Abstract
Following observations of survival of microbes and other life forms in deep subsurface environments it is necessary to understand their biological functioning under high pressure conditions. Key aspects of biochemical reactions and transport processes within cells are determined by the intracellular water dynamics. We studied water diffusion and rotational relaxation in live Shewanella oneidensis bacteria at pressures up to 500 MPa using quasi-elastic neutron scattering (QENS). The intracellular diffusion exhibits a significantly greater slowdown (by −10–30%) and an increase in rotational relaxation times (+10–40%) compared with water dynamics in the aqueous solutions used to resuspend the bacterial samples. Those results indicate both a pressure-induced viscosity increase and slowdown in ionic/macromolecular transport properties within the cells affecting the rates of metabolic and other biological processes. Our new data support emerging models for intracellular organisation with nanoscale water channels threading between macromolecular regions within a dynamically organized structure rather than a homogenous gel-like cytoplasm.
Similar content being viewed by others
Introduction
Understanding the behavior of water inside cells is a long-standing problem in molecular biochemistry and cell biology1,2,3,4,5,6,7,8. The influence of high hydrostatic pressure on cellular structure and dynamics is important for areas ranging from biology in deep planetary environments to food preservation and biotechnology9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24. Investigating pressure effects on intracellular water dynamics forms a key aspect of this research and it leads to new insights into biological functioning of organisms at ambient and under hyperbaric conditions25. Water management inside prokaryotic organisms is still not completely understood. It has long been assumed that the internal structure of bacterial cells consisted of unencapsulated nucleic acid material suspended in a homogeneous gel-like mixture of proteins and other macromolecules along with aqueous electrolyte solutions forming the cytosol. Within the context of that model, main questions related to water mobility evolved around the “crowded” nature of the macromolecular environment, and how dissolved ions, metabolites and macromolecular species would affect the relaxation dynamics and transport properties. Recent discussions now suggest a more structured internal environment for the prokaryotic cell8,26,27,28,29. The new models envisage a dynamically changing but cooperatively well organized “superclustered” arrangement of proteins and macromolecular complexes, separated by channels containing the aqueous electrolyte solution28. Our new results determining intracellular water transport dynamics at high pressure support this view.
Nuclear magnetic resonance (NMR) and quasi-elastic neutron scattering (QENS) studies at ambient pressure have demonstrated diffusional dynamics for intracellular water that are indistinguishable from bulk water or aqueous solutions over length scales extending from localized environments up to cellular dimensions30,31. 2H NMR spin relaxation data also reveal ~15% water molecules that are strongly bound to internal surfaces, or buried within protein complexes30. Our QENS results at high pressure probe the dynamics of the main fraction of the mobile aqueous species that are responsible for transport and reaction processes within the bacterial cells, over real space correlation lengths extending between ~3 to 60 Å.
QENS provides information on diffusive and rotational relaxation rates in water and aqueous solutions over picosecond (ps) to nanosecond (ns) timescales, and techniques have now been developed for in situ QENS studies at high pressures extending into the GigaPascal range32,33. In a previous study we applied high-P QENS combined with H/D isotopic substitution of live organisms to investigate water dynamics in wild type (WT) Shewanella oneidensis at ambient and P = 200 MPa25. These results provided a first indication of diffusional slowdown occurring for intracellular water compared with aqueous hydrogenated or perdeuterated (Hb, Db) buffer solutions. However, it is known that H2O self-diffusion in aqueous media is also affected by pressure (P) as well as temperature (T) variables in this pressure range, and also depends on the presence of dissolved ions34,35. Here we extended the pressure range of our investigations of intracellular water dynamics up to 500 MPa to fully demonstrate the occurrence of the effect, obtaining QENS data at three separate neutron facilities and QENS instruments with different energy- and momentum-exchange and resolution characteristics. The consistency of the results clearly establishes the pressure induced changes in translational diffusion and rotational relaxation of intracellular water molecules, relative to bulk aqueous solutions used as the suspension media for the microbial samples. As part of our study, we also obtained data for pressure-resistant (PR) survivor populations that had been previously exposed to pressures of 500 (in a single compression step) and 750 MPa (via sequential intermediate treatments at 250 and 500 MPa, followed by resuscitation and re-compression) and compared the results with those for wild-type (WT) S. oneidensis samples36,37 (Table 1).
Results
In Fig. 1, we show results of typical QENS datasets and their analyses for WT and PR (500 MPa and 750 MPa) S. oneidensis bacteria at ambient (0.1 MPa), 200 MPa and 500 MPa, obtained at TOFTOF (FRM-II, Germany), IN6 (ILL, France), and IRIS (ISIS, UK)). Details of the three instruments and data analysis procedures are provided below in Methods and the full set of datasets is provided in Supplementary Information (SI). Further details of the analysis are given in our previous communication25. The data presented here correspond to water dynamics in the intracellular medium (Im) obtained from the isotopic contrast dataset combination Hc/Db − Db, using data recorded from “normal” hydrogenated cells (Hc) resuspended in perdeuterated buffer (Db), after subtracting the Db buffer contribution (Fig. 1). The neutron scattering (S(Q,w)) intensities are shown using a logarithmic scale to highlight the QENS contributions. After subtraction of the central Gaussian elastic line with its width determined by the instrumental resolution the QENS data were modelled by a narrow Lorentzian identified with translational diffusion, along with a broader second Lorentzian component from which rotational relaxation times could be extracted (Fig. 1 and Supplementary Figs 1–5).
