Abstract
Solid organ transplant (SOT) recipients are especially at risk of developing infections by multidrug resistant bacteria (MDR). In this study, the biofilm-forming capability of 209 MDR strains (Escherichia coli n = 106, Klebsiella pneumoniae n = 78, and Enterobacter spp. n = 25) isolated from rectal swabs in the first 48 hours before or after kidney (93 patients), liver (60 patients) or kidney/pancreas transplants (5 patients) were evaluated by using a microplate assay. Thirty-nine strains were isolated before transplant and 170 strains were isolated post-transplant. Overall, 16% of E. coli strains, 73% of K. pneumoniae strains and 4% Enterobacter strains showed moderate or strong biofilm production. Nine strains isolated from infection sites after transplantation were responsible of infections in the first month. Of these, 4 K. pneumoniae, 1 E. coli and 1 Enterobacter spp. strains isolated pre-transplant or post-transplant as colonizers caused infections in the post-transplant period. Our results suggest that in vitro biofilm formation could be an important factor for adhesion to intestine and colonization in MDR K. pneumoniae strains in SOT recipients, but this factor appears to be less important for MDR E. coli and Enterobacter spp.
Similar content being viewed by others
Introduction
Solid organ transplant (SOT) recipients have an increased risk of developing bacterial infections because they receive long immunosuppressive therapy to avoid rejection. Furthermore, SOT recipients are especially at risk of developing infections by bacteria with intrinsic or acquired antimicrobial resistance as they are frequently exposed to antibiotics in healthcare settings1,2,3. Currently, the increasing prevalence of multi-drug resistant (MDR) Gram-negative pathogens, such as extended-spectrum β-lactamase-producing (ESBLs) Enterobacteriaceae strains, strains with overproduction of intrinsic chromosomal AmpC β-lactamases4,5, and carbapenem-resistant K. pneumoniae and E. coli strains, are of particular concern in SOT recipients.
Important risk factors for nosocomial infections in SOT recipients are, among others, the use of catheters, hospitalization in intensive care units, and pre-transplant faecal carriage of multidrug-resistant isolates6,7.
The Enterobacteriaceae family is a heterogeneous group of Gram-negative bacteria, whose natural habitat is the intestinal tract of humans and animals. Among the hospital-acquired infections due to Enterobacteriaceae, urinary tract infections (UTIs) are the most common and lower respiratory tract and bloodstream infections are the most lethal8,9,10. Virulence factors in Enterobacteriaceae include different adhesins, hemolysin production, serum resistance, and biofilm formation. These factors, especially the ability to form biofilms in the human intestine may contribute to gut colonization and have a great impact on the function of the intestinal microbiome and its interactions with the host11,12,13,14,15,16,17,18,19.
A biofilm is often defined as a structured community of microorganisms enclosed in a self-produced matrix and adherent to an inert or living surface. Growth in biofilms enhances the survival of bacterial populations in hospital settings and inside patients, increasing the probability of causing nosocomial infections20,21,22. Also, several investigations have revealed that horizontal gene transfer is connected with biofilm formation23,24.
Previous studies showed a relationship between the presence of antimicrobial resistance genes and biofilm formation in E. coli and Klebsiella strains isolated from patients with different infections14,25,26, however, the correlation between antibiotic resistance and biofilm formation for most pathogens (i.e. Enterobacter spp.) remains unclear. The goal of present study, which is part of a wider research project (The ENTHERE study), was to analyse the capacity to form biofilm of 209 Enterobacteriaceae strains isolated from pre- or post-transplant patients. This investigation could contribute to a better understanding the relationship between adherence capability, antimicrobial resistance, and pathogenicity of MDR bacteria in SOT recipients.
Results
Antibiotic resistance genes
The most common genes found in the strains isolated in this study were extended-spectrum β-lactamases, followed by carbapenemases, being blaCTXM and blaSHV most prevalent in E. coli, blaCTXM and blaOXA-48 in K. pneumoniae and blaCTXM and AmpC hyperproduction in Enterobacter spp. Distribution of antibiotic resistance genes in Enterobacteriaceae strains isolated from transplant patients is shown in Table 1.
