Abstract
Members of the family Araneidae are common orb-weaving spiders, and they produce several types of silks throughout their behaviors and lives, from reproduction to foraging. Egg sac, prey capture thread, or dragline silk possesses characteristic mechanical properties, and its variability makes it a highly attractive material for ecological, evolutional, and industrial fields. However, the complete set of constituents of silks produced by a single species is still unclear, and novel spidroin genes as well as other proteins are still being found. Here, we present the first genome in genus Araneus together with the full set of spidroin genes with unamplified long reads and confirmed with transcriptome of the silk glands and proteome analysis of the dragline silk. The catalogue includes the first full length sequence of a paralog of major ampullate spidroin MaSp3, and several spider silk-constituting elements designated SpiCE. Family-wide phylogenomic analysis of Araneidae suggests the relatively recent acquisition of these genes, and multiple-omics analyses demonstrate that these proteins are critical components in the abdominal spidroin gland and dragline silk, contributing to the outstanding mechanical properties of silk in this group of species.
Similar content being viewed by others
Introduction
Large nocturnal spider, Araneus ventricosus (family: Araneidae, superfamily: Araneoidea), is a common orb-weaving spider found throughout Japan and East Asia that builds vertical webs (perpendicular to the ground). Silks of A. ventricosus have served high extensibility, toughness, and strength1,2; there is considerable interest in industrial applications of synthetic spider silks.
Araneoids have seven specialised types of abdominal silk glands and use them differently in various situations throughout their lives3,4,5,6,7,8,9. Interestingly, many of the silk proteins produced in each gland are encoded by different orthologue groups of the spidroin gene family10,11 that likely diverged before the divergence of spider families. Furthermore, paralogues within these orthologue groups have been reported. For instance, two types of tubuliform genes (CySp or TuSp) used as the outer shell of the egg case were found in Argiope bruennichi12. Moreover, eight types of dragline silk genes, major ampullate spidroin (MaSp), were reported in Nephila clavipes13. Proteomic studies of spider silks are also identifying protein constituents other than spidroins, and the full catalogue of silk-related genes is yet to be uncovered.
The main difficulty in the study of spidroins is due to the unique organisation of these genes. Spidroin genes are very long, typically on the order of 10 k bp, and are almost entirely comprised of repetitive sequences between conserved non-repetitive N/C-terminal domains11,14,15,16,17,18. Such highly repetitive sequence structure poses a great challenge in sequence assembly based on short reads (including Sanger sequencing), and PCR amplification often results in chimeric artefacts. A common approach of target capture-based sequencing avoids misamplification, but is not optimal in finding novel spidroins. Previous approaches in genomic sequencing used PCR amplification, which can obtain a comprehensive list of spidroins, but the sequences tend to be partial, incomplete, or chimeric. Babb and colleagues exemplified the importance of genomic data to fully understand the diversity of spidroin genes13, using the draft genome data of N. clavipes and long read sequencing of spidroins but with long-range PCR amplicons. Many of their sequences still contain gaps and are thus not complete, and even if a spidroin-like gene sequence is obtained, the expression and presence of the protein products in actual silks should be further confirmed. Hence, the finding or isolation of the new spidroin gene requires multi-omics confirmation based on a high quality genome assembled with unamplified single molecule long read sequencing.
To this end, here, we present the draft genome of A. ventricosus, including full spidroin gene sets with a hybrid sequencing approach. These data sets will be a powerful reference to study the full extent of spidroin diversity and evolution. Using the draft genome, and transcriptomic as well as proteomic analyses, we reveal the unexpected complexity of A. ventricosus spider silk genetics.
Results
Genome sequence of A. ventricosus
We report the genome of A. ventricosus sequenced using a hybrid sequencing with a combination of Nanopore, 10x GemCode and Illumina technologies. Nanopore sequencing produced approximately 5.5 million long reads with a N50 length of 7.4 kbp (Table S1), and the latter produced over 500 million GemCode barcoded 150-bp paired-end reads. These sequenced reads were assembled into 300,730 scaffolds (Longest 9.34 Mbp, N50 scaffold size: 59,619 bp) comprising a 3.66 Gb genome (Table 1). The genome size estimated from the kmer distribution was 2.16 Gbp, with 37.4% repeat length and 2.6% heterozygosity with GenomeScope19 (Table S2). The extent of the repeat and thus the total genome size seems to be underestimated since our repeat analysis identified 51.1% to be the total repetitive content (Table S3). Although the genome seems to still contain at maximum 11.2% of uncollapsed heterozygosity as suggested by the BUSCO duplication rate, we consider the genome assembly to be comprehensive, in light of the cDNA-seq mapping rate (96.8% ± 0.7).
The gene content within the A. ventricosus genome was analyzed using cDNA sequencing. The cDNA was constructed from RNA samples from five independent whole bodies and six silk abdominal silk glands (Table S4). Approximately 35 million 150-bp paired-end reads were sequenced in each sample. Based on a gene model constructed using cDNA sequencing data, 277,986 open reading frames (ORFs) were predicted, and up to 29,380 (conservatively 14,767) protein-coding genes were estimated based on the expression level and functional annotation (Figs S1, S2). The quality of the predicted gene set was estimated by the BUSCO completeness score, and the test with the Arthropoda gene model showed 93.06% (Table 1).
Full spidroin gene set in A. ventricosus
First candidates of the spidroin gene were computationally screened based on sequence similarity. Candidates were then manually curated using the cDNA (see methods), unamplified long nanopore genomic DNA reads, and direct-RNA sequencing20 without reverse transcription or amplification steps. The final gene set was summarised into eleven spidroin genes belonging in seven orthologue groups (Fig. 1 and Table S5).
Catalogue of spidroins in A. ventricosus. This summary table shows the spidroin genic characters and structures obtained from the A. ventricosus genome. The icons in the first column represent spidroin type, and the specific colour is used for each type. The colour panel at second column represents the motif variety in the repetitive domain. The motif box includes β-sheet ((GA)n and An), β-turn (GPGGX, GPGQQ and XQQ), 310 helix (GGX), and spacer. Sequence structure column shows the N/C-terminal and repetitive domains, and each size is drawn to scale. The number of stripe in the repetitive domain also reflects the number of repeats.
With the exception of Flag and AgSp gene, the full-length of all spidroin genes without gaps were newly determined. Identified spidroins were highly diverse in the sequence length, exon-intron architecture, and repetitive structures (Fig. 1). The majority of the spidroin genes contained intronic regions, and linker domains were also found between repetitive units. The longest spidroin gene is the Flag gene constituted by three contigs, and the length of the total exonic regions is approximately 22.5 kb. Even the shortest MiSpB gene is 7.5 kb in length. Two paralogues were found for both MaSp2 and MiSp, with different gene structures and the number of repetitive units (Fig. 1). Furthermore, MaSp2A and MaSp2B were tandemly arranged within the same contig (Fig. S3). As previously described21,22,23, distinct repetitive motifs were reconfirmed in each spidroin gene (Fig. 1). There were no common repetitive motifs in all spidroin gene family, and this variety presumably reflects the diverse functionality of different spidroins.
