Abstract
Because of its multifactorial nature, predicting the presence of cancer using a single biomarker is difficult. We aimed to establish a novel machine-learning model for predicting hepatocellular carcinoma (HCC) using real-world data obtained during clinical practice. To establish a predictive model, we developed a machine-learning framework which developed optimized classifiers and their respective hyperparameter, depending on the nature of the data, using a grid-search method. We applied the current framework to 539 and 1043 patients with and without HCC to develop a predictive model for the diagnosis of HCC. Using the optimal hyperparameter, gradient boosting provided the highest predictive accuracy for the presence of HCC (87.34%) and produced an area under the curve (AUC) of 0.940. Using cut-offs of 200 ng/mL for AFP, 40 mAu/mL for DCP, and 15% for AFP-L3, the accuracies of AFP, DCP, and AFP-L3 for predicting HCC were 70.67% (AUC, 0.766), 74.91% (AUC, 0.644), and 71.05% (AUC, 0.683), respectively. A novel predictive model using a machine-learning approach reduced the misclassification rate by about half compared with a single tumor marker. The framework used in the current study can be applied to various kinds of data, thus potentially become a translational mechanism between academic research and clinical practice.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is one of the commonest cancers and is the leading cause of cancer-related deaths worldwide1. Despite recent improvements in therapeutic interventions2,3,4,5, HCC is still associated with a poor prognosis in patients with an advanced disease stage6. Previous studies have reported the beneficial influence of regular surveillance for HCC in high-risk populations to detect HCC at an early stage7,8,9.
Regarding the suggested guidelines for HCC surveillance, whether tumor markers should be included in a surveillance program, along with ultrasonography, remains controversial, since the sensitivity or specificity of alpha-fetoprotein (AFP), which has been the most widely used tumor marker for HCC, by itself is unsatisfactory10,11,12,13. Because of the multifactorial nature of HCC14, it is difficult to predict its presence using a single biomarker. Therefore, combining multiple biomarkers to improve diagnostic accuracy is important. To date, other tumor markers, such as des-gamma-carboxyprothrombin (DCP)15,16 and the Lens culinaris agglutinin-reactive fraction of AFP (AFP-L3)17,18, have been proposed to complement the diagnostic accuracy of AFP. In addition to information on tumor markers, data on biomarkers of liver inflammation (aspartate aminotransferase [AST] and alanine aminotransferase [ALT]), fibrosis (platelet count)19, liver function (total bilirubin [TB] and albumin)20, and the hepatitis virus status are commonly available in daily clinical practice. These biomarkers alter the pretest probability for a diagnosis of HCC using tumor marker and thus are useful for predicting the presence of HCC.
Machine learning is a multidisciplinary field combining computer science and mathematics and focused on implementing computer algorithms capable of maximizing predictive accuracy from static or dynamic data sources using analytic or probabilistic models21. Combining clinical data using this analytical tool can enable the development of a novel model for HCC prediction. The aims of the present study are (1) to develop a machine-learning framework to establish the most appropriate model depending on the applied data, and (2) to apply this framework to existing data from HCC patients to develop an appropriate model for HCC prediction.
Materials and Methods
Patients
From all the patients who visited the liver clinic at the University of Tokyo Hospital between January 1997 and May 2016, we extracted 4242 patients (1311 HCC patients and 2931 non-HCC patients) for whom information on the presence (or absence) of HCC was available and who had undergone laboratory testing on at least one occasion. All the patients in the HCC-positive group had been diagnosed as having HCC at the time of their first visit and had received initial treatment at our institution. Patients who subsequently developed HCC during the follow-up period for chronic liver disease were included in the HCC-negative group in the current study. Patients for whom information on AFP, AFP-L3, DCP, AST, ALT, platelet count, alkaline phosphatase (ALP), gamma-glutamyl transferase (GGT), albumin, TB, age, sex, height, body weight, hepatitis B surface (HBs) antigen, and hepatitis C virus (HCV) antibody status were available were selected. Finally, we included 539 HCC patients and 1043 non-HCC patients with the required information in the current analysis.