Hemagglutination in E. coli strains
Hemagglutination (HA) was observed in 23 strains (21.7%). HA was mannose sensitive (MS) in 9 strains (8.5%) and mannose resistant (MR) in 14 strains (13.2%) (Supplementary Table 1). We observed an inverse relationship between hemagglutinating strains and biofilm formation, because the isolates that show either sensitivity or resistance to the hemagglutination in presence of mannose are non- or weak biofilm former strains.
Biofilm formation by Enterobacteriaceae strains
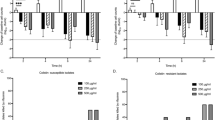
The ability of Enterobacteriaceae strains to form biofilms was quantified by crystal violet (CV) staining after 48 h at 37 °C. Bacterial growth was, in general, not strongly influenced by the culture conditions, as estimated by the colony forming units (CFU) of planktonic cells in 4 strains of each species, with numbers in the range from 1.95 × 108 to 6.45 × 108 CFUs/well. All the biofilms were found at the liquid-air interface. Overall, 16% of E. coli strains and 73% of K. pneumoniae strains showed moderate or strong biofilm production while only one Enterobacter strain showed moderate biofilm production (Fig. 1).
Seventy-two percent of 67 K. pneumoniae isolates shown moderate or strong biofilm formation (23 from renal transplants and 25 from hepatic transplants) whereas only 16 post-transplant E. coli isolates (19.5%) were biofilm forming strains, mostly from renal transplant recipients (n = 11).
Morphology of biofilms was found to be variable, even for biofilms formed by the same species. Six representative infection-related isolates of the three species were selected for confocal microscopy analysis. These strains showed poor, weak, moderate/strong biofilm formation. All strains formed similar biofilms before and after re-emergence as infection related strains in transplant patients (Fig. 2). K. pneumoniae strains 25/27 and 158/163 formed biofilms with firmly packed, mushroom-shaped microcolony structures. Large red masses can be seen inside these structures, possibly due to the early formation of these microcolonies. These structures did not appear in the biofilms formed by E. coli (weak biofilm formation) which showed a thinner and less compact structure. The moderate biofilm producer K. pneumoniae 168/169 formed flat biofilms with loosely packed irregular microcolony structures and composed mainly by dead cells, as clearly showed by measurements of red/green pixel intensities inside biofilms (Supplementary Fig. 1). Moderate or poor biofilm producer strains (according to CV data) K. pneumoniae 171/172 and Enterobacter 120/121 respectively, showed few attached cells with loosely packed irregular microcolony structures with little heterogeneity.
Biofilm formation by the six Enterobacteriaceae strains responsible of infections. These strains were compared with their respective isolates pre-transplant (Pre-T) or isolates from rectal swabs post-transplant (colonizing strains, C-S). Up: wells were stained with Crystal Violet inside 24-well plates. Source of isolation during infections is indicated. Down: representative examples of CLSM images of selected strains after biofilm formation. Bacteria were stained with the BacLight LIVE/DEAD viability kit. Live cells fluoresce in green with Syto 9 dye and dead cells are stained red with propidium iodide. Original magnification: ×200.
Infections by MDR in SOT recipients
Transplant recipients were colonized with MDR Enterobacteriaceae because the number of MDRE isolates post-transplant was 4.3 times the number of MDRE isolates pre-transplant (E. coli 81 vs. 25, K. pneumoniae 67 vs. 11, and Enterobacter 22 vs. 3).
