Abstract
Citrate synthase (CS, EC 2.3.3.1) catalyses the initial reaction of the tricarboxylic acid (TCA) cycle. Although CSs from heterotrophic bacteria have been extensively studied, cyanobacterial CSs are not well-understood. Cyanobacteria can produce various metabolites from carbon dioxide. Synechocystis sp. PCC 6803 (Synechocystis 6803) is a cyanobacterium used to synthesize metabolites through metabolic engineering techniques. The production of acetyl-CoA-derived metabolites in Synechocystis 6803 has been widely examined. However, the biochemical mechanisms of reactions involving acetyl-CoA in Synechocystis 6803 are poorly understood. We characterised the CS from Synechocystis 6803 (SyCS) and compared its characteristics with other bacterial CSs. SyCS catalysed only the generation of citrate, and did not catalyse the cleavage of citrate. It is suggested that SyCS is not related to the reductive TCA cycle. The substrate affinity and turnover number of SyCS were lower than those of CSs from heterotrophic bacteria. SyCS was activated by MgCl2 and CaCl2, which inhibit various bacterial CSs. SyCS was not inhibited by ATP and NADH; which are typical feedback inhibitors of other bacterial CSs. SyCS was inhibited by phosphoenolpyruvate and activated by ADP, which has not been reported for CSs from heterotrophic bacteria. Thus, SyCS showed unique characteristics, particularly its sensitivity to effectors.
Similar content being viewed by others
Introduction
Citrate synthase (CS, EC 2.3.3.1) catalyses the first reaction of the tricarboxylic acid (TCA) cycle: oxaloacetate + acetyl-CoA + H2O → citrate + CoA-SH. Different bacterial CSs have been characterised, and studies have shown that the biochemical properties of CSs from Gram-positive bacteria and Gram-negative bacteria exhibit some differences. CSs from Gram-negative bacteria are inhibited by NADH, whereas CSs from Gram-positive bacteria are not1,2. However, the biochemical properties of cyanobacterial CSs have not been well-characterised.
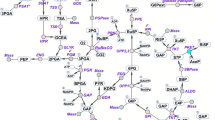
Photoautotrophic cyanobacteria have been evaluated for use in the sustainable production of bioplastics and biofuels from carbon dioxide using light energy3. Synechocystis sp. PCC 6803 (hereafter, Synechocystis 6803) is a unicellular cyanobacterium that has been widely studied for use in metabolite production. Synechocystis 6803 exhibits suitable characteristics as a host for metabolite production such as rapid growth and natural transformation capability4. Synechocystis 6803 can be repeatedly genetically manipulated5. Various genetic toolsets have been developed for Synechocystis 68036. This freshwater cyanobacterium can also grow in artificial seawater supplemented with nitrogen and phosphorus7. Under dark and anaerobic conditions, Synechocystis 6803 produces organic acids such as d-lactate and succinate, which are used in bioplastics production (Fig. 1)8. In Synechocystis 6803, biochemical analyses of a phosphoenolpyruvate carboxylase (encoded by pps, sll0920) being the rate-limiting enzyme in succinate production9,10 and d-lactate dehydrogenase (encoded by ddh, slr1556) catalysing the final reaction in d-lactate production have been performed (Fig. 1)11,12. These reports identified the enzymatic properties and the critical amino acid residues regulating the enzymatic activities, and emphasised the importance of biochemical studies of this cyanobacterium11,12. For example, we have previously indicated that lower activity of d-lactate dehydrogenase (Ddh) from Synechocystis 6803 compared to other bacterial Ddh is one of the weak point of d-lactate production using Synechocystis 680312. We have indicated that Ddh from Synechocystis 6803 also catalyses the reaction from oxaloacetate to malate and an amino acid substitution alters the substrate specificity of Ddh12.
Metabolic map around the TCA cycle in Synechocystis sp. PCC 6803. The information of genes encoding enzymes in Synechocystis sp. PCC 6803 were obtained by Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.genome.jp/kegg/). The green squares represent the useful metabolites having been reported previously. The blue dotted lines represent the synthetic reactions being catalysed by foreign enzymes. 3HB: 3-hydroxybutyrate, 3HP: 3-hydroxypropionic acid, PHB: polyhydroxybutyrate.
Characterising acetyl-CoA metabolism and the enzymes involved is crucial for understanding the basic science of cyanobacteria and its potential practical applications. The intracellular acetyl-CoA concentration of Synechocystis 6803 is much higher compared to those of other cyanobacteria13. The production of polyhydroxybutyrate (PHB)14,15,16, 3-hydroxybutyrate (3HB)17, n-butanol18, and 3-hydroxypropionic acid (3HP)19,20 from acetyl-CoA in Synechocystis 6803 has been studied (Fig. 1). Ethylene production from the TCA cycle intermediates in Synechocystis 6803 has also been demonstrated (Fig. 1)21,22. Despite these metabolic engineering applications, few studies have conducted biochemical analysis using acetyl-CoA as a substrate in cyanobacterium. In this study, we performed biochemical analyses of SyCS (encoded by gltA, sll0401) (Fig. 1) and compared its properties with those of other bacterial CSs.
Results
Purification and kinetic analyses of SyCS
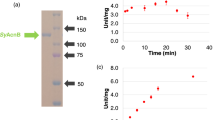
For biochemical analysis of SyCS, glutathione-S-transferase (GST)-tagged SyCS was purified from Escherichia coli by affinity chromatography (Fig. 2a). The optimal conditions at which SyCS showed maximal activity