Abstract
Recent studies have realized the link between gut microbiota and human health and diseases. The question of diet, environment or gene is the determining factor for dominant microbiota and microbiota profile has not been fully resolved, for these comparative studies have been performed on populations of different ethnicities and in short-term intervention studies. Here, the Southern Chinese populations are compared, specifically the children of Guangzhou City (China), Penang City (west coast Malaysia) and Kelantan City (east coast Malaysia). These Chinese people have similar ancestry thus it would allow us to delineate the effect of diet and ethnicity on gut microbiota composition. For comparison, the Penang and Kelantan Malay children were also included. The results revealed that differences in microbiota genera within an ethnicity in different cities was due to differences in food type. Sharing the similar diet but different ethnicity in a city or different cities and living environment showed similar gut microbiota. The major gut microbiota (more than 1% total Operational Taxonomy Units, OTUs) of the children population are largely determined by diet but not ethnicity, environment, and lifestyle. Elucidating the link between diet and microbiota would facilitate the development of strategies to improve human health at a younger age.
Similar content being viewed by others
Introduction
The gastrointestinal tract contains the greatest number of colonized microbes, mostly anaerobes, estimated to cover 70 percent of all the microbes in the human body1. Existing at such high concentrations in our body, it has been found that these gut microbes play important roles in modulating our health and diseases2,3,4. Thus, it is most relevant to understand factors determining gut microbiota profile.
Diet is an important factor determining the dominant gut microbiota1,5,6. A western diet consists of high fat, refined carbohydrates, and low fiber, resulting in European having high Bacteroides abundance and an underrepresented Prevotella genus7. Whereas the Africans have high levels of Prevotella, resulting from the high fiber content in their diet7. This is due to the Prevotella genus containing many species with high fiber degrading potential8. However, in this and other studies on long-term habitual dietary consumption in distant regions and close vicinity of different ethnicity9,10,11, the contribution of human genes, living environment and lifestyle in the microbiota profile could not be verified.
Many intervention studies had demonstrated that short-term diet changes12 and seasonal variation in dietary habit13 can affect the gut microbiota, but the magnitude of change did not result in a permanent alteration of enterotype. They also could not verify the factor(s) that determine the stability of the microbiome. A recent publication reported that individuals with distinct ancestral origins who shared a relatively common environment showed significant similarities in the compositions of the microbiomes14.
In our project, we focused on three Asian cities, Guangzhou China, Penang West coast Malaysia, and Kelantan East coast Malaysia. The two Malaysian cities are populated by Chinese of Southern China ancestry. All three cities also have the same tropical climate and are all surrounded by sea (food source). Whilst all three population consume Chinese food, Malaysia Chinese also consume the local diet (Malay foods) and local ingredients. For instance, the Peranakans of Penang consume Chinese-Malay fusion food.
This study aims to compare the Chinese (CN) population of Penang (PN), Kelantan (KL), and Guangzhou (GZ) based on their microbiota and diet. Having the same ancestors, it would be interesting to find out if their gut microbiota will be similar due to their race or will it segregate based on their different locations and dietary habits. The proportion of Chinese in the two Malaysian cities is different. The percentage of Malays (ML) and Chinese in Penang are 41.6% and 41.5% respectively, which is approximately equal. As for Kelantan, the percentage of Malays and Chinese in the region is 92.3% and 3.2% respectively, making the Chinese population a minority race in Kelantan15. The difference in the proportion of Chinese in the two cities will also result in the difference in the level of influence in the food culture. Hence, the food consumed by the Chinese in Kelantan would have a higher level of influence by the Malay culture than that in Penang. If the hypothesis that diet is the most important influence on gut microbiota, then based on the level of influence in food culture, the gut microbiota of the Penang Chinese should be more similar to that of the Guangzhou Chinese than that of the Kelantan Chinese. The highest difference in gut microbiota should be the highest between that of Guangzhou Chinese and Kelantan Chinese. Besides, Penang is an urban modern city, while Kelantan is rural and largely agricultural. Children were chosen in this study, for they consumed more uniform and traceable home cooked food, and at a critical age whereby the microbiota profile has established to that of the adult-like16 and may determine later life health status.