Eight out of 158 (5.1%) patients enrolled in the project developed infection and six of the strains isolated from these patients (4%) were re-emergent strains detected in pre-transplant rectal swab samples or colonizers strains post-transplant (Fig. 3). All infection related strains were isolated during the first month post-transplant. These strains caused urinary infections (K. pneumoniae strain 172 and E. coli strain 199), bacteraemia (Enterobacter cloacae strain 121) and abdominal abscesses (K. pneumoniae strain 163) in renal transplant receptors, and skin abscess (K. pneumoniae strain 27) and cholecystitis (K. pneumoniae strain 169) in hepatic transplant receptors (Supplementary Table 1). Two other strains showing distinct pulsed-field gel electrophoresis (PFGE) patterns were not genetically related (not shown) and caused urinary tract infections (E. coli strain 228 and K. pneumoniae strain 239) in renal and hepatic transplant recipients, respectively. Patients infected with K. pneumoniae strains 27, 169 and 172 or E. coli strains 199 and 228 required additional antimicrobial treatment whereas transplant recipients infected with E. cloacae 121, and K. pneumoniae 163 and 239 did not. These infection related strains from hepatic transplant recipients formed moderate or strong biofilms, whereas strains from renal transplant recipients produced the full range of biofilms (from weak to strong biofilm production). Data on antibiotic resistance, biofilm formation and HA for the eight strains responsible for infection of transplant recipients are summarized in Table 2.
Discussion
Postoperative care of transplant recipients implies frequent manipulations, the use of invasive medical devices, and the frequent use of extensive antibiotic therapy, which further contributes to the selection of multidrug-resistant bacteria. This cycle is difficult to avoid and leads to a scenario with narrowed therapeutic options. Moreover, patients awaiting transplantation could also become colonized with these bacteria.
Biofilm formation is one important characteristic of E. coli and other Enterobacteriaceae27. Biofilm producing bacteria may be responsible for many nosocomial infections. Biofilm characteristics protect bacteria from the host immune system along with the antibiotics, therefore, the role of biofilms during bacterial infections now constitutes an active field of research.
In this study, the ability of biofilm formation in vitro varied extensively among the E. coli isolates, and was independent of the type of transplant. The percentage of strains of K. pneumoniae producing biofilm was much higher than in E. coli or Enterobacter strains and was similar in kidney and liver transplants. In fact, the rate of biofilm formation by Enterobacter strains was very low (4%). Other studies showed similar or slightly higher proportion of Enterobacter strains that can form biofilms28,29. The high rate of biofilm-producing K. pneumoniae and the variability among E. coli isolates found in this study are in concordance with other studies evaluating MDR strains14,30,31.
We evaluated the possible relationship between biofilm formation and the presence of antimicrobial resistance genes. No relationship was found between strong biofilm formation and the presence of resistance genes in the three bacterial groups; but among the non-biofilm producers of K. pneumoniae (OD620 ≤ 0.05, n = 16) 11 strains were positive for the combination of CTXM-G1 + OXA-48 and other 3 carried the Verona integrin-encoded metallo-β-lactamase (VIM). Only 3 K. pneumoniae strains which were unable to form biofilms carried out other different resistant genes (strains 80, 83, and 102).
Looking at the type of transplant, there is no significant difference between strains isolated in liver transplantation with respect to kidney transplantation (unpaired t-test, p > 0.05).
K. pneumoniae, E. coli and Enterobacter strains isolated from rectal swabs prior to transplant or isolated from rectal swabs during the first weeks after transplantation were found to cause infections in 6 patients. These infection-related strains showed the same PFGE pattern and formed similar biofilms that their homologous colonizing strains. However, two K. pneumoniae strains (27 and 163) formed more biofilm that their corresponding pre-transplant isolates (25 and 158, paired t-test: p = 0.008 and p = 0.014 respectively). Although infections can occur at any time after transplantation, their incidence is highest in the first post-operative month32,33,34. In our study, these reemergent strains cause infections during the first month post-transplant, indicating that early detection and decolonization of these MDR on time is of critical importance to avoid further complications. However, enhanced surveillance monitoring for longer periods regarding opportunistic infections must also be addressed.