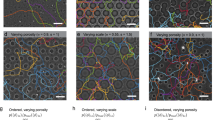
Left: Cartoon depicting isotopic QENS dataset subtractions applied to highlight intracellular water dynamics in different spatial regions. (A) Intracellular medium (Im) dynamics via the isotopic contrast [Hc/Db − Db] (i.e., bacterial cells minus buffer contribution). We note that this dataset subtraction provides information on the dynamics of all species contained within the cell envelope: i.e., membrane structures, proteins, DNA, RNA, etc., as well as the cytoplasm that dominates the mobile species on the QENS timescale. (B) Intracellular medium: cytoplasm only (Imc) dynamics obtained via the isotopic contrast [(Dc/Hb − Dc/Db) − Hb]. This isotopic subtraction gives information on the cytoplasm dynamics only. The two exhibit minor differences in the diffusion coefficients (Table 1). Right: (C) Representative datasets and analysis of the neutron dynamic scattering function S(Q,ω) for the intracellular medium Im (isotopic dataset contrast [Hc/Db − Db]). Data are shown here for a pressure-resistant S. oneidensis population that had been previously exposed to to 500 MPa (500 MPa-PR) and subsequently cultured at ambient conditions, then examined by neutron scattering using the IN6 instrument at ILL (instrumental resolution 85 μeV; λ = 5.12 Å). Data were recorded at room temperature at pressures of 0.1 and 500 MPa for momentum transfers between 0.2–1.7 Å−1. The raw data and lineshape fits are shown with a logarithmic intensity scale to highlight QENS contributions. The central line (grey) due to elastic scattering is modelled by a delta function convoluted with the instrumental resolution giving rise to a Gaussian. The narrow QENS Lorentzian signal (blue line) indicates the translational diffusion component with a linewidth (Γ) that varies as a function of Q. The broader (olive-green) Lorentzian represents the faster rotational relaxation that is Q-independent. The global fit (red continuous curve) is overlain on the data points (black open squares). A full set of datasets obtained and their analyses is provided in Supplementary Figs 1–5.
Water diffusion coefficients (DT) were determined by plotting the QENS half width at half maximum (HWHM, ΓT) for the narrow Lorentzian contribution associated with translational relaxation against the square of the reciprocal scattering vector (Q2) (Fig. 2, Table 2; Supplementary Fig. 6), and fitting the data using a jump model between sites separated by an average distance (l) with a mean residence time (τ0) during which the molecules undergo oscillatory motions38:
ΓT(Q2) data extracted from fitting QENS data of the narrow Lorentzian contribution (purely translational) from QENS data for intracellular medium (Im = Hc/Db − Db) for S. oneidensis WT (top panel), 500 MPa-PR (middle panel) and 750 MPa-PR (bottom panel) recorded at room temperature at 0.1, 200 and 500 MPa. Upward pointing triangles: data obtained at IRIS (ISIS, UK) using an instrumental resolution of 17.5 μeV: square symbols: data from TOFTOF (FRMII, Germany) at an instrumental resolution 60 μeV; downward pointing symbols: data from IN6 (ILL, France) at an instrumental resolution 85 μeV (λ = 5.12 Å). The symbols represent measured values and the solid lines show the results of fitting using a jump diffusion model with DT as a fitted parameter (Table 1).
The Hc/Db − Db isotopic contrast applied to most of our samples reveals contributions from water motions within the intracellular medium (Im), without distinguishing between the cytoplasm and cell envelope. The larger number of experimental runs that we could conduct at TOFTOF (FRMII) allowed us to examine a wider range of isotopic contrasts for WT samples at 1 atm and 200MPa25. Those data allowed construction of a further [(Dc/Hb − Dc/Db) − Hb] contrast that distinguishes dynamics within the cytoplasm alone (Imc) from those occurring within the cell envel’ (Fig. 1). That work allowed us to correlate the “dip” observed near 1 Å−2 in some of the ΓT(Q2) datasets with transmembrane water transport via Aquaporin channels25 (Fig. 2). However, the overall DT values for intracellular water motions extracted from the different datasets were identical within the systematic errors associated with each measurement (Table 2). We do note, however, that the DT values for the cytoplasm only (Imc) appear to be systematically lower (by ~5–6%) than those recorded for the full set of species contained within the cell envelope, including contributions from the membrane structures, proteins, nucleic acids etc, represented by the Im isotopic contrast subtraction (Table 1). We do not yet fully understand this result, that seems to imply that H2O dynamics in the cytoplasm are slower than those obtained by considering the entirety of the bacterial cell. Our DT values for the various bacterial vs buffer solution samples are summarized in Table 2 and compared with a compilation of literature data for diffusion in H2O and aqueous solutions in Fig. 3.
