Abstract
The cucurbit powdery mildew elicited by Podosphaera xanthii is one of the most important limiting factors in cucurbit production. Our knowledge of the genetic and molecular bases underlying the physiological processes governing this disease is very limited. We used RNA-sequencing to identify differentially expressed genes in leaves of Cucumis melo upon inoculation with P. xanthii, using RNA samples obtained at different time points during the early stages of infection and their corresponding uninfected controls. In parallel, melon plants were phenotypically characterized using imaging techniques. We found a high number of differentially expressed genes (DEGs) in infected plants, which allowed for the identification of many plant processes that were dysregulated by the infection. Among those, genes involved in photosynthesis and related processes were found to be upregulated, whereas genes involved in secondary metabolism pathways, such as phenylpropanoid biosynthesis, were downregulated. These changes in gene expression could be functionally validated by chlorophyll fluorescence imaging and blue-green fluorescence imaging analyses, which corroborated the alterations in photosynthetic activity and the suppression of phenolic compound biosynthesis. The powdery mildew disease in melon is a consequence of a complex and multifaceted process that involves the dysregulation of many plant pathways such as primary and secondary metabolism.
Similar content being viewed by others
Introduction
The Cucurbitaceae or cucurbit family includes many economically important species, particularly those with edible fruits such as cucumber, melon, watermelon, zucchini and pumpkin1. Unfortunately, phytopathogens hinder the production of cucurbits, leading to over 200 known diseases with diverse aetiologies2. Cucurbit powdery mildew is the primary fungal disease that affects cucurbits under both greenhouse and open field conditions. Like other powdery mildew diseases, it is easily recognizable by characteristic symptoms such as the whitish, talcum-like, powdery fungal growth that develops on leaf surfaces, petioles and stems, and rarely on fruits2,3. There are two species that can cause powdery mildew in cucurbits around the world, namely, Golovinomyces orontii and Podosphaera xanthii. The latter is considered to be the main causal agent of powdery mildew in cucurbits, and it is one the most important limiting factors for cucurbits production4,5,6.
The arrival of a conidium to the leaf of a susceptible host and its adhesion, penetration, nutrition and proliferation as well as plant defence response suppression, are necessary steps for the establishment of a compatible interaction7. Like most powdery mildew fungi, P. xanthii grows on the foliar surface, taking nutrients from the epidermal host cells through the development of a specialized parasitism-related structure called the haustorium8. The epiphytic transcriptome of P. xanthii is available9, several molecular tools have been developed for the specific functional analysis of P. xanthii genes such as host-induced gene silencing and transient transformation10,11 and the transcriptomes and genomes of major crops such as cucumber, melon, watermelon and zucchini are also available12,13,14,15,16,17,18. However, the genetic and molecular bases of the physiological processes governing this intimate plant-fungus interaction remain largely unknown.
RNA-seq is a revolutionary tool for transcriptomics which has altered our view of the extent and complexity of eukaryotic transcriptomes19. RNA-seq analysis has been demonstrated to be an effective approach to decipher the primary changes in gene expression, providing a far more precise measurement of transcript levels and their isoforms than other methods and allowing researchers to detect transcripts with low abundance20,21. Fungal diseases of plants remain a major challenge in agriculture. Hence, an understanding of disease mechanisms at the molecular level is of paramount importance for identifying possible intervention points for their control22. For this reason, several studies have used RNA-seq analysis to obtain a more comprehensive view of the primary molecular mechanisms that are dysregulated in plants upon pathogen inoculation23,24,25,26,27. However, whole-transcriptome changes during early disease stages in susceptible plant species are less well-documented than those of resistant ones. This is the case for cucurbit-powdery mildew interactions. There are only two reports that describe the transcriptome profiles of pumpkin and melon lines resistant to powdery mildew and their comparison with susceptible cultivars23,28.
Different imaging techniques are currently widely used in plant physiology to assess the impact of biotic stress on host plants since they reveal the metabolic gradients that pathogens usually induce in infected leaves29,30. Examples of these techniques include chlorophyll fluorescence imaging (Chl-FI) and multicolour fluorescence imaging (MCFI). Studying the red chlorophyll fluorescence (Chl-F) emitted by photosystem II (PSII) provides information on the photosynthetic performance of plants in terms of activity31 and indirect information on the CO2 assimilation rate32. Similarly, MCFI is a very useful technique for monitoring the plant health status. It is based on recording the blue (F440), green (F520), red (F680) and far red (F740) fluorescence that leaves emit when they are excited with UV light. Particularly, the so-called blue-green fluorescence (BGF) is a valuable technique to study secondary metabolism, since phenolic compounds from the phenylpropanoid pathway are the primary emitters of that fluorescence33,34. To date, many plant-pathogen interactions have been subject to analysis using imaging techniques29,35 and most of them have the primary aim of identifying specific disease identity marks.
This study is intended to elucidate the primary alterations that take place in the physiology and metabolism P. xanthii-inoculated melon plants during the first stages of infection. For this purpose, RNA-seq analysis was used to search for changes in leaf gene expression after pathogen inoculation. In parallel, plants were phenotypically characterized by different imaging techniques to provide experimental support for the physiological changes anticipated by RNA-seq analysis. Our findings show that photosynthesis and secondary metabolism are the primary physiological processes of the host that are altered during the initial infection stages. This study is the first step in looking for effectors of P. xanthii specifically aimed at manipulating these central functions of the host physiology. Unraveling these mechanisms would be desirable as a preliminary step in the development of new melon varieties that are resistant to P. xanthii.
Results
Visualization of the development of P. xanthii structures during the first stages of infection
In parallel with the sampling of plant material for RNA-seq analysis, leaf disks were collected to visualize the development of P. xanthii infection structures during the period of time analyzed by RNA-seq, that is, the first 72 h of interaction (Fig. 1). At 0 hpi (hours post-inoculation), only the conidia were visible. At 24 hpi, most of the spores had germinated. At this stage, although not observed in the pictures, the first haustoria had already developed8. At 48 hpi, the primary hyphae were very abundant, and the initial formation of secondary hyphae could also be detected. Finally, at 72 hpi, extensive branching was observed in the secondary hypha.
Time-course analysis on the development of P. xanthii on the leaves of melon plants used for RNA-seq analysis by CLSM. Fungal structures were stained with propidium iodide. The pictures were taken at 0, 24, 48 and 72 h post-inoculation (hpi). (A–C), an un-germinated conidium on the leaf surface. (D,F), a conidium with a germ tube (arrowhead). (G–I), a germinated spore (arrowhead) with primary hyphae and initial development of secondary hyphae. (J–L), Weft of secondary hyphae. Bars: (A–F) 10 µm; (G–L) 100 µm.