The comparison with previously isolated full or partial sequences in A. ventricosus supported the accuracy of our spidroin gene set. The AcSp gene sequence almost entirely matched the previously reported one24 (Accession no. MG021196; Fig. S4). Regarding the Flag gene, the only known C-terminus region25 (Accession no. EF025541) was clearly aligned with our isolated gene sequence (Fig. S5). The CySp and MiSp genes had slightly longer sequences than known sequences obtained from the PCR approach26,27 (Accession no. MF192838 for CySp; Fig. S6, JX513956 for MiSp; Fig. S7), reconfirming the problem of PCR-based amplicon sequencing of spidroins, and the advantage of unamplified single molecule approach.
Full length sequence of a novel spidroin gene MaSp3
In addition to the above classical spidroin genes, the first full length of MaSp3 gene, originally named by Collin and colleagues28 partially reported in Argiope argentata and L. hesperus was also isolated.
The N/C-terminus domain sequences of the MaSp3 gene show only limited homology to other MaSp family genes in A. ventricosus, suggesting distinct divergence in this paralogue (Fig. 2a). When the terminal domains were clustered among the published MaSp genes in the family Araneidae, the N-terminus domains clearly cluster into three MaSp paralogs, independent of taxonomy (MaSp1, MaSp2, and MaSp3, Fig. 2b). Such distinct clustering was not observed for the C-terminus domain of all three paralogues, not just MaSp3. Although previous reports suggested the lack of common MaSp motifs (An and GPG) in the MaSp3 repetitive domain in A. argentata28, the A. ventricosus MaSp3 actually possesses these motifs. In contrast, its repetitive domain has a highly frequent arginine motif “GGR”, and the motif has never been reported as a spidroin motif (Fig. 2a).
Characteristics of MaSp3 gene in various scales from gene family to global spidroin category. (a) Alignment result and the sequence logo among MaSp subsets. (b) Clustering of the N/C-terminus among MaSp subsets in closely related spiders. The similarity of N-terminus demonstrated clearer clusters than did the C-terminus domain. (c) Global spidroin category in the superfamily Araneoidea. This spectral clustering was performed using every partial and complete sequence of the spidroin genes. Node colours represent the spidroin gene subsets.
To further confirm the distinction of MaSp3 as a paralog of MaSp, we prepared a global view of spidroins in the superfamily Araneoidea. A full length of our spidroin gene set was clustered with previously reported spidroin genes (Table S6) by spectral clustering29 based on a combination of multiple local sequence similarities to capture the combined sequence similarity/divergence of the N-terminus, repeats, and C-terminus regions (Table S7). Again, the result of spectral clustering confirms that MaSp3 is a subset of the MaSp category (Fig. 2c).
Phylogenetic origin of MaSp3
To investigate when the MaSp3 gene was evolutionarily acquired, we implemented a phylogenomic conservation analysis. A phylogenetic tree, including our A. ventricosus genome, was constructed based on a core orthologue gene set30, which was identified from assembled contigs with Araneids transcriptome data obtained from the NCBI SRA database (http://www.ncbi.nlm.nih.gov/sra). Moreover, to achieve higher resolution in the family Araneidae, we performed additional transcriptome analyses for five other spiders belonging to the family Araneidae (Tables S8–S10). We then constructed a phylogenetic tree expanding the family Araneidae and properly reflecting various previously reported trees30,31 (Fig. 3). The phylogenetic conservation of MaSp3 was then mapped on the phylogenetic tree using a homology search of 170 amino acids of the N/C-terminus region. Furthermore, a conservation analysis among 163 spider transcriptome data showed that the MaSp3 presents only in a part of Araneidae (Table S11), as the result of phylogenomic analysis, and strikingly, the MaSp3 is only conserved in a subset clade within the family Araneidae. Therefore, the MaSp3 gene in Araneidae seems to be acquired after the branching event from Nephila, Micrathena, and Verrucossa (Fig. 3). In light of these phylogenetic conservation patterns and the lack of GGR motif, partial sequence previously reported to be MaSp3 in L. hesperus (family Theridiidae) is likely to be a different paralogue of MaSp family.
Phylogenetic location of MaSp3 gene around Araneoidea. Phylogenetic tree based on the protein sequence of 4,934 orthologous genes in closely related spiders in superfamily Araneoidea. A scorpion was used as root. The e-values at the head and tail represent the result of BLAST search using N-terminus (head) and C-terminus (tail) 170 residues of MaSp3 as the query. The orange boxes represent e-value < 1.0e-5.
Proteome and expression profiling within dragline silk
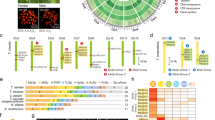
The contribution of MaSp3 to the composition of dragline silk was investigated by transcriptomic and proteomic approaches. The expression profiling was performed from cDNA-Seq samples in five whole bodies and abdominal silk glands (major ampullate, minor ampullate, and others). As expected, the MaSp3 gene was highly expressed in the major ampullate gland but is hardly observed in other glands (Figs 4a and S8, S9). This expression pattern was very similar to other profiles in MaSp family genes (MaSp1, MaSp2A, and MaSp2B).
Expression and proteome analysis in dragline silk. (a) Gene expression level of the spidroin genes in the whole body and each abdominal silk gland with three biological replicates per sample. The pictures in each graph are representative images of the samples. Other glands include multiple silk glands other than major ampullate and minor ampullate. The expression profiles of other spidroin genes were described at Fig. S8. (b) The left heat map of proteome of dragline silk in A. ventricosus. Orange arrows indicate the spidroin proteins. Right: heat map of expression of corresponding genes. Blue arrows indicate the SpiCEs.
The proteomic analysis was performed using dragline silks directly collected from five female A. ventricosus (Table S4). The spectra obtained from nanoLC-MS/MS analysis were annotated by a MASCOT search against our A. ventricosus genome database. Proteome analysis revealed that dragline silk yielded approximately 100 proteins on average (Table S12). Proteins shared in all samples were summarised at Fig. 4b. As expected from a previous study32, we confirmed the presence of the MaSp and AcSp proteins in dragline silk (Fig. 4b). Proteome analysis demonstrated that the peptides from MaSp3 specific linker domain were detected (Fig. S10), and the MaSp3 peptides were the most abundant in the dragline silks (Fig. 4b).