Compilation of translational self-diffusion coefficients (DT) extracted from QENS data obtained at the three spectrometers and neutron scattering facilities. Data from TOFTOF are shown as square symbols, IRIS as upward pointing triangles and IN6 as downward pointing triangles. Panel (A) DT values for the H-buffer (Hb); panel (B) DT values for intracellular [Hc/Db − Db; blue symbols] and the cytoplasm [(Dc/Hb) − (Dc/Db) − Hb; purple] medium for Shewanella oneidensis WT cells; panel (C) DT values for intracellular [Hc/Db − Db; olive] and the cytoplasm [(Dc/Hb) − (Dc/Db) − H]; light green] medium for S. oneidensis 500 MPa PR cells; panel (D) DT values for intracellular [Hc/Db − Db; red] medium for S. oneidensis 750 MPa PR cells. The dashed lines enclosing the light blue shaded area correspond to the range of literature values reported for bulk water and aqueous solutions between 298–308 K5,31,39,40,41,42,61,68,69,70.
Rotational relaxation times (τR = ħ/ΓR; ΓR = 2 ħDR)5,31,38,39 extracted from the broad Lorentzian contributions are also reported in Table 2 and in Fig. 4. In addition, we analysed EISF data for our samples using a roto-translational model with the O-H distance fixed at that for water (0.98 Å) (Fig. 5). Those results revealed up to ~30% reduction in the concentration of mobile protons occurring on the ps timescale by 500 MPa.
Rotational relaxation (ΓR(Q2)) data extracted from fitting QENS data for the broad Lorentzian contribution from QENS data for the dynamics of the intracellular medium (Im = [Hc/Db − Db]) for wild type S. oneidensis (top panel), 500 MPa pressure resistant (PR) survivors (middle panel) and 750 MPa PR survivors (bottom panel) recorded at room temperature at 0.1, 200 and 500 MPa. Upward pointing triangles: data obtained at IRIS (energy resolution 17.5 μeV). Square symbols: data from TOFTOF (resolution 60 μeV). Downward pointing triangles: data from IN6 (resolution 85 μeV).
Top panels: EISF for Hb buffer solution, calculated using Eq. S7 (SI). The data were modelled considering roto-translational movement of H2O molecules with radius 0.98 Å. The inserts indicate the fraction of mobile protons detected in each experiment, on a ps timescale within the instrumental resolution window. Upward pointing triangles: data obtained at IRIS (ISIS, UK) (resolution 17.5 μeV); square symbols: data from TOFTOF (FRMII, Germany) at instrumental resolution 60 μeV. Downward triangles: data from IN6 (ILL, France) at instrumental resolution 85 μeV (λ = 5.12 Å). Bottom panels: EISF for the intracellular medium [Hc/Db − Db] for S. oneidensis 500 MPa PR. The solid black line represents the model fit for Hb at 0.1 MPa: i.e., roto-translational movement of a molecule with radius 0.98 Å. Upward pointing triangles: data: data from IRIS (ISIS, UK): downward pointing triangles from IN6 (ILL, France).
Discussion
The intracellular water diffusion rates determined for WT bacteria at ambient pressure obtained at the three neutron scattering facilities overlapped closely with DT = 2.43–2.57 × 10−5 cm2s−1 (Fig. 2, Table 1). The consistency between these results provides confidence in our analysis and interpretation of the QENS datasets. The DT values obtained coincide with the lower limit of measurements recorded for bulk aqueous phases, and are similar to data obtained for nanoconfined water in porous inorganic matrices40,41,42,43,44,45,46 (Fig. 3). We do note that one early QENS result reported a slowdown in water dynamics for the extreme halophile Haloarcula marismortua with high intracellular Cl− content47. However, that result was not supported by more recent 2H NMR spin relaxation rate studies30.
As the pressure was increased to 200 and then 500 MPa in our experiments, the intracellular DT values became increasingly lowered relative to those for bulk aqueous solutions, including the Hb buffer medium used to resuspend microbial samples in our experiments, decreasing to 2.00–2.11 × 10−5 cm2s−1 at 200 MPa and 1.76 × 10−5 cm2s−1 at 500 MPa (Fig. 3). The extended pressure range of our new QENS data allows to demonstrate that the intracellular water self-dynamics do exhibit significantly greater slowdown compared with bulk water and aqueous electrolyte solutions comparable to cytoplasmic compositions, with implications for bacterial functioning under highly compressive conditions relevant to deep planetary biology (Fig. 3; Tables 1, 2). We suggest that this enhanced DT slowdown can be linked to the presence of spatially restricted pathways for intracellular water diffusion, leading to barriers to H2O mobility as the cells are compressed.
Classic interpretations of prokaryotes typically present the intracellular cytoplasm as a homogenous gel-like medium containing dissolved or suspended macromolecular and ionic components. However, recent models rather suggest a more structured internal environment, with cooperatively organized “superclustered” arrangements of proteins and macromolecular complexes, separated by channels enabling transport of the aqueous electrolyte solution8,26,27,28,29. Arguments for and against each view of the bacterial intracellular organisation at ambient conditions have been presented by Shepherd, who concluded that it was most useful to consider “the cytomatrix as a cooperative system of macromolecular and water networks“8. The significant slowdown in water mobility observed as a function of pressure, combined with the corresponding increase in rotational relaxation lifetimes, could then be associated with closing down bottlenecks, combined with increased interactions with macromolecules lining the walls of nanoscale intracellular diffusional channels26,27,28,29, rather than interpreted more generally as non-specific “crowding effects” within a gel-like model for the bacterial cytoplasm. Data on bulk water and dilute aqueous electrolyte solutions up to very high density show that water rotational dynamics are indeed almost unaffected by steric effects, and rather depend only on the strength and number of hydrogen bonds of the water molecule with its first neighbors32. We suggest that our QENS results, obtained at high pressure, are thus more consistent with channel closure within the emerging interpretation of a dynamically structured “supercluster” arrangement of macromolecular components within the cytoplasm, leaving channels for water diffusion to transport ions and hydrophilic molecules within the cell and enable reactions to occur. Our data will provide a reference point for future modelling studies.