Illumina sequencing, mapping and identification of DEGs
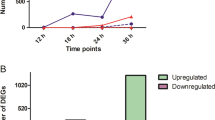
High-throughput sequencing produced a total of 1661.7 million short reads, of which 86.41% (1435.9 million) passed the quality control thresholds. To calculate the expression profile, clean reads were mapped onto the melon reference transcriptome with the Bowtie2 algorithm. The alignment results were reported in SAM/BAM format. The mapping rates for various samples are summarized in Table S1. The expression estimates are reported in total counts for each gene. DESeq. 2 software was employed to analyse the differentially expressed genes at three time points (24, 48 and 72 hpi) in infected plants relative to un-inoculated control plants. Differentially Expressed Genes (DEGs) were reported in log2FoldChange (FC) with a corresponding p-value for each gene. An MA plot was used to represent the DEGs of infected plants relative to control plants at 24, 48 and 72 hpi (Fig. 2A). Among all the DEGs, 538 were common at three time points, while 237, 1870 and 2448, were exclusively exhibited at 24 hpi, 48 hpi and 72 hpi, respectively (Fig. 2B). Furthermore, the heatmap of expression levels of all these melon DEGs showed that the expression profiles of them varied significantly in response to P. xanthii infection (Fig. 2C). A total of 1114 genes showed differential expression patterns at 24 hpi, from which 863 were found to be upregulated and 251 downregulated. The number of DEGs at 48 hpi was 3785, with 2781 upregulated and 1004 downregulated genes. At 72 hpi 4226 DEGs were detected. Of these, 2697 exhibited upregulation and 1529 showed downregulation (Fig. 2D).
Summary of sequence data and the number of differentially expressed genes determined by RNA-seq in melon plants infected with P. xanthii versus non-infected control plants. Data were collected at 24, 48 and 72 h post-inoculation (hpi). (A) log2-fold change versus a mean expression scatter plot of the RNA-seq results. (B) A Venn diagram displaying the distribution of DEGs (genes with >1 or > −1 log2-fold change in expression) at each time point. (C) Hierarchical clustering of DEGS at each time point. Rows are clustered using distance and average linkage. Changes of gene expression are displayed from red (lower expression) to green (higher expression). (D) Number of genes up- and downregulated at each time point in infected plants compared to non-infected control plants.
Gene functional enrichment analysis of DEGs and metabolism overview
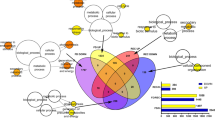
To understand the primary functions of melon DEGs in response to P. xanthii infection, a gene functional enrichment analysis was performed using GENECODIS. A singular enrichment analysis of the GO biological process revealed a total of 33, 108 and 83 significantly enriched (adj. p-val. < 0.05) Gene Ontology (GO) terms at 24, 48 and 72 hpi, respectively (Table S2). These GO terms were ordered according to their hypergeometric p-value (Hyp), with the first positions being those with minor Hyp values. Subsequently, the GO terms were filtered to select the top 20 to reduce the amount of data and to facilitate the representation of the most enriched terms at each time point (Table 1). Among the first 20 GO terms over the three analysed time points, photosynthesis-related terms such as photosynthesis (GO:0015979), response to red light (GO:0010114), response to blue light (GO:0009637), response to far red light (GO:0010218) or response to light stimulus (GO:0009416) were common. Furthermore, GO terms for secondary metabolism-related genes such as response to wounding (GO:0009611), defence response to fungus (GO:0050832) or lignin biosynthesis process (GO:0009809) were also highly represented, with these GO terms being especially important at 72 hpi. In addition, other terms related to hormone signalling such as response to jasmonic acid stimulus (GO:0009753), response to abscisic acid stimulus (GO:0009757) or response to salicylic acid stimulus (GO:0009751) were also present as biological process functional enrichment.
Since the main dysregulated processes were similar at the three time points analyzed, to obtain a more comprehensive overview of the primary process dysregulation that occurred in melon plants during the first stages of P. xanthii infection, we focused our study at 24 and 72 hpi and performed a MapMan analysis of DEGs at those time points (Fig. 3). In general terms, at 24 hpi, primary metabolism-related genes such as light reaction genes, Calvin cycle genes and photorespiration genes were upregulated, whereas some secondary metabolism-related genes such as phenylpropanoid and flavonoid biosynthesis genes were slightly downregulated. Furthermore, cell wall modification-related genes were generally upregulated (Fig. 3A). At 72 hpi, the light reaction genes, Calvin cycle genes and photorespiration genes were also upregulated. Similarly, certain secondary metabolism related genes were downregulated, and a few of them were strongly repressed. Additionally, some starch synthesis and degradation-related genes were also down-regulated. However, compared to 24 hpi, the cell wall modification-related genes were mostly downregulated at 72 hpi (Fig. 3B). The MapMan analysis allowed us to validate that the most important GO terms were identified, completing the information and specifying the relative expression level of the primary dysregulated processes. Therefore, in general, photosynthesis and related processes were upregulated, whereas the major secondary metabolism pathways such us the phenylpropanoid biosynthesis pathway were downregulated.
MapMan overview of plant metabolism in P. xanthii-infected melon leaves showing all the DEGs at 24 (A) and 72 hpi (B). The analysis was performed using MapMan v.3.5.0. Individual genes are represented by small squares. The colour key represents the RPKM (Reads Per Kilobase Million)-normalized log2-transformed counts. Red represents the downregulation and green represents the upregulation of melon genes from infected plants compared to uninfected controls.
P. xanthii infection induces changes in the expression of melon genes involved in photosynthesis and secondary metabolism
To more thoroughly visualize the genes subjected to gene expression changes in some of the primary process that were dysregulated in melon plants infected by P. xanthii, a MapMan analyses specific for photosynthesis and secondary metabolism were performed at 24 and 72 hpi (Fig. 4). Among the 28 DEGs involved in photosynthesis and related processes at 24 hpi, 26 were upregulated, whereas only 2 were downregulated (Table S3). For example, several lhc genes that encode Lhcb and Lhca antenna proteins of PSII and photosystem I (PSI), respectively, or petE2 that encodes the predominant plastocyanin isoform, were upregulated. In addition, some Calvin cycle genes such as those encoding ribulose bisphosphate carboxylase small chain (rbcS-1a, rbcS-2b and rbcS-3b), phosphoribulokinase (prk) and rubisco activase (rca) were also overexpressed. The only two suppressed genes were psbT and OHP1, which encode the PSII 5 kDa protein subunit PSII-T and one helix protein 1, respectively (Fig. 4A). At 72 hpi, 50 photosynthesis-related genes were DEGs. Forty-one of these genes were upregulated and 9 downregulated (Table S3). Among the upregulated genes, we note the other lhc genes (lhcb5 and lhca3), psb genes of PSII (psbO-2, psbW, psbX, and psbY) or petE1 and pgr5, codifying isoform 1 of plastocyanin and proton gradient regulation 5 protein, respectively. Although the psbT gene was not repressed, psbQ and psb28 were downregulated at 72 hpi (Fig. 4B).
MapMan overview of primary photosynthetic metabolism in P. xanthii-infected melon leaves showing all the DEGs at 24 (A) and 72 hpi (B). The analysis was performed using MapMan v.3.5.0. Individual genes are represented by small squares. The colour key represents the RPKM-normalized log2-transformed counts. Red represents the downregulation and green represents the upregulation of melon genes from infected plants compared to uninfected controls.