Relationship between the mechanical property and protein composition
We then investigated the impact of dragline components on dragline mechanical properties. Dragline silks used for mechanical property testing were obtained from 10 specimens of female A. ventricosus with various nutrition conditions. In this study, the following mechanical properties of the silks were measured: thread diameter (µm), strain at breaking (%), ultimate strength (MPa), and toughness (MJ/m3). These physical properties of dragline silk were measured on an Instron 3342 materials tester (Instron, Inc.) and summarised in Fig. 5a. As shown in Fig. 3b, the average values were diameter = 4.47 ± 0.96 µm, strain at breaking = 19.55 ± 5.02%, ultimate strength = 906.9 ± 100.9 MPa, and toughness = 84.28 ± 31.91 MJ/m3. However, there was high variance between the samples. Previous studies have reported that prey variation alters the silk composition15,33,34. Therefore, we first examined the effect of the nutrition condition on silk mechanics (Fig. S12), but the mechanical properties did not seem to be affected by these nutrition conditions (Fig. 5a and Tables S12, S13). Furthermore, the nutritional manipulations did not have an effect on protein composition in dragline silk (Table S12). Since the silk mechanical properties nor its components were independent of the drastic change in the nutrition conditions from being starved or directly after feeding, we investigated the direct relationship between silk components and mechanical properties.
Mechanical properties of dragline silk in A. ventricosus and phylogenetic trait. (a) Relationship between the mechanical properties and nutrition conditions. There are no significant differences. (b) PCA score plot of the mechanical properties (Strain at break, toughness, and tensile strength) and proteins included in dragline silk. (c) Phylogenetic position of MaSp3 and four genes especially associated with the mechanical property. Coloured boxes represent MaSp3 and SpiCEs found in transcriptome data of organisms.
The MaSp3 was constitutively the most abundant in dragline silk among six spidroins (represented by orange arrow in Fig. 4b), and PCA did not show direct contribution of MaSp3 to the mechanical properties of dragline silk (Fig. 5b). Therefore, genetic approaches to knock down MaSp3 expression or synthetic approaches would be necessary to elucidate the specific contribution of MaSp3 within A. ventricosus, due to the little intraspecies differences of MaSp3 abundances. On the other hand, among family Araneidae species, dragline silks in genus Araneus and Argiope with MaSp3 have higher toughness and tensile strength than the one in genus Nephila without it35, thus it is suggested that the MaSp3 may account for the interspecies differences. On the other hand, low molecular weight novel spider silk-constituting element, named SpiCE (Figs 4 and 5), was shown to be pivotal in intraspecies differences. SpiCE proteins contributing to toughness or tensile strength were found, and these four proteins (coded by g22833.t1, g160600.t1, g149801.t1, and g149799.t1) were highly expressed exclusively in the major ampullate gland (Fig. 4b, blue arrows). The conservation pattern of these silk-related proteins was investigated among the superfamily Araneidae based on sequence homology. Although g160600.t1 and g149801.t1 were widely conserved, two other genes were not uniformly conserved in the whole body transcriptomes of closely related spiders. Of note, the conservation pattern of g22833.t1 gene was very similar to that of the MaSp3 gene (Fig. 5c), suggesting a possible association between the two proteins.
Discussion
The extreme length and repetitive structure of the spidroin genes posed a challenge for comprehensively sequencing of these genes within a genome due to limitations in the short read based assembly and difficulty in correct amplification of long repeats with PCR. By combining multiple sequencing approaches, including nanopore long reads of unamplified DNA and RNA single molecules, this paper first demonstrated a working strategy to obtain the full spidroin gene set. Using the obtained genomic information, we successfully identified the first full length sequence of MaSp3 and several other SpiCE proteins that possibly contribute to the mechanical properties of dragline silk through a multiple omics approach.
The A. ventricosus genome data enabled comparative genomics in Araneoids. Our hybrid sequencing could present the approximately 3.5 Gb A. ventricosus genome with the full spidroin gene set, at high BUSCO coverage and comprehensive cDNA-Seq mapping rates. The 10X GemCode barcoded synthetic long read assembly provided accurate and comprehensive foundations of genome assembly, while this technology alone was not able to complete the repetitive spidroin regions. Hybrid sequencing using nanopore long reads from unamplified single molecule genomic DNA and direct-RNA sequencing20 without reverse transcription finally allowed the completion of the full length spidroin genes (Table S1). Current molecular biology techniques on DNA mostly rely on PCR; however, PCR amplification of long repetitive spidroins very often result in chimeric sequences or amplicons of different lengths. Comparison of previously reported sequences (CySp and MiSp of A. ventricosus) using spidroin amplicons with our unamplified single molecule sequences clearly shows the difficulty of obtaining accurate full-length sequence by amplicon-based approach. We believe it is critical to use single molecule long read sequencing techniques without amplification for such difficult sequences, including but not limited to spidroins. Genome sequencing and gene prediction analysis revealed that there are seven spidroin gene orthologue groups in common with other Araneoids. Because of the better continuity of our assembly, we can accurately identify the existence of multiple paralogs, locate the introns within the spidroins, and study the gene order of the spidroins. Interestingly, the two paralogs of MaSp2 are tandemly co-localised within the A. ventricosus genome, and such localisation may have implications on spidroin expression regulation.
One of the key findings from the genome sequencing was the identification of new paralogs of MaSp type spidroin MaSp3. The existence of this paralogue has been suggested from partial terminal domains obtained from target capture sequencing36, but the full length sequence was not previously reported. According to the phylogenetic analysis with N/C-terminus domains (Fig. 2b), while the N-terminus domains were distinct in MaSp1-3, the C-terminus domains were not clearly separated. Therefore, it had been difficult to recognise the MaSp3 gene and its N-terminus through partial sequencing of target captured cDNA. Phylogenetic analysis around the superfamily Araneoidea based on orthologous genes, with novel transcriptome data of five spiders generated in this work to increase the resolution of the phylogeny, revealed that MaSp3 might be relatively recently acquired after the branching event from genus Nephila. The genus Nephila was formerly classified into another family (Nephilidae), and Blackledge and colleagues previously showed that the genus Araneus and Nephila were strictly categorised into different clades in the aspect of web architecture31. The conservation pattern of MaSp3 in a subclade of Araneidae excluding Nephila mirrors such observation, and this synapomorphy may provide clues to the different orb web characters and mechanical properties in Araneidae. Furthermore, our proteome and transcriptome analyses showed that MaSp3 is one of the most important constituents of dragline silk (Figs 2 and 4). A detailed biochemical analysis regarding the unique repeat structure of MaSp3 would be interesting future work in this direction.
