Abstract
Arbuscular mycorrhizal fungi (AMF) can be beneficial for improving restoration of radioactive-cesium (137Cs)-contaminated soils through soil remediation. However, there has been no information on species diversity and the composition of AMF communities in 137Cs-contaminated soil after the Fukushima-Daiichi Nuclear Power Plant (NPP) disaster. We examined the community dynamics of indigenous AMF colonizing roots of napiergrass (Pennisetum purpureum) in two different 137Cs-contaminated land-use fields (grassland and paddy field) by an Illumina MiSeq sequencing investigation within a 30-km radius around the Fukushima-Daiichi NPP in 2013 (sampling year 1) and 2014 (sampling year 2). We found nine AMF families, including Glomeraceae, Gigasporaceae, Paraglomeraceae, Claroideoglomeraceae, Acaulosporaceae, Archeosporaceae, Ambisporaceae, Diversisporaceae and uncultured Glomeromycotina in roots. Glomeraceae was the most abundant in both grassland and paddy field, followed by Paraglomeraceae. The diversity of AMF in grassland and paddy field was higher in 2014 than in 2013. Furthermore, the AMF community structure was impacted by sampling year and land-use type. The AMF community structures colonizing napiergrass roots were also significantly impacted by land-use type and year throughout the 2-year investigation. To our knowledge, our results are the first report to reveal the community dynamics of indigenous AMF in the 137Cs-contaminated fields around NPP.
Similar content being viewed by others
Introduction
A catastrophic earthquake and tsunami occurred in Japan on March 11, 2011, which severely damaged the Fukushima-Daiichi Nuclear Power Plant (NPP). The disaster also triggered the worst nuclear accident in world history since Chernobyl. This disaster led to emissions of radioactive materials such as, radiocesium (137Cs and 134Cs), strontium-90 (90Sr), iodine-131 (131I), tellurium-132 (132Te) and xenon-133 (133Xe), from the NPP1. Among these materials, 137Cs especially provokes the largest concern because of its devastating effect on agriculture and stock farming for decades. For 137Cs, a total emission of 36.6 (20.1–53.1) PBq was released into the atmosphere during the NPP disaster2, and this contaminated the soil, over a vast area. In the current 137Cs-removal approach at Fukushima Daiichi NPP, Kang et al.3,4 reported that napiergrass (Pennisetum purpureum Schum.), which has a large aboveground biomass and a high ability to accumulate Cs in its shoots, as a highly suitable potential phytoremediation crop3,4,5. Addtionally, the same author suggested the possibility of increasing Cs transfer from the Cs-contaminated soil around the NPP using napiergrass3.
Arbuscular mycorrhizal fungi (AMF) can increase host plant phosphorus (P) uptake and growth, and AMF may especially improve plant micronutrients uptake6. Additionally, the role of AMF in radionuclide-contaminated soils and their effects on radioactive-Cs uptake and transfer to plants have been examined with a possible way to develop the use of AMF for soil remediation strategies7,8,9. Hammer et al.10 indicated that AMF selectively absorb potassium (K) which is an analog for Cs. In addition, Cs is a weak analog of K, but is a better analog of rubidium (Rb). These analogs are accumulated by fruiting bodies of mushrooms, such as saprotrophic and ectomycorrhizal fungi11. Previous studies have shown no correlation between 137Cs and K uptake in mushrooms12. On the contrary, 137Cs and 133Cs uptake have been positively correlated with Rb uptake in fruiting bodies of mushrooms, such as saprotrophic and ectomycorrhizal fungi11. Likewise, AMF can also take up, accumulate, and translocate 137Cs in their external hyphae13,14. These results indicate a potential involvement of AMF in the Cs biogeochemical cycle and in plant Cs accumulation. However, previous studies have also demonstrated that AMF either facilitate, or have no effect on 137Cs uptake by host plants. Thus, the actual role of AMF in plant uptake of Cs, and the capacity and AMF species to accumulate is not fully understood and remains challenging because there is no ecological and functional information of the AMF in 137Cs-contaminated soil due to the Fukushima-Daiichi NPP accident.
Furthermore, AMF community structure as well as nutritional improvement by AMF can play an important role in ecosystem restoration and sustainability15,16. Many studies have reported AMF communities can impact a number of important ecosystem processes, including plant productivity, plant diversity and soil structure17,18. To promote and evaluate efficient characterization of AMF communities by sequencing techniques, current studies have relied on next-generation sequencing technologies, such as Illumina Miseq platform19,20. Therefore, understanding AMF diversity and communities can be important to explore the ecological environment21.