Analysis of our EISF data for the intracellular medium (Fig. 5; Supplementary Fig. 8) compared with model fits for Hb at 0.1 MPa show the existence of an “extra structural contribution” to the static vs dynamic scattering signal. This is not surprising considering that a typical bacterial cell contains ~30% of structural macromolecules containing strongly bound H atoms (e.g., lipids, DNA, RNA, proteins etc.), that renders a fully quantitative analysis of the EISF data unfeasible. However, a qualitative comparison demonstrates an overall lowering of intensity in the high-Q range that points to a greater reduction in the concentration of mobile protons compared with the pure Hb buffer solution (i.e., ~40% vs. ~30%, respectively). To further investigate this effect we also examined our data for the isotopic contrast (Dc/Hb − Dc/Db) − Hb, that distinguishes between cytoplasmic vs cell envelope water dynamics25. This analysis was possible only for data obtained at IRIS (ISIS)48,49 as biomolecular dynamics recorded in the nanosecond range fall inside the energy resolution of the lower resolution spectrometers (IN6 and TOFTOF). We obtained the same result as before, with a higher proportion of mobile protons (~40%) present in the cytoplasm compared with the Hb buffer solution at high pressure (Supplementary Fig. 9). This observation is correlated with the greater slowdown in self-diffusion coefficient (~30 vs. ~20%) observed for the bacterial cells compared with bulk aqueous solutions, occurring as a result of densification.
The PR survivor populations that had been previously exposed to 500 and 750 MPa exhibited DT values between 2.04–2.24 × 10−5 cm2s−1 at ambient pressure, that were even lower than those recorded for the WT samples. These PR samples also exhibited a greater diffusional slowdown than the buffer media as the pressure was increased to 200 and 500 MPa (Fig. 3, Table 1). The changes in DT were accompanied by an average increase of ~27% in the H2O rotational relaxation time (τR) over the same pressure ranges (Tables 1, 2; Fig. 4; Supplementary Fig. 7). This is slightly greater than ΔτR ~24% observed for the Hb (and Db) buffer media, as the pressure was raised from 0.1 to 200 and then 500 MPa. Both results contrast with the relatively P-independent rotational correlation times observed for pure bulk H2O over the same pressure range31. Previous QENS studies of live bacteria including E. coli have found that intracellular diffusion rates are close to those of bulk aqueous solutions at ambient pressure31. However, our new data show that the intracellular DT values occur at the lowest end of the range of previous measurements for bulk water, although they do overlap with water nanoconfined in porous environments40,41,42,43,44,45,46,50. In addition, we find that the rotational relaxation times for intracellular H2O differs significantly from those observed for bulk H2O as a function of pressure (Fig. 4, Table 1).
The pressure-induced lowering in the self-diffusion coefficient for H2O molecules is correlated with an increase in viscosity (η) and it has implications for diffusion of ions, metabolite species, biological macromolecules as well as larger nanoparticles within the cytoplasm. We can apply the Stokes-Einstein relation that links DT and η for Newtonian fluids to estimate changes in intracellular cytosolic viscosity at the pressures and temperatures considered in this study51,52. At ambient P and T conditions, pure H2O exhibits a viscosity of 8.9 × 10−4 Pa s. Our results indicate that this value should increase by 20–40% as the cells are exposed to pressures of 200 and 500 MPa. The diffusion of ions through aqueous media is expected to take place on a similar timescale to DT for molecular H2O, whereas metabolites and proteins diffuse approximately 10–100 times slower. The relative rates of diffusion and the details of intracellular dynamics among these different species determine metabolic function as well as other important biological processes. Parry et al. have investigated the transport of macromolecules and other nanoscale objects through the bacterial cytoplasm and observed a size-dependent ergodicity breaking analogous to that found at the glass transition in glass-forming liquids53. They found that the dynamic transition between diffusive displacements and cage-like motions occurred as the particle size exceeded approximately 30 nm. The large displacement excursions of the probe molecules correlated with the metabolic activity. Although the authors discussed the cytosol in terms of a homogeneous gel-like transport medium, the arguments can be equally well applied to aqueous channels within a more highly organized model of the cellular interior structure. In that case, we might expect the “blocking” transition to be strongly pressure dependent as the nanoscale channels become constricted, and carrying out the particle tracking experiments using optical probes under high pressure conditions could be used to reveal the channel dimensions. We note that if the channels and their constrictions exhibit a range of sizes, this would have an internal filtering effect on the transport dynamics of large macromolecules and potentially foreign bodies within the bacterial cells.