New spidroin subgroup candidates have been reported in previous studies13,28,37, but thus far, the classifications have been based on inter-species comparison of terminal domains or repeats. Although many unknown spidroin-like genes were also found in N. clavipes13, many remained unknown due to very weak conservation of the N-terminal domain, and the observation is based on amplicon-based sequencing. Since we now have the first plausible complete sequence set of a single species, we can utilise these data for global clustering, considering local sequence similarity information in both terminal sequences, repeats, and linkers by means of spectral clustering (Fig. 2c). This clustering approach clearly distinguishes different spidroin subtypes and demonstrates that some subtypes such as AcSp tend to subcluster among taxonomic neighbours, i.e., clade-specific adaptation is observed. Additionally, MaSp, MiSp, and CySp are more conserved. The new spidroin found in this work, MaSp3, was classified into the MaSp group but was sufficiently distant from the MaSp1 and MaSp2 paralogue subclusters. Based on this clustering, previously reported unknown fibroin-like proteins (Sp-907 and Sp-74867)13 in N. clavipes did not belong to any categories. This global clustering approach seems to clearly show the subtypes of spidroins and may be utilised as a complemental method to existing classifications.
Our high quality genome sequence also served as a reference for sensitive and reproducible proteome analysis. This study detected and confirmed many proteins within dragline that have been previously reported, such as alpha-2-macroglobulin 2, which was found as an exclusive protein in major ampullate silk glands in a western black widow spider (L. hesperus)32. Peroxidasin has been found at the boundary of a peripheral layer and the silk fibre core in the caddisfly silk fibre38, and its presence in spider dragline may serve similar functionality. On the other hand, we did not confirm the contribution of CRP (Cysteine Rich Protein), which has been reported as a protein to form structural constituents of fibres in a cobweb spider (family Theridiidae)39,40. Four CRP-like proteins were found in our A. ventricosus genome, and they were similar to CRP5 protein (Accession ADV40350.1). Although they were expressed in the whole body or abdominal spidroin glands (Fig. S12), the peptides were not found in dragline silk.
The presence of our curated spidroins was clearly confirmed in the proteome analysis, and we identified new genes possibly associated with the extraordinary mechanical properties of dragline silk. These genes are much shorter than the spidroins, and they are not similar to spidroins (Fig. S13). However, these genes were both highly expressed specifically in the major ampullate gland and are highly abundant as protein constituents within the dragline. The abundance of these proteins showed correlation with the toughness of the dragline, and one of them, g22833.t1, showed similar clade-specific conservation patterns with MaSp3 (Fig. 5c). The abundance estimate of the protein in this work was based only on the calculated emPAI values of the nanoLC-MS/MS analysis and should be further confirmed by other forms of direct quantification such as Western blotting. However, spider silk may be more highly complex than previously expected, with clade-specific duplication, the divergence of spidroin paralogues and the presence of other SpiCEs.
Our Araneus genome assembled by the hybrid of synthetic and single molecule long reads revealed the full length of a new spidroin. Due to the recent development of genomic technologies, many unknown spidroin genes have started to be discovered from spider genome or transcriptome data. We consider that such novel spidroin finding may occur among all spider clades. The MaSp3 found in genus Araneus and closely related spiders is a clade-specific spidroin, presumably correlating with the ability of these clade of spiders to produce large orb webs, requiring the extra toughness they exhibit. We have also identified non-canonical silk constituents that do not show homology to existing spidroins that we termed SpiCE, and according to our mechanical property analysis (Fig. 5), these proteins also contribute to the overall toughness of the silk. It is also interesting that the conservation patterns of many SpiCE proteins mirror that of MaSp3. This perspective suggests the possibility for a greater diversity of spidroin evolution and their related proteins that may be clade-specific to suit specific ecological adaptations (Fig. S14). Although MaSp is especially apt to be the research target and many paralogues have been observed13,28, the variety is also observed in prey capture thread. Some cribellate spiders have characteristic prey capture threads such as cribellate silk41 and pseudoflagelliform silk42, and recent studies have shown that these genes are categorized into specific spidroin types43,44. Therefore, the spidroin repertoire represents the behavioral and ecological variety of spiders, and more new spidroins or other silk constituents are likely to be found in the future.
Methods
Spider sample preparation
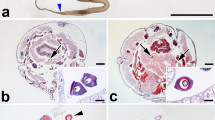
Spider specimens were initially identified based on morphological characteristics, and further confirmed by the transcriptome assembly of cytochrome c oxidase subunit 1 (COX1) in the Barcode of Life Data System (BOLD: http://www.barcodinglife.org). A. ventricosus (L. Koch, 1878) samples were collected from Akita, Yamagata, and Kumamoto Prefecture, Japan (December 2015). The samples were stored in a centrifuge tube and transported live back to the laboratory. A. ventricosus was kept in plastic containers PAPM340 (RISUPACK CO., LTD.) inside the laboratory with an average room temperature of 25.1 °C and 57.8% of humidity for approximately 2 weeks before the experiment. Light was controlled by an automatic system under a 12-h light/dark cycle. A. ventricosus was fed one cricket (Gryllus bimaculatus - commercially purchased from mito-korogi farm) once every 2 days. Water was provided once every day by softly spraying inside the plastic container. According to a previously reported standardised protocol of field sampling45, immediately upon arrival at the laboratory, A. ventricosus were immersed in liquid nitrogen (LN2) for whole body cDNA, RNA, genome sequencing and stored at −80 °C. Each gland tissue sample was dissected after anaesthetising with CO2, washed with phosphate buffered saline (PBS) and stabilised in RNAlater (Life Technologies). Photos of the dissected glands were taken immediately using VHX-5000 (Keyence), and the samples were flash frozen at −80 °C. Three biological replicates were separately prepared for all gland samples. Neoscona theisi, Gasteracantha kuhli, Argiope aemula, Cyrtophora unicolor, Acanthepeira sp., and Zygiella dispar samples were used only for cDNA sequencing. Sampling location data are described in Table S4.
HMW (high molecular weight) gDNA extraction
gDNA was extracted from four adult A. ventricosus whole bodies using Genomic-tip 20/G (QIAGEN) basically following the manufacturer’s protocol. To keep the HMW quality, every step was performed as gently as possible. Flash frozen spider specimens were separated into each body segment, and gDNA was extracted from the cephalothorax and legs. The specimens with the abdomen removed were homogenised using BioMasher II (Funakoshi) and mixed with 2 ml of Buffer G2 (QIAGEN), including 200 µg/ml RNase A. After addition of 50 µL Proteinase K (20 mg/mL), the lysate was incubated at 50 °C for up to 12 h on a shaker (300 rpm). The lysate was centrifuged at 5,000 × g for 5 min at 4 °C to pellet the debris, and the aqueous phase was loaded onto a pre-equilibrated QIAGEN Genomic-tip 20/G (QIAGEN) by gravity flow. The QIAGEN Genomic-tip 20/G (QIAGEN) was then washed three times and the DNA was eluted with high-salt buffer (Buffer QF) (QIAGEN). The eluted DNA was desalted and concentrated by isopropanol precipitation and resuspended in 10 mM Tris-HCl (pH 8.5). Extracted gDNA was quantified using a Qubit Broad Range (BR) dsDNA assay (Life Technologies) and qualified using TapeStation 2200 with genomic DNA Screen Tape (Agilent Technologies).
