Abstract
The leucine-rich repeat receptor-like protein kinase (LRR-RLK) plays an important role in plant development and disease defence. Although genome-wide studies of LRR-RLKs have been performed in several species, a comprehensive analysis, including evolutionary, structural and expressional analyses and their relationships to function, has not been carried out in the radish (Raphanus sativus L.). In this study, we identified 292 LRR-RLK genes in the R. sativus genome and classified them into 23 subgroups. The subgroups containing genes involved in defence were more likely to evolve from tandem duplication rather than whole genome triplication (WGT), had lower expression profiles and were expressed in fewer tissues than the subgroups related to development. Gene structures and conserved domains did not differ in the defence-related or development-related subgroups, but they were distinct in each subgroup. This study sheds light on the evolutionary and expressional relationships with the functions of R. sativus LRR-RLKs and provides an integrated framework for additional investigation into these functions.
Similar content being viewed by others
Introduction
Cell surface receptors play important roles in perceiving and processing external and internal signals that arrive at the cell surface in both plants and animals. Receptor-like protein kinases (RLKs) are one large family of cell surface receptors1. A typical RLK usually consists of three distinct domains: an extracellular N-terminal domain, a transmembrane domain, and a C-terminal intracellular kinase domain (KD)2. KDs, which contain 250 to 300 amino acid residues that fold into a three-dimensional catalytic core, are fairly conserved3,4. The extracellular domains are highly divergent, and they usually consist of different protein domains2,5. Based on the structures of the extracellular domains, RLKs can be further classified into 17 subfamilies6, including the LRR-RLK family and the lectin-RLK family. The LRR-RLK family, which contains leucine-rich-repeat (LRR) domains within the extracellular domain, is the largest subgroup in the RLK superfamily5. Based on the structures of the KD, LRR-RLKs have been further classified into approximately 23 subgroups in Arabidopsis thaliana5,7,8.
The biological functions of LRR-RLKs have been classified into two main categories: defence against pathogens and development9,10,11. For example, the SIF2, IOS1 and FRK genes, as members of subgroup I, are involved in defence signalling12,13. The subgroup XII-1 members FLS2 and EFR can mediate plant resistance against bacterial pathogens14,15 in A. thaliana as well as in rice Xa3/Xa26 and Xa21, which mediate race-specific resistance to Xanthomonas oryzae pv. oryzae16. In addition, NIK, which is contained in subgroup II17,18 and PEPR2, which is a member of subgroup XI, contribute to defence responses19. Alternatively, RUL1, which is contained in subgroup III, is involved in secondary growth20. Subgroup V includes the SRF gene, which is involved in cell wall biology, and the SCM gene, which is related to root hair specification21,22. In addition, subgroup XIIIa contains two FEI genes that regulate cell wall development via signalling pathways23, while subgroup XIIIb includes ERECTA and ERECTA-LIKE genes that are related to stomatal development and organ size regulation24. In addition, CLV125, BRI126 and HAESA27, which are distributed in subgroups II and XI, are involved in plant development. Several LRR-RLKs play roles in both development and defence because of the crosstalk between these two signalling pathways or the recognition of multiple ligands by the same receptor28. One example is BAK1, which is involved in developmental regulation via the interaction with BRI1 and is involved in defence via the interaction with FLS226.
Because of the complexity of plant development as well as the responses to environmental and complementary functions between different proteins, the function of very few LRR-RLKs has been studied, especially in non-model plants29. Therefore, using bioinformatic tools to systematically study the evolution, expression and potential functions of the differentiation of LRR-RLKs helps to improve our understanding of their functions in efficiently regulating networks during plant growth. Several studies have investigated the membership and evolution of the LRR-RLK gene family in plant species, including the soybean6, two Citrus species30, four Rosaceae genomes29 and various species11,31. The radish (Raphanus sativus L., 2n = 2 × = 18), which is a member of the Brassiceae tribe in the plant family Brassicaceae and a relative of Brassica rapa and Brassica oleracea, has a long cultivation history in the world and is an economically important crop grown worldwide32. However, no systematic analysis of this gene family has been carried out in R. sativus. Thus, a comprehensive investigation of the LRR-RLK genes in the R. sativus genome is needed.