The current study was performed in accordance with the ethical guidelines of the Declaration of Helsinki. This research project was approved by the ethics committee of the University of Tokyo (approval number, 11474). Informed consent was obtained in the form of an opt-out on the website. Patients who rejected participation in our study were excluded. The study design was also included in a comprehensive protocol for retrospective studies and was approved by the ethics committee of the University of Tokyo (approval number, 2058).
Diagnosis of HCC
Hepatocellular carcinoma was diagnosed using dynamic computed tomography (CT) imaging, with hyper-attenuation during the arterial phase and washout during the late phase regarded as a definite sign of HCC22. When a definite diagnosis of HCC could not be made using CT, an ultrasound-guided tumor biopsy was performed and the pathological diagnosis was based on the Edmondson-Steiner criteria23.
Development of graphical user interface machine-learning framework
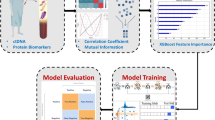
To establish a predictive model, we developed a graphical user interface machine-learning framework using R version 3.4.3 (http://www.r-project.org) and the Shiny and Caret packages. The model had two main components. The first component consisted of the establishment of an algorithm. Comma-separated values (CSV) dataset files with a labeled variable were dragged and dropped onto a dashboard, and the framework automatically implemented supervised learning and developed optimized classifiers and their respective hyperparameters, depending on the nature of the data, using a grid-search method (Fig. 1). We used a linear logistic regression model for the linear classification. The Akaike information criterion was used for variable selection in this model. Algorithms including support vector machines using an RBF kernel, gradient boosting, random forests, neural networks, and deep learning were also used for a non-linear classification model. The classifiers and their respective hyperparameters are shown in Table 1. For deep learning model, we defined two dense layers using ReLU activation function with drop-out ratio of 0.5, and then added output layer with the sigmoid activation function. We compiled the model using binary cross entropy as the loss function. An RMS prop optimizer was used as a hyperparameter for the optimization of deep neural network. The framework automatically selected the best classifier and its respective hyperparameter for the prediction model based on a grid search. The detailed process of searching for the optimal hyperparameters was shown in Supplementary Table 1. Algorithm optimization (e.g., a heatmap of predictive accuracy in a support vector machine [SVM]) or materials to compare the accuracies among the classifiers (confusion matrix or receiver operating characteristic curve) were automatically created.
The concept of graphical user interface machine learning framework. Comma-separated values (CSV) dataset files with a labeled variable were dragged and dropped onto a dashboard, and the framework automatically implemented supervised learning and developed optimized classifiers and their respective hyperparameters.
The second component consisted of the application of the developed model to a new dataset of interest. The CSV dataset of interest was dragged and dropped onto a dashboard, and the software applied the optimized classifiers and hyperparameters developed in the first component and outputted the probabilities of the respective labels.
Statistical analysis
Continuous variables were expressed as the medians with the first and third quartiles, while categorical variables were expressed as frequencies (%). Comparisons were performed using the Wilcoxon rank-sum or chi-square test for quantitative and categorical variables, respectively. We adopted the approaches used in the developed framework described above to predict the presence of HCC. To evaluate the accuracy of the model, we randomly split a total of 1582 patients into three parts: (i) the training set (80%), which was used to build the model, (ii) the development set, which was used for tuning the model parameters, and (iii) the test set, which was used to evaluate the performance of each classifier and assessed the predictive accuracy of the developed model. We then used a receiver-operation characteristics (ROC) curve analysis to assess the predictive accuracy of our classifier. The area under the curve (AUC) was evaluated as the ability to predict the presence of HCC. The variable importance for class discrimination in the predictive model was assessed using the mean decrease in the Gini impurity24.