Library preparation for genome sequencing
For synthetic long-read sequencing, 10 ng purified HMW gDNA was used. The library preparation was performed with GemCode using a Chromium instrument and Genome Reagent Kit v2 (10X Genomics) following the manufacturer’s protocol. Library quality was estimated by TapeStation 2200 with D1000 Screen Tape (Agilent Technologies).
For nanopore long-read sequencing, the libraries were completed following the 1D library protocol (SQK-LSK108, Oxford Nanopore Technologies). The HMW gDNA applied to library preparation was purified by >10 kb size selection using a BluePippin (Sage Science) with 0.75% Agarose Gel Cassette.
Total RNA extraction
Total RNA was extracted using a spider transcriptome protocol, as previously described45. Flash frozen spider specimens were immersed in 1 mL TRIzol Reagent (Invitrogen) and homogenised with a metal cone using the Multi-Beads Shocker (Yasui Kikai). Following phase separation with the addition of chloroform, the upper aqueous phase containing extracted RNA was further purified using a RNeasy Plus Mini Kit (Qiagen) automated with QIACube (Qiagen). The quantity of purified total RNA was measured with NanoDrop 2000 (Thermo Scientific) and Qubit Broad Range (BR) RNA assay (Life Technologies), and the integrity was estimated by electrophoresis using TapeStation 2200 with RNA Screen Tape (Agilent Technologies).
Library preparation for cDNA and direct-RNA sequencing
The cDNA library was constructed using a standard protocol of the NEBNext Ultra RNA Library Prep Kit for Illumina (New England BioLabs). Approximately 100 µg total RNA was used for mRNA isolation by NEBNext Oligo d(T)25 beads (skipping wash step with Tris buffer). The first and second strands of cDNA were synthesized using ProtoScript II Reverse Transcriptase and NEBNext Second Strand Synthesis Enzyme Mix. Synthesized double-stranded cDNA was end-repaired using NEBNext End Prep Enzyme Mix and ligated with a NEBNext Adaptor for Illumina. After the USER enzyme treatment, cDNA was amplified by PCR with the following conditions (20 μL cDNA, 2.5 μL Index Primer, 2.5 μL Universal PCR Primer, 25 μL NEBNext Q5 Hot Start HiFi PCR Master Mix 2X; 98 °C for 30 s and 12 cycles each of 98 °C for 10 s, 65 °C for 75 s and 65 °C for 5 min). When the total RNA volume was less than 10 ng, the library was prepared using SMART-Seq v4 Ultra Low Input RNA Kit for Sequencing (Clontech) according to the manufacturer’s protocol, with subsequent fragmentation and Illumina library preparation with Hyper Plus Kit (Kapa Biosystems). For direct-RNA sequencing, 500 ng of mRNA was prepared using the NucleoTrap mRNA Mini Kit (Clonetech) and the libraries were completed following manufacturer’s protocol (SQK-RNA001, Oxford Nanopore Technologies).
Sequencing
The GemCoded genome library was prepared with Chromium (10X Genomics), and cDNA sequencing was performed with a NextSeq 500 instrument (Illumina, Inc.) using a 150-bp paired-end read with a NextSeq 500 High Output Kit (300 cycles). Sequenced reads were assessed with FastQC (v0.10.1: http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc/).
Nanopore genome and direct-RNA sequencing was performed using a MinION device with a total of eight v9.4 SpotON MinION flow cells (FLO-MIN106, Oxford Nanopore Technologies). The data sets obtained from this study were deposited and are available at the DNA Data Bank of Japan (DDBJ: http://www.ddbj.nig.ac.jp/) Sequence Read Archive with Accession no. DRA006821 and DRA006933.
De novo genome assembly and error correction
The NextSeq reads prepared by Chromium were assembled with Supernova (v. 2.0.0). Supernova assembly was further scaffolded and gap closed using the MinION reads with PBJelly46 and corrected using the NextSeq reads with two rounds of Pilon47.
To validate the genome assembly, we calculated genomic coverage and genomic completeness. First, the DNA-Seq data was mapped to the genome with BWA MEM (Burrows-Wheeler Alignment v0.7.12-r1039)48, and after Sequence Alignment/Map (SAM) to BAM conversion with SAMtools (v 1.3)49, the genome coverage was calculated with QualiMap bamqc50 v2.2. Second, the genomic completeness of the Supernova assembly was validated with BUSCO (Benchmarking Universal Single-Copy Ortholog, Eukaryote and Arthropoda lineage gene set, -m genome) version 2.0151.
Gene prediction and annotation
The gene model created by the cDNA-seq data mapping with HISAT2 version 2.1.052 and BRAKER version 1.953 was used for gene prediction. To annotate the predicted gene models, we submitted the amino acid sequences to similarity searches using BLAST against UniProt (Swiss-Prot and TrEMBL)54, and HMMER version 3.1b255 searches against Pfam-A56. The protein-coding gene number was estimated using the intersection or union of transcript abundance (see below) and the functional annotations of UniProt and Pfam (Fig. S1). The tRNA and rRNA genes were also predicted with tRNAscan-SE version 1.3.157 and Barrnap (https://github.com/tseemann/barrnap), and conducted repeat identification with RepeatModeler (http://www.repeatmasker.org/RepeatModeler/) and RepeatMasker (http://www.repeatmasker.org).
Spidroin gene curation based on the hybrid assembly
The spidroin gene curation was carried out by the hybrid assembly with the short and long reads. The conceptual diagram is shown in Fig. S15. Short reads obtained by Illumina sequencing is typically assembled using a de Bruijn graph, but such assembly is not feasible with the repetitive region. Therefore, we developed an original SMoC (Spidroin Motif Collection) algorithm. The SMoC algorithm first picks up the spidroin gene N/C-terminus candidates (non-repetitive region) with BLAST search from assembled genomic contigs, and repetitive regions from transcriptome assembly. These candidates are used as seed sequences for a screening of the short reads harboring an exact match of extremely large k-mer (approximately 100) up to the 5′-end, and the obtained short reads are aligned to constructs a PWM (Position Weight Matrix) on the 3′-side of the matching k-mer. Using very strict thresholds, seed sequence is extended based on the PWM until there is a split in the graph; i.e., neighboring repeat is not resolvable. By repeating this overlap-based extension algorithm, we can obtain the full length subsets of the repeat units. Finally, these pre-assembled repeat units are mapped onto error-corrected long reads obtained from the direct sequencing of the genomic DNA or RNA.
Expression analyses
Transcript abundances were estimated by kallisto version 0.42.2.158 in transcripts per million (TPM)59. Each transcriptome data set was obtained from the whole body and individual abdominal silk glands, and our A. ventricosus genome and predicted genes were used as the references.