A similar approach was performed to analyse the changes in the expression of secondary metabolism-related genes in P. xanthii-infected melon plants (Fig. 5). Among the 20 DEGs involved in secondary metabolism and detected at 24 hpi, 12 were downregulated and 8 were upregulated (Table S3). Several genes involved in the phenylpropanoid and shikimate pathways, which have a key role in the biosynthesis of different defence compounds against pathogens, were repressed, including the phenylalanine ammonia lyase isoform genes pal1 and pal4, which are involved in the first step of phenylpropanoids biosynthesis. By contrast, pal2 was upregulated. Other important genes in the phenylpropanoids pathway such as c4h and 4cl3 were downregulated. Moreover, adt6, a gene related to phenylalanine biosynthesis, was also downregulated (Fig. 5A). At 72 hpi, 62 genes involved in secondary metabolism were DEGs, of which 28 were downregulated and 34 upregulated (Table S3). For example, pal4 was strongly repressed, whereas pal2 was overexpressed. In addition, although 4cl3 remained downregulated, 4cl1 and 4cl2 were overexpressed. Furthermore, adt6 experienced an increase repression level and other genes in the shikimate pathway that trigger phenylalanine biosynthesis, such as dhsp1, dhsp2, dhsp3, sk1 and sk2, were downregulated (Fig. 5B). In summary, at 24 hpi, the phenylpropanoid pathway seems to be affected at the phenylalanine and p-coumarate biosynthesis levels, whereas at 72 hpi, the inhibition of this pathway is upstream, at the first steps of shikimate synthesis (Fig. S1).
MapMan overview of secondary metabolism in P. xanthii-infected melon leaves showing all DEGs at 24 (A) and 72 hpi (B). The analysis was performed using MapMan v.3.5.0. Individual genes are represented by small squares. The colour key represents the RPKM-normalized log2-transformed counts. Red represents the downregulation and green represents the upregulation of melon genes from infected plants compared to uninfected controls.
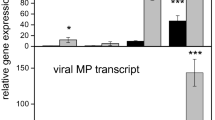
The results obtained by RNA-Seq were validated by qRT-PCR analysis. Twelve DEGs involved in photosynthesis and secondary metabolism were selected, subjected to expression analysis by qRT-PCR at the same time points and compared to the RNA-seq data. In general terms, the expression levels observed by qRT-PCR were very similar to those obtained by RNA-seq, with the only exceptions being the OHP1 and pgr5 at 48 hpi. These results allowed us to confirm the reliability of the RNA-seq data (Fig. 6).
Validation of RNA-Seq data from P. xanthii-melon compatible interactions by qRT-PCR analysis of selected primary and secondary metabolism DEGs. The relative expression (log2-fold change) of twelve genes at 24, 48, and 72 hpi is shown. The transcript abundance was normalized to the transcription of the endogenous control β-actin gene (XM_008462689.2) and the relative expression of each gene was calibrated to the uninfected control at the corresponding time point. The qRT-PCR data (white bars) are expressed as the mean values of three experimental replicates from three independent experiments, with error bars depicting the standard error. The RNA-seq data (black bars) are also shown for comparison.
Fluorescence imaging analysis data from melon leaves in response to P. xanthii infection are consistent with the RNA-seq results
To functionally validate the expression data revealed by RNA-seq analysis, the impact of the fungal infection on melon physiology was investigated by different fluorescence imaging techniques. The impact on leaf photosynthesis was investigated in terms of PSII efficiency and Calvin cycle activity (Fig. 7). The maximum quantum yield of PSII as measured as the FV/FM, did not show significant differences between infected and uninfected leaves. However, a consistent inhibition of the PSII (at 24 and 72 hpi) could be measured as a decrease in the ΦPSII of infected leaves, correlating with a reduction in the net CO2 fixation rate (PN) of infected leaves at 72 hpi. Notably, the NPQ was significantly higher in infected leaves at 24 and 72 hpi. In addition, all these changes in the Chl-FI parameters could not be attributed to alterations in the contents of pigments such as chlorophylls, carotenoids or xanthophylls induced by the fungus, since no significant differences were found between infected and uninfected plants (Table S4).
Impact of P. xanthii infection on the primary metabolism of melon leaves. (A) Standard images of the RGB, FV/FM, ΦPSII and NPQ from non-infected and P. xanthii-infected melon leaves at 24 and 72 hpi. A false colour scale was applied for each parameter. Images from a representative experiment are shown. (B) The average values of FV/FM, ΦPSII, and NPQ (n = 6) and the net photosynthesis rate (n = 10) for non-infected and infected melon leaves at 24 and 72 hpi are shown, with bars representing the standard error. The asterisks indicate a statistically significant difference between the samples according to a Student’s t-test (∗P < 0.1, ∗∗P < 0.01, and ∗∗∗P < 0.001). The abbreviations are: FV/FM, maximum quantum yield of PSII; ΦPSII, effective quantum yield of PSII; NPQ, non-photochemical quenching; and PN, net photosynthesis rate.
The accumulation of secondary metabolites in response to fungal infection was analysed by BGFI (Fig. 8). The F440 and F520 emitted by the infected leaves were comparable to that released by the uninfected controls at 24 and 72 hpi. This result indicated that there was no alteration in the accumulation of phenolic compounds in response to P. xanthii infection. In conclusion, and as revealed by RNA-seq analysis, this fungal infection seems to alter the photosynthesis efficiency of susceptible melon plants and manipulate the activation of the phenylpropanoid pathway.
Impact of P. xanthii infection on the secondary metabolism of melon leaves. (A) Standard images of the RGB, F440 and F520 emitted by phenolic compounds from non-infected and P. xanthii-infected melon leaves at 24 and 72 hpi. A false colour scale was applied for each parameter. Images from a representative experiment are shown. (B) The average values of F440 and F520 (n = 6) for non-ninfected and infected melon leaves at 24 and 72 hpi are shown, with bars representing the standard error. No statistically significant differences between the samples were obtained according to a Student’s t test. The abbreviations are F440, blue fluorescence and F520, green fluorescence.
Discussion
The cucurbit powdery mildew disease elicited by P. xanthii is one of the most important limiting factors in cucurbit production worldwide1. The molecular mechanisms underlying this pathogenic interaction remain largely unknown, with only a few studies addressing the transcriptomic analysis in this fungus9 and in resistant pumpkin and melon lines23,28. For that reason, this study is intended to decipher the primary molecular and physiological changes taking place in melon plants during the first stages of the compatible interaction with P. xanthii. To this end, we analyzed this interaction using a combination of RNA-seq with different imaging techniques.
RNA-seq is a sensitive and effective approach for identifying the primary changes in gene expression20,21, but it generates a large amount of data that are difficult to process. To visualize the data and to perform a more efficient analysis, tools such as GO terms enrichment or MapMan analyses were used23,36,37. Although some terms obtained by GO enrichment analysis were different at the three time points analyzed here, most of them were related to two major physiological processes, photosynthesis and secondary metabolism. Therefore, to yield a simpler and comprehensive study, the subsequent analyses were performed using 24 and 72 hpi data points, which, on the other hand, were the time points from which image analysis data were obtained. The metabolism overview performed by MapMan yielded very similar results relative to the ones obtained through GO terms enrichment analysis, but the first approach allowed us obtain more visual results and to identify the primary genes dysregulated in each pathway. Therefore, by combining both analyses, it has been possible to detect different functional processes and related pathways that were being dysregulated in the melon plants infected with P. xanthii, with photosynthesis and secondary metabolism being the stand-outs.