Considering the above facts, combining napiergrass as a candidate plant with high potential for 137Cs-remediation and the 137Cs uptake function by AMF in the 137Cs-contaminated soil in Fukushima may be a better technique to remove 137Cs from the 137Cs-contaminated soil. However, the role of AMF combined with napiergrass in processes of 137Cs uptake are unclear and not fully understood. Therefore, we investigated the distribution of indigenous AMF colonizing roots of napiergrass in the 137Cs-contaminated soil around the NPP as a preliminary investigation. This study was undertaken as a first investigation of the distribution of indigenous AMF colonizing roots of napiergrass that grew in two high 137Cs-contaminated soils within a 30-km radius around the Fukushima-Daiichi NPP. Additionally, the indigenous AMF communities in roots of naipergrass from 137Cs-contaminated soils were investigated in detail using a next-generation sequencing of the Illumina Miseq platform for the first time.
Materials and Methods
Experimental design
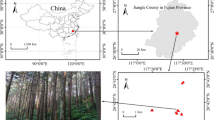
We conducted a field study with two objectives. First objective was to understand whether different plant densities in naipergrass increase the 137Cs removal from the 137Cs-contaminated soil based on a study by Kang et al.3. Second objective according to our main purpose of the study was to determine the distribution of indigenous AMF colonizing roots of napiergrass that grew in two high 137Cs-contaminated soils within a 30-km radius around the Fukushima-Daiichi NPP. Thus, a field experiment was conducted in two different land-use types (lowland and upland soils) in Fukushima, Japan. This experiment was performed in collaboration with Namie town and with the permission of the Ministry of Environment, Japan. Two experimental sites were used based on their different land-use histories before the NPP accident. Both the lowland and upland fields were used as a paddy field (37.486381N, 140.959085E) and grassland (37.489980N, 140.949368E), respectively, before the disaster of the Fukushima-Daiichi NPP on March 11, 2011. Physicochemical properties of the paddy field and grassland at the beginning of the field experiment in 2013 are shown in Table S1, which were adapted from Kang et al.3. Further details on the methods of soil physicochemical properties regarding each land-use type are presented in Kang et al.3.
To determie the communities of AMF colonizing roots of napiergrass (P.purpureum Schum., var. Merkeron), three of the 2.0 m × 2.0 m plots were established in both the paddy field and grassland. The planting density in plot for napiergrass at each land-use type were low density (4 plants plot−1), medium density (16 plants plot−1), and high density (44 plants plot−1), which were randomly distributed. Four-week-old napiergrass nursery plants were transplanted into plots with 0.3-m (high density), 0.5-m (medium density) or 1-m (low density) interrow spacing between each plot on May 27, 2013 and on May 17, 2014. The amount of N- and P-applied rates were 10 and 10 g m−2, respectively. No K fertilizer was applied to the two fields during this 2-year investigation. Further details on the information of plant growth parameters and uptake of Cs regarding each land-use type are presented in Kang et al.3.
Root sampling and staining
Root samples in napiergrass were manually collected from three plants in each plot (to a depth of 15 cm, with a diameter of 20 cm), resulting in three independent root samples products per plot. A total of nine independent root samples of napiergrass in each land-use type (a total of 18 root samples in each year) were analyzed for morphological traits and for DNA extraction on October 29, 2013 (year 1), and October 28, 2014 (year 2). The root samples were stained with 5% (w/v) black ink-vinegar solution22, and the AMF root colonization was measured according to Giovannetti and Mosse23.
DNA extraction and PCR amplification
Genomic DNA was taken from 50 mg of fresh root samples using the DNeasy Plant Mini Kit (Qiagen, GmbH, Germany) according to the manufacturer’s instructions. The DNA pellet was resuspended in 50 μl of TE buffer (pH 8.0) and stored at −30 °C until use for polymerase chain reaction (PCR). In total, 36 samples from the grassland field and a paddy field in the 2-year investigation were subjected to DNA extraction. The DNA samples extracted from roots were used as PCR templates. To decrease variations in the PCR process, samples were amplified in triplicate24 using the fusion primer set in a PCR at 10 μl per subsample, resulting in three independent PCR products per plant. Primers AMV4.5NF (forward) and AMDGR (reverse)25 were used for the amplification of AMF 18 S rRNA gene fragments by Mastercycler ep gradient (Eppendorf, Hamburg, Germany). PCR was performed in 10 μl reaction mixtures containing each 2X of reaction buffer, 0.3 μM of forward and reverse primers (10 μM), 1 U of Taq DNA polymerase (KOD multi & Epi, Toyobo, Japan), and 1 µl of template DNA. The PCR protocol was composed of initial treatment at 94 °C for 2 min; 45 cycles of treatments at 98 °C for 10 s and at 60 °C for 10 s.
Molecular diversity of AMF communities in roots
To reduce potential early-round PCR errors, three independent PCR products per plant were pooled together and purified using NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel, Duren, Germany), and quantified using UV spectrophotometry (DS-11 NanoPad, DeNovix Inc., USA). The purified PCR amplicons were normalized before pyrosequencing. The purified amplicons were paired-end (PE) sequenced on an Illumina MiSeq platform (Bioengineering Lab Co., Ltd, Kanagawa, Japan). In total, 36 sequencing libraries were constructed and independently sequenced. Sequence read processing was performed using QIIME version 1.9.126.