Other than comparing the Chinese population, the Penang and Kelantan Malay samples were also collected for comparison analysis. This is to investigate if the gut microbiota of the Penang and Kelantan Chinese would be similar to that of other Chinese populations from another country (China) or to the Malays who reside in the same city sharing many food types, living environment, and lifestyle.
Results
Effect of ethnicity on microbiota
To access the impact of ethnicity on gut microbiota, (1) two different ethnicities; namely, the Malay and Southern Chinese living in Penang and Kelantan, and (2) the Chinese children from Guangzhou, Penang, and Kelantan were compared.
Firstly, the unweighted Principle Coordinates Analysis (PCoA) plot of the Chinese population showed a separation between the Guangzhou and the Kelantan children (Fig. 1) at a p-value of 0.001 statistically and comparatively high R-value of 0.517 (Supplementary Table S1). There was also a separation between the Chinese children of Guangzhou and Penang (p = 0.001, R = 0.328), as well as between the Kelantan and Penang (p = 0.001, R = 0.380). Similarly, a separation for the Malay population between Kelantan and Penang children (p = 0.001, R = 0.249) was observed. The separation is significant by the statistical analysis of Analysis of Similarities (ANOSIM) and can also be clearly seen in the PCoA plots (Fig. 1a,c).
Unweighted and weighted UniFrac Principle Coordinates Analysis (PCoA) plots of microbiota among the same ethnicity. Two different races (Chinese and Malay) of children from Guangzhou, Kelantan, and Penang were compared against each other and visualized in the unweighted (left) and weighted (right) PCoA plots (a,c and b,d). The statistical analysis of ANOSIM was shown in Supplementary Table S1.
As for the weighted PCoA plots (Fig. 1b,d), there was a separation between the Guangzhou and Kelantan Chinese (p = 0.001, R = 0.260) but there was no separation between the Guangzhou and Penang Chinese, as well as between the Penang and Kelantan Chinese. On the other hand, there was a separation between the Kelantan and Penang Malay children at p-value of 0.001 and R-value of 0.235 (Table S1).
The results of the unweighted PCoA plots show that the types of bacteria present in the microbiota of the children population from the separate locations are different from each other, as evident from the separation for the pairwise comparisons (Table S1). On the other hand, the results of the weighted PCoA plots show that the relative abundance of some bacterial types in the microbiota is significantly different between the Guangzhou and Kelantan Chinese children, as well as between the Penang and Kelantan Malay children. However, there was no difference in the relative bacterial abundances for the other two pairs.
The children of different ethnicities living in the same location were compared to each other (Fig. 2). For the Kelantan Malay and Chinese population, there was a separation between them in the unweighted PCoA plot (p = 0.001, R = 0.164) but no separation for the weighted PCoA plot (Fig. 2a,b and Supplementary Table S2). As for the Penang Malay and Chinese population, there was also no separation between them for both the unweighted and weighted PCoA plots (Fig. 2c,d and Table S2).
Unweighted and weighted UniFrac PCoA plots of microbiota between two ethnicities in the same location. Chinese and Malay population in two countries (Kelantan and Penang) comparison were separately plotted for race comparison in unweighted PCoA (a,c) and weighted (b,d). The statistical analysis of ANOSIM was shown in Supplementary Table S2.
The results of the unweighted PCoA plots show that the types of bacteria present in the microbiota of Kelantan Chinese and Malay children are slightly different from each other as there was a separation in the unweighted PCoA plot. On the other hand, the microbiota of the Penang Chinese and Malay children are very similar as there was no separation between their microbiota. As for the weighted PCoA plots, the results show that the microbiota of the two different ethnicities in the same locations are very similar to each other as there was no separation for both the weighted PCoA plots.