HA in E. coli and other Enterobacteriaceae species is mediated by fimbriae. Whereas in chronic conditions like urolithiasis biofilm plays an important role in persistence of infection, hemagglutination mediated by type I fimbriae, which bind to a mannose-containing receptor, are found in most urinary isolates where plays an important role in acute urinary tract infections31,35. The expression of type I fimbriae is indicated by MS hemagglutination while MR hemagglutination can be mediated by several types, such as P-fimbriae and DR fimbriae. Thus, MRHA-positive isolates can be considered most likely having P or DR fimbriae instead of type I fimbriae. Several authors have found a positive correlation between biofilm formation and the presence of type I pili in Enterobacteriaceae strains, including E. coli36,37. In this study, the number of E. coli strains that showed MRHA (n = 14) was higher than those that showed MSHA (n = 9). Interestingly, the 2 E. coli strains (199 and 228) recovered from infections were both MRHA and produced a very fast hemagglutination of human red blood cells. Of note, E. coli hemagglutinating strains but these two, were all poor biofilm producers; therefore, there was no correlation between MR or MS hemagglutination of human erythrocytes and biofilm formation.
In this report, we have demonstrated that E. coli and K. pneumoniae isolates display a high degree of phenotypic variability, being K. pneumoniae strains the greatest biofilm producers, and that Enterobacter isolates not form even moderate biofilms in vitro. Investigation of the mechanisms responsible for strong biofilm formation in K. pneumoniae is needed and would clarify some pathogenicity attributes of these bacteria in transplant recipients. On the other hand, as some isolates of the three genera analysed in this study cause infections but did not produce biofilms in vitro, it would be worthwhile to identify the presence and significance of other virulence factors responsible for the development of recurrent infections after SOT.
Methods
Study population and setting
The present prospective cohort study was conducted between August 2014 and April 2018 in seven University Hospitals from five Spanish regions. This national project (ENTHERE study) focused on the study of intestinal colonization and infections with MDRE in patients with kidney, liver, and kidney/pancreas transplants. Transplant patients who have been treated with antibiotics active against MDR Enterobacteriaceae (in the 30 days prior to inclusion in the study) such us colistin, carbapenems, amikacin or tigecycline, or patients who are participating in another study or clinical trial that includes active antibiotic treatment were not included in this study.
Bacterial strains
Over a period of 33 months, from October 2014 to June 2017, a total of 209 MDRE isolates defined as AmpC-hyperproducing and/or extended-spectrum β-lactamases (ESBLs) or carbapenemases producers, were included in this study. MDRE were obtained from 158 patients with kidney (n = 93) or liver transplant (n = 60) or both kidney and pancreatic transplant (n = 5) from the ENTHERE study. Among these, 106 were Escherichia coli, 78 were Klebsiella pneumoniae, and 25 were Enterobacter spp. (Supplementary Table 1). Bacteria were isolated from rectal swabs (in the 48 h previous to transplant and weekly samples till 4–6 weeks after transplantation) and directly inoculated onto chromID® ESBL and chromID® CARBA chromogenic agar plates (bioMérieux, Marcy L’Etoile, France). Colonies were subcultured on blood agar and MacConkey agar and preliminary identification and antimicrobial susceptibility testing the Vitek 2 compact system (bioMérieux, France) was used. Identification was confirmed by MALDI-TOF using the Vitek MS (bioMérieux) system, in accordance with manufacturer’s instructions. Stock cultures were frozen at −80 °C with 20% (vol/vol) glycerol.
Molecular characterization of resistance genes
Standard PCR were used to amplify several genes encoding extended-spectrum β-lactamases (blaTEM, blaSHV and blaCTXM) and carbapenemases (blaKPC, blaVIM, blaIMP, blaNDM and blaOXA-48). PCR multiplex plasmid-mediated AmpC (blaCIT, blaFOX, blaMOX, blaDHA, blaACC and blaEBC) was performed as described elsewhere38.
Genomic DNA was extracted using an InstaGeneTM Matrix Kit (Bio-Rad, Madrid, Spain) according to the manufacturer’s instructions. Then, 1 µL was added to a reaction mixture containing 1× KAPA Taq ReadyMix (KapaBiosystems, Wilmington, USA) and 0.5 µM of each primer. Amplification conditions were 94 °C for 5 min, followed by 30 cycles of 94 °C for 30 s, primer annealing at 55, 60 or 64 °C for 30 s and 72 °C for 1 min, and a final elongation at 72 °C for 7 min. PCR products were analysed on 2% (wt/vol) agarose gels stained with ethidium bromide 10 µg/mL and visualized by UV transillumination.