We recognise that our results concern the effect of pressure on the intracellular water dynamics at ambient temperature, whereas organisms exposed to deep subsurface conditions experience a range of variable T environments. Early studies of bacterial metabolism under extreme high P were conducted at ambient T54,55,56. However, our work on survival of E. coli and S. oneidensis considered both P and T effects36,37. Despite the fact that S. oneidensis grows optimally at 30 °C at ambient pressure57, within the Shewanella genus, several strains grow and thrive in high pressure environments and at cold temperatures (e.g. S. benthica −50MPa and 10 °C58 whereas S. frigidimarina at ambient P and 10 °C59). We could not reproduce such a wide range of P and T conditions in our neutron studies. However, in our previous study, we examined the survival and colony-forming characteristics of WT and P-resistant populations at temperatures between 8 and 37 °C36. We did not observe any clear cross-correlation between T- and P-response effects affecting survival in S. oneidensis. In our neutron scattering studies, we examined the intracellular water dynamics for both wild-type (WT) and pressure-tolerant strains of S. oneidensis cultured following exposure to pressures extending up to 750 MPa. Specifically accounting for both P and T effects on intracellular water dynamics in different organisms is not yet possible. The temperatures in the deepest oceans and oceanic trenches where organisms have been found to survive at pressures up to 110 MPa range between −1 to + 4 °C60. Bulk H2O exhibits a density maximum and rapidly increasing viscosity throughout this range61, that may be reflected in the bacterial cytoplasmic dynamics, and that may influence the intracellular transport and reaction mechanisms and kinetics. This should be explored in future work combining specific targets for experimental studies of metabolic pathways linked to aqueous transport inside the cells, directed at specific model organisms. The situation in rock-hosted suboceanic or continental environments is different. Away from tectonic plate boundaries or other main thermal sources, the geothermal gradient is ~25/30 °C/Km. Once pressures of 200–500 MPa are achieved, measurements on bulk water indicate that DT is almost halved (i.e., to ~1.5/1.2 10−5 cm2s−1, respectively)61. However, experiments under nanoscale confinement conditions have shown that liquid water adopts a different and thermodynamically more stable structure in the vicinity of the walls, and its dynamics change dramatically such that freezing is no longer observed: i.e., liquid water enters a “super-cold” state40,41,42,44,62. Due to the complexity of the bacterial internal structure and composition, we can not yet extend these arguments to the biophysical behavior of living organisms, but they do lead to cause for reflection.
Conclusions
Our results demonstrate that average H2O diffusion rates are lower and rotational relaxation times are longer for intracellular water within the prokaryotic organism Shewanella oneidensis than for bulk aqueous solutions at ambient pressure. The dynamic slowdown is reflected in a decrease in the intracellular water diffusion coefficient that becomes increasingly enhanced as pressure is increased. These effects can be associated with increased constriction within nanoscale channels and interactions with the channel walls, in accord with currently emerging models of a prokaryotic cell structure with a high degree of internal organization. Our pressure-resistant bacterial survivor populations seem to exhibit slightly slower intracellular water diffusion rates than the WT samples, that could indicate more restricted pathways for H2O diffusion than within the general WT population. The H2O diffusion results are expected to correlate with increased internal viscosity of the cytosolic medium, as well as a reduction in diffusion of ions and metabolites through it. Larger macromolecules and foreign particles within the cells might become selectively blocked as the transport channels become restricted at high pressure. Our findings correlate with the observed reduction in the rates of internal metabolic processes as the pressure is increased54,55,56.
Methods
S. oneidensis samples (CIP 106686) obtained from the Collection Institut Pasteur (Paris, France) were grown at 30 °C in lysogeny broth (LB) at 180 rpm, after selecting a single colony from an LB agar plate harvested at stationary phase (108cells/mL). These constituted the wild type (WT) samples investigated in this study. Pressure resistant (PR) sample populations were cultured from survivors following sequential pressurization experiments in Teflon sample containers using a piston cylinder apparatus36,37. The PR samples selected for these QENS studies included one that had been treated in a single step at 500 MPa, and another that had been subjected to sequential pressurization-recovery steps at 250, 500 and then 750 MPa, after cultivation of the survivors from each run36. Our neutron scattering experiments examined cells with normal isotopic composition (Hc) as well as perdeuterated bacteria (Dc) that were re-suspended in normal (Hb) or perdeuterated (Db) buffer media at a concentration of 50 mg mL−1. We used phosphate buffer solution (PBS) at 10 mM concentration and pH7.4, containing 2.7 mM KCl and 137 mM NaCl. Perdeuterated cells (Dc) were prepared in the Deuteration Facility within the Life Sciences Group at the Institut Laue Langevin (ILL)63. The D-silantes medium used to grow the bacteria led to >98% labeling64. Samples were exposed to the neutron beams for 6–10 h at each pressure point. Following each experiment, recovered samples were cultured on agar plates in order to establish the survival rate through colony formation by plate counting. For the WT and PR samples, the survival rate following exposure at ambient pressure was ~80% compared with the initial concentration (Supplementary Fig. 10A). For WT bacteria exposed to 500 MPa in the beam, the survival rate was 0.05%, while that for PR samples that had been previously exposed to 500 MPa in a single step was 2.5%. The PR material that had been cultured after exposure to 750 MPa in 3 steps (0.1-250-500-750 MPa) represented 7.5% of the starting population. Survival rates following exposure to the neutron beam were comparable to those observed during ex situ pressurisation experiments carried out at UCL36,37. Complementary small angle neutron scattering (SANS) studies were carried out to ensure that extended exposure to high pressure did not result in structural modification in the bacterial samples. The similarity between the scattering profiles suggests that both WT and PR populations maintained a rod-like structure throughout these experiments (Supplementary Fig. 10B).