In order to confirm the separation of sample clusters by the race factor, a Partitioning Around Medoids (PAM) Clustering approach and a Distance-based Redundancy Analysis (db-RDA) involving 175 identifiable genera were performed (Fig. 3a,b). It was interesting to observe that two optimized clusters (1 and 2) were found among each race in each city validated by a mean silhouette width of 0.356 (Fig. 3a). 68.16% and 31.84% of people belong to cluster 1 and 2 respectively. The total variation observed was 13.97% and the Monte Carlo test was significant (p = 0.002) at 499 permutations (Fig. 3b). Nevertheless, separation of Guangzhou Chinese from that of Penang and Kelantan Chinese could be clearly seen within the same cluster (Fig. 3, Supplementary Tables S3 and S4), particularly in Cluster 1. Moreover, the Chinese and Malay population of Penang and Kelantan within the same cluster are closer to each other and separation is not obvious as that of Guangzhou Chinese (Fig. 3, Supplementary Tables S3 and S4). Both methods suggested that the relative abundance of the bacteria reflects the distance between the cities and the types of bacteria determine the different clusters.
PCoA plot of PAM clustering using Jensen-Shannon distance (JSD) and db-RDA using Bray-Curtis distance comparing similarities between the two races from the three cities. Samples were revealed into two clusters clearly using PAM methods based on JSD distance matrix and bacterial genus abundances. (a) The two clusters were circled and within each cluster, the different races from the three cities were described in the different colors of centroid connectors. The Calinski-Harabasz (CH) index including the average silhouette (Si) coefficient was also shown. db-RDA plot was illustrated using Bray-Curtis distance and grouped into two clusters which estimated from PAM clustering method. (b) The percentage of variation explained for each axis is shown and the Monte Carlo test was significant at p = 0.002 and 499 permutations.
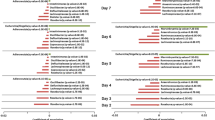
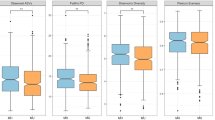
In comparing Operational Taxonomy Units (OTUs) above 1% of the total population of bacterial genera, total 17 bacterial genera were found significantly different between cluster 1 and 2 using the Mann-Whitney U test (Fig. 4 and Supplementary Table S5). Cluster 1 was dominated by 14 bacteria (Fig. 4a) and Bacteroides was the most abundant, whereas Cluster 2 was dominated by 3 bacteria and led by Prevotella (Fig. 4b). The ratio of both bacteria was found comparable within each cluster, but significantly different between two clusters across the different ethnicities and cities (Fig. 4c). Those major bacteria were found in the abundances order in a heatmap (d). Bacteroides, Prevotella, Fecalibacterium, and Bifidobacterium were the top 4 bacteria and the respective significant bacteria in pairs were found (Supplementary Tables S6–S8).
Relative abundance of major bacteria among the two ethnicities and the three different cities. Individual top 1% of the bacterial genera were plotted as box-plots. 20 bacteria were abundant in Cluster 1 (a) and 7 bacteria were rich in Cluster 2 (b). The ratio of Bacteroides and Prevotella abundances were compared (c). The symbol keys will be represented for (a–c) Overall major bacteria profile was shown as a heatmap (d). Relative abundances of OTU were illustrated as a spectrum of colors. The significant bacteria were marked with asterisks in a, b and c. Two-tailed ****p < 0.0001, ***p ≥ 0.0001–<0.001, **p ≥ 0.001–<0.01, *p < 0.05.
The predominant bacteria, with abundances above 1% of the total bacterial population in the two clusters are shown in Fig. 4d. Kruskal Wallis statistical test followed by Dunn’s post-hoc multiple comparison tests were performed (Supplementary Tables S6–S8). In a pairwise comparison of each of the bacterium between the two clusters, in the same city and race, the abundance of Bacteroides, Prevotella, Parabacteroides, and Streptococcus was significantly different and may have contributed to the differentiation of the clusters (Supplementary Table S6). Bacteroides, Parabacteroides, and Streptococcus were predominant members of Cluster 1, whereas, Prevotella was the most abundant in Cluster 2.
Within the same city, the abundant of the four predominant bacteria (Bacteroides, Prevotella, Oscillospira, Dorea, and Odoribacter) were found to be significantly different between the two ethnicities (Supplementary Table S7). However, between the same ethnicity in the three different cities, numerous bacteria (18 out of the 27 of those above 1% of bacterial population) showed differences in their abundance, which suggests that geographical location-associated factor(s) but not the ethnicity is the major determinant of the bacterial abundance (Supplementary Table S8).