The reads were truncated at any site that received an average quality score <20 over a 40 bp sliding window, and the truncated reads shorter than 40 bp were discarded using FASTX-Toolkit. Then, PE reads were assembled according to their overlap sequence with a minimum overlap length of 10 bp, while reads that could not be assembled were discarded. The clean sequences were analyzed using the FLASH (Fast length adjustment of short reads). Chimeric sequences were identified and removed using UCHIME in USEARCH. Representative sequences were checked against the MaarjAM AMF database27 and NCBI GenBank. Phylogenetic analyses were performed using the Neighbor-joining (NJ) phylogenic trees (Tamure-Nei model) and maximum likelihood (TN93 + G) algorithms implemented in the program MEGA 7.028. Bootstrap values were estimated from 1,000 replicates. Additionally, the AMF OTU groups were classified according to Redecker et al.29. Representative sequence OTUs, defined as groups of closely related sequences with a high level of bootstrap support in the phylogenetic analysis, were selected. If more than one sequence from our study was present in the same OTU cluster, one was chosen as the representative sequence. Following the same similar approach as in Sosa-Hernández et al.30, we considered matches with ≥97% similarity a species level match, ≥90% a genus level match, and ≥80% a family level match. A species level match refers to how confidently we assigned a name to the OTU based on known sequences, and did not imply that these OTUs are to be considered equivalent to those species. The raw sequence data are available in the the DNA Data Bank of Japan (DDBJ) (DDBJ Sequence Read Archive, DRA006649, BioProject Accession: SSUB009022). Additionally, we calculated measures of sample coverage for all taxa using the R package vegan.
Statistical analysis
An arcsine-square root transformation was used for normalization of the data of AMF colonization in the roots of napiergrass. We determined significant differences between land-use types and sampling years were assessed using two-way analysis of variance (two-way ANOVA). In addition, Hill numbers (or the effective number of species) have been increasingly used to quantify the species/taxonomic diversity of an assemblage because they represent an intuitive and statistically rigorous alternative to other diversity indices31. Thus, diversity of AMF OTU communities were measured based on the first three Hill numbers, such as species richness, Shannon diversity (the exponential of Shannon entropy), and Simpson diversity (the inverse Simpson concentration) using the package iNEXT in R 3.4.2 (http://www.r-project.org/)31. The AMF communities among land-use types and sampling years were assessed using venn diagram analysis with the R package gplots.
We conducted permutational multivariate analysis of variance (PERMANOVA) using the vegan package in R 3.4.2 to examine the effect of land-use type and sampling year on AMF community structure32. To determine the relationship of land-use type and sampling year with respect to AMF communities, we performed redundancy analysis (RDA) as a multivariate analysis using the package vegan in R 3.4.2. The data of AMF communities were log-transformed. The sequences data matrix was composed of the abundance of AMF families and land-use type or sampling year. The environmental variable of land-use type and sampling year on the AMF communities were estimated according to Higo et al.33,34. This analysis was carried out using the vegan package in R 3.4.2 with 999 permutations of the Monte-Carlo test. To investigate if AMF community structure differed significantly between land-use type and sampling year, the PERMANOVA was performed with 9999 permutations using the adonis function in the package vegan in R 3.4.2.
Results
AMF colonization in the two different land-use types
Overall, AMF colonization in the napiergrass regardless of land-use type was never greater than 20% throughout the 2-year field study (Fig. 1). The AMF colonization ranged from 3.9% to 6.0% in grassland and from 4.5% to 4.7% in paddy field in sampling year 1 (2013). In sampling year 2 (2014), the AMF colonization ranged from 8.7% to 10.8% in the grassland, and from 10.8% to 11.7% in the paddy field. The AMF colonization in the napiergrass was influenced by sampling year, but was not influenced by land-use type.
Boxplots illustrating differences in group averages of root colonization of arbuscular mycorrhizal fungi (AMF) in napiergrass at two land-use types in sampling years 1 and 2. Bold horizontal lines represent median values; box margins ± SE and vertical lines represent minimum and maximum values of the groups. Error bars indicate standard errors of the means (n = 9). GL = grassland, PF = paddy field.
General sequencing information and taxonomic richness
In this study, a total of 3,861,816 paired-end sequences were obtained from the 36 libraries using the AMV4.5NF/AMDGR primer set. Of these, 3,264,994 sequences belonged to Glomeromycotina (corresponding to 84.5% of the total). We showed the taxonomic distributions of the obtained sequences in root samples (Table S2). We found a total of 230 OTUs in the AMF communities based on AMF family level (Table S2). The phylogenetic placement of OTUs in different land-use types was determined using the phylogenetic tree. The AMF OTUs were classified into one of nine AMF families including Glomeraceae (average relative abundance: 39.97%), Gigasporaceae (22.03%), Paraglomeraceae (33.06%), Claroideoglomeraceae (1.32%), Acaulosporaceae (1.06%), Archeosporaceae (0.90%), Ambisporaceae (0.38%), Diversisporaceae (0.18%) and unknown Glomeromycotina in roots (4.37%) (Fig. S1).