Room temperature QENS experiments were carried out at: IRIS (inverted time-of-flight (TOF) spectrometer)48,49 at ISIS (Rutherford-Appleton Laboratory, Harwell Science and Innovation campus, UK), with energy resolution 17.5 μeV; IN6 monochromatic TOF at the Institut Laue Langevin (ILL, Grenoble, France) with energy resolution 85 μeV (λ = 5.12 Å) and an extended energy range, and TOFTOF (monochromatic TOF)65 at the FRM-II reactor source (Garching, Germany) with an intermediate energy resolution 60 μeV (λ = 6.0 Å). The energy resolution that determined the full width at half maximum (FWHM, Γ) of the central elastic Gaussian peak was fixed by the instrument parameters and incident neutron wavelength (Supplementary Table 1). HP experiments at TOFTOF used a flat plate cell constructed from Al7075 alloy that permitted studies up to 200MPa25,66. The 200 and 500 MPa experiments at ISIS used a HP coil cell with internal diameter 0.5 mm, and those at ILL used a cylindrical cell; both of these were connected to a manual high pressure generator. Data were also collected at 0.1 MPa at ILL-IN6 using a flat plate Al liquid cell. Diagrams and photographs of the HP cell arrangements are given in Supplementary Fig. 11.
QENS data obtained over momentum transfers of between approximately 0.1 and 1.8 Å−1 provided information on diffusional and rotational water relaxation dynamics over real space correlation lengths ~3–60 Å. The large difference in incoherent neutron scattering cross section between H and D (80.27 vs 2.05barn, respectively) meant that D scattering was almost invisible relative to H when both were present. In our previous report of work during several experimental sessions at TOFTOF (FRM-II) we were able to examine a wide range of Hc, Dc, Hb, Db isotopic contrast combinations and separate out contributions from transmembrane vs purely intracellular (Imc) water dynamics. In this study we focused mainly on water dynamics within the overall intracellular environment derived from the (Hc/Db − Db) isotopic contrast combination (Im), that did not distinguish between contributions from the cytoplasm and cell envelope25.
Our initial experiments focused on obtaining data for Hb and Db aqueous buffer solutions, followed by QENS studies of Hc cells resuspended in perdeuterated buffer (Db). The QENS datasets were typically fit using two Lorentzian components. In agreement with previous work, we assigned the narrow component to translational diffusion with a HWHM linewidth ΓT(Q), and the broader (faster relaxing) component to a convolution of translational and rotational contributions (ΓT + ΓR)5,25,31,38,39,67. The relative amplitudes of the two components were taken as free parameters to fit the lineshapes. This approach was applied to both the aqueous buffer and the Im, determined by the isotopic contrast (Hc/Db − Db). In cases where we had sufficient data to implement the isotopic contrast (Dc/Hb − Dc/Db) − Hb that focused on the cytoplasm alone (Imc), the datasets only supported fitting a single Lorentzian, that was mainly associated with the translational diffusion contribution.
References
Watterson, J. G. A role for water in cell structure. Biochem. J. 248, 615–617 (1987).
Chaplin, M. Do we underestimate the importance of water in cell biology? Nature Rev. Molecular Cell Biol. 7, 861–866 (2006).
Ball, P. Water is an active matrix of life for cell and molecular biology. Proc. Natl. Acad. Sci. USA 114, 13327–13335 (2017).
Clegg, J. S. Intracellular water and the cytomatrix: some methods of study and current views. J Cell Biol. 99, 167s–171s (1984).
Jasnin, M. et al. Down to atomic-scale intracellular water dynamics. EMBO Rep. 9, 543–547 (2008).
Ball, P. Water as an active constituent in cell biology. Chem. Rev. 108, 74–108 (2008).
Dellerue, S. & Bellisent-Funel, M.-C. Relaxational dynamics of water molecules at protein surface. Chem. Phys. 258, 315–325 (2000).
Shepherd, V. A. The cytomatrix as a cooperative system of macromolecular and water networks. Curr. Topics Developmental Biol. 75, 171–223 (2006).
Bartlett, D. H. Pressure efects on in vivo microbial processes. Biochim. Biophys. Acta 1595, 367–381 (2002).
Heremans, K. High pressure effects on proteins and other biomolecules. Ann. Rev. Biophys. Bioeng. 11, 1–21 (1982).
Bartlett, D. H. Microbial adaptations to the psychrosphere/piezosphere. J. Molec. Microbiol. Biotechnol. 1, 93–100 (1999).
Patterson, M. F. Microbiology of pressure-treated foods. J. Appl. Microbiol. 98, 1400–1409 (2005).
Oger, P. M. & Jebbar, M. The many ways of coping with pressure. Res. Microbiol. 161, 799–809 (2010).
Silva, J. L. et al. High-pressure chemical biology and biotechnology. Chem. Rev. 114, 7239–7267 (2014).
Huang, Q. et al. Extreme biophysics: enzymes under pressure. J. Comput. Chem. 38, 1174–1182 (2017).
Harrison, J. P. et al. The limits for life under multiple extremes. Trends Microbiol. 21, 204–212 (2013).
Abe, F., Kato, C. & Horikoshi, K. Pressure-regulated metabolism in microorganisms. Trends Microbiol. 7, 447–453 (1999).