Effect of diet on microbiota
As diet could be a geolocation-associated causative factor in influencing gut microbiota, the effects of the diet were investigated. The correlation between the portion of 15 food types (complex carbohydrates enriched foods, oily foods, vegetables, refined sugars enriched foods, dairy products, fruits, meats, protein enriched plants, eggs, seafood, caffeinated drinks, biscuits/pastries/cakes, yoghurt, preserved foods and curry) and bacterial abundances at the genus level is shown in Fig. 5.
Correlation between the quantity of the various food types consumed in the fraction and bacterial abundances at the genus level. The food types and bacterial abundances were described in different axes. The Spearman rho (r) values were reflected as a color gradient and positively and negatively correlations are found in red and blue spectrums. The significant correlation is shown as asterisks * two-tailed p < 0.05.
Out of the 27 bacteria compared, while there are many significant correlations, there are four genera that stood out with high rho values (≥±0.3) among the 15 food types. The four genera are Bacteroides, Fecalibacterium, Bifidobacterium, and Collinsella. (Supplementary Table S9). Interestingly, all correlated with oily foods, vegetables, refined sugars enriched foods, fruits, meats, and curry and caffeinated drinks. Looking into the correlation between bacteria and the foods, Bifidobacterium and Collinsella are positively correlated with refined sugars enriched foods, while the latter is also positively correlated with fruits and curry foods. However, Bacteroides, Faecalibacterium, Bifidobacterium, and Collinsella are negatively correlated with caffeinated drinks, curry, oily foods, and Southeast Asian vegetables respectively.
Discussion
The results show one main point, which is that the difference in microbiota composition is mainly due to geographical location, be it locational dietary habit, ethnicity or environmental influence. When compared the different ethnicities living in the same location, there was no significant difference in the microbiota composition within the same cluster according to both PAM Clustering and dbRDA analysis. On the other hand, when children of same ethnicity living in separate locations, in particular, Chinese in Guangzhou and the two Malaysian cities within Cluster 1 were compared, there was a significant difference in the microbiota composition and abundance according to dbRDA analysis. Thus, the microbiota profile is not to any significant extent determined by ethnicity among the children. In a previous study on dichorionic triplet sets have shown that genetics do affect gut microbiota differences, but only at one month of age and after which, environmental factors play a larger role17.
It is reasonable to assume that the subjects of separate locations are being exposed to different environments or lifestyles (cultures) and have access to different food types and hence will have different microbiota. A recent study done in mice has shown that mice in individually ventilated cages have different microbiota as compared to mice who live together in the same cage when they were all initially implanted with the same gut microbiota18. It was also shown that the microbiome similarity was found among the genetically unrelated individuals who shared household items14. This suggests that the environment can affect gut microbiota and those in the same environment will have similar gut microbiota types.
However, in our study among the children, microbiota profiles of an ethnicity (e.g. Chinese) resided in the same city (e.g. Guangzhou or Kelantan) are differentiated into the two clusters; while the same ethnicity in the same cluster (e.g. Chinese or Malay) living in distant cities (e.g. Penang and Kelantan) with different living environment and lifestyle (Penang being urban, while the Kelantan is rural) have similar microbiota profiles. This strongly suggested microbiota profile is to a little extent influenced by the living environment and further confirmed that ethnicity is not the major determining factor.
Despite different ethnicities, their food culture might have become more similar for the children in the same city (both Cluster 1 and 2). Malaysian cities are populated by Chinese and Malay, and thus most people consumed both Chinese and Malay types and fusion foods. Hence, it is no surprise that the microbiota of the Malaysian Chinese and Malay are very similar. On the other hand, the preferred food type is a personal choice and may explain the segregation in the microbiota between the clusters even within a city.
Among the three cities, higher consumption of refined sugar enriched foods, fruit, and curry led to a higher abundance of Bifidobacterium and Collinsella (r ≥±0.3); whereas higher consumption of curry led to a higher abundance of Faecalibacterium and Collinsella. On the other hand, oily food and vegetable consumption was negatively associated with Bifidobacterium and Collinsella respectively. Bacteroides, Faecalibacterium and Bifidobacterium are the predominant bacteria in Cluster 1. The relative proportion of refined sugar, fruit, curry, oily food and vegetable, and perhaps not a single food item in the diet may have determined Cluster 1 profile. High fat consumption has been reported to associate with Bacteroides7,19,20,21. Cluster 2 is dominated by Prevotella and was not found to be strongly associated, positively or negatively with any of the food types in this study. Dietary fiber has been reported to promote Prevotella abundnace7. Nevertheless, Prevotella is negatively associated with Bacteroides, as observed in this study and others7,19,20,21.
