for practical large-scale integration into glass microfluidic devices. Sensors and Actuators B: Chemical 89, 315–323, https://doi.org/10.1016/S0925-4005(02)00468-9 (2003).
Crites, B. et al. ParchMint: A Microfluidics Benchmark Suite. In 2018 IEEE International Symposium on Workload Characterization (IISWC), 78–79, https://doi.org/10.1109/IISWC.2018.8573487 (2018).
Thorsen, T., Roberts, R. W., Arnold, F. H. & Quake, S. R. Dynamic Pattern Formation in a Vesicle-Generating Microfluidic Device. Physical Review Letters 86, 4163–4166, https://doi.org/10.1103/PhysRevLett.86.4163 (2001).
Kobayashi, J. et al. A Microfluidic Device for Conducting Gas-Liquid-Solid Hydrogenation Reactions. Science 304, 1305–1308, https://doi.org/10.1126/science.1096956 (2004).
Tan, Y.-C., Cristini, V. & Lee, A. P. Monodispersed microfluidic droplet generation by shear focusing microfluidic device. Sensors and Actuators B: Chemical 114, 350–356, https://doi.org/10.1016/j.snb.2005.06.008 (2006).
Balagadde, F. K., You, L., Hansen, C. L., Arnold, F. H. & Quake, S. R. Long-Term Monitoring of Bacteria Undergoing Programmed Population Control in a Microchemostat. Science 309, 137–140, https://doi.org/10.1126/science.1109173 (2005).
Wang, C. J., Bergmann, A., Lin, B., Kim, K. & Levchenko, A. Diverse Sensitivity Thresholds in Dynamic Signaling Responses by Social Amoebae. Sci. Signal. 5, ra17–ra17, https://doi.org/10.1126/scisignal.2002449 (2012).
Fang, C. et al. Integrated Microfluidic and Imaging Platform for a Kinase Activity Radioassay to Analyze Minute Patient Cancer Samples. Cancer Research 70, 8299–8308, https://doi.org/10.1158/0008-5472.CAN-10-0851 (2010).
Liu, W. et al. An integrated microfluidic system for studying cell-microenvironmental interactions versatilely and dynamically. Lab on a Chip 10, 1717–1724, https://doi.org/10.1039/C001049A (2010).
Lee, C.-C., Snyder, T. M. & Quake, S. R. A microfluidic oligonucleotide synthesizer. Nucleic acids research 38, 2514–2521 (2010).
Wen, J., Legendre, L. A., Bienvenue, J. M. & Landers, J. P. Purification of Nucleic Acids in Microfluidic Devices. Analytical Chemistry 80, 6472–6479, https://doi.org/10.1021/ac8014998 (2008).
Huang, H. Fluigi: An end-to-end Software Workflow for Microfluidic Design. Ph.D. thesis, Boston University (2015).
Silva, R. et al. Rapid prototyping and parametric optimization of plastic acoustofluidic devices for blood–bacteria separation. Biomedical Microdevices 19, 70, https://doi.org/10.1007/s10544-017-0210-3 (2017).
Lashkaripour, A., Rodriguez, C., Ortiz, L. & Densmore, D. Performance tuning of microfluidic flow-focusing droplet generators. Lab on a Chip 19, 1041–1053, https://doi.org/10.1039/C8LC01253A (2019).
Taguchi, G. et al. Taguchi methods: design of experiments, vol. 4 (Amer Supplier Inst, 1993).
Wang, J., Brisk, P. & Grover, H. Â. W. Random design of microfluidics. Lab on a Chip 16, 4212–4219, https://doi.org/10.1039/C6LC00758A (2016).
Wang, J., Rodgers, V. G. J., Brisk, P. & Grover, W. H. Instantaneous simulation of fluids and particles in complex microfluidic devices. PLOS ONE 12, e0189429, https://doi.org/10.1371/journal.pone.0189429 (2017).
McCalla, W. J. Fundamentals of Computer-Aided Circuit Simulation. The Springer International Series in Engineering and Computer Science (Springer US, 1988).