The amplified products were purified with a QIAquick PCR Purification Kit (QIAGEN Inc., Barcelona, Spain). DNA sequences on both strands were determined using an external resource (Macrogen Inc., Amsterdam, The Netherlands). Primers used in the detection of antimicrobial resistance genes, expected amplicon sizes, annealing temperatures, and references are shown in Supplementary Table 2.
Molecular fingerprinting
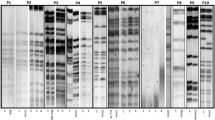
Repetitive extragenic palindromic PCR (REP-PCR) typing was performed on all isolates as described elsewhere39. Amplicons were run in a 1.5% agarose gel for 100 min, stained with ethidium bromide, and photographed. When at least two different bands were observed among isolates clonal relationship was also determined by PFGE. For this purpose, bacterial DNA embedded in agarose plugs was digested with the restriction enzyme XbaI and DNA separation was then performed in a CHEF-DRIII variable angle system (Bio-Rad, California, USA). Finally, the PFGE patterns were analysed with Fingerprinting II v4.5 software (Bio-Rad) and interpreted according to the criteria established by Tenover et al.40.
Biofilm formation
Biofilm formation was estimated in 24-well polystyrene plates (Corning, Costar) as previously described41. Briefly, Enterobacteriaceae strains were grown in Luria broth medium for 24 h at 37 °C with shaking, and a 1:1,000 dilution was prepared in Luria broth. Twenty-five microliters were placed in each well containing 1.5 ml of culture medium. The plates were incubated for 48 h at 37 °C in static. Planktonic cells were removed and wells containing biofilms were rinsed three times with distilled water and the remaining adherent bacteria were stained with 2 ml/well of crystal violet (0.7% [wt/vol] solution; Sigma-Aldrich) for 12 min. Excess stain was removed by washes with distilled water. CV was extracted by acetic acid (33% [vol/vol]), and the plates were incubated at room temperature in a bench orbital shaker for 1 min at 400 rpm to release the dye into the solution. Then, two samples of 100 µl from each well were transferred to a 96-well flat-bottom plate, and the amount of dye was determined at 620 nm using a microplate reader (Multiskan FC; Thermo Fisher). In each experiment, results were corrected for background staining by subtracting the value for crystal violet bound to uninoculated controls. The assays were performed 4 times for each isolate and the mean ± SD was reported. Strains were classified as non-adherent, weak, moderate or strong biofilm producers using the following criteria: OD ≤ 0.05, non-biofilm producer; OD > 0.05–0.1 weak biofilm producer; OD > 0.1–0.3 moderate biofilm producer; and OD > 0.3 strong biofilm producer.
Confocal laser scanning microscopy (CLSM)
Biofilm architecture of selected strains was studied in 4-well µ-slides (Ibidi, Martinsried, Germany) as previously described41. Briefly, the slides were placed inclined (~45°) into an incubator to form a liquid-air interface and after 48 h at 37 °C, unfixed planktonic cells were removed by rinsing with saline (0.85% NaCl), and bacterial viability within biofilms was determined by using the BacLight LIVE/DEAD bacterial viability kit (Molecular Probes Inc.). A series of optical sections was obtained with a Nikon A1R confocal microscope; the excitation wavelengths were 488 nm (green) and 561 nm (red), and 500 to 550 nm and 570 to 620 nm emission filters were used, respectively. Images at the liquid-air interface were captured at random with a 20× Plan Apo (numerical aperture [NA], 0.75) objective. Reconstructions of confocal sections and quantitative measurements were performed using NIS-Elements software, version 3.2.