A study done on Malaysian children of three ethnic groups namely Chinese, Malay, and aboriginal Orang Asli showed that the Chinese and Malay children share greater bacterial genetic lineages then when compared to the Orang Asli22. While one of the reasons could be the difference in dietary habit due to socioeconomic status, another reason they suggested was that the Chinese and Malay children were exposed to similar food types22. This agrees with the results of this study that diet is the main determinant of gut microbiota.
Interestingly, the predominant microbiota detected among Southern Han Chinese (Guangzhou in this study), are different from those found among the central and Northern Han Chinese (Wuxi, Chengdu, Zhengzhou and Harbin), where the gut microbiota of the youngsters was largely populated by Phascolarctobacterium, Roseburia, Bacteroides, Blaustia, Faecalibacterium and Clostridium in that order23. The gut microbiota profile of Guangzhou children is different from that of the central and Northern Chinese, but closer to that of the Southeast Asian.
Another interesting observation is that the Guangzhou children have almost no Bifidobacterium in their microbiota, as it is inversely proportional to the abundance of Bacteroides. This is of concern as many of the Bifidobacterium species are known to have health benefits. While B. adolescentis have been used for manufacturing of functional dairy products, B. longum lowers the risk of diarrhea and allergies in infants24,25, and several other Bifidobacterium species show benefits to gastrointestinal health as well. Hence, it would be relevant to investigate the gut microbiota health of Guangzhou children to verify if the lack of Bifidobacterium indeed resulted in a higher occurrence of gut-related diseases.
Diet is a stronger influencing factor that can affect the changes in the relative abundance of gut microbes. From the correlation tables, it can be seen that the consumption of one food type can correlate with many genera and a single genus can also correlate with many food types. This shows that the food consumed is not exclusively used by one type of bacteria and the bacteria also do not exclusively feed on one type of food. The correlation between diet and microbiota is multidimensional, future metatranscriptomic and metabolomic studies would provide further understanding in their complex interrelation and may allow better modulation of microbiota through dietary intervention.
Conclusion
In summary, differences in the gut microbiota are largely due to dietary habit. Living environmental condition, and ethnicity are not determining factors for Southern Chinese and Southeast Asian children.
Methods
Ethics approval and consent to participate
The respective Institutional Review Board (IRB) at Guangzhou, Penang, Kelantan universities reviewed and approved the informed consent and the study protocols and other documents. The Joint Ethics Committee on Clinical Studies of School of Pharmaceutical Sciences, Universiti Sains Malaysia and Hospital Lam Wah Ee (JEC-SPS, USM & HLWE) (USM-HLWE/IEC/2014-0006) confirmed that the study was in compliance with the declaration of Helsinki and Malaysia Good Clinical Practice guidelines.
Study Design
A total of 201 local-born healthy children between 7 and 12 years old of both genders (male n = 110, female n = 91) were recruited at the respective schools from three cities: Guangzhou Southern Han Chinese (n = 81), Penang Southern Han Chinese (n = 21), Penang Malay (n = 21), Kelantan Southern Han Chinese (n = 45) and Kelantan Malay (n = 33). The participants who were healthy 7–12 years old children should not consume any pre/pro-biotics and antibiotics 2 weeks and 3 weeks prior respectively and have any recent illness that required to get medical attention last 3 months. All the participants had been informed and the consented documents were received from a parent upon the collection of samples.
Dietary intake questionnaires
Subjects were given the food frequency questionnaire (FFQ) requesting them to fill in the number of times they consume a specific type of food on a daily, weekly or monthly basis. We adopted the validated FFQ developed by the Singapore Health Promotion Board. The complete FFQs (Supplementary Text S12) and the raw data can be found. The questionnaire focuses on types, frequency, and the portion of foods and ingredients being consumed. We characterized the food items into 15 food types such as complex carbohydrates enriched foods, oily foods, vegetables, refined sugars enriched foods, dairy products, fruits, meats, protein enriched plants, eggs, seafood, caffeinated drinks, biscuits/pastries/cakes, yoghurt, preserved foods, and curry.