In this study, we systematically identified the R. sativus LRR-RLK genes and classified them into 23 subgroups. A detailed analysis of the genome organization, sequence phylogeny, gene structure, conserved domains, evolutionary patterns, and expression profiling was carried out. Moreover, we investigated the evolutionary and expressional relationships in the function of these subgroups. Our results provide a framework for the further functional characterization of LRR-RLK genes in R. sativus.
Methods
Identification and classification of LRR-RLK genes
The A. thaliana genome and annotation data were downloaded from the TAIR10 database (http://www.arabidopsis.org/)33. The genome resources of R. sativus34, B. rapa35 and B. oleracea36 were downloaded from the BRAD database (http://brassicadb.org/brad/)37.
To identify LRR-RLK genes, a set of proteins from the draft genome of R. sativus was scanned using hmmsearch from the HMMER v3.1 suite38 using a Hidden Markov Model (HMM) corresponding to the Pfam database 31.039 pkinase domain model (PF00069) and pkinase_Tyr domain model (PF07714). The credible RLK proteins were generated with an E-value cutoff of 1 × 10−5 and coverage of the Pfam domain models of at least 50% from the raw screening proteins7,29.
To classify the RLKs identified into subfamilies, previously defined HMMs of different RLK subfamilies (https://github.com/lileiting/Plant_Pkinase_fam.hmm) were used to scan the RLK proteins7,29. The proteins were assigned to subgroups according to their best matched HMMs with an E-value cutoff of 1 × 10−5, which was referred to Fischer et al.11 and Shiu et al.9 The LRR-RLK genes of A. thaliana, B. rapa and B. oleracea were also identified using the same methods.
Localization of LRR-RLK genes in the R. sativus genome
To construct physical maps indicating the distribution of LRR-RLK genes, genome localization details for the R. sativus LRR-RLK genes were collected from the annotation information. The MG2C (http://mg2c.iask.in/mg2c_v2.0/) was used to visualize the LRR-RLK genes on nine chromosomes40.
The phylogenetic tree construction of LRR-RLKs
Multiple sequence alignment of complete amino acid sequences was performed using MUSCLE software41. The MEGA-X program42 was used to construct a Neighbour-joining (NJ) phylogenetic tree using the Jones-Taylor-Thornton (JTT) model with 500 bootstrap replicates. The uniform rates and homogeneous lineages were adopted, and the partial deletion with a site coverage cutoff of 70% was used for gaps/missing data treatment. The online tool iTol43 was used to clearly show the subfamily branches after the classification of all proteins.
Conserved motifs and gene structure analysis
The motifs for LRR-RLK genes were searched using MEME software44. The default number of motifs to be found was set to 15, with a motif window length of 6 to 100 bp. The global perspective of the motifs in each subgroup was presented as a heatmap using the R package pheatmap45. The presence of transmembrane domains and signal peptides was predicted using TMHMM v2.0 (http://www.cbs.dtu.dk/services/TMHMM/)46 and Signalp v4.147, respectively.
Tandem duplications and syntenic analysis of LRR-RLK genes
To detect the generated mechanism of LRR-RLK genes, we analysed the tandem duplications and syntenic regions of the R. sativus and A. thaliana LRR-RLK genes. Tandem genes in R. sativus and A. thaliana were defined as those genes that (1) were separated by ten or fewer genes, (2) were within 200 kb, and (3) were orthologous genes, which were identified by the BLASTP program48 with an E-value cutoff < =1e-20 and coverage > =60%8. We identified syntenic orthologues using SynOrths software49, which is based on both sequence similarity and the collinearity of the flanking genes.