Hemagglutination and HA inhibition tests
To identify other factors associated with biofilm formation in E. coli strains, we studied their hemagglutinating activities with human group A erythrocytes and the mannose sensitivities of these agglutinations as previously described41. Human erythrocytes were obtained from healthy volunteers after informed consent. Hemagglutination tests were performed on microscope slides using 10% suspensions of human group A erythrocytes. E. coli were cultured at 37 °C for 24 h in Luria medium, washed, and suspended in PBS to a concentration of ~5 × 109 CFUs per ml. Twenty-five microliters of cultures were mixed with 50 µl of erythrocytes with or without 1% D-mannose (Sigma). Agglutination of erythrocytes was examined visually after a short period (up to 1 min) of rocking at RT. Hemagglutination was considered resistant to mannose when it occurred despite the presence of mannose and sensitive to mannose when it was inhibited by the presence of this carbohydrate.
Statistical analysis
Data were described with means and standard deviation. Comparisons of the quantitative data was carried out by comparing means with the paired or unpaired t-test as corresponding. The alpha error was set at 0.05, and all p values were bilateral.
Ethical approval and informed consent
The study was performed in accordance with the Declaration of Helsinki. The protocol was reviewed and approved by the Institutional Ethics Committee at all seven participating hospitals according to local standards (EUDRAT-). The participating hospitals were: Hospital Universitario Marqués de Valdecilla (Santander), Coordinating Center; Hospital Universitario Cruces (Baracaldo); Hospital Clínic Universitari (Barcelona); Hospital General Universitario Gregorio Marañón (Madrid); Hospital Universitario 12 de Octubre (Madrid); Hospital Universitario Reina Sofía (Córdoba) and Hospital Universitario Ramón y Cajal (Madrid). Informed consent was obtained from each patient.
Data Availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
Change history
29 April 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Patel, R. & Paya, C. V. Infections in solid-organ transplant recipients. Clinical microbiology reviews 10, 86–124 (1997).
Mlynarczyk, G. et al. A threat of the Klebsiella pneumoniae carbapenemase-producing strains among transplant recipients. Transplantation proceedings 43, 3135–3136 (2011).
Paterson, D. L. & Bonomo, R. A. Extended-spectrum beta-lactamases: a clinical update. Clinical microbiology reviews 18, 657–686 (2005).
Poole, K. Resistance to beta-lactam antibiotics. Cell Mol Life Sci 61, 2200–2223 (2004).
Wiedemann, B. Genetic and biochemical basis of resistance of Enterobacteriaceae to beta-lactam antibiotics. The Journal of antimicrobial chemotherapy (18)Suppl B, 31–38 (1986).
Aguado, J. M. et al. Management of multidrug resistant Gram-negative bacilli infections in solid organ transplant recipients: SET/GESITRA-SEIMC/REIPI recommendations. Transplantation reviews (Orlando, Fla 32, 36–57 (2018).
Grim, S. A. & Clark, N. M. Management of infectious complications in solid-organ transplant recipients. Clinical pharmacology and therapeutics 90, 333–342 (2011).
Peleg, A. Y. & Hooper, D. C. Hospital-acquired infections due to gram-negative bacteria. The New England journal of medicine 362, 1804–1813 (2010).
Organization, W. H. Antimicrobial Resistance. Global Report on surveillance. WHO (2014).
Magiorakos, A. P. et al. Infection prevention and control measures and tools for the prevention of entry of carbapenem-resistant Enterobacteriaceae into healthcare settings: guidance from the European Centre for Disease Prevention and Control. Antimicrobial resistance and infection control 6, 113 (2017).
Luthje, P. & Brauner, A. Virulence factors of uropathogenic E. coli and their interaction with the host. Advances in microbial physiology 65, 337–372 (2014).
Murphy, C. N. & Clegg, S. Klebsiella pneumoniae and type 3 fimbriae: nosocomial infection, regulation and biofilm formation. Future microbiology 7, 991–1002 (2012).
Emody, L., Kerenyi, M. & Nagy, G. Virulence factors of uropathogenic Escherichia coli. International journal of antimicrobial agents 22 Suppl 2, 29–33 (2003).
Vuotto, C. et al. Biofilm formation and antibiotic resistance in Klebsiella pneumoniae urinary strains. Journal of applied microbiology 123, 1003–1018 (2017).