Collection of samples and DNA extraction
Approximately 10 g of stool was collected in 2 ml of RNAlater® (Ambion, Inc., Texas, USA) and 0.2 ml of fecal homogenate was further washed with 1x Phosphate-buffered saline (PBS). The DNA extraction was done using the TIANamp Stool DNA kit (TIANGEN Biotech, Co., Ltd., Beijing, China) according to the manufacturer’s protocol. Detail protocol (Supplementary Text 10) could be accessed.
Next-generation Sequencing
Once the concentration of the extracted double-stranded DNA had been measured using the Quant-iTTM Picrogreen® kit (Invitrogen, Inc., Carlsbad, USA), it was amplified with a Nextera® transposase sequences (details at S11 Text in supplementary information) and KAPA HiFiTM PCR kit (Roche life science, Inc., Indiana, USA). The PCR products were then purified by Agencourt® AMPure XP beads (Beckman Coulter, Inc., Fullerton, USA). In the index PCR, the reaction mixture was attached with the Illumina indexes (Illumina, Inc., San Diego, USA) using the KAPA HiFiTM PCR kit The DNA libraries produced were purified again using the Agencourt® AMPure XP beads. After the quantity of each library had been checked, it was normalised and pooled as Pooled Amplicon Library (PAL). Then, the PAL was then re-quantified with qPCR using the KAPA Library Quantification Kit (Roche life science, Inc., Indiana, USA).
The PAL and the internal control library (Illumina, Inc., San Diego, USA) were denatured with sodium hydroxide (NaOH) and diluted with a pre-chilled hybridization buffer (HT1) to the appropriate final concentration. The solution was sequenced on the Illumina® Miseq Desktop Sequencer (Illumina, Inc., San Diego, USA) The detailed protocol of the 16S rRNA sequencing preparation was described in Supplementary Text 11.
Data analysis
Bioinformatics analysis
The sequence data obtained were analyzed using QIIME (Quantitative Insights Into Microbial Ecology) version 1.9.126. Using the QIIME the forward and reverse reads of the same sample were first joined. The paired reads were demultiplexed and quality filtered at Q-score of 25. Chimeric sequences were filtered out and Non-chimera were selected by USEARCH v6.127. After which, using Greengenes v13.8 reference database, open-reference operational taxonomic units (OTUs) were picked out from the non-chimeric sequences at 97% similarity. The OTUs were summarized into taxonomy profiles of the bacteria in each sample. The bacterial genus data was provided. Beta diversity was calculated using the UniFrac distance matrices, generating the two- dimensional Principal Coordinate Analysis plots (PCoA). Weighted and unweighted UniFrac distance matrices were used to derive the beta diversity of the samples28 which data could be accessed.
The relative abundance of genus data was filtered because the unassigned fraction was not considered in the calculation of clustering, distance-based redundancy analysis (db-RDA), the taxonomy classification and their statistical analysis. In total 175 genera were identified. A constrained distance-based redundancy analysis (db-RDA) was performed with the software CANOCO 5 (Microcomputer Power, USA). A relative abundance of bacterial genus level OTU data was applied and a classified triplot was drawn using Bray Curtis distance according to the manufacturer’s instructions29.
Clustering
Applying the approach of Enterotyping30,31 using R version 3.5.1, the Jensen-Shannon distance (JSD) matrix was calculated based on the 175 relative abundances. The matrix was clustered by the Partitioning Around Medoids (PAM) algorithm using cluster package. Calinski-Harabasz (CH) index allowed estimation of the optimal number of clusters. The cluster number was validated using the individual and average silhouette coefficient (Si) and visualized by PCoA plots using the JSD matrix and adegraphics R package.
Statistical analysis
ANOSIM (Analysis of similarities) in QIIME was used to test for degree of separation between groups compared by using beta diversity Unifrac distance matrices as input. R-value and p-value shown in Table S1 and Table S2 were derived from Anosim test on the pairwise comparison based on unweighted and weighted Unifrac distance matrices. P-values < 0.01 are indicated with a * and shows significant separation.