Oh, K. W., Lee, K., Ahn, B. & Furlani, E. P. Design of pressure-driven microfluidic networks using electric circuit analogy. Lab on a Chip 12, 515–545, https://doi.org/10.1039/C2LC20799K (2012).
Qin, D., Xia, Y. & Whitesides, G. M. Soft lithography for micro- and nanoscale patterning. Nature Protocols 5, 491–502, https://doi.org/10.1038/nprot.2009.234 (2010).
Thies, W., Urbanski, J. P., Thorsen, T. & Amarasinghe, S. Abstraction layers for scalable microfluidic biocomputing. Natural Computing 7, 255–275, https://doi.org/10.1007/s11047-006-9032-6 (2008).
Hu, K., Yu, F., Ho, T. Y. & Chakrabarty, K. Testing of Flow-Based Microfluidic Biochips: Fault Modeling, Test Generation, and Experimental Demonstration. IEEE Transactions on Computer-Aided Design of Integrated Circuits and Systems 33, 1463–1475, https://doi.org/10.1109/TCAD.2014.2336215 (2014).
Huang, H. & Densmore, D. Fluigi: Microfluidic Device Synthesis for Synthetic Biology. J. Emerg. Technol. Comput. Syst. 11(26), 1–26:19, https://doi.org/10.1145/2660773 (2014).
Lake, M. et al. Microfluidic device design, fabrication, and testing protocols. Nature Protocols (2015).
Kong, D. S. et al. Open-source, community-driven microfluidics with Metafluidics. Nature Biotechnology 35, 523–529, https://doi.org/10.1038/nbt.3873 (2017).
DeChiara, N. S., Wilson, D. J. & Mace, C. R. An open software platform for the automated design of paper-based microfluidic devices. Scientific Reports 7, 16224, https://doi.org/10.1038/s41598-017-16542-8 (2017).
Lashkaripour, A., Silva, R. & Densmore, D. Desktop micromilled microfluidics. Microfluidics and Nanofluidics 22, 31, https://doi.org/10.1007/s10404-018-2048-2 (2018).
Walsh, D. I., Kong, D. S., Murthy, S. K. & Carr, P. A. Enabling Microfluidics: from Clean Rooms to Makerspaces. Trends in Biotechnology 35, 383–392, https://doi.org/10.1016/j.tibtech.2017.01.001 (2017).
Acknowledgements
We want to thank Dr. Phillip Brisk’s group at UC Riverside for their joint effort in designing the standard interchange format. Swapnil Bhatia for his feedback on the engineering methodology. Dylan Samperi for his initial work on SS-Analyzer and the Microfluidic Building Blocks. Prashant Vaidyanathan for his extensive feedback that helped shape this manuscript. Finally, we would like to thank Aaron Heuckroth for the inception of 3DμF. This work was funded by the National Science Foundation’s Award #1522074.
Author information
Authors and Affiliations
Contributions
R.S.– interchange standard, demonstration software, 3DμF development, design, fabrication, writing, and organization of manuscript. J.L.– interchange standard and 3DμF development. D.S. & S.N. - microfluidic building blocks. D.D.– overarching design paradigms, hardware, and software access philosophy, funding of equipment, personnel/laboratory resources.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sanka, R., Lippai, J., Samarasekera, D. et al. 3DμF - Interactive Design Environment for Continuous Flow Microfluidic Devices. Sci Rep 9, 9166 (2019). https://doi.org/10.1038/s41598-019-45623-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45623-z
This article is cited by
-
Open-source interactive design platform for 3D-printed microfluidic devices
Communications Engineering (2024)
-
Design automation of microfluidic single and double emulsion droplets with machine learning
Nature Communications (2024)
-
Microfluidics-driven high-throughput phenotyping and screening in synthetic biology: from single cells to cell-free systems
Biotechnology and Bioprocess Engineering (2024)
-
Finger-operated pumping platform for microfluidic preparation of nanoparticles
Microfluidics and Nanofluidics (2024)
-
Biofoundries: Bridging Automation and Biomanufacturing in Synthetic Biology
Biotechnology and Bioprocess Engineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.