Among all the photosynthesis-related DEGs, only ohp1 was downregulated at both 24 and 72 hpi. The Ohp1 protein is required for PSII function38. Therefore, the downregulation of this gene could be part of a mechanism to decrease ΦPSII in a regulated manner. However, lhcb6 was upregulated upon inoculation at all the data points. Lhcb6 is a subunit of the minor light harvesting complex of PSII that is involved directly in the thermal dissipation of excess energy39,40. Furthermore, psbS was up-regulated at 48 hpi, coinciding with the maximum expression of lhcb6. The interaction of PsbS with the minor antenna, including Lhcb6, is crucial for the regulation of NPQ41,42. Moreover, the expression of petE2 and pgr5 was also upregulated. Both genes encode key players in the cyclic electron flow around PSI, which dissipates the ΔpH across the thylakoid43,44,45, and it is essential for the photoprotection of the thylakoids46. These transcriptomics data are consistent with those of other plants infected with powdery mildew, in which several photosynthesis-related genes were found to be upregulated23,47,48.
The photosynthesis-related RNA-seq results were validated by different functional approaches such as chlorophyll fluorescence imaging (Chl-FI) and the net photosynthesis rate (PN). Compared to the control leaves, the P. xanthii-infected melon leaves showed photosynthesis inhibition at two levels, in terms of the electron transport rate (decrease in ΦPSII) and carbon fixation (decrease in PN) from 24 hpi onwards. Moreover, in the infected leaves, the mechanism of energy dissipation, as measured as the NPQ, was induced, although the FV/FM was not affected. The photosynthesis was previously shown to be inhibited, as measured as a decrease in the ΦPSII and PN, while the NPQ increased during the fungal infection35,49. These photosynthetic changes were not attributed to alterations in the chlorophyll contents of melon leaves. Those data are in accordance with previously obtained results in cucumber leaves infected with P. xanthii, which displayed a decrease in the ΦPSII several days before a reduction in the chlorophyll content could be detected50. Hence, the changes in the ΦPSII and NPQ could be caused by a modification in the expression of genes such as ohp1, lhcb6 and psbS. Swarbrick et al.51 demonstrated downregulation in the expression of photosynthetic genes such as rcbS and Chl-a/b-binding proteins (cab) related to a decrease in the PN of barley leaves infected by powdery mildew. However, during P. xanthii-melon interaction, there were increases in the expression levels of important photosynthetic genes such as rbcS 1-a, 2-b, 3-b, rca and prk. This finding could be the result of the manipulation of gene regulation by P. xanthii to increase the photosynthesis and photosynthates in melon plants for its own benefit; nevertheless, the plant effectively inhibited the photosynthetic activity, as a part of the plant defence programme to limit carbon source availability for the pathogen52. Cyclic electron flow around PSI, for which petE and pgr5 genes were found to be upregulated, could contribute to the inhibition of the Calvin cycle.
It is well known that phenylpropanoid compounds play a key role in plant defence against pathogens53. Therefore, the repression of different DEGs related to this pathway and the shikimate pathway could be important for the establishment of compatible interactions. Whereas c4h, a key gene in lignin synthesis54, was downregulated at 24 hpi, did not show expression changes at 72 hpi, suggesting that it could be affecting the phenylalanine synthesis only at very early stages of the interaction. However adt6, pal1, pal4 and 4cl3 were downregulated at 24 and 72 hpi. The arogenate pathway has been described as the predominant pathway for phenylalanine synthesis55,56, hence, enzymes such as prephenate aminotransferase (PAT) or arogenate dehydratase (ADT) become key action points. In relation to this situation, the observed downregulation of adt6 could interfere with phenylalanine synthesis. The repression of pal genes could be even more important. This gene encodes enzymes that catalyse the first step in the phenylpropanoid pathway and are considered a key point of regulation between primary and secondary metabolism57. Between the pal1 and pal2 genes, Rohde et al.58 qualified pal1 as being more important for phenylpropanoid biosynthesis. Moreover, different studies indicate that pal1 expression is induced by wounding and pathogen attack59,60,61,62. In addition, pal4 has been shown to act as an important gene in lignin biosynthesis58,63,64. In our study, both the pal1 and pal4 genes showed increased repression at 72 hpi, when the fungal development was more evident. The 4cl3 repression does not seem very important, since its function is related to flavonoid biosynthesis, but not to lignin biosynthesis, whose function is performed primarily by 4cl1 and 4cl265, both of which are upregulated at 72 hpi. However, the repression of several genes in the shikimate pathway at 72 hpi, such as dahps1, dahps2, dahps3, sk1 and sk2, was more important since they act as shikimate pathway regulators65. The dahps (3-Deoxy-D-arabinoheptulosonate 7-phosphate synthase) genes act as a branch point to convert primary carbon source into the shikimate pathway and could act as a shikimate pathway key point to compete for phosphoenolpyruvate and erythrose 4-phosphate with the glycolysis and the pentose phosphate pathway65. Moreover, dahps genes are induced by wounding and fungal elicitors, which induce the synthesis of phenylpropanoids metabolites66,67. On other hand, several studies have suggested that sk genes have an important role in plant defense response, increasing their expression in the presence of fungal elicitors68. The repression of key genes related directly to plant defense such as pal or, dahps and sk when the development of P. xanthii increases, suggests that they are manipulated by the fungus as a strategy to favour infection development. Our RNA-seq data suggest that the phenylpropanoid synthesis is not totally blocked but is affected enough to avoid the accumulation of compounds necessary for plant defense, such as lignin or phytoalexins, resulting in a compatible interaction as described in other pathosystems24,69. Similarly, secondary metabolism-related genes have been found to be commonly repressed in other plant-powdery mildew compatible interactions26,48,70.
BGF experiments were conducted to validate the RNA-seq results on secondary metabolism. The accumulation of plant defence compounds is frequently found in diseased plants. This accumulation of host secondary metabolites could be measured as an increase in F440 and F550 in bacterial, viral and fungal infections29. However, in P. xanthii infection, the F440 and F550 parameters did not display significant differences between uninfected and infected melon leaves, pointing to no changes in the accumulation of secondary metabolites in response to P. xanthii. This result is in accordance with a previous study on the barley powdery mildew Blumeria graminis in susceptible barley plants51. Furthermore, P. xanthii infection in zucchini plants was related to an increase of blue-green fluorescence from the leaves71. However, in that case, the fluorescence was shown to be emitted by components of the fungal cell walls and not by plant secondary metabolites.
But what is the situation in cucurbit cultivars resistant to P. xanthii? Two transcriptomic studies carried out with pumpkin and melon lines resistant to P. xanthii showed that different genes of photosynthesis and secondary metabolism were differentially expressed23,28. Thus, most of photosynthesis related genes such as psb27, psbP, psaK, psaE, psaH, psaG or psaN were up-regulated in the pumpkin inbred resistant line “112-2”23, whereas in the melon resistant cultivar MR-1, carbon fixation related genes were up-regulated at the early stages of the interaction, but they were down-regulated at 72 hpi28. However, in the melon susceptible cultivar “Rochet” examined in this work, other photosynthesis related genes were either down-regulated such as ohp1 or up-regulated such as lhcb6, petE and pgr5. In addition, the carbon fixation related genes were up-regulated at all time points analyzed. With respect to the secondary metabolism related genes, in the pumpkin resistant line 4cl3 or adt6 were up-regulated23. Aligned to this, in the melon resistant cultivar most of the pathogen-induced DEGs were enriched in pathways related to flavonoids, phytoalexins and phenylpropanoid biosynthesis, such as, for example, the pal genes that increased their expression after exposure to the pathogen28. In contrast, in the melon susceptible cultivar, most of the secondary metabolism genes were down-regulated, including the pal genes. In other words, the genes of photosynthesis and secondary metabolism of susceptible and resistant plants are expressed differentially and in opposite ways in response to the attack of P. xanthii.