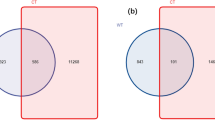
Diversity of AMF communities in the two different land-use types
A plateau was reached by all rarefaction curves (Fig. 2), which shows that high enough sequenced reads could identify most sequence types at a 97% similarity level (Table S3). The samples from roots in the sampling year 2 (2014), regardless of land-use type, had a larger number of OTUs (49–61) compared with those from roots in sampling year 1 (26–38OTUs). Additionally, we found 38, 26, 49 and 61 AMF OTUs in the sampling year-1 grassland, year-1 paddy field, year-2 grassland and year-2 paddy field, respectively (Fig. 3). The number of total OTUs in this experiment was 111, of which only 10.8% OTUs were common in all treatments. OTUs that occurred specifically in only the sampling year-1 grassland, year-1 paddy field, year-2 grassland and year-2 paddy field land-use types were 11.7%, 4.5%, 22.5% and 32.4%, respectively. In addition, the richness of root OTUs from the paddy field in sampling year 2 had a larger number of OTUs compared with those from the paddy field in sampling year 1 (Fig. 4A). No similar tendency in the OTU richness, Shannon diversity (H′) and Simpson index (1/D) was found in grassland. Additionally, no significant difference in the OTU richness between paddy field and grassland was found in both sampling years (Fig. 4A–C).
Boxplots illustrating differences in group averages regarding (A) operational taxonomic unit (OTU) richness, (B) Shannon index (H′) and (C) Simpson index (1/D) in the roots of napiergrass at two land-use management types in sampling years 1 and 2. Bold horizontal lines represent median values; box margins ± SE and vertical lines represent minimum and maximum values of the groups. n.s. = not significant by two-way ANOVA. Error bars indicate standard errors of the means (n = 9). GL = grassland, PF = paddy field. Circles show outliers.
Dynamics of AMF communities in the two different land-use types
Land-use type and sampling year strongly impacted AMF communities by influencing the proportions of three dominant AMF families (Fig. 5). Glomeraceae was predominant and detected at a much higher frequency in both grassland and paddy field regardless of sampling year. Gigasporaceae was more abundant in sampling year 2 compared with in sampling year 1 regardless of land-use type, and Paraglomeraceae in the paddy field in sampling year 2 had a larger number of abundance compared with in sampling year 1. No similar tendency was found in grassland. The distribution of each AMF family was different in each land-use type during the 2-year experiment.
Boxplots illustrating differences in group averages of each family of arbuscular mycorrhizal fungi (AMF) in napiergrass roots at two land-use management in the sampling years 1 and 2. Bold horizontal lines represent median values; box margins ± SE and vertical lines represent minimum and maximum values of the groups. GL = grassland, PF = paddy field. Circles show outliers.
Dynamics of AMF communities in the two different land-use types
The RDA trends clearly showed that the land-use type and sampling year significantly altered the AMF community structure in the roots (Fig. 6). As shown in Fig. 6, the AMF communities in the sampling year-2 plots were in the first (paddy field) and fourth (grassland and paddy field) quadrants, while most of the AMF communities in the sampling year-1 plots were in the second (paddy field) and third (grassland) quadrants. The ordination diagram indicates that paddy field and grassland (R2 = 0.756, P = 0.001), and the sampling year-1 and year-2 plots (R2 = 0.801, P = 0.001) contributed significantly to the variation in AMF root communities (Fig. 6). We also conducted PERMANOVA to determine the effect of each land-use type and sampling year to the AMF root communities. The result of PERMANOVA showed that land-use type (F = 2.577, P = 0.027) and sampling year (F = 3.763, P = 0.002) significantly affected the AMF root community structure, but the AMF root communities in the roots were not affected by the interaction of land-use and sampling year (F = 1.708, P = 0.1117) (Fig. 6).
Redundancy analysis (RDA) of communities of arbuscular mycorrhizal fungi (AMF) in roots sampled from two land-use types and sampling year. The environmental variables explain 18.4% and 14.6% of the total variations in the first two axes. Variables with significant effects in Monte Carlo tests (P < 0.05) are shown. Solid lines indicate effects of land-use types, and dashed lines indicate the effects of sampling year.
Discussion
Our study is the first investigation to reveal the distribution of indigenous AMF colonization and communities in the agricultural field around the Fukushima-Daiichi NPP after the March 11 disaster.