Heremans, K. Biology under extreme conditions. High Press. Res. 24, 57–66 (2010).
Somkuti, J. & Smeller, L. High pressure effects on allergen food proteins. Biophys Chem 183, 19–29 (2013).
Picard, A. & Daniel, I. Pressure as an environmental parameter for microbial life - a review. Biophys. Chem. 183, 30–41 (2013).
Meersman, F. et al. High-pressure biochemistry and biophysics. Rev. Mineral. Geochem. 75, 607–648 (2013).
Hazael, R. et al. Pressure as a limiting factor for life. Life 6, 34 (2016).
Meersman, F. & McMillan, P. F. High hydrostatic pressure: a probing tool and a necessary parameter in biophysical chemistry. Chem. Commun. 50, 766–775 (2014).
Aertsen, A. et al. Biotechnology under high pressure: applications and implications. Trends Biotechnol. 27, 434–441 (2009).
Foglia, F. et al. Water dynamics in Shewanella oneidensis at high pressure. Sci Rep 6, 18862 (2016).
Hunter, P. Not so simple after all. EMBO Rep. 9, 224–226 (2008).
Spitzer, J. From water and ions to crowded biomacromolecules: in vivo structuring of a prokaryotic cell. Microbiol. Mol. Biol. Rev. 75, 491–506 (2011).
Spitzer, J. & Poolman, B. How crowded is the prokaryotic cytoplasm? FEBS Lett. 587, 2094–2098 (2013).
Cossins, B. P., Jacobson, M. P. & Guallar, V. A new view of the bacterial cytosol environment. PLoS Comput. Biol. 7, e1002066 (2011).
Persson, E. & Halle, B. Cell water dynamics on multiple time scales. Proc. Natl. Acad. Sci. USA 105, 6266–6271 (2008).
Jasnin, M. et al. In vivo measurement of internal and global macromolecular motions in Escherichia coli. Biophys. J 95, 857–864 (2008).
Bove, L. E. et al. Translational and rotational diffusion in water in the Gigapascal range. Phys Rev Lett 111, 185901 (2013).
Klotz, S., Strässle, T. & Bove, L. E. Quasi-elastic neutron scattering in the multi-GPa range and its application to liquid water. Appl. Phys. Lett. 103, 193504 (2013).
Krynicki, K., Green, C. D. & Sawyer, S. W. Pressure and temperature dependence of self-diffusion in water. Faraday Disc. Chem. Soc. 66, 199–208 (1978).
Harris, K. R. & Woolf, L. A. Pressure and temperature dependence of the self diffusion coefficient of water and oxygen-18 water. J. Chem. Soc. Faraday I 76, 377–385 (1980).
Hazael, R. et al. Laboratory investigation of high pressure survival in Shewanella oneidensis MR-1 into the GigaPascal pressure range. Frontiers Microbiol. 5, 612 (2014).
Vanlint, D. et al. Rapid acquisition of gigapascal high pressure resistance by Escherichia coli. mBio 2, e00130 (2011).
Bée, M. Quasielastic Neutron Scattering: Principles and Applications in Solid State Chemistry, Biology and Material Science. Bristol: Adam Hilger (1988).
Jasnin, M. Atomic-scale dynamics inside living cells explored by neutron scattering. J. Royal Soc. Interface 6, S611–S617 (2009).
Bellissent-Funel, M.-C., Chen, S. H. & Zanotti, J.-M. Single-particle dynamics of water molecules in confined space. Phys. Rev. E 59, 4558–4569 (1995).
Bellissent-Funel, M.-C., Lal, J. & Bosio, L. Structural study of water confined in porous glass by neutron scattering. J. Chem. Phys. 98, 4246–4252 (1993).
Teixeira, J. The “puzzle” of water behaviour at low temperature. Water 2, 702–710 (2010).
Levinger, N. E. Water in confinement. Science 298, 1722–1723 (2002).
Yoshida, K. et al. Thermodynamic, structural, and dynamic properties of supercooled water confined in mesoporous MCM-41 studied with calorimetric, neutron diffraction, and neutron spin echo measurements. J. Chem. Phys. 129, 054702 (2008).
Osti, N. C. et al. Characteristic features of water dynamics in restricted geometries investigated with quasi-elastic neutron scattering. Chem. Phys. 465-466, 1–8 (2016).
Gallo, P., Rovere, M. & Spohr, E. Supercooled confined water and the mode coupling crossover temperature. Phys. Rev. Lett. 85, 4317 (2000).
Tehei, M. et al. Neutron scattering reveals extremely slow cell water in Dead Sea organism. Proc. Natl. Acad. Sci. USA 104, 766–771 (2007).
Campbell, S. I., Telling, M. T. F. & Carlile, C. J. The optimisation of analyser geometry in near-backscattering spectrometers – IRIS and ISIS-pulsed source. Physica B: Cond. Matter 276–278, 206–207 (2000).
Carlile, C. J. & Adams, M. A. The design of the IRIS inelastic neutron spectrometer and improvements to its analyser. Physica B: Cond. Matter 182, 431–440 (1992).
Zanotti, J.-M. & Bellisent-Funel, M.-C. Relaxational dynamics of supercooled water in porous glass. Phys. Rev. E 59, 3084–3093 (1999).