Results
Patient characteristics
Finally, we extracted 1582 patients from our database (539 HCC and 1043 non-HCC patients). The dataset did not contain any missing data. The patient characteristics are shown in Table 2. The proportions of patients with a male sex, HCV antibody-positivity, and HBs antigen-negativity were significantly higher among the HCC patients, compared with the non-HCC patients. The serum levels of AFP, AFP-L3, DCP, AST, ALP, GGT, and TB, and the patient age were also significantly higher among the HCC patients, whereas the serum ALT level, platelet count, and albumin level were lower.
Predictive accuracy for HCC of each classifier
Table 3 shows the predictive accuracy for HCC presence for each classifier using the optimum hyperparameter that provided the highest predictive value in each procedure. We assessed the predictive accuracy of the developed model in the test set. The predictive accuracy for HCC presence provided by gradient boosting was 87.34%, which was the highest among all the classifiers in our framework. The optimal hyperparameters of this classifier for the data used in the present study were eta = 0.08, gamma = 0.02, max depth = 1, min_child_ weight = 1.5, nround = 300, subsample = 0.5, and colsample_bytree = 0.9. An ROC analysis showed that the AUC, sensitivity, and specificity for this optimal classifier were 0.940, 93.27%, and 75.93%, respectively (Fig. 2). Deep learning was not an optimal classifier for the current data.
Assessment of variable importance for class discrimination of the predictive model
We then investigated the variable importance of the optimal predictive model using the gradient boosting developed in the current study. Figure 3 shows the mean decrease in the Gini impurity of this model. Patient age followed by three tumor markers and albumin level were the most important variables for HCC prediction.
Predictive accuracy for HCC of single tumor markers
We also investigated the diagnostic accuracy of models using a single tumor marker. Using cut-offs of 200 ng/mL for AFP, 40 mAu/mL for DCP, and 15% for AFP-L325, the accuracies of AFP, DCP, and AFP-L3 for HCC presence were 70.67%, 74.91%, and 71.05%, respectively. We also plotted the ROC curves for the prediction of HCC for three tumor markers (Supplementary Fig. 1). The AUCs for the prediction of HCC for AFP, DCP, and AFP-L3 were 0.766, 0.644, and 0.683, respectively.
Discussion
In addition to tumor marker levels, biomarkers of liver inflammation, liver fibrosis, liver function, and the hepatitis virus status are commonly measured in daily clinical practice. These biomarkers can be used to predict the presence of HCC. Ideally, all clinically available information should be used for such predictions. In the current study, we developed a graphical user interface framework to establish the most appropriate model automatically depending on the applied data using a machine-learning approach and then assessed the accuracy of the model.
Model fitting is important for a successful predictive method. If the data is linearly separable, a linear model will fit the data26,27. However, if the data is linearly inseparable, a non-linear model will fit the data better. Therefore, classifiers should be selected depending on the nature of the data. Also, the learning parameters of each classifier should be tuned properly using a grid search method28,29 to obtain the ideal hyperparameters providing the highest predictive values. Using the optimal hyperparameter, gradient boosting (non-linear model) provided the highest accuracy (87.34%) for the data used in the current study. This model reduced the misclassification rate by about half, compared with a single tumor marker.
Personalization is one of the ultimate goals of modern medicine30. Predictive models provide a personalized assessment of the probability of a clinical event using patient-specific characteristics and have increasingly been incorporated into practice in the field of cancer medicine31,32,33. The framework developed in the current study can be used to identify optimal classifiers easily and can be applied to new datasets of interest containing various kinds of data, thus potentially becoming a translational mechanism between academic research and clinical practice.
Deep learning has enabled major breakthroughs in the processing of images, video, speech, and audio34. However, deep learning was not the optimal classifier in the current study. Deep learning requires a large polynomial sample size in terms of the dimensions of the input and an exponential sample in terms of the depth of the network to obtain ideal convergence boundaries35, which may be unrealistic requirements in clinical settings for ethical or methodologic reasons. Instead, identifying optimal classifiers and hyperparameters depending on the available data is important. The framework developed in the current study may help to provide optimal models.