Transcriptional profile analysis
For the expression profiling analysis of LRR-RLK genes, we used RNA-seq data from R. sativus tissues including flowers, siliques, leaves, stems, callus, and roots that were generated earlier and submitted to the NCBI database32,34. RNA-seq reads were aligned to the R. sativus genome using TopHat250 software. Following the alignments, we calculated transcripts abundance on the basis of fragments per kilobase of transcript per million mapped reads (FPKM) values using Cufflinks software51.
Results
Identification, classification and distribution of LRR-RLK genes in R. sativus
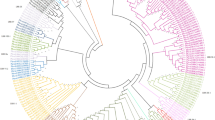
We searched all annotated genes in the R. sativus genome for putative RLKs and identified 1540 typical RLKs in R. sativus (Supplementary Table S1). Using the HMM search approach, we classified the RLKs into 119 subfamilies, which resulted in 292 genes that were classified into the LRR-RLK family (Fig. 1 and Supplementary Table S2). We used the same method identified 225, 305 and 225 LRR-RLK genes in A. thaliana, B. rapa and B. oleracea respectively (Supplementary Table S3). The R. sativus LRR-RLK family genes were further classified into 23 subgroups, and the numbers of the subgroup genes ranged from 1 in Xb-2 to 59 in III (Supplementary Table S4). Of these subgroups, III, I-1 and XI-1 were the three largest subgroups, with 59, 44 and 44 gene numbers, respectively. The other subgroups had no more than 30 members, and Xb-2 was the smallest subgroup, with only one gene. We found that R. sativus and A. thaliana had the same subgroups; however, the ratios of the LRR-RLK gene numbers between R. sativus and A. thaliana in each subgroup ranged from 0.92 to 3. Of these subgroups, only I-1 contained fewer gene members in R. sativus than that in A. thaliana.
Compared with B. rapa and B. oleracea, most subgroups contained similar numbers of genes. While, the genes number of subgroups I-1 and XII-1 in B. oleracea were far less than that of R. sativus and B. rapa. Besides, the gene number of Xb-1 in R. sativus is less than B. rapa and B. oleracea. Although R. sativus, B. rapa and B. oleracea have same ancestral species and shared similar evolutionary history, several subgroups of LRR-RLKs have varied greatly.
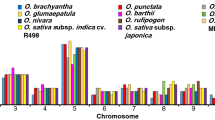
We mapped 288 R. sativus LRR-RLK genes on 9 chromosomes, while the remaining 5 genes were assigned to unassembled genomic sequence scaffolds (Fig. 2). The distribution of these genes appeared to be uneven, with the distribution ratio for each chromosome ranging from 4.78% (14 genes on chromosome 6) to 15.36% (45 members on chromosomes 5). This distribution pattern is similar to that of the LRR-LRK genes in other plant species1,6.
Phylogenetic analysis of LRR-RLKs
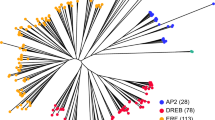
To study the evolutionary relationships of the LRR-RLK genes in R. sativus, the LRR-RLKs from R. sativus and A. thaliana were used to construct a phylogenetic tree (Fig. 3 and Supplementary Table S5). The phylogenetic tree showed that most LRR-RLK genes in the same subfamily were classified in combination with their A. thaliana homologues. The phylogenetic tree confirmed the classification of the subgroups of R. sativus LRR-RLKs.