Occurrence of AMF colonization in roots
We observed the AMF colonization of napiergrass in Cs-contaminated soil (Fig. 1), which had slightly lower AMF colonization than that observed in other studies regardless of land-use type and sampling year7,35. A previous study reported that AMF root colonization in plants was negatively affected by increased soil Cs concentrations36, in agreement with the findings of our study. Thus, higher soil 137Cs concentration may inhibit the activity of the indigenous AMF population in the soil to promote mycorrhization in the 137Cs-contaminated soil. On the contrary, others studies have shown that soil Cs concentration did not inhibit AMF root colonization of leek, ryegrass, barely, cucumber, and sunflower7,35. Furthermore, it is unclear whether napiergrass increased Cs uptake from Cs-contaminated soil in terms of AMF root colonization. This is because our study did not examine the transfer of Cs uptake by napiergrass with AMF. However, Gyuricza et al.8 reported the capacity of AMF to transport radioactive Cs under in vitro conditions. These authors demonstrated that AMF could take up, translocate and transfer radioactive Cs to their host through their extraradical hyphal network. Thus, further investigation into the functional aspects of AMF root colonization on 137Cs uptake would provide support to elucidating the benefit of AMF on 137Cs removal from Cs-contaminated soil around the Fukushima-Daiichi NPP.
Distribution of AMF taxa in roots
Here, for the first time, we used an Illumina Miseq Platform of an approximately 250 bp SSU rDNA amplicon together with a high throughput phylogenetic annotation method to investigate AMF taxa in the 137Cs-contaminated soil. Overall, the most frequently detected families or OTUs in the roots of napiergrass between land-use type and sampling year belonged to Glomeraceae (Figs 5 and 6), which are commonly found in grasslands and arable lands37,38,39,40. Glomeraceae are especially predominant among arable fields because they adapt well to disturbed environments compared with other families as well as have high sporulation rates for rapid recovery41. Furthermore, previous studies have shown that Glomeraceae including the genus of Glomus, Funneliformis, Rhizophagus, and Paraglomus can form mycorrhizae in plant roots via their fragments of mycelium or colonized root fragments, rapidly constructing a anastomosis of hyphae42. However, Gigasporaceae including the genus of Gigaspora, Racocetra, and Scutellospora spreads and promotes via the dispersal of spore or infection from an intact mycelium43,44. Additionally, some other studies have indicated that Glomeraceae and Glomus-like genus (Paraglomus genus) have a certain resistance in complex environments45,46. As a result, these characteristics help with the survival and spread of Glomus-like genus including Paraglomus in a Cs-contaminated soil. Furthermore, the differences between Gigasporaceae and Glomeraceae clearly contribute to the dominance of the Glomeraceae OTUs over Gigasporaceae OTUs in roots from each land-use type. However, our results showed that Gigasporaceae OTUs were also relatively abundant among other detected AMF families. This result suggests that the emergence of the taxa can result from adaptation to the Cs-contaminated ecological environment.
Furthermore, previous studies Oehl et al.47 and Alguacil et al.48 reported that soil type strongly influenced AMF community structure as well as the prevalence and presence of many AMF. The AMF community structures differed between the soil characteristics of paddy field and that of grassland (Figs 5 and 6). Zhao et al.49 showed that AMF communities were highly affected by soil texture distribution, and their results indicated a significant relationship between the AMF community and the content of silt and sand. In this study, the soil characteristics such as the content of silt and sand were different between the soil of paddy field and that of grassland. These differences in content, and therefore, differences in aeration may be important for plant root growth and soil humus decomposition. Future studies examining the functional aspects of Glomeraceae or Gigasporaceae, and the impact of soil texture and plant root growth would help to elucidate the advantages of specific AMF on 137Cs uptake by AMF to plants in 137Cs-contaminated soils.
Dynamics of AMF communities in roots
Year-round shifts in composition of AMF communities in soils and colonizing roots have been reported among diverse plant species under various environmental conditions33,50,51,52,53,54. The year-round dynamics of AMF communities showed different structures between land-use type and sampling year (Figs 5 and 6). Similar results have been obtained in some other studies55,56,57,58. Some AMF OTUs were either replaced or disappeared in the 2-year investigation (Fig. 5). These shifts in OTU abundance and distribution, however, did not lead to changes to overall OTU richness among the AMF communities for either land-use type or sampling year. Thus, longer-term vs short-term trials could determine relationships among land-use type, host crops and AMF. When choosing AMF host crops or land-use type, AMF diversity and abundance should be prioritized to remove 137Cs and to ensure a healthy ecosystem in the 137Cs-contaminated field around the Fukushima-Daiichi NPP.