Our RNA-seq and fluorescence imaging results suggest that during the first stages of infection, P. xanthii manipulates plant gene expression. Because of the plant-pathogen interaction, melon leaf physiology and especially photosynthesis and secondary metabolism were altered. Host manipulation is believed to be carried out by the panel of effectors secreted by the pathogen72. During host colonization, P. xanthii secretes an important number of proteins, most of which have an unknown function9,11. We believe that some of these proteins could be responsible for reprogramming the melon physiology to allow disease establishment. The catalogues of P. xanthii effector candidates, both epiphytic9 and haustorial (Polonio et al., unpublished results), are known. To identify P. xanthii effectors that specifically target photosynthesis or secondary metabolism, different approaches such as yeast two-hybrid screening73,74,75 or dual RNA-seq analysis76,77 can be undertaken. Currently, we are resolving the melon-P. xanthii interactions through a dual RNA-seq approach.
Methods
Growth conditions of plants and fungi
Melon plants (Cucumis melo L.) cv Rochet (Semillas Fitó, Barcelona, Spain) susceptible to the P. xanthii isolate 2086 were used in all the experiments. The plants were cultivated in a growth chamber with a 16 h: 8 h, light: dark cycle at 25 °C for one month. The powdery mildew isolate was cultured on previously disinfected zucchini (Cucurbita pepo L.) cotyledons cv Negro Belleza (Semillas Fitó) and maintained in Bertrand medium under a 16 h: 8 h, light: dark cycle at 22 °C for one week78.
Experimental design
To elucidate the changes in the gene expression levels in these melon plants during the first stages of their compatible interactions with P. xanthii, we performed an RNA-seq analysis at 24 h, 48 h and 72 h post-inoculation (hpi). For the inoculation, P. xanthii conidia were collected by immersing infected zucchini cotyledons in 50 ml of a 0.01% Tween-20/distilled water solution. The third and fourth true leaves of 1-month-old melon plants were inoculated with a spore suspension at 1 × 106 conidia ml−1 (infected plants) or with 0.01% Tween-20 in distilled water (control plants). We used the reference transcriptome of C. melo to define the differentially expressed genes (DEGs) and later orthologous annotation with Arabidopsis thaliana to perform GO enrichment and MapMan analysis. In parallel, the changes in the physiology of the melon plants induced by the infection were analysed using different imaging techniques and other experimental approaches as described below.
Confocal laser scanning microscopy (CLSM)
To visualize the development of the fungal structures during the initial stages of the infection process, a CLSM analysis was performed. For this purpose, leaf discs were taken at the same time points selected for RNA-seq analysis and the fungal structures were stained with an aqueous solution of 100 µg mL−1 propidium iodide79. The samples were observed using a Leica SP5 II confocal microscope (Leica Microsystems, Wetzlar, Germany). The samples were excited with a 488 nm laser line and their fluorescence was detected over a 510–570 bandpass range. Bright field images were taken using the transmission channel. All the images were observed using a 40× oil-immersion objective and were processed with Leica LAS AF software (LCS Lite, Leica Microsystems).
RNA isolation, cDNA library construction and Illumina sequencing
For RNA isolation, melon leaves were collected, immediately frozen in liquid nitrogen and stored at −80 °C until use. The frozen leaves were ground with a mortar and pestle and the total RNA was extracted using TRI Reagent (Sigma-Aldrich, Saint Louis, USA) according to the manufacturer’s instructions. The total RNA was quantified using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The quality and quantity of the RNA were measured on an Agilent Bioanalyzer 2100 using an RNA Pico 6000 chip (Agilent Technologies, Santa Clara, CA, USA). Approximately 1 μg of each sample was used for cDNA library construction using an Illumina Stranded mRNA Sample Preparation Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions. Finally, the cDNA libraries were sequenced by Illumina NextSeq 550 system (Illumina).
RNA-Seq data analysis
The detailed strategy for the differential expression analysis is depicted in Fig. S2. The analysis process was automatized using Autoflow80. The raw reads were pre-processed using the SeqTrimNext pipeline81 (http://www.scbi.uma.es/seqtrimnext) available at the Plataforma Andaluza de Bioinformática (University of Málaga, Spain) using the specific NGS technology configuration parameters. This pre-processing removes low quality, ambiguous and low complexity stretches, linkers, adaptors, vector fragments, organelle DNA, polyA, polyT tails, and contaminated sequences while keeping the longest informative part of the read. SeqTrimNext also discarded sequences below 25 bp. Subsequently, the clean reads were aligned with the melon transcriptome sequence82 (http://melonomics.cragenomica.es/files/Transcriptome/) with Bowtie283 in BAM files, which were then sorted and indexed using SAMtools v1.484. The reads containing discordant alignments were rejected due to their ambiguous location. Uniquely localized reads were used to calculate the read number value for each gene. Differentially expressed genes (DEGs) between two samples were analyzed using DESeq. 285, one of the R packages. For each gene, a P-value < 0.05 and log2-fold change >1 or < −1 were considered the significance threshold. All the DEGs were annotated with orthologous genes in A. thaliana (https://plants.ensembl.org/Arabidopsis_thaliana/Info/Index) using Full-LengtherNext (http://www.scbi.uma.es/fulllengthernext) and they were hierarchichally clustered by heatmap representation.
Gene functional enrichment and MapMan analysis of DEGs
The DEGs annotated with orthologous genes in A. thaliana were used to identify the Gene Ontology functional categories using web-based GENECODIS software86,87,88 (http://genecodis.cnb.csic.es/). MapMan software v. 3.5.089 was used to provide a graphical overview of the metabolic and regulatory pathways for the detected DEGs.
Quantitative reverse transcription (qRT)-PCR
The gene expression for the selected plant genes was quantified by qRT-PCR. The total RNA was isolated from the melon plants as described above. First-strand cDNA synthesis was performed using Invitrogen Superscript III Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA) with random primers according to the manufacturer’s instructions. The qRT-PCR reactions were conducted in a CFX384 Touch Real-Time PCR detection system (Bio-Rad, Hercules, CA, USA) using SsoFast EvaGreen Supermix according to the manufacturer’s recommendations (Bio-Rad). Gene-specific primers (Table S5) were designed using Primer390. The C. melo β-actin gene (XM_008462689.2) was used as a reference gene91. The qRT-PCR conditions were as follows: enzyme activation step at 95 °C for 30 s, followed by 40 cycles of 5 s at 95 °C and 5 s at 55 °C. After the amplifications, the data were analysed using CFX Manager Software (Bio-Rad). Additionally, the amplicon sizes were confirmed by visualization on 1% agarose gels.
Analysis of photosynthetic activity
The photosynthetic performance in terms of PSII activity was studied by variable Chl-FI, using an Open FluorCam 700 MF (Photon System Instruments, Brno, Czechia). The applied protocol was the one described as number 1 by Pineda et al.92. Black and white images corresponding to transient chlorophyll fluorescence values were collected and used to calculate several fluorescence parameters, using FluorCam software version 5.0. The images obtained for these parameters (FV/FM: maximum quantum yield of PSII, ΦPSII: effective quantum yield of PSII and NPQ: non-photochemical quenching), were calculated according to Maxwell & Johnson93. A false colour scale was applied to the calculated images using the Fluorcam software.