Sahly, H. et al. Increased serum resistance in Klebsiella pneumoniae strains producing extended-spectrum beta-lactamases. Antimicrobial agents and chemotherapy 48, 3477–3482 (2004).
Costerton, J. W., Stewart, P. S. & Greenberg, E. P. Bacterial biofilms: a common cause of persistent infections. Science 284, 1318–1322 (1999).
Schroll, C., Barken, K. B., Krogfelt, K. A. & Struve, C. Role of type 1 and type 3 fimbriae in Klebsiella pneumoniae biofilm formation. BMC microbiology 10, 179 (2010).
Rossi, E. et al. “It’s a gut feeling” - Escherichia coli biofilm formation in the gastrointestinal tract environment. Critical reviews in microbiology 44, 1–30 (2017).
Sicard, J. F., Le Bihan, G., Vogeleer, P., Jacques, M. & Harel, J. Interactions of Intestinal Bacteria with Components of the Intestinal Mucus. Frontiers in cellular and infection microbiology 7, 387 (2017).
Lebeaux, D., Ghigo, J. M. & Beloin, C. Biofilm-related infections: bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol Mol Biol Rev 78, 510–543 (2014).
Percival, S. L., Suleman, L., Vuotto, C. & Donelli, G. Healthcare-associated infections, medical devices and biofilms: risk, tolerance and control. Journal of medical microbiology 64, 323–334 (2015).
Greene, C., Vadlamudi, G., Newton, D., Foxman, B. & Xi, C. The influence of biofilm formation and multidrug resistance on environmental survival of clinical and environmental isolates of Acinetobacter baumannii. American journal of infection control 44, e65–71 (2016).
Madsen, J. S., Burmolle, M., Hansen, L. H. & Sorensen, S. J. The interconnection between biofilm formation and horizontal gene transfer. FEMS immunology and medical microbiology 65, 183–195 (2012).
Burmolle, M. et al. Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Applied and environmental microbiology 72, 3916–3923 (2006).
Soto, S. M. et al. Biofilm formation in uropathogenic Escherichia coli strains: relationship with prostatitis, urovirulence factors and antimicrobial resistance. The Journal of urology 177, 365–368 (2007).
Alcantar-Curiel, M. D. et al. Association of Antibiotic Resistance, Cell Adherence, and Biofilm Production with the Endemicity of Nosocomial Klebsiella pneumoniae. BioMed research international 2018, 7012958 (2018).
Nielsen, D. W. et al. The Impact of Media, Phylogenetic Classification, and E. coli Pathotypes on Biofilm Formation in Extraintestinal and Commensal E. coli From Humans and Animals. Frontiers in microbiology 9, 902 (2018).
Nyenje, M. E., Green, E. & Ndip, R. N. Evaluation of the effect of different growth media and temperature on the suitability of biofilm formation by Enterobacter cloacae strains isolated from food samples in South Africa. Molecules (Basel, Switzerland) 18, 9582–9593 (2013).
Kovacs, B. et al. Is biofilm formation related to the hypermutator phenotype in clinical Enterobacteriaceae isolates? FEMS microbiology letters 347, 116–122 (2013).
Ostria-Hernandez, M. L. et al. Nosocomial, Multidrug-Resistant Klebsiella pneumoniae Strains Isolated from Mexico City Produce Robust Biofilms on Abiotic Surfaces but Not on Human Lung Cells. Microbial drug resistance (Larchmont, N.Y 24, 422–433 (2018).
Reisner, A. et al. Type 1 fimbriae contribute to catheter-associated urinary tract infections caused by Escherichia coli. Journal of bacteriology 196, 931–939 (2014).
Losada, I., Cuervas-Mons, V., Millan, I. & Damaso, D. Early infection in liver transplant recipients: incidence, severity, risk factors and antibiotic sensitivity of bacterial isolates. Enfermedades infecciosas y microbiologia clinica 20, 422–430 (2002).