Gene structure and conserved motif analysis of R. sativus LRR-RLKs
To gain insight into the structural diversity of the R. sativus LRR-RLK genes, we analysed their intron numbers (Supplementary Tables S2 and S6). The results showed that the numbers varied widely, ranging from 0 to 37. A total of 111 genes of R. sativus LRR-RLKs (37.88%) had only one intron, while 26 genes had no introns. Furthermore, a total of 86 genes had more than ten introns, and 70 genes had more than one and less than ten introns. Most of the R. sativus LRR-RLK genes in the same subgroups were conserved in terms of the intron number, while different subgroups had different patterns of intron numbers. For instance, all of the R. sativus LRR-RLK genes in subgroups III, VII, Xa and Xb contained zero, one, and two introns. In addition, all of the R. sativus LRR-RLK genes in subgroups V, VIII and XIIIb contained more than ten introns. In addition, the members of subgroups I-1, VI-2 and VIII-1 displayed a large variability in the number of introns. Most interestingly, the members of subgroup Xb-1 contained zero introns, while the members of subgroup XIIIb contained 22–25 introns. The exon/intron number indicated the conservation of gene structure within subgroups and the divergence among the different subgroups of R. sativus LRR-RLKs.
To further understand the potential functions and diversification of the LRR-RLK genes in R. sativus, a total of 15 conserved motifs were identified and numbered 1–15 using the MEME programme44. The results suggested that most members in one subgroup have the same motif compositions (Fig. 4 and Supplementary Table S2). However, in terms of types and numbers, most different subgroups have different motif compositions (Fig. 4 and Supplementary Table S2). For example, the MEME-6 motif was absent in subgroups VI-1, XIV, III, V, VII-1 and VII-2, and the MEME-13 motif was absent in the other 8 subgroups. In addition, the search for the possible transmembrane (TM) domain in all the R. sativus LRR-RLKs showed that a total of 257 members had at least one TM domain, while 36 members had no TM domain, and 73 members had at least two TM domains. In addition, signal peptides were also predicted with SignalP47, which showed that 210 members had signal peptides (Supplementary Table S2).
Tandem and syntenic analysis of LRR-RLK genes
Whole genome and tandem duplications provide critical sources of raw genetic material for genome complexity and evolutionary novelty52. In the R. sativus genome, 32 of 292 identified LRR-RLK genes, 10.92%, were demonstrated to be tandem duplications and were, distributed in 12 tandem arrays of 2–4 genes. Furthermore, 14.15% (32 out of 225) of A. thaliana LRR-RLK genes were located in tandem duplicated regions, which comprised 11 clusters in total (Supplementary Table S7). Tandem duplication clusters were distributed only among five subgroups in R. sativus. Remarkably, subgroup I-1 contained the most clusters and genes in both A. thaliana and R. sativus.
The ancestor of the diploid Brassica and Raphanus species has undergone a Brassiceae-lineage-specific whole-genome triplication (WGT) since its divergence from the A. thaliana lineage approximately 20 millions of years ago (MYA)35,53. The WGT was followed by diploidization that involved substantial genome reshuffling and gene losses54,55. We investigated the syntenic relationship of LRR-RLK genes between A. thaliana and R. sativus to trace the evolutionary history of LRR-RLK genes during the WGT. The genome-wide comparative analysis of the syntenic regions showed that 156 LRR-RLK genes in the A. thaliana genome had 225 corresponding genes in the R. sativus genome, indicating that 76.79% of the R. sativus LRR-RLK genes were retained after the WGT (Supplementary Table S8). Of the 156 A. thaliana LRR-RLK genes, 96 were shown to retain one copy, 55 retained two or three copies, and 3 genes had four copies in R. sativus. In addition, only two clusters of the 11 A. thaliana tandem duplication clusters had retained genes in R. sativus, and the others were lost during the WGT.
We also found that different subgroups had different percentages of tandem duplications and syntenic genes. For example, less than 50% of the genes in subgroups I-1, XII-1 and VIII-1 were retained during the WGT; however, in the other 12 subgroups, all the genes were retained. In addition, 40.91%, 40.00%, 33.33%, 25.00% and 17.39% of the genes in subgroups I-1, VIII-1, XI-2, VI-2 and XII-1 were present in tandem duplications, respectively (Supplementary Table S9).