Phylogenetic analyses
The phylogenetic trees for the N, C-terminus domains of spidroin genes in the family Araneidae (Fig. 2b) were constructed using known domains (Table S6). N, C-terminus domains were determined by BLASTP. The phylogenetic tree in Fig. 3 was constructed using the existing transcriptome data (Table S8) collected via the NCBI SRA database (http://www.ncbi.nlm.nih.gov/sra), in addition to newly sequenced transcriptome data in Neoscona theisi (DRR129306), Gasteracantha kuhli (DRR129307), Argiope aemula (DRR129308), Cyrtophora unicolor (DRR129309), Zygiella dispar (DRR129310), Araneus seminiger (DRR129311), and Cyclosa octotuberculata (DRR129312) according to transcriptome analysis method as described above. Furthermore, in addition to the above samples, the 156 spider transcriptome data sets collected by Fernandez and colleagues60 were assembled and used for the comprehensive MaSp3 gene conservation analysis (Table S11). The de novo transcriptome assembly was performed using Bridger61, with the following options: pair_gap_length = 0 and k-mer = 31. The assembled contigs were validated with BUSCO51. The 4,934 spider-specific gene set previously used in spider phylogenetic tree30 was obtained from assembled transcriptome contigs using HMMER version 3.1b255.
Collected orthologue genes were aligned with MAFFT version 7.30962 (mafft -auto–localpair–maxiterate 1,000) and then trimmed with trimAl version 1.2rev5963. Bootstrap analysis was performed using RAxML version 8.2.1164, and the phylogenetic tree was drawn using FigTree version 1.4.3 (http://tree.bio.ed.ac.uk/software/figtree/).
Dragline silk collection and proteome analysis
Dragline silk was reeled directly from adult A. ventricosus restrained using two pieces of sponge and locked with rubber bands (avoiding any kind of harm to the spider). First, dragline silk was meticulously removed from the spinnerets of the spider with a couple of tweezers and this silk was reeled using a reeling machine developed by Spiber Inc. in an aluminium bobbin at a constant speed of 1.28 m/min for 1 h.
A silk sample was gently washed with 100 µL Base buffer [50 mM Ammonium carbonate in distilled water] with 0.1% SDS per 0.5–1.0 mg of silk at RT for 1–2 min. After the supernatant removal, silk samples immersed into 46 µL Base buffer and 4 µL 500 mM DTT (Dithiothreitol) mixture. The silk solution was incubated for 1 h at 60 °C and left to stand until cool at RT. The supernatant was discarded, and 46 µL Base buffer and 4 µL 500 mM IAA (Iodoacetamide) mixture were added. The silk solution was incubated for 30 min at RT in dark, and the supernatant was discarded. Spider silk was washed with Base buffer three times. Peptide digestion of the washed silk sample was performed by 50 µL trypsin (10 ng/µL) at 37 °C overnight. Digested peptides were mixed with 250 µL of 0.5% formic acid and incubated at RT for 15 min with a rotator. Peptide samples were desalted using MonoSpin C18 (GL Sciences) and dried at RT.
Starvation test
The investigation of the impact of nutrition condition on the mechanical property was implemented by controlling the timing of the feeding. Over the course of two weeks, spiders were fed at day 1, day 4, and day 7, and dragline silks were sampled at day 2 (control 1: for 1 day after feeding), day 5 (control 2: for 1 day after feeding), and day 14 (starvation: for 1 week after feeding). This experiment was then replicated with 10 individuals (Tables S12, S13).
Mechanical property of dragline silks
The silk was carefully removed from the aluminium bobbins (without adding too much stress to the silk). Samples of 2 cm were taken and located into a paper template where the silk was attached with cyanoacrylate CA-156 (CEMEDINE CO., LTD.). For each specimen used, 10–15 testing pieces of silk were made. To determine the mechanical properties, we determined the diameter of the testing pieces by microscopic observation (Nikon eclipse LV100ND, lens 150x0). Three regions of the sample silk were selected and measured using NIS-Elements D 4.20.00 64-bit (Nikon). Tensile strength was measured using an Instron 3342 machine (Analysis program Bluehill lite Version 2.32 Instron 2005). The length of the testing pieces was set to 20 mm, and the testing speed was set to 10 mm/sec.
Liquid chromatography mass spectrometry analysis
Each sample for proteome analysis was dissolved with 12 µL of 0.5% acetic acid 5% acetonitrile, and 5 µL of the solution was loaded on hand-made spray needle column (Reprosil-Pur C18 materials, 100 µm i.d. Dr. Maisch GmbH, Germany, 5 µm tip i.d., 130 mm length) using a HTC-PAL autosampler (CTC Analytics, Zwingen, Switzerland). The peptide fragments in the samples were separated through the column by reversed phase chromatography of linear gradient mode using UltiMate 3000 nanoLC Pump (Dionex Co., Sunnyvale, CA, USA). As the mobile phases, (A) acetic acid/water (0.5:100, v/v), (B) acetic acid/acetonitrile (0.5:100, v/v) and (C) acetic acid/dimethyl sulfoxide (0.5:100, v/v) were mixed keeping the flow rate of 500 nL/min. The composition was changed as follow: (A) + (B) = 96%, (C) = 4%, (B) 0–4% (0–5 min), 4–24% (5–65 min), 24–76% (65–70 min), 76% (70–80 min), and 0% (80.1–120 min). The separated peptides were ionized at 2400 V by positive electrospray method, injected into LTQ orbitrap XL ETD (Thermo Electron, San Jose, CA, USA) and detected as peptide ions (scan range: m/z300–1500, mass resolution: 60000 at m/z 400). Top 10 peaks of multiple charged peptide ions were subjected to collision-induced dissociation (isolation width: 2, normalized collision energy: 35 V, activation Q: 0.25, activation time: 30 s) to identify the amino acid sequence.
Database search for protein identification
The peak lists were created from LC-MS raw data files with msconvert.exe provided from ProteoWizard65, and analyzed with Mascot server version 2.5 (Matrix Science, Boston, MA, USA)66 for identification of peptides and proteins in each samples. For the analysis, our A. ventricosus genome sequence was used with the following conditions: Precursor mass tolerance; 6 ppm, Product ion mass tolerance; 0.5 Da, Enzyme; Trypsin, Max missed coverages; 2, Fixed modification; carbamidomethylation at Cys, Variable modification; N-acetylation at protein N-term and oxidation at Met, Criteria for identification; p < 0.05 (MS/MS ion search).