Conclusions
We found that the members of the Glomeraceae, Paraglomerace and Gigasporaceae were predominant in all land-use types. Our results revealed a temporal shift in the AMF communities in the 137Cs-contaminated soil around the Fukushima-Daiichi NPP over the 2-year period. Moreover, in different land-use types, diversity and structure of AMF communities also differed. Thus, different land-use types are potentially important to AMF communities. However, it remains unknown whether the AMF communities can be appropriate candidates for the phytoremediation technique for 137Cs removal in the long run. Therefore, it is important to better understand the relationships of 137Cs in soil and the functions of indigenous AMF communities in terms of ecological aspects to remove 137Cs from 137Cs-contaminated soil around the area of the Fukushima-Daiichi NPP. This information will provide a better understanding of the interaction between plants and AMF, for future revegetation and phytoremediation studies in polluted sites.
References
Yoshida, N. & Kanda, J. Tracking the Fukushima radionuclides. Science 336, 1115–1116 (2012).
Stohl, A. et al. Xenon-133 and caesium-137 releases into the atmosphere from the Fukushima Dai-ichi nuclear power plant: determination of the source term, atmospheric dispersion, and deposition. Atmos. Chem. Phys. 12, 2313–2343 (2012).
Kang, D. J. et al. Remediation of radiocesium-137 affected soil using napiergrass under different planting density and cutting frequency regimes. Water, Air, Soil Pollut. 228, 268 (2017).
Kang, D. J., Seo, Y. J., Saito, T., Suzuki, H. & Ishii, Y. Uptake and translocation of cesium-133 in napiergrass (Pennisetum purpureum Schum.) under hydroponic conditions. Ecotoxicol. Environ. Saf. 82, 122–126 (2012).
Kang, D. J., Tazoe, H., Yamada, M. & Ishii, Y. Differences in remediation effect of 137Cs in napiergrass (Pennisetum purpureum Schum.) under different land-use soil and cutting frequency conditions. Water, Air, Soil Pollut. 225, 2022 (2014).
Smith, S. E. & Read, D. J. Arbuscular mycorrhizaes. In Smith SE, Read DJ (Eds), Mycorrhizal symbiosis 3rd Edition. Academic Press, London, pp.13–187 (2008).
Rosén, K., Weiliang, Z. & Mårtensson, A. Arbuscular mycorrhizal fungi mediated uptake of 137Cs in leek and ryegrass. Sci. Total Environ. 338, 283–290 (2005).
Dubchak, S., Ogar, A., Mietelski, J. W. & Turnau, K. Influence of silver and titanium nanoparticles on arbuscular mycorrhiza colonization and accumulation of radiocaesium in Helianthus annuus. Spa. J. Agric. Res. 8, 103–108 (2010).
Gyuricza, V., Declerck, S. & Dupré de Boulois, H. Arbuscular mycorrhizal fungi decrease radiocesium accumulation in Medicago truncatula. J. Environ. Radio. 101, 591–596 (2010).
Hammer, E. C., Nasr, H., Pallon, J., Olsson, P. A. & Wallander, H. Elemental composition of arbuscular mycorrhizal fungi at high salinity. Mycorrhiza 21, 117–129 (2011).
Vinichuk, M., Rosen, K., Johanson, K. J. & Dahlberg, A. Correlations between potassium, rubidium and cesium (133Cs and 137Cs) in sporocarps of Suillus variegatus in Swedish boreal forest. J. Environ. Radioact. 102, 386–392 (2011).
Yoshida, S. & Muramatsu, Y. Concentration of alkali and alkaline earth elements in mushrooms and plants collected in a Japanese pine forest, and their relationship with 137Cs. J. Environ. Radioact. 41, 183–205 (1998).
Declerck, S., Dupré de Boulois, H., Bivort, C. & Delvaux, B. Extraradical mycelium of the arbuscular mycorrhizal fungus Glomus lamellosum can take up, accumulate and translocate radiocaesium under root-organ culture conditions. Environ. Microbiol. 5, 510–516 (2003).
De Boulois, H. D., Voets, L., Delvaux, B., Jakobsen, I. & Declerck, S. Transport of radiocaesium by arbuscular mycorrhizal fungi to Medicago truncatula under in vitro conditions. Environ. Microbiol. 8, 1926–1934 (2006).
Herder, G. D., Van Isterdael, G., Beeckman, T. & De Smet, I. The roots of a new green revolution. Trends Plant Sci. 15, 600–607 (2010).
Verbruggen, E., van der Heijden, M. G. A., Weedon, J. T., Kowalchuk, G. A. & Röling, W. F. M. Community assembly, species richness and nestedness of arbuscular mycorrhizal fungi in agricultural soils. Mol. Ecol. 21, 2341–2353 (2012).
van der Heijden, M. G. A. et al. The mycorrhizal contribution to plant productivity, plant nutrition and soil structure in experimental grassland. New Phytol. 172, 739–752 (2006).