Expression profiles of LRR-RLK genes in R. sativus
To further explore the expression patterns of LRR-RLK genes in R. sativus, the transcriptomes of six different tissues (i.e., flowers, siliques, leaves, stem, callus, and roots) were collected from previous reports32,34 (Supplementary Table S2). In general, we found that 20.89% (61 out of 292 genes) genes were not expressed in any tissues with FPKM >1. These unexpressed genes were found in 11 subgroups (I-1, III, IX, V, VI-2, VIII-1, Xa, Xb-1, XI-1, XI-2 and XII-1) (Supplementary Fig. S1). Approximately half of the genes in several subgroups, such as I-1 and XII-1, were not expressed or were only expressed in a few tissues, and the expression profiles were very low in these subgroups (Fig. 5 and Supplementary Table S10).
Discussion
LRR-RLKs, which are one of the largest gene families in plants, are primarily involved in plant growth and immunity in plants56. In this study, we identified 292 LRR-RLK genes in the genome of R. sativus and classified them into 23 subgroups. To confirm the accuracy of the identification and classification results, we used the same method to identify the LRR-RLK genes in A. thaliana as a reference. The LRR-RLK genes and their subgroups in A. thaliana identified in this study were identical with those of previous reports7,29, which indicates that our results were reliable. Compared with the ratio of the genome gene numbers of the two species (43,240/25,498 = 1.70)34,57, that of the LRR-RLK genes in R. sativus have not expanded from a global perspective, which is a result similar to that of its close species B. rapa11.
We analysed the retention/expansion of each subgroup and found that the ratios varied in different subgroups, which indicates that the evolutionary histories of the subgroups are different. The syntenic and tandem duplication analyses suggested that there are at least two different evolutionary mechanisms of these subgroups. One mechanism is that the LRR-RLK genes were tripled in the WGT and quickly lost in the following fractionation process, and the LRR-RLK genes were expanded via tandem duplication, such as in subgroups I-1, XII-1 and VIII-1. Another mechanism is that the LRR-RLK genes were also tripled during the WGT but were slowly lost and were retained as syntenic genes, such as in subgroups III, V and X. For subgroups I-1 and XII-1, most of the genes involved in the responses to biotic stress were currently reported in A. thaliana11. In addition, subgroups III and V usually contain genes involved in development58,59. These observations confirmed that the RLK genes involved in the stress responses were mostly duplicated by tandem duplication9, although few LRR-RLK genes have been functionally characterized. The recent tandem duplication generated functional redundancy in the LRR-RLK genes that can allow positive selection or relaxed negative selection of their function9, providing a reservoir to generate new disease resistant specificities against the fast-evolving effector genes of plant pathogens. Besides, we have found that the gene numbers of I-1, XII-1 varied greatly among the closely related species B. rapa, B. oleracea and R. sativus, which indicated that the subgroups involved in the responses to biotic stress employed different evolutionary process in different species.
Plant resistance genes need to be blocked in the absence of pathogens to avoid auto-immunity, which is detrimental to plant growth and development60. Notably, the genes in the defence-related subgroups, such as I-1 and XII-1, had a low expression quantity and were expressed in only a few tissues, which confirms that these subgroups are involved in the response to disease. However, no stress-treated RNA-Seq data were collected in this study. Contrasting results were observed in the developmentally related subgroups III and V, which had high expression profiles, and most of the genes were expressed in more than two tissues. Previous studies have demonstrated that introns always play an important role in cellular and developmental processes via alternate splicing or gene expression regulation61, and the protein domains or motifs decide the biological function of the genes. Thus, we found that each subgroup of R. sativus LRR-RLKs can differ from the others, either in the gene structure or conserved domains. However, the gene structure and conserved domains do not distinguish between defence-related or developmental-related subgroups.