The net photosynthesis rate (PN) was determined using an infrared gas analyser (IRGA LI-6400, Li-Cor Inc., Lincoln, NB, USA) on a 100 mm2 melon leaf area. For the measurements, radiation was supplied by a Qbeam solid state LED lighting system attached to the leaf cuvette (6400-02B LED, Li-Cor Inc.).
Pigment content quantification
The pigment content was determined spectroscopically according to Lichtenthaler and Buschmann94. For each treatment, three leaf disks of 78 mm2 each were isolated and immediately frozen under liquid nitrogen. Quantifications were perfomed on acetone 80% (v/v in water) leaf extracts prepared by the homogenization of the tissue in liquid nitrogen and a subsequent centrifugation at 16000 g to remove the insoluble material. The absorbance at 470, 647 and 663 nm was measured using 80% acetone as a blank. The chlorophyll a (Chl a), chlorophyll b (Chl b), total chlorophyll (Chl T), and total xanthophyll and carotenoid (Xanth + Car) contents were determined according to the equations described in Lichtenthaler and Buschmann94.
Multicolour fluorescence imaging (MCFI)
To further investigate the plant health status, and with special interest in plant secondary metabolism, the multicolour fluorescence emission from the adaxial side of melon leaves was recorded using an Open FluorCam FC 800-O (Photon Systems Instruments), according to Pérez-Bueno et al.95. Images of F680, F740 and particularly, BGF (F440 and F520 emitted by phenolic compounds from the phenylpropanoid pathway) from both uninfected and P. xanthii-infected melon leaves were captured by FluorCam software version 7.1.0.3, which also applied a false colour scale to the black and white images recorded here.
Data Avaliability
The complete RNA-seq sequencing data for all the samples were deposited in the NCBI Sequence Read Archive and are accessible under the accession number PRJNA434538.
References
Pérez-García, A. et al. The powdery mildew fungus Podosphaera fusca (synonym Podosphaera xanthii), a constant threat to cucurbits. Mol. Plant Pathol. 10, 153–160 (2009).
Zitter, T. A., Hopkins, D. L. & Tomas, C. E. Compendium of cucurbits diseases. St Paul, MN, USA: APS Press (1996).
Sitterly W.P. Powdery mildew of cucurbits. In: Spencer DM, ed. Powdery mildews. London, UK: Academic Press, 359–379 (1978).
del Pino, D. et al. Occurrence of races and pathotypes of cucurbit powdery mildew in southeastern Spain. Phytoparasitica 30, 459–466 (2002).
Fernández-Ortuño, D. et al. Occurrence and distribution of resistance to QoI fungicides in populations of Podosphaera fusca in south central Spain. Eur. J. Plant Pathol. 115, 215–222 (2006).
Bellón-Gómez, D., Vela-Corcía, D., Pérez-García, A. & Torés, J. A. Sensitivity of Podosphaera xanthii populations to anti-powdery-mildew fungicides in Spain. Pest Manag. Sci. 71, 1407–1413 (2015).
Spanu, P. D. Why do some fungi give up their freedom and become obligate dependants on their host? New Phytol. 171, 447–450 (2006).
Martínez-Cruz, J., Romero, D., Dávila, J. C. & Pérez-García, A. The Podosphaera xanthii haustorium, the fungal Trojan horse of cucurbit-powdery mildew interactions. Fungal Genet. Biol. 71, 21–31 (2014).
Vela-Corcía, D., Bautista, R., De Vicente, A., Spanu, P. D. & Pérez-García, A. De novo analysis of the epiphytic transcriptome of the cucurbit powdery mildew fungus Podosphaera xanthii and identification of candidate secreted effector proteins. PLoS One 11, 1–21 (2016).
Martínez-Cruz, J., Romero, D., de Vicente, A. & Pérez-García, A. Transformation of the cucurbit powdery mildew pathogen Podosphaera xanthii by Agrobacterium tumefaciens. New Phytol. 213, 1961–1973 (2017).
Martínez-Cruz, J. et al. The functional characterization of Podosphaera xanthii candidate effector genes reveals novel target functions for fungal pathogenicity. Mol. Plant-Microbe Interact. 31, 914–931 (2018).
Huang, S. et al. The genome of the cucumber, Cucumis sativus L. Nat. Genet. 41, 1275–1281 (2009).
Guo, S. et al. Characterization of transcriptome dynamics during watermelon fruit development: Sequencing, assembly, annotation and gene expression profiles. BMC Genomics 12, 454 (2011).
Ando, K., Carr, K. M. & Grumet, R. Transcriptome analyses of early cucumber fruit growth identifies distinct gene modules associated with phases of development. BMC Genomics 13, 518 (2012).
Blanca, J. et al. Transcriptome sequencing for SNP discovery across Cucumis melo. BMC Genomics 13, 280 (2012).
Garcia-Mas, J. et al. The genome of melon (Cucumis melo L.). Proc. Natl. Acad. Sci. 109, 11872–11877 (2012).
Xu, Y. et al. The draft genome of watermelon (Citrullus lanatus) and resequencing of 20 diverse accessions. Nat. Genet. 45, 51–58 (2013).
Montero-Pau, J. et al. De novo assembly of the zucchini genome reveals a whole-genome duplication associated with the origin of the Cucurbita genus. Plant Biotechnol. J. 16, 1161–1171 (2017).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
’t Hoen, P. A. C. et al. Deep sequencing-based expression analysis shows major advances in robustness, resolution and inter-lab portability over five microarray platforms. Nucleic Acids Res. 36, e141 (2008).
Marguerat, S. & Bähler, J. RNA-seq: From technology to biology. Cell. Mol. Life Sci. 67, 569–579 (2010).
Rezzonico, F., Rupp, O. & Fahrentrapp, J. Pathogen recognition in compatible plant-microbe interactions. Sci. Rep. 7, 1–12 (2017).
Guo, W. et al. Transcriptome profiling of pumpkin (Cucurbita moschata Duch.) leaves infected with powdery mildew. PLoS One 13, e0190175 (2018).
Weng, K. et al. Transcriptome of Erysiphe necator-infected Vitis pseudoreticulata leaves provides insight into grapevine resistance to powdery mildew. Hortic. Res. 1, 1–12 (2014).
Xin, M. et al. Transcriptome comparison of susceptible and resistant wheat in response to powdery mildew infection. Genomics, Proteomics Bioinforma. 10, 94–106 (2012).
Borges, A. F., Ferreira, R. B. & Monteiro, S. Transcriptomic changes following the compatible interaction Vitis vinifera-Erysiphe necator. Paving the way towards an enantioselective role in plant defence modulation. Plant Physiol. Biochem. 68, 71–80 (2013).
Fung, R. W. M. et al. Powdery mildew induces defense-oriented reprogramming of the transcriptome in a susceptible but not in a resistant grapevine. Plant Physiol. 146, 236–249 (2007).
Zhu, Q. et al. Comparative transcriptome profiling of genes and pathways related to resistance against powdery mildew in two contrasting melon genotypes. Sci Hortic 227, 169–180 (2018).
Barón, M., Pineda, M. & Pérez-Bueno, M. L. Picturing pathogen infection in plants. Zeitschrift fur Naturforsch. C 71, 355–368 (2016).
Mahlein, A. K. Plant disease detection by imaging sensors - parallels and specific demands for precision agriculture and plant phenotyping. Plant Dis. 100, 241–251 (2016).