Vogelsang, K. M., Reynolds, H. L. & Bever, J. D. Mycorrhizal fungal identity and richness determine the diversity and productivity of a tallgrass prairie system. New Phytol. 172, 554–562 (2006).
Lekberg, Y. et al. 454-sequencing reveals stochastic local reassembly and high disturbance tolerance within arbuscular mycorrhizal fungal communities. J. Ecol. 100, 151–160 (2011).
Öpik, M. et al. Global sampling of plant roots expands the described molecular diversity of arbuscular mycorrhizal fungi. Mycorrhiza 23, 411–430 (2013).
Chaudhry, V., Rehman, A., Mishra, A., Chauhan, P. S. & Nautiyal, C. S. Changes in bacterial community structure of agricultural land due to long-term organic and chemical amendments. Microb. Ecol. 64, 450–460 (2012).
Vierheilig, H., Coughlan, A. P., Wyss, U. & Piché, Y. Ink and vinegar, a simple staining technique for arbuscular-mycorrhizal fungi. Appl. Environ. Microbiol. 64, 5004–5007 (1998).
Giovannetti, M. & Mosse, B. An evaluation of techniques for measuring vesicular arbuscular mycorrhizal infection in roots. New Phytol. 84, 489–500 (2006).
Polz, M. F. & Cavanaugh, C. M. Bias in template-to-product ratios in multitemplate PCR. Appl. Environ. Microbiol. 64, 3724–3730 (1998).
Lumini, E., Orgiazzi, A., Borriello, R., Bonfante, P. & Bianciotto, V. Disclosing arbuscular mycorrhizal fungal biodiversity in soil through a land-use gradient using a pyrosequencing approach. Environ. Microbiol. 12, 2165–2179 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Öpik, M. et al. The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol. 188, 223–241 (2010).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Redecker, D. et al. An evidence-based consensus for the classification of arbuscular mycorrhizal fungi (Glomeromycota). Mycorrhiza 23, 515–531 (2013).
Sosa-Hernández, M. A., Roy, J., Hempel, S. & Rillig, M. C. Evidence for subsoil specialization in arbuscular mycorrhizal fungi. Front. Ecol. Evol. 6, 67 (2018).
Hsieh, T. C., Ma, K. H., Chao, A. & McInerny, G. iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol. Evol. 7, 1451–1456 (2016).
Hammer, Ø., Harper, D. A. T. & Ryan, P. D. PAST: Paleontological Statistics Software Package for Education and Data Analysis. 9 (2001).
Higo, M. et al. Can phosphorus application and cover cropping alter arbuscular mycorrhizal fungal communities and soybean performance after a 5-year phosphorus-unfertilized crop rotational system? PeerJ 6, e4606 (2018).
Higo, M., Tatewaki, Y., Gunji, K., Kaseda, A. & Isobe, K. Cover cropping can be a stronger determinant than host crop identity for arbuscular mycorrhizal fungal communities colonizing maize and soybean. PeerJ 7, e6403 (2019).
Vinichuk, M., Mårtensson, A. & Rosén, K. Inoculation with arbuscular mycorrhizae does not improve 137Cs uptake in crops grown in the Chernobyl region. J. Environ. Radio. 126, 14–19 (2013).
Wiesel, L., Dubchak, S., Turnau, K., Broadley, M. R. & White, P. J. Caesium inhibits the colonization of Medicago truncatula by arbuscular mycorrhizal fungi. J. Environ. Radio. 141, 57–61 (2015).
Higo, M. et al. Impact of a 5-year winter cover crop rotational system on the molecular diversity of arbuscular mycorrhizal fungi colonizing roots of subsequent soybean. Biol. Fertil. Soils 50, 913–926 (2014).
Higo, M. et al. Molecular diversity and distribution of indigenous arbuscular mycorrhizal communities colonizing roots of two different winter cover crops in response to their root proliferation. J. Microbiol. 54, 86–97 (2016).
Xu, M. et al. Land use alters arbuscular mycorrhizal fungal communities and their potential role in carbon sequestration on the Tibetan Plateau. Sci. Rep. 7, 3067 (2017a).
Xu, X. et al. The influence of environmental factors on communities of arbuscular mycorrhizal fungi associated with Chenopodium ambrosioides revealed by MiSeq sequencing investigation. Sci. Rep. 7, 45134 (2017).
Oehl, F. et al. Impact of land use intensity on the species diversity of arbuscular mycorrhizal fungi in agroecosystems of central Europe. Appl. Environ. Microbiol. 69, 2816–2824 (2003).
Giovannetti, M., Azzolini, D. & Citernesi, A. S. Anastomosis formation and nuclear and protoplasmic exchange in arbuscular mycorrhizal fungi. Appl. Environ. Microbiol. 65, 5571–5575 (1999).
Biermann, B. & Linderman, R. G. Use of vesicular-arbuscular mycorrhizal roots, intraradical vesicles and extraradical vesicles as inoculum. New Phytol. 95, 97–105 (2006).