Previous studies have illustrated the phylogenetic relationship, expression profiling, orthologous relationships and other characteristics of LRR-RLKs in several plants6,29,59, and have systematic analyzed the LRR-RLKs in plants and demonstrated that positive selection and gene expansion should be two major factors that result in differences between defence-related and development-related subgroups10,11. In this study, we supplemented the different evolutionary mechanisms and expressional profiles between the two groups. In total, our structural, evolutionary, and expressional analysis suggested the divergence of R. sativus subfamilies and their relationships to function. The results of this study provide insights into the evolution, expression and function of the R. sativus LRR-RLK gene family and provide a framework for the further functional investigation of these genes.
References
Yang, Y. et al. Genome-wide analysis of lectin receptor-like kinases in Populus. BMC Genomics 17, 699 (2016).
Shiu, S. H. & Bleecker, A. B. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl. Acad. Sci. USA 98, 10763–10768 (2001).
Hanks, S. K. & Hunter, T. Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J. 9, 576–596 (1995).
Liu, P. L. et al. Duplication and diversification of lectin receptor-like kinases (LecRLK) genes in soybean. Sci Rep. 8, 5861 (2018).
Shiu, S. H. & Bleecker, A. B. Plant receptor-like kinase gene family: diversity, function, and signaling. Sci. Signal 2001 (2001).
Zhou, F., Guo, Y. & Qiu, L. Genome-wide identification and evolutionary analysis of leucine-rich repeat receptor-like protein kinase genes in soybean. BMC Plant Biol. 16, 58 (2016).
Lehtishiu, M. D. & Shiu, S. H. Diversity, classification and function of the plant protein kinase superfamily. Philos. T. R. Soc. B 367, 2619–2639 (2012).
Lehtishiu, M. D., Zou, C., Hanada, K. & Shiu, S. H. Evolutionary history and stress regulation of plant receptor-like kinase/pelle genes. Plant Physiol. 150, 12–26, https://doi.org/10.1104/pp.108.134353 (2009).
Shiu, S. H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004).
Tang, P. et al. Disease resistance signature of the leucine-rich repeat receptor-like kinase genes in four plant species. Plant Sci. 179, 399–406 (2010).
Fischer, I., Dievart, A., Droc, G., Dufayard, J. F. & Chantret, N. Evolutionary dynamics of the leucine-rich repeat receptor-like kinase (LRR-RLK) subfamily in angiosperms. Plant Physiol. 170, 1595–1610 (2016).
Yuan, N. et al. Stress Induced Factor 2, a leucine-rich repeat kinase regulates basal plant pathogen defense. Plant Physiol. 176, 3062–3080 (2018).
Porter, K., Shimono, M., Tian, M. & Day, B. Arabidopsis Actin-Depolymerizing Factor-4 links pathogen perception, defense activation and transcription to cytoskeletal dynamics. PLoS Path. 8 (2012).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Gomezgomez, L. & Boller, T. FLS2: An LRR receptor–like kinase involved in the perception of the bacterial Elicitor Flagellin in. Arabidopsis. Mol. Cell 5, 1003–1011 (2000).
Zhao, J., Fu, J., Li, X., Xu, C. & Wang, S. Dissection of the factors affecting development-controlled and race-specific disease resistance conferred by leucine-rich repeat receptor kinase-type R genes in rice. Theor. Appl. Genet. 119, 231–239 (2009).
Fontes, E. P. B., Santos, A. A., Luz, D. F., Waclawovsky, A. J. & Chory, J. The geminivirus nuclear shuttle protein is a virulence factor that suppresses transmembrane receptor kinase activity. Genes Dev. 18, 2545–2556 (2004).
Santos, A. A., Lopes, K. V. G., Apfata, J. A. C. & Fontes, E. P. B. NSP-interacting kinase, NIK: a transducer of plant defence signalling. J. Exp. Bot. 61, 3839–3845 (2010).
Yamaguchi, Y., Huffaker, A., Bryan, A. C., Tax, F. E. & Ryan, C. A. PEPR2 is a second receptor for the pep1 and pep2 peptides and contributes to defense responses in Arabidopsis. Plant Cell 22, 508–522 (2010).