Murchie, E. H. & Lawson, T. Chlorophyll fluorescence analysis: A guide to good practice and understanding some new applications. J. Exp. Bot. 64, 3983–3998 (2013).
Baker, N. R. Chlorophyll fluorescence: A probe of photosynthesis in vivo. Annu. Rev. Plant Biol. 59, 89–113 (2008).
Buschmann, C. & Lichtenthaler, H. K. Principles and characteristics of multi-colour fluorescence imaging of plants. J. Plant Physiol. 152, 297–314 (1998).
Cerovic, Z. G., Samson, G., Morales, F., Tremblay, N. & Moya, I. Ultraviolet-induced fluorescence for plant monitoring: present state and prospects. Agronomie 19, 543–578 (1999).
Rolfe, S. A. & Scholes, J. D. Chlorophyll fluorescence imaging of plant-pathogen interactions. Protoplasma 247, 163–175 (2010).
Gao, Y., Xu, H., Shen, Y. & Wang, J. Transcriptomic analysis of rice (Oryza sativa) endosperm using the RNA-Seq technique. Plant Mol. Biol. 81, 363–378 (2013).
Shi, H. et al. Comparative physiological, metabolomic, and transcriptomic analyses reveal mechanisms of improved abiotic stress resistance in bermudagrass [Cynodon dactylon (L). Pers.] by exogenous melatonin. J. Exp. Bot. 66, 681–694 (2015).
Beck, J. et al. Small one-helix proteins are essential for photosynthesis in Arabidopsis. Front. Plant Sci. 8, 1–14 (2017).
Passarini, F., Wientjes, E., Hienerwadel, R. & Croce, R. Molecular basis of light harvesting and photoprotection in CP24. J. Biol. Chem. 284, 29536–29546 (2009).
de Bianchi, S., Ballottari, M., Dall’Osto, L. & Bassi, R. Regulation of plant light harvesting by thermal dissipation of excess energy. Biochem. Soc. Trans. 38, 651–660 (2010).
Pérez-Bueno, M. L., Johnson, M. P., Zia, A., Ruban, A. V. & Horton, P. The Lhcb protein and xanthophyll composition of the light harvesting antenna controls the ΔpH-dependency of non-photochemical quenching in Arabidopsis thaliana. FEBS Lett 582, 1477–1482 (2008).
Sacharz, J., Giovagnetti, V., Ungerer, P., Mastroianni, G. & Ruban, A. V. The xanthophyll cycle affects reversible interactions between PsbS and light-harvesting complex II to control non-photochemical quenching. Nat. Plants. 3, 16225 (2017).
Munekage, Y. et al. Cyclic electron flow around photosystem I is essential for photosynthesis. Nature 429, 579–582 (2004).
Miyake, C. Alternative electron flows (water-water cycle and cyclic electron flow around PSI) in photosynthesis: Molecular mechanisms and physiological functions. Plant Cell Physiol. 51, 1951–1963 (2010).
Rumeau, D., Peltier, G. & Cournac, L. Chlororespiration and cyclic electron flow around PSI during photosynthesis and plant stress response. Plant, Cell Environ. 30, 1041–1051 (2007).
Munekage, Y. et al. PGR5 is involved in cyclic electron flow around photosystem I and is essential for photoprotection in Arabidopsis. Cell 110, 361–371 (2002).
Zhang, H. et al. Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genomics 15, 898 (2014).
Fu, Y. et al. Quantitative proteomics reveals the central changes of wheat in response to powdery mildew. J. Proteomics 130, 108–119 (2016).
Repka, V. Chlorophyll-deficient mutant in oak (Quercus petraea L.) displays an accelerated hypersensitive-like cell death and an enhanced resistance to powdery mildew disease. Photosynthetica 40, 183–193 (2002).
Berdugo, C. A., Zito, R., Paulus, S. & Mahlein, A. K. Fusion of sensor data for the detection and differentiation of plant diseases in cucumber. Plant Pathol 63, 1344–1356 (2014).
Swarbrick, P. J., Schulze-Lefert, P. & Scholes, J. D. Metabolic consequences of susceptibility and resistance (race-specific and broad-spectrum) in barley leaves challenged with powdery mildew. Plant, Cell Environ. 29, 1061–1076 (2006).
Bolton, M. D. Primary metabolism and plant defense - fuel for the fire. Mol. Plant-Microbe Interact. 22, 487–497 (2009).
Dixon, R. A. et al. The phenylpropanoid pathway and plant defence - a genomics perspective. Mol. Plant Pathol 3, 371–390 (2002).
Yang, F. et al. Engineering secondary cell wall deposition in plants. Plant Biotechnol. J. 11, 325–335 (2013).
Maeda, H. et al. RNAi suppression of arogenate dehydratase1 reveals that phenylalanine is synthesized predominantly via the arogenate pathway in petunia petals. Plant Cell 22, 832–849 (2010).
Maeda, H., Yoo, H. & Dudareva, N. Prephenate aminotransferase directs plant phenylalanine biosynthesis via arogenate. Nat. Chem. Biol. 7, 19–21 (2011).
Olsen, K. M., Lea, U. S., Slimestad, R., Verheul, M. & Lillo, C. Differential expression of four Arabidopsis PAL genes; PAL1 and PAL2 have functional specialization in abiotic environmental-triggered flavonoid synthesis. J. Plant Physiol. 165, 1491–1499 (2008).
Rohde, A. et al. Molecular phenotyping of the pal1 and pal2 mutants of Arabidopsis thaliana reveals far-reaching consequences on phenylpropanoid, amino acid, and carbohydrate metabolism. Plant Cell 16, 2749–2771 (2004).
Mauch-Mani, B. Production of salicylic acid precursors is a major function of phenylalanine ammonia-lyase in the resistance of Arabidopsis to Peronospora parasitica. Plant Cell 8, 203–212 (1996).
Ohl, S., Hedrick, S. A., Chory, J. & Lamb, C. J. Functional properties of a phenylalanine ammonia-lyase promoter. Plant Cell 2, 837–848 (1990).
Cochrane, F. C., Davin, L. B. & Lewis, N. G. The Arabidopsis phenylalanine ammonia lyase gene family: Kinetic characterization of the four PAL isoforms. Phytochemistry 65, 1557–1564 (2004).
Rookes, J. E. & Cahill, D. M. A PAL1 gene promoter – green fluorescent protein reporter system to analyse defence responses in live cells of Arabidopsis thaliana. Eur. J. Plant Pathol. 109, 83–94 (2003).
Raes, J. Genome-wide characterization of the lignification toolbox in. Arabidopsis. Plant Physiol. 133, 1051–1071 (2003).
Li, Y., Kim, J. I., Pysh, L. & Chapple, C. Four isoforms of Arabidopsis thaliana 4-coumarate: CoA ligase (4CL) have overlapping yet distinct roles in phenylpropanoid metabolism. Plant Physiol. 169, pp.00838.2015 (2015).
Tzin, V. & Galili, G. The biosynthetic pathways for shikimate and aromatic amino acids in Arabidopsis thaliana. Arab. B. 8, 1–18 (2010).
McCue, K. F. & Conn, E. E. Induction of 3-deoxy-D-arabino-heptulosonate-7-phosphate synthase activity by fungal elicitor in cultures of Petroselinum crispum. Proc. Natl. Acad. Sci. 86, 7374–7377 (1989).