The previous studies which compared the predictive performance of tabular data also showed the highest predictive performance of gradient boosting in the medical fields (e.g, urinary tract infections36, hip fractures37, sepsis38, or bioactive molecules39). Notably, Chiew et al. showed outstanding performance of gradient boosting compared to other machine learning algorithms for the risk prediction of suspected sepsis patients in the emergency department using relatively small number of sample38. Gradient boosting may be the best algorithm for the analysis of tabular data especially in the medical field where it is difficult to collect a large amount of data for ethical or methodologic reasons.
In the future, predictive models using machine learning approach may be implemented in electronic medical record system and may offer decision support to improve patient outcomes and reduce clinical diagnosis error in daily medical practice. The accuracy of diagnostic algorithm based on machine learning approach depends on the number of samples for training40. Larger quantities of multidimensional medical data will be stored in the future, potentially improve the accuracy of machine learning based classifier. Discrepancy of disease distribution between train and test samples is also an important factor for the performance of each classifier. The predictive model developed in the current study is based on the data of tertiary referral center requires. Therefore, further study with external validation in a community-and clinic based population is needed to assess the practical performance of the current model.
In conclusion, the framework developed in the current study provided a novel predictive model of HCC, producing an area under the curve of 0.943. This model reduced the misclassification rate by about half, compared with that for a single tumor marker. The current framework can be applied to various kinds of data, and thus could potentially become a translational mechanism between academic research and clinical practice.
Data Availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Parkin, D. M., Bray, F., Ferlay, J. & Pisani, P. Global cancer statistics, 2002. CA: a cancer journal for clinicians 55, 74–108 (2005).
Shiina, S. et al. Radiofrequency ablation for hepatocellular carcinoma: 10-year outcome and prognostic factors. The American journal of gastroenterology 107, 569–577; quiz 578, https://doi.org/10.1038/ajg.2011.425 (2012).
Kudo, M. et al. Survival Analysis over 28 Years of 173,378 Patients with Hepatocellular Carcinoma in Japan. Liver cancer 5, 190–197, https://doi.org/10.1159/000367775 (2016).
Kitai, S. et al. Survival Benefit of Locoregional Treatment for Hepatocellular Carcinoma with Advanced Liver Cirrhosis. Liver cancer 5, 175–189, https://doi.org/10.1159/000367765 (2016).
Llovet, J. M. & Bruix, J. Systematic review of randomized trials for unresectable hepatocellular carcinoma: Chemoembolization improves survival. Hepatology (Baltimore, Md.) 37, 429–442, https://doi.org/10.1053/jhep.2003.50047 (2003).
Trevisani, F., Cantarini, M. C., Wands, J. R. & Bernardi, M. Recent advances in the natural history of hepatocellular carcinoma. Carcinogenesis 29, 1299–1305, https://doi.org/10.1093/carcin/bgn113 (2008).
Bruix, J. & Sherman, M. Management of hepatocellular carcinoma: an update. Hepatology (Baltimore, Md.) 53, 1020–1022, https://doi.org/10.1002/hep.24199 (2011).
Omata, M. et al. Asian Pacific Association for the Study of the Liver consensus recommendations on hepatocellular carcinoma. Hepatology international 4, 439–474, https://doi.org/10.1007/s12072-010-9165-7 (2010).
EASL-EORTC clinical practice guidelines: management of hepatocellular carcinoma. Journal of hepatology 56, 908–943, https://doi.org/10.1016/j.jhep.2011.12.001 (2012).
Tandon, P. & Garcia-Tsao, G. Prognostic indicators in hepatocellular carcinoma: a systematic review of 72 studies. Liver international: official journal of the International Association for the Study of the Liver 29, 502–510, https://doi.org/10.1111/j.1478-3231.2008.01957.x (2009).
Singal, A. et al. Meta-analysis: surveillance with ultrasound for early-stage hepatocellular carcinoma in patients with cirrhosis. Alimentary pharmacology & therapeutics 30, 37–47, https://doi.org/10.1111/j.1365-2036.2009.04014.x (2009).