Agusti, J., Lichtenberger, R., Schwarz, M., Nehlin, L. & Greb, T. Characterization of transcriptome remodeling during cambium formation identifies MOL1 and RUL1 as opposing regulators of secondary growth. PLoS Genet. 7 (2011).
Dolan, L. Positional information and mobile transcriptional regulators determine cell pattern in the Arabidopsis root epidermis. J. Exp. Bot. 57, 51–54 (2006).
Eyuboglu, B. et al. Molecular characterisation of the strubbelig-receptor family of genes encoding putative leucine-rich repeat receptor-like kinases in Arabidopsis thaliana. BMC Plant Biol. 7, 16–16 (2007).
Xu, S., Rahman, A., Baskin, T. I. & Kieber, J. J. Two leucine-rich repeat receptor kinases mediate signaling, linking cell wall biosynthesis and ACC synthase in Arabidopsis. Plant Cell 20, 3065–3079 (2008).
Uchida, N., Shimada, M. & Tasaka, M. ERECTA-family receptor kinases regulate stem cell homeostasis via buffering its cytokinin responsiveness in the shoot apical meristem. Plant Cell Physiol. 54, 343–351 (2013).
Sijacic, P. & Liu, Z. Novel insights from live-imaging in shoot meristem development. J. Integr. Plant Biol. 52, 393–399 (2010).
Chinchilla, D., Shan, L., He, P., De Vries, S. C. & Kemmerling, B. One for all: the receptor-associated kinase BAK1. Trends Plant Sci. 14, 535–541 (2009).
Jinn, T., Stone, J. M. & Walker, J. C. HAESA, an Arabidopsis leucine-rich repeat receptor kinase, controls floral organ abscission. Genes Dev. 14, 108–117 (2000).
Afzal, A. J., Wood, A. J. & Lightfoot, D. A. Plant receptor-like serine threonine kinases: roles in signaling and plant defense. Mol. Plant-Microbe Interact. 21, 507–517 (2008).
Sun, J., Li, L., Wang, P., Zhang, S. & Wu, J. Genome-wide characterization, evolution, and expression analysis of the leucine-rich repeat receptor-like protein kinase (LRR-RLK) gene family in Rosaceae genomes. BMC Genomics 18, 763 (2017).
Magalhaes, D. M. et al. LRR-RLK family from two Citrus species: genome-wide identification and evolutionary aspects. BMC Genomics 17, 623 (2016).
Dievart, A. et al. Leucine-Rich repeat receptor kinases are sporadically distributed in eukaryotic genomes. BMC Evol. Biol. 11, 367–367 (2011).
Wang, J. et al. Insights into the species-specific metabolic engineering of glucosinolates in radish (Raphanus sativus L.) based on comparative genomic analysis. Sci Rep. 7 (2017).
Huala, E. et al. The Arabidopsis Information Resource (TAIR): a comprehensive database and web-based information retrieval, analysis, and visualization system for a model plant. Nucleic Acids Res. 29, 102–105 (2001).
Zhang, X. et al. A de novo genome of a Chinese radish cultivar. Hortic. Plant J. 1, 155–164 (2015).
Wang, X. et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43, 1035–1039 (2011).
Liu, S. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun 5, 3930–3930 (2014).
Cheng, F. et al. BRAD, the genetics and genomics database for Brassica plants. BMC Plant Biol. 11, 136–136 (2011).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, 29–37 (2011).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, 222–230 (2014).
Jiangtao, C. et al. MapGene2Chrom, a tool to draw gene physical map based on Perl and SVG languages. Hereditas 37, 91 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol (2018).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44 (2016).
Bailey, T. L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, 202–208 (2009).
Kolde, R. pheatmap: Pretty Heatmaps (2015).