Computational analysis and statistics
All computational data curation, treatment, and basic analysis were performed using Perl custom scripts with the G-language Genome Analysis Environment version 1.9.167. Statistical analyses were implemented using R package version 3.2.1. For the global spidroin category, the networks were constructed based on the sequence similarity among all the spidroin genes. The sequence similarity was calculated as a bit score with all-against-all BLASTP. The scores were normalised to 0.0–1.0 using a previously described normalisation method68. Using the normalised scores, all spidroin genes were clustered by spectral clustering with clusterx version 0.9.869, and the clustering results were drawn by Cytoscape (v. 3.5.1), with a force-directed layout. Sequence logo was constructed by WebLogo 370. PCA (principal component analysis) was calculated based on the correlation matrix and performed using JMP software version 13.2.0 (SAS Institute).
Data Access
Raw sequence reads used for genome assembly and expression analysis have been submitted to DDBJ SRA (sequence read archive). Accession numbers of the whole body transcriptome are DRR129306 (Neoscona theisi), DRR129307 (Gasteracantha kuhli), DRR129308 (Argiope aemula), DRR129309 (Cyrtophora unicolor), DRR129310 (Zygiella dispar), DRR129311 (Araneus seminiger), DRR129312 (Cyclosa octotuberculata), and DRR129313-DRR129317 (Araneus ventricosus). Accession numbers of silk gland transcriptome in Araneus ventricosus are DRR138403-DRR138405 (major ampullate), DRR138406-138408 (minor ampullate), and DRR138409-138411 (other silk glands). Accession numbers of MinION sequencing for direct-RNA is DRR138400 (Araneus ventricosus) and direct-DNA is DRR138402 (Araneus ventricosus). Accession number of GemCoded sequencing in Araneus ventricosus is DRR138401 (Tables S4, S8). Assembled files have been submitted to figshare.com (Table S8). The whole genome sequence is available at the Whole-Genome Shotgun (WGS) database in DDBJ under accession number of BGPR01000001-BGPR01300721.
References
Blackledge, T. A. et al. Sequential origin in the high performance properties of orb spider dragline silk. Sci. Rep. 2, 782 (2012).
Omenetto, F. G. & Kaplan, D. L. New opportunities for an ancient material. Science 329, 528–531 (2010).
Lucas, F. Spiders and their silks. Discovery 25, 20–26 (1964).
Gosline, J. M., DeMont, M. E. & Denny, M. W. The structure and properties of spider silk. Endeavour 10, 31–43 (1986).
Gosline, J. M., Guerette, P. A., Ortlepp, C. S. & Savage, K. N. The mechanical design of spider silks: from fibroin sequence to mechanical function. J. Exp. Biol. 202, 3295–3303 (1999).
Lazaris, A. et al. Spider silk fibers spun from soluble recombinant silk produced in mammalian cells. Science 295, 472–476 (2002).
Lewis, R. V. Spider silk: ancient ideas for new biomaterials. Chem. Rev. 106, 3762–3774 (2006).
Rainer, F. Biology of spiders. 3rd edn, (Oxford University Press, 2011).
Vollrath, F. Spider Webs and Silks. Sci. Am. 266, 70–76 (1992).
Guerette, P. A., Ginzinger, D. G., Weber, B. H. & Gosline, J. M. Silk properties determined by gland-specific expression of a spider fibroin gene family. Science 272, 112–115 (1996).
Gatesy, J., Hayashi, C., Motriuk, D., Woods, J. & Lewis, R. Extreme diversity, conservation, and convergence of spider silk fibroin sequences. Science 291, 2603–2605 (2001).
Zhao, A. C. et al. Novel molecular and mechanical properties of egg case silk from wasp spider, Argiope bruennichi. Biochemistry 45, 3348–3356 (2006).
Babb, P. L. et al. The Nephila clavipes genome highlights the diversity of spider silk genes and their complex expression. Nat. Genet. 49, 895–903 (2017).
Ayoub, N. A., Garb, J. E., Tinghitella, R. M., Collin, M. A. & Hayashi, C. Y. Blueprint for a high-performance biomaterial: full-length spider dragline silk genes. PLoS One 2, e514 (2007).
Hayashi, C. Y., Blackledge, T. A. & Lewis, R. V. Molecular and mechanical characterization of aciniform silk: uniformity of iterated sequence modules in a novel member of the spider silk fibroin gene family. Mol. Biol. Evol. 21, 1950–1959 (2004).
Hayashi, C. Y. & Lewis, R. V. Molecular architecture and evolution of a modular spider silk protein gene. Science 287, 1477–1479 (2000).
Perry, D. J., Bittencourt, D., Siltberg-Liberles, J., Rech, E. L. & Lewis, R. V. Piriform spider silk sequences reveal unique repetitive elements. Biomacromolecules 11, 3000–3006 (2010).
Xu, M. & Lewis, R. V. Structure of a protein superfiber: spider dragline silk. Proc. Natl. Acad. Sci. USA 87, 7120–7124 (1990).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat Methods 15, 201–206 (2018).
Hayashi, C. Y. & Lewis, R. V. Evidence from flagelliform silk cDNA for the structural basis of elasticity and modular nature of spider silks. J. Mol. Biol. 275, 773–784 (1998).
Malay, A. D., Arakawa, K. & Numata, K. Analysis of repetitive amino acid motifs reveals the essential features of spider dragline silk proteins. PLoS One 12, e0183397 (2017).
Teule, F. et al. A protocol for the production of recombinant spider silk-like proteins for artificial fiber spinning. Nat. Protoc. 4, 341–355 (2009).
Wen, R. et al. Molecular cloning and analysis of the full-length aciniform spidroin gene from Araneus ventricosus. Int. J. Biol. Macromol (2017).
Lee, K. S. et al. Molecular cloning and expression of the C-terminus of spider flagelliform silk protein from Araneus ventricosus. J. Biosci. 32, 705–712 (2007).
Chen, G. et al. Full-length minor ampullate spidroin gene sequence. PLoS One 7, e52293 (2012).
Wen, R., Liu, X. & Meng, Q. Characterization of full-length tubuliform spidroin gene from Araneus ventricosus. Int. J. Biol. Macromol. 105, 702–710 (2017).
Collin, M. A., Clarke, T. H. 3rd, Ayoub, N. A. & Hayashi, C. Y. Genomic perspectives of spider silk genes through target capture sequencing: Conservation of stabilization mechanisms and homology-based structural models of spidroin terminal regions. Int. J. Biol. Macromol. 113, 829–840 (2018).
Paccanaro, A., Casbon, J. A. & Saqi, M. A. Spectral clustering of protein sequences. Nucleic Acids Res. 34, 1571–1580 (2006).
Garrison, N. L. et al. Spider phylogenomics: untangling the Spider Tree of Life. PeerJ 4, e1719 (2016).
Blackledge, T. A. et al. Reconstructing web evolution and spider diversification in the molecular era. Proc. Natl. Acad. Sci. USA 106, 5229–5234 (2009).