Lok, A. S. et al. Des-gamma-carboxy prothrombin and alpha-fetoprotein as biomarkers for the early detection of hepatocellular carcinoma. Gastroenterology 138, 493–502, https://doi.org/10.1053/j.gastro.2009.10.031 (2010).
Giannini, E. G. et al. Alpha-fetoprotein has no prognostic role in small hepatocellular carcinoma identified during surveillance in compensated cirrhosis. Hepatology (Baltimore, Md.) 56, 1371–1379, https://doi.org/10.1002/hep.25814 (2012).
Sanyal, A. J., Yoon, S. K. & Lencioni, R. The etiology of hepatocellular carcinoma and consequences for treatment. The oncologist 15(Suppl 4), 14–22, https://doi.org/10.1634/theoncologist.2010-S4-14 (2010).
Liebman, H. A. et al. Des-gamma-carboxy (abnormal) prothrombin as a serum marker of primary hepatocellular carcinoma. The New England journal of medicine 310, 1427–1431, https://doi.org/10.1056/nejm198405313102204 (1984).
Okuda, H., Obata, H., Nakanishi, T., Furukawa, R. & Hashimoto, E. Production of abnormal prothrombin (des-gamma-carboxy prothrombin) by hepatocellular carcinoma. A clinical and experimental study. Journal of hepatology 4, 357–363 (1987).
Aoyagi, Y. et al. Fucosylation of serum alpha-fetoprotein in patients with primary hepatocellular carcinoma. Biochimica et biophysica acta 830, 217–223 (1985).
Sato, Y. et al. Early recognition of hepatocellular carcinoma based on altered profiles of alpha-fetoprotein. The New England journal of medicine 328, 1802–1806, https://doi.org/10.1056/nejm199306243282502 (1993).
Poynard, T. & Bedossa, P. Age and platelet count: a simple index for predicting the presence of histological lesions in patients with antibodies to hepatitis C virus. METAVIR and CLINIVIR Cooperative Study Groups. Journal of viral hepatitis 4, 199–208 (1997).
Johnson, P. J. et al. Assessment of liver function in patients with hepatocellular carcinoma: a new evidence-based approach-the ALBI grade. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 33, 550–558, https://doi.org/10.1200/jco.2014.57.9151 (2015).
Wang, S. & Summers, R. M. Machine learning and radiology. Medical image analysis 16, 933–951, https://doi.org/10.1016/j.media.2012.02.005 (2012).
Torzilli, G. et al. Accurate preoperative evaluation of liver mass lesions without fine-needle biopsy. Hepatology (Baltimore. Md.) 30, 889–893, https://doi.org/10.1002/hep.510300411 (1999).
Edmondson, H. A. & Steiner, P. E. Primary carcinoma of the liver: a study of 100 cases among 48,900 necropsies. Cancer 7, 462–503 (1954).
Menze, B. H. et al. A comparison of random forest and its Gini importance with standard chemometric methods for the feature selection and classification of spectral data. BMC bioinformatics 10, 213 (2009).
Tateishi, R. et al. Diagnostic accuracy of tumor markers for hepatocellular carcinoma: a systematic review. Hepatology international 2, 17–30, https://doi.org/10.1007/s12072-007-9038-x (2008).
Wu, X., Zhu, X., Wu, G.-Q. & Ding, W. Data mining with big data. IEEE transactions on knowledge and data engineering 26, 97–107 (2014).
Hengl, S., Kreutz, C., Timmer, J. & Maiwald, T. Data-based identifiability analysis of non-linear dynamical models. Bioinformatics (Oxford, England) 23, 2612–2618, https://doi.org/10.1093/bioinformatics/btm382 (2007).
Djebbari, A., Liu, Z., Phan, S. & Famili, F. An ensemble machine learning approach to predict survival in breast cancer. International journal of computational biology and drug design 1, 275–294 (2008).
Friedrichs, F. & Igel, C. Evolutionary tuning of multiple SVM parameters. Neurocomputing 64, 107–117 (2005).