Ranieri, U. et al. Dynamical crossover in hot dense water: the hydrogen bond role. J. Phys. Chem. B 120(34), 9051–9 (2016).
Chen, S. H. et al. The violation of the Stokes-Einstein relation in supercooled water. Proc. Natl. Acad. Sci. USA 103, 12974–12978 (2006).
Parry, B. R. et al. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 156, 183–194 (2014).
Sharma, A. et al. Microbial activity at gigapascal pressures. Science 295, 1514–1516 (2002).
Picard, A. et al. Monitoring microbial redox transformations of metal and metalloid elements under high pressure using in situ X-ray absorption spectroscopy. Geobiol. 9, 196–204 (2011).
Picard, A. et al. Iron reduction by the deep-sea bacterium Shewanella profunda LT13a under subsurface pressure and temperature conditions. Frontiers Microbiol. 5, 796 (2015).
Venkateswaran, K. et al. Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp.nov. Int. J. Syst. Bacteriol. 49, 705–724 (1999).
Lauro, F. M. et al. Draft genome sequence of the deep-sea bacterium Shewanella benthica strain KT99. Genome A 1, e00210 (2013).
Bozal, N. et al. Shewanella frigidimarina and Shewanella livingstonensis sp. nov isolated from Antarctic coastal areas. ISEM 52, 195–205 (2002).
Yasuhara, M. & Danovaro, R. Temperature impacts on deep-sea biodiversity. Biol. Rev. 91, 275–287 (2016).
Mallamace, F., Corsaro, C. & Stanley, H. E. A singular thermodynamically consistent temperature at the origin of the anomalous behavior of liquid water. Sci. Rep. 2, 993 (2012).
Taschin, A. et al. Evidences of two distinct local structures of water from ambient to supercooled conditions. Nat. Commun. 4, 2401 (2013).
Haertlein, M. et al. Biomolecular deuteration for neutron structural biology and dynamics. Methods in Enzymology 566, 113–157 (2016).
Unruh, T., Neuhaus, E. & Petry, W. The high-resolution time-of-flight spectrometer TOFTOF. Nucl. Instr. Meth. Phys. Res. A 580, 1414–1422 (2007).
Appavou, M.-S. et al. The influence of 2 kbar pressure on the global and internal dynamics of human haemoglobin observed in quasielastic neutron scattering. Eur. J. Biophys. 40, 705–714 (2011).
Jasnin, M. et al. Specific cellular water dynamics observed in vivo by neutron scattering and NMR. Phys. Chem. Chem. Phys. 12, 10154–10160 (2010).
Longeville, S. & Lechner, R. E. Light and heavy water dynamics. Physica B 276, 534–535 (2000).
Prielmeier, X. et al. The pressure dependence of self diffusion in supercooled light and heavy water. Ber Bunsenges Phys. Chem. 92, 1111–1117 (1998).
Teixeira, J. et al. Experimental determination of the nature of diffusive motions of water molecules at low temperatures. Physical Review A 31(3), 1913–1917 (1985).
Acknowledgements
This study was supported by grants from the Leverhulme Trust (RPG-350) and the Deep Life program of the Deep Carbon Observatory (Sloan Foundation) to PFM. VTF acknowledges UK EPSRC support via grants GR/R99393/01 and EP/C015452/1 for creation of the Deuteration Laboratory in ILL’s Life Sciences Group and EU contracts RII3-CT-2003-505925 and NMP4-CT-2006-033256. We thank Mohamed Zbiri at ILL for help with beamline support. This project has received funding from the European Union’s Seventh Framework Programme for research, technological development and demonstration under the NMI3-II Grant number 283883 to provide access to experiments at FRM-II. LEB gratefully acknowledges financial support from the Swiss National Science Foundation under grant number 200021-149847 and from the French ANRwithin the Blanc International programme PACS under reference ANR-13-IS04-0006-01. We also thank the sample environment teams at all neutron facilities. FF also thanks ISIS (Didcot, UK: grant RB1410406) for beamtime (doi: 10.5286/ISIS.E.RB1410406) and for financial support for experimental consumables (Grant RB1410406). We are grateful to the Institut Laue Langevin (Grenoble, France) for neutron beamtime (doi:10.5291/ILL-DATA.8-04-785). The open access fee was covered by FILL2030, a European Union project within the European Commision's Horizon2020 Research and Innovation programme under grant agreement No 731096.
Author information
Authors and Affiliations
Contributions
P.F.M., F.M., F.F. and R.H. devised the overall study, and prepared WT and PR samples at UCL. D/H isotopic growth/exchange experiments were carried out at the ILL-PSB Deuteration Lab by M.M., M.H. and V.T.F., F.F., R.H. and F.M. participated in previously reported neutron experiments at FRM-II; F.M., L.E.B., M.M.K. and M.C.W. participated in obtaining new data at ILL, and F.F. with assistance from V.G.S. and S.R. at ISIS. Data analysis was conducted by F.F. with assistance from L.E.B., M.C.W. and P.F.M. The results were collected and the manuscript developed by F.F., P.F.M. and L.E.B. with contributions from all authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Foglia, F., Hazael, R., Meersman, F. et al. In Vivo Water Dynamics in Shewanella oneidensis Bacteria at High Pressure. Sci Rep 9, 8716 (2019). https://doi.org/10.1038/s41598-019-44704-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44704-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.