Krogh, A., Larsson, B., Von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Petersen, T. N., Brunak, S., Von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Feng, C., Jian, W., Lu, F. & Wang, X. Syntenic gene analysis between Brassica rapa and other Brassicaceae species. Front. Plant Sci. 3, 198 (2012).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and cufflinks. Nat Protoc. 7, 562–78 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Yu, J. et al. Genome evolutionary dynamics followed by diversifying selection explains the complexity of the Sesamum indicum genome. BMC Genomics 18, 257 (2017).
Jeong, Y. et al. Elucidating the triplicated ancestral genome structure of radish based on chromosome-level comparison with the Brassica genomes. Theor. Appl. Genet. 129, 1357–1372 (2016).
Cheng, F. et al. Deciphering the diploid ancestral genome of the mesohexaploid Brassica rapa. Plant Cell 25, 1541–1554 (2013).
Mun, J. et al. Genome-wide comparative analysis of the Brassica rapa gene space reveals genome shrinkage and differential loss of duplicated genes after whole genome triplication. Genome Biol. 10, 1–18 (2009).
Wu, Y. & Zhou, J. Receptor-like kinases in plant innate immunity. J. Integr. Plant Biol. 55, 1271–1286 (2013).
Initiative, A. G. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Liu, P., Du, L., Huang, Y., Gao, S. & Yu, M. Origin and diversification of leucine-rich repeat receptor-like protein kinase (LRR-RLK) genes in plants. BMC Evol. Biol. 17, 47 (2017).
Zhu, H. et al. Genome-wide identification and characterization of the LRR-RLK gene family in two vernicia species. Comp. Funct. Genomics 2015, 823427–823427 (2015).
Marone, D., Russo, M. A., Laido, G., De Leonardis, A. M. & Mastrangelo, A. M. Plant nucleotide binding site-leucine-rich repeat (NBS-LRR) genes: active guardians in host defense responses. Int. J. Mol. Sci. 14, 7302–7326 (2013).
Roy, S. W. & Gilbert, W. The evolution of spliceosomal introns: patterns, puzzles and progress. Nat. Rev. Genet. 7, 211–221 (2006).
Acknowledgements
This work was supported by grants from the National technology system of commodity vegetable industry-radish (CARS-23-A-14), and New variety breeding project of the Major Science and Technology Projects of Zhejiang (2016C02051-2).
Author information
Authors and Affiliations
Contributions
C.B. conceived the research and designed the experiments. J.W. conducted the experiments, analysed the data and wrote the paper. T.H., W.W., H.H. and Q.W. participated in the experiment and involved in manuscript preparation. All of the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41598_2019_43516_MOESM2_ESM.docx
Investigation of evolutionary and expressional relationships in the function of the leucine-rich repeat receptor-like protein kinase gene family (LRR-RLK) in the radish (Raphanus sativus L.)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Hu, T., Wang, W. et al. Investigation of evolutionary and expressional relationships in the function of the leucine-rich repeat receptor-like protein kinase gene family (LRR-RLK) in the radish (Raphanus sativus L.). Sci Rep 9, 6937 (2019). https://doi.org/10.1038/s41598-019-43516-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-43516-9
This article is cited by
-
Genomic investigation of duplication, functional conservation, and divergence in the LRR-RLK Family of Saccharum
BMC Genomics (2024)
-
Massive expansion of P-selectin genes in two Venerida species, Sinonovacula constricta and Mercenaria mercenaria: evidence from comparative genomics of Bivalvia
BMC Genomics (2022)
-
Identification and characterization of plant resistance genes (R genes) in sorghum and their involvement in plant defense against aphids
Plant Growth Regulation (2022)
-
OsSRK1, a lectin receptor-like kinase, controls plant height by mediating internode elongation in Oryza sativa L
Molecular Breeding (2022)
-
Expression, purification, and phylogenetic analysis of MDIS1-INTERACTING RECEPTOR-LIKE KINASE1 (MIK1)
The Protein Journal (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.