Daniell, T. J., Husband, R., Fitter, A. H. & Young, J. P. W. Molecular diversity of arbuscular mycorrhizal fungi colonising arable crops. FEMS Microbiol. Ecol. 36, 203–209 (2001).
Bever, J. D., Richardson, S. C., Lawrence, B. M., Holmes, J. & Watson, M. Preferential allocation to beneficial symbiont with spatial structure maintains mycorrhizal mutualism. Ecol. Lett. 12, 13–21 (2009).
Barto, E. K. et al. The fungal fast lane: common mycorrhizal networks extend bioactive zones of allelochemicals in soils. PLoS One 6, e27195 (2011).
Oehl, F. et al. Soil type and land use intensity determine the composition of arbuscular mycorrhizal fungal communities. Soil Biol. Biochem. 42, 724–738 (2010).
Alguacil, M. M., Torres, M. P., Montesinos-Navarro, A. & Roldán, A. Soil characteristics driving arbuscular mycorrhizal fungal communities in semiarid Mediterranean soils. Appl. Environ. Microbiol. 82, 3348–3356 (2016).
Zhao, H. et al. Species diversity and drivers of arbuscular mycorrhizal fungal communities in a semi-arid mountain in China. PeerJ 5, e4155 (2017).
vandenkoornhuyse, P. et al. Arbuscular mycorrhizal community composition associated with two plant species in a grassland ecosystem. Mol. Ecol. 11, 1555–1564 (2002).
vandenkoornhuyse, P., Ridgway, K. P., Watson, I. J., Fitter, A. H. & Young, J. P. W. Co-existing grass species have distinctive arbuscular mycorrhizal communities. Mol. Ecol. 12, 3085–3095 (2003).
Higo, M. et al. Temporal variation of the molecular diversity of arbuscular mycorrhizal communities in three different winter cover crop rotational systems. Biol. Fertil. Soils 51, 21–32 (2015).
Higo, M., Isobe, K., Matsuda, Y., Ichida, M. & Torigoe, Y. Influence of sowing season and host crop identity on the community structure of arbuscular mycorrhizal fungi colonizing roots of two different gramineous and leguminous crop species. Adv. in Microbiol. 5, 107–116 (2015).
Higo, M., Takahashi, Y., Gunji, K. & Isobe, K. How are arbuscular mycorrhizal associations related to maize growth performance during short-term cover crop rotation? J. Sci. Food Agric. 98, 1388–1396 (2018).
Higo, M. et al. Diversity and vertical distribution of indigenous arbuscular mycorrhizal fungi under two soybean rotational systems. Biol. Fertil. Soils 49, 1085–1096 (2013).
Wang, C. et al. Differences in arbuscular mycorrhizal fungal community composition in soils of three land use types in subtropical hilly area of southern China. PLoS One 10, e0130983 (2015).
Xiang, D. et al. Land use influences arbuscular mycorrhizal fungal communities in the farming–pastoral ecotone of northern China. New Phytol. 204, 968–978 (2014).
Senés-Guerrero, C. & Schüßler, A. A conserved arbuscular mycorrhizal fungal core-species community colonizes potato roots in the Andes. Fungal Divers. 77, 317–333 (2016).
Acknowledgements
Thanks are expressed to Ayli Chong at Edanz Group for proofreading our manuscript and constructive criticism of the manuscript. This study was financially supported by a Grant-in-Aid for Scientific Research (C) Grant Numbers 25450495 and 17K08163. Additionally, this study was financially supported by a research grant from Life Science Research Center, College of Bioresource Sciences, Nihon University (S1391007).
Author information
Authors and Affiliations
Contributions
M.H. and D.K. designed the research; M.H., D.K. and K.I. performed the research and participated in the sampling. M.H. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Higo, M., Kang, DJ. & Isobe, K. First report of community dynamics of arbuscular mycorrhizal fungi in radiocesium degradation lands after the Fukushima-Daiichi Nuclear disaster in Japan. Sci Rep 9, 8240 (2019). https://doi.org/10.1038/s41598-019-44665-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44665-7
This article is cited by
-
Radiotolerant endophytic bacteria and analysis of the effects of 137Cesium on the metabolome of Pantoea sp.
Brazilian Journal of Microbiology (2024)
-
Arbuscular mycorrhizal fungi community composition, richness and diversity on enset (Ensete ventricosum (Welw.) Cheesman) in Ethiopia is influenced by manure application intensity in low-input farming systems
Plant and Soil (2022)
-
High-throughput sequencing analysis of the rhizosphere arbuscular mycorrhizal fungi (AMF) community composition associated with Ferula sinkiangensis
BMC Microbiology (2020)
-
Amplicon sequencing analysis of arbuscular mycorrhizal fungal communities colonizing maize roots in different cover cropping and tillage systems
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.