For the bacteria at the genus level, only the top 1% highest abundance for each population were used for statistical analysis by GraphPad Prism 7 (GraphPad Software Inc., San Diego, USA). The non-parametric ANOVA; Kruskal-Wallis test, Mann-Whitney U test, and Multiple comparison post-hoc Dunn’s test were performed to check for significant differences in the distribution of bacteria between the groups compared.
As for the diet data, the types of food consumed were classified into 15 different groups according to the FFQs, based on the frequency and portion of the food types consumed per month for every subject, and expressed as a fraction of total foods consumed. In addition, the bivariate Spearman rank correlation was done for correlation between the abundance of bacteria present and consumption of a food type.
A Permutational multivariate analysis of variance (PERMANOVA) test was performed for the JSD and the Bray Curtis distance matrices using pairwiseAdonis (pairwise comparison of vegan R package) followed by Bonferroni post-hoc test. For db-RDA model, Monte Carlo permutation test with 499 random permutations was also performed and pseudo F values, pseudo F statistics, and p values were generated.
Data Availability
The dataset generated or analysed during this study are available in the KNB repository, doi:10.5063/F12B8W9B and the rest of the data and materials are provided as the Supplementary Information Files.
References
Sekirov, I., Russell, S., Antunes, L. & Finlay, B. Gut microbiota in health and disease. Physiological Reviews 90(3), 859–904 (2010).
Quigley, L. et al. The complex microbiota of raw milk. FEMS Microbiol Rev 37(5), 664–98 (2013).
Donia, M. S. & Fischbach, M. A. Human Microbiota. Small molecules from the human microbiota. Science 349(6246), 1254766 (2015).
Riedl, R. A. et al. The biomass and composition of the gut microbiota modify anaerobic metabolism. The FASEB Journal 31(1), 890.2 (2017).
Chung, W. S. F. et al. Modulation of the human gut microbiota by dietary fibers occurs at the species level. BMC Biology 14(1), 3 (2016).
Graf, D. et al. Contribution of diet to the composition of the human gut microbiota. Microbial Ecology in Health and Disease 26, 26164 (2015).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proceedings of the National Academy of Sciences of the United States of America 107(33), 14691–6 (2010).
Scott, K. P., Gratz, S. W., Sheridan, P. O., Flint, H. J. & Duncan, S. H. The influence of diet on the gut microbiota. Pharmacological Research 69(1), 52–60 (2013).
Chen, J. et al. Impact of demographics on human gut microbial diversity in a US Midwest population. PeerJ 10, 1–20 (2016).
Mai, V., Mccrary, Q. M., Sinha, R. & Glei, M. Associations between dietary habits and body mass index with gut microbiota composition and fecal water genotoxicity: an observational study in African American and Caucasian American volunteers. Nutrition Journal 8(49), 1–10 (2009).
Deschasaux, M. et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nature Medicine 24, 1526–1531 (2018).
Brüssow, H. M. and healthy aging: Observational and nutritional intervention studies. Microbial Biotechnology 6(4), 326–334 (2013).
Davenport, E. R. et al. Seasonal variation in human gut microbiome composition. Plos One 9(3), e90731 (2014).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Population 2015. Malaysia. 2015, https://web.archive.org/web/20160212125740/http://pmr.penerangan.gov.my/index.php/info-terkini/19463-unjuran-populasi-penduduk-2015.html. 11 May 2015. Accessed on 25 Nov 2017.
Hollister, E. B. et al. Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome 3, 36 (2015).
Murphy, K. et al. The gut microbiota composition in dichorionic triplet sets suggests a role for host genetic factors. Plos One 10(4), e0122561 (2015).
Lundberg, R., Bahl, M. I., Licht, T. R., Toft, M. F. & Hansen, A. K. Microbiota composition of simultaneously colonized mice housed under either a gnotobiotic isolator or individually ventilated cage regime. Scientific Reports 7, 42245 (2017).
Nakayama, J. et al. Diversity in gut bacterial community of school-age children in Asia. Scientific Reports 5, 8397 (2015).
Nakayama, J. et al. Impact of Westernized diet on gut microbiota in children on Leyte island. Frontiers in Microbiology 14(8), 197 (2017).