Chaw, R. C., Correa-Garhwal, S. M., Clarke, T. H., Ayoub, N. A. & Hayashi, C. Y. Proteomic Evidence for Components of Spider Silk Synthesis from Black Widow Silk Glands and Fibers. J. Proteome Res. 14, 4223–4231 (2015).
Tso, I. M., Wu, H. C. & Hwang, I. R. Giant wood spider Nephila pilipes alters silk protein in response to prey variation. J. Exp. Biol. 208, 1053–1061 (2005).
Craig, C. L. et al. Evidence for diet effects on the composition of silk proteins produced by spiders. Mol. Biol. Evol. 17, 1904–1913 (2000).
Swanson, B., Blackledge, T. A., Beltrán, J. & Hayashi, C. Variation in the material properties of spider dragline silk across species. Applied Physics A 82, 213–218 (2006).
Kallal, R. J., Fernandez, R., Giribet, G. & Hormiga, G. A phylotranscriptomic backbone of the orb-weaving spider family Araneidae (Arachnida, Araneae) supported by multiple methodological approaches. Mol. Phylogenet. Evol. 126, 129–140 (2018).
Gaines, W. A. T. & Marcotte, W. R. Jr. Identification and characterization of multiple Spidroin 1 genes encoding major ampullate silk proteins in Nephila clavipes. Insect Mol. Biol. 17, 465–474 (2008).
Wang, C. S., Ashton, N. N., Weiss, R. B. & Stewart, R. J. Peroxinectin catalyzed dityrosine crosslinking in the adhesive underwater silk of a casemaker caddisfly larvae, Hysperophylax occidentalis. Insect Biochem. Mol. Biol. 54, 69–79 (2014).
Clarke, T. H. et al. Evolutionary shifts in gene expression decoupled from gene duplication across functionally distinct spider silk glands. Sci. Rep. 7, 8393 (2017).
Pham, T. et al. Dragline silk: a fiber assembled with low-molecular-weight cysteine-rich proteins. Biomacromolecules 15, 4073–4081 (2014).
Vollrath, F. et al. Compounds in the Droplets of the Orb Spiders Viscid Spiral. Nature 345, 526–528 (1990).
Coddington, J. A. Cladistics and Spider Classification: Araneomorph Phylogeny and the Monophyly of Orbweavers (Araneae: Araneomorphae, Orbiculariae). Acta Zoologica Fennica 190, 75–87 (1987).
Correa-Garhwal, S. M. et al. Silk genes and silk gene expression in the spider Tengella perfuga (Zoropsidae), including a potential cribellar spidroin (CrSp). PLoS One 13, e0203563 (2018).
Garb, J. E., Dimauro, T., Vo, V. & Hayashi, C. Y. Silk genes support the single origin of orb webs. Science 312, 1762 (2006).
Kono, N., Nakamura, H., Ito, Y., Tomita, M. & Arakawa, K. Evaluation of the impact of RNA preservation methods of spiders for de novo transcriptome assembly. Mol. Ecol. Resour. 16, 662–672 (2016).
English, A. C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS One 7, e47768 (2012).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Okonechnikov, K., Conesa, A. & Garcia-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12, 357–360 (2015).
Hoff, K. J., Lange, S., Lomsadze, A., Borodovsky, M. & Stanke, M. BRAKER1: Unsupervised RNA-Seq-Based Genome Annotation with GeneMark-ET and AUGUSTUS. Bioinformatics 32, 767–769 (2016).
UniProt Consortium, T. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Eddy, S. R. Accelerated Profile HMM Searches. PLoS Comput. Biol. 7, e1002195 (2011).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44, D279–285 (2016).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Fernandez, R. et al. Phylogenomics, Diversification Dynamics, and Comparative Transcriptomics across the Spider Tree of Life. Curr. Biol. 28, 2190–2193 (2018).
Chang, Z. et al. Bridger: a new framework for de novo transcriptome assembly using RNA-seq data. Genome Biol. 16, 30 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Chambers, M. C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Arakawa, K. et al. G-language Genome Analysis Environment: a workbench for nucleotide sequence data mining. Bioinformatics 19, 305–306 (2003).
Matsui, M., Tomita, M. & Kanai, A. Comprehensive computational analysis of bacterial CRP/FNR superfamily and its target motifs reveals stepwise evolution of transcriptional networks. Genome Biol. Evol. 5, 267–282 (2013).
Nepusz, T., Sasidharan, R. & Paccanaro, A. SCPS: a fast implementation of a spectral method for detecting protein families on a genome-wide scale. BMC Bioinformatics 11, 120 (2010).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
The authors thank Akio Tanikawa for morphological identification of spiders, and for helpful comments about phylogenetic discussion along with Akira Shinkai. Hitoshi Kawakami provided photographs of A. ventricosus, and Yuki Takai, Nozomi Abe, and Yuki Onozawa provided technical support in sequencing and proteome analysis. This work was funded by the ImPACT Program of Council for Science, Technology and Innovation (Cabinet Office, Government of Japan) and in part by research funds from the Yamagata Prefectural Government and Tsuruoka City, Japan.
Author information
Authors and Affiliations
Contributions
K.A. designed the entire project and performed genome sequencing and assembly. D.A.P.M., A.S., R.O. and H.N. collected spider samples and examined the mechanical properties. N.K., Y.Y. and K.A. analysed and curated the genome data. N.K. and K.A. performed expression analyses. N.K., M.F. and M.M. performed proteome analyses. N.K., M.T. and K.A. managed the computer resources. N.K. and K.A. wrote the manuscript. All authors contributed to editing and revising the manuscript.
Corresponding author
Ethics declarations
Competing Interests
Four of the authors are employees of Spiber Inc., a venture company selling artificial spider silk products. However, all study design was made by Nobuaki Kono and Kazuharu Arakawa of Keio University, and Spiber Inc. had no role in study design, data analysis, data interpretation, or writing of the manuscript.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kono, N., Nakamura, H., Ohtoshi, R. et al. Orb-weaving spider Araneus ventricosus genome elucidates the spidroin gene catalogue. Sci Rep 9, 8380 (2019). https://doi.org/10.1038/s41598-019-44775-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44775-2
This article is cited by
-
Lampshade web spider Ectatosticta davidi chromosome-level genome assembly provides evidence for its phylogenetic position
Communications Biology (2023)
-
Spiders’ digestive system as a source of trypsin inhibitors: functional activity of a member of atracotoxin structural family
Scientific Reports (2023)
-
Regionalization of cell types in silk glands of Larinioides sclopetarius suggest that spider silk fibers are complex layered structures
Scientific Reports (2023)
-
A molecular atlas reveals the tri-sectional spinning mechanism of spider dragline silk
Nature Communications (2023)
-
NMR assignment and dynamics of the dimeric form of soluble C-terminal domain major ampullate spidroin 2 from Latrodectus hesperus
Biomolecular NMR Assignments (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.