Balkan, A., Balkan, Y. & Mete, A. O. A Case of Early detected Multidrug-resistant Acinetobacter baumannii Infection after Liver Transplantation. Euroasian journal of hepato-gastroenterology 6, 170–172 (2016).
Hashimoto, M. et al. Acquisition of methicillin-resistant Staphylococcus aureus after living donor liver transplantation: a retrospective cohort study. BMC infectious diseases 8, 155 (2008).
Connell, I. et al. Type 1 fimbrial expression enhances Escherichia coli virulence for the urinary tract. Proceedings of the National Academy of Sciences of the United States of America 93, 9827–9832 (1996).
Shanks, R. M. et al. A Serratia marcescens OxyR homolog mediates surface attachment and biofilm formation. Journal of bacteriology 189, 7262–7272 (2007).
Schembri, M. A. & Klemm, P. Biofilm formation in a hydrodynamic environment by novel fimh variants and ramifications for virulence. Infection and immunity 69, 1322–1328 (2001).
Perez-Perez, F. J. & Hanson, N. D. Detection of plasmid-mediated AmpC beta-lactamase genes in clinical isolates by using multiplex PCR. Journal of clinical microbiology 40, 2153–2162 (2002).
Vila, J., Marcos, M. A. & Jimenez de Anta, M. T. A comparative study of different PCR-based DNA fingerprinting techniques for typing of the Acinetobacter calcoaceticus-A. baumannii complex. Journal of medical microbiology 44, 482–489 (1996).
Tenover, F. C. et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. Journal of clinical microbiology 33, 2233–2239 (1995).
Remuzgo-Martinez, S. et al. Biofilm Formation and Quorum-Sensing-Molecule Production by Clinical Isolates of Serratia liquefaciens. Applied and environmental microbiology 81, 3306–3315 (2015).
Acknowledgements
The authors thank Dr. Fidel Madrazo (Electron Microscopy Unit, Technology Support Services, IDIVAL) for helping with confocal microscopy. This research was supported by ‘Plan Nacional de I + D + i and Instituto de Salud Carlos III (Fondo de Investigaciones Sanitarias PI 13/01191 to MCF and PI 16/01103 to JRV), Subdirección General de Redes y Centros de Investigación Cooperativa, Spanish Ministry of Economy and Competitiveness, Spanish Network for Research in Infectious Diseases (REIPI RD12/0015) and (REIPI RD16/0016) co-financed by the European Development Regional Fund “A way to achieve Europe” ERDF.
Author information
Authors and Affiliations
Author notes
A comprehensive list of consortium members appears at the end of the paper
Consortia
Contributions
J.R.-V., I.C.-G., M.F.-M., C.G.-R. and M.C.F. conceived and designed the experiments, J.R.-V., I.C.-G., M.F.-M. and C.G.-R. performed the experiments. All authors have contributed strains. J.R.-V., I.C.-G., M.F.-M., C.G.-R. and M.C.F. analysed the data, J.R.-V. and M.C.F. wrote the paper. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ramos-Vivas, J., Chapartegui-González, I., Fernández-Martínez, M. et al. Biofilm formation by multidrug resistant Enterobacteriaceae strains isolated from solid organ transplant recipients. Sci Rep 9, 8928 (2019). https://doi.org/10.1038/s41598-019-45060-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45060-y
This article is cited by
-
Probiotic cell-free supernatant as effective antimicrobials against Klebsiella pneumoniae and reduce antibiotic resistance development
International Microbiology (2024)
-
Probiotic lactic acid bacteria as a means of preventing in vitro urinary catheter colonization and biofilm formation
Journal of the Egyptian Public Health Association (2023)
-
Investigation of antimicrobial resistance patterns and molecular typing of Pseudomonas aeruginosa isolates among Coronavirus disease-19 patients
BMC Microbiology (2023)
-
Characterization of virulence determinants and phylogenetic background of multiple and extensively drug resistant Escherichia coli isolated from different clinical sources in Egypt
Applied Microbiology and Biotechnology (2022)
-
Shewanella spp. from wastewater treatment plant-affected environment: isolation and characterization
Environmental Science and Pollution Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.