Mussa Kisuse, J. M. et al. Urban Diets Linked to Gut Microbiome and Metabolome Alterations in Children: A Comparative Cross-sectional Study in Thailand. Frontiers in Microbiology 9, 1345 (2018).
Chong, C. W. et al. Effect of ethnicity and socioeconomic variation to the gut microbiota composition among pre-adolescent in Malaysia. Scientific Reports 5, 13338 (2015).
Zhang, J. C. et al. A phylo-functional core of gut microbiota in healthy young chinese cohorts across lifestyles, geography and ethnicities. The ISME 9, 1979–1990 (2015).
Gomes, A. M. P. & Malcata, F. X. Bifidobacterium spp. and Lactobacillus acidophilus: Biological, biochemical, technological and therapeutical properties relevant for use as probiotics. Trends in Food Science and Technology 4-5, 139–157 (1999).
Szymański, H., Armańska, M., Kowalska-Duplaga, K. & Szajewska, H. Bifidobacterium longum PL03, Lactobacillus rhamnosus KL53A, and Lactobacillus plantarum PL02 in the prevention of antibiotic-associated diarrhoea in children: a randomized controlled pilot trial. Digestion 78(1), 13–17 (2008).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7(5), 335–6 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19), 2460–2461 (2010).
Lozupone, C. & Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Applied and Environmental Microbiology 71(12), 8228–8235 (2005).
Braak, C. J. F. t. & Šmilauer, P. Canoco reference manual and user’s guide: software for ordination (version 5.0). Microcomputer Power (2012).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Costea, P. I. et al. Enterotypes in the landscape of gut microbial community composition. Nature Microbiology 3(1), 8–16 (2018).
Acknowledgements
We would like to acknowledge all the individual people involved in samples collection, preparation and studies arrangement to Penang’s team: Lew L. Ching, Hor Y. Yan, Amy L.S. Yik, Ong J. Sin, Wong C. Boon, Yong C. Chung, Tan P. Lei, Kelantan’s team: Dr. Nur A.B. Muhammad and Madam N.A Abdullah and Guangzhou’s team: Li Zhou, Xiaolu Shi all the volunteers who contributed the samples from three countries. We are greatly thankful to Prof. Chong F.F. Mary and her team providing the food questionnaire including the photographs of foods portion size. The samples processing, sequencing, and data analysis were funded by the National University of Singapore grant R-571-000-241-592. The samples collection and subjects consenting were funded by the respective universities. For Kelantan’s samples, Fundamental Research Grant Scheme (203/PPSP/6171192), Research University Grant for the individual (1001/PPSP/812151) and Short-term grant (304/PPSP/6313278).
Author information
Authors and Affiliations
Contributions
W.W.T.K. prepared study design, was responsible for research coordination, samples processing, performed data analysis, generated figures and edited the manuscript. Y.Z. contributed to Guangzhou and Penang samples processing and drafted the manuscript. G.J.Y.G. contributed to Kelantan samples processing. M.S.W. co-ordinated Kelantan’s samples. M.T.L. performed on Penang study collaboration, coordination and edited the manuscript. Y.Y.L. contributed to study design, Kelantan study collaboration and helped co-ordination. H.C. supported the Guangzhou samples preparation and coordination. Y.K.L. led the study design, overall research collaboration, coordination, and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khine, W.W.T., Zhang, Y., Goie, G.J.Y. et al. Gut microbiome of pre-adolescent children of two ethnicities residing in three distant cities. Sci Rep 9, 7831 (2019). https://doi.org/10.1038/s41598-019-44369-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44369-y
This article is cited by
-
Characterization of gut microbiota in the Uyghur osteopenia population
Scientific Reports (2024)
-
Dysbiosis of gut microbiota due to diet, alcohol intake, body mass index, and gastrointestinal diseases in India
Applied Microbiology and Biotechnology (2023)
-
Early onset of immune-mediated diseases in minority ethnic groups in the UK
BMC Medicine (2022)
-
A single serving of mixed spices alters gut microflora composition: a dose–response randomised trial
Scientific Reports (2021)
-
Ethnicity influences the gut microbiota of individuals sharing a geographical location: a cross-sectional study from a middle-income country
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.