Blaha, M. J. & Blumenthal, R. S. Risk factors: new risk-assessment guidelines—more or less personalized? Nature Reviews Cardiology 11, 136 (2014).
Ganne-Carrie, N. et al. Nomogram for individualized prediction of hepatocellular carcinoma occurrence in hepatitis C virus cirrhosis (ANRS CO12 CirVir). Hepatology (Baltimore, Md.) 64, 1136–1147, https://doi.org/10.1002/hep.28702 (2016).
Yang, H. I. et al. Nomograms for risk of hepatocellular carcinoma in patients with chronic hepatitis B virus infection. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 28, 2437–2444, https://doi.org/10.1200/jco.2009.27.4456 (2010).
Graefen, M. et al. International validation of a preoperative nomogram for prostate cancer recurrence after radical prostatectomy. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 20, 3206–3212, https://doi.org/10.1200/jco.2002.12.019 (2002).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. nature 521, 436 (2015).
Zhang, C., Bengio, S., Hardt, M., Recht, B. & Vinyals, O. Understanding deep learning requires rethinking generalization. Proceedings of the International Conference of Learning Representations (ICLR) (2017).
Taylor, R. A., Moore, C. L., Cheung, K. H. & Brandt, C. Predicting urinary tract infections in the emergency department with machine learning. PloS one 13, e0194085, https://doi.org/10.1371/journal.pone.0194085 (2018).
Kruse, C., Eiken, P. & Vestergaard, P. Machine Learning Principles Can Improve Hip Fracture Prediction. Calcified tissue international 100, 348–360, https://doi.org/10.1007/s00223-017-0238-7 (2017).
Chiew, C. J. et al. Heart rate variability based machine learning models for risk prediction of suspected sepsis patients in the emergency department. Medicine 98, e14197, https://doi.org/10.1097/md.0000000000014197 (2019).
Babajide Mustapha, I. & Saeed, F. Bioactive Molecule Prediction Using Extreme Gradient Boosting. Molecules (Basel, Switzerland) 21, https://doi.org/10.3390/molecules21080983 (2016).
Zacksenhouse, M., Braun, S., Feldman, M. & Sidahmed, M. Toward helicopter gearbox diagnostics from a small number of examples. Mechanical Systems and Signal Processing 14, 523–543 (2000).
Acknowledgements
This research was supported by AMED under Grant Number JP17fk0210304, JP18fk0210040, and 19fk0210040h0002.
Author information
Authors and Affiliations
Contributions
The seven authors are justifiably credited with authorship, according to the authorship criteria. In detail: M.S. – conception, design, analysis and interpretation of data, drafting of the manuscript, final approval given; K.M.-analysis and interpretation of data, final approval given; S.K.-analysis and interpretation of data, final approval given; R.T. – analysis and interpretation of data, critical revision of manuscript, final approval given; S.S. – collection and assembly of data, final approval given; K.K. – critical revision of manuscript, final approval given; Y.Y.– critical revision of manuscript, final approval given.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sato, M., Morimoto, K., Kajihara, S. et al. Machine-learning Approach for the Development of a Novel Predictive Model for the Diagnosis of Hepatocellular Carcinoma. Sci Rep 9, 7704 (2019). https://doi.org/10.1038/s41598-019-44022-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44022-8
This article is cited by
-
Development of a transformer model for predicting the prognosis of patients with hepatocellular carcinoma after radiofrequency ablation
Hepatology International (2024)
-
A novel predictive model of microvascular invasion in hepatocellular carcinoma based on differential protein expression
BMC Gastroenterology (2023)
-
Diagnosis of autism spectrum disorder based on functional brain networks and machine learning
Scientific Reports (2023)
-
Deep learning–assisted LI-RADS grading and distinguishing hepatocellular carcinoma (HCC) from non-HCC based on multiphase CT: a two-center study
European Radiology (2023)
-
Hepatocellular carcinoma: molecular mechanism, targeted therapy, and biomarkers
Cancer and Metastasis Reviews (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.