Herrmann, K. M. & Weaver, L. M. The shikimate pathway. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 473–503 (1999).
Görlach, J. et al. Temporally distinct accumulation of transcripts encoding enzymes of the prechorismate pathway in elicitor-treated, cultured tomato cells. Proc. Natl. Acad. Sci. USA 92, 3166–3170 (1995).
Xu, L. et al. Lignin metabolism has a central role in the resistance of cotton to the wilt fungus Verticillium dahliae as revealed by RNA-Seq-dependent transcriptional analysis and histochemistry. J. Exp. Bot. 62, 5607–5621 (2011).
Jambagi, S. & Dunwell, J. M. Global transcriptome analysis and identification of differentially expressed genes after infection of Fragaria vesca with powdery mildew (Podosphaera aphanis). Transcriptomics 03, 1 (2015).
Pineda, M., Pérez-Bueno, M. L., Paredes, V. & Barón, M. Use of multicolour fluorescence imaging for diagnosis of bacterial and fungal infection on zucchini by implementing machine learning. Funct. Plant Biol 44, 563–572 (2017).
Lo Presti, L. et al. Fungal effectors and plant susceptibility. Annu. Rev. Plant Biol. 66, 513–545 (2015).
Schmidt, S. M. et al. Interaction of a Blumeria graminis f. sp. hordei effector candidate with a barley ARF-GAP suggests that host vesicle trafficking is a fungal pathogenicity target. Mol. Plant Pathol. 15, 535–549 (2014).
Weßling, R. et al. Convergent targeting of a common host protein-network by pathogen effectors from three kingdoms of life. Cell Host microbe 16, 364–375 (2015).
Pennington, H. G. et al. Interactions between the powdery mildew effector BEC1054 and barley proteins identify candidate host targets. J. Proteome Res. 15, 826–839 (2016).
Alkan, N., Friedlander, G., Ment, D., Prusky, D. & Fluhr, R. Simultaneous transcriptome analysis of Colletotrichum gloeosporioides and tomato fruit pathosystem reveals novel fungal pathogenicity and fruit defense strategies. New Phytol. 205, 801–815 (2015).
Meyer, F. E. et al. Dual RNA-Sequencing of Eucalyptus nitens during Phytophthora cinnamomi challenge reveals pathogen and host factors influencing compatibility. Front. Plant Sci. 7, 1–15 (2016).
Álvarez, B. & Torés, A. Cultivo in vitro de Sphaerotheca fuliginea (Schlecht. ex Fr.), efecto de diferentes fuentes de carbono sobre su desarrollo. Boletín Sanid. Veg. Plagas 23, 283–288 (1997).
Zhang, N. et al. Cell permeability and nuclear DNA staining by propidium iodide in basidiomycetous yeasts. Appl. Microbiol. Biotechnol. 102, 4183–4191 (2018).
Seoane, P. et al. AutoFlow, a versatile workflow engine illustrated by assembling an optimised de novo transcriptome for a non-model species, such as faba bean (Vicia faba). Curr. Bioinform. 11, 440–450 (2016).
Falgueras, J. et al. SeqTrim: A high-throughput pipeline for pre-processing any type of sequence read. BMC Bioinformatics 11, 38 (2010).
Blanca, J. M. et al. Melon transcriptome characterization: Simple sequence repeats and single nucleotide polymorphisms discovery for high throughput genotyping across the species. Plant Genome 4, 118–131 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinforma. Apl. note 25, 2078–2079 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Carmona-saez, P., Chagoyen, M. & Tirado, F. GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome Biol. 8, 1–8 (2007).
Nogales-Cadenas, R. et al. GeneCodis: Interpreting gene lists through enrichment analysis and integration of diverse biological information. Nucleic Acids Res. 37, 317–322 (2009).
Tabas-Madrid, D., Nogales-Cadenas, R. & Pascual-Montano, A. GeneCodis3: A non-redundant and modular enrichment analysis tool for functional genomics. Nucleic Acids Res. 40, 478–483 (2012).
Thimm, O. et al. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 37, 914–939 (2004).
Koressaar, T. & Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 23, 1289–1291 (2007).
Wu, T. & Cao, J. Molecular cloning and expression of a bush related CmV1 gene in tropical pumpkin. Mol. Biol. Rep. 37, 649–652 (2010).
Pineda, M., Soukupová, J., Matouš, K., Nedbal, L. & Barón, M. Conventional and combinatorial chlorophyll fluorescence imaging of tobamovirus-infected plants. Photosynthetica 46, 441–451 (2008).
Maxwell, K. & Johnson, G. N. Chlorophyll fluorescence - a practical guide. J. Exp. Bot. 51, 659–668 (2000).
Lichtenthaler, H. K. & Buschmann, C. Chlorophylls and carotenoids: measurement and characterization bu UV–vis spectroscopy. Protein Food Anal. Chem. 1, F4.3.1–F4.3.8 (2001).
Pérez-Bueno, M. L., Pineda, M., Díaz-Casado, E. & Barón, M. Spatial and temporal dynamics of primary and secondary metabolism in Phaseolus vulgaris challenged by Pseudomonas syringae. Physiol. Plant. 153, 161–174 (2015).
Acknowledgements
We thank Irene Linares (University of Malaga, Spain) for excellent technical assistance. We also thank Dr Josefa Gómez from “Edificio de Bioinnovación” (University of Malaga) for the excellent work done in the Illumina sequencing process. We also thankfully acknowledge the computer resources and the technical support provided by the “Platafoma Andaluza de Bioinformática” of the University of Málaga. This study was supported by grants from the “Agencia Estatal de Investigación (AEI)” (AGL2013-41939-R; AGL2016-76216-C2-1-R) of the former “Ministerio de Economía y Competitividad (MINECO)”, MINECO-CSIC and ERDF funds (RECUPERA 2020/20134R060) and a grant from CEICE-Junta de Andalucía (P12-AGR-0370), all of them co-financed by FEDER funds (European Union). A.P. was supported by a PhD fellowship (BES-2014-068602) from MINECO.
Author information
Authors and Affiliations
Contributions
A.P., M.P., M.P.-B., M.B. and A.P.-G. planned and designed the research; A.P., M.P., J.M.-C. and M.P.-B. performed the experiments; A.P., R.B., M.P., M.P.-B., M.B. and A.P.-G. analyzed the data and A.P., M.P., M.P.-B., M.B. and A.P.-G. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Polonio, Á., Pineda, M., Bautista, R. et al. RNA-seq analysis and fluorescence imaging of melon powdery mildew disease reveal an orchestrated reprogramming of host physiology. Sci Rep 9, 7978 (2019). https://doi.org/10.1038/s41598-019-44443-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44443-5
This article is cited by
-
Regulation of hormone pathways in wheat infested by Blumeria graminis f. sp. tritici
BMC Plant Biology (2023)
-
Lighting up plants with near-infrared fluorescence probes
Science China Chemistry (2023)
-
Sensor-based phenotyping of above-ground plant-pathogen interactions
Plant Methods (2022)
-
Comparative transcriptome analysis of melon (Cucumis melo L.) reveals candidate genes and pathways involved in powdery mildew resistance
Scientific Reports (2022)
-
Controllable synthesis and stabilization of Tamarix aphylla-mediated copper oxide nanoparticles for the management of Fusarium wilt on musk melon
3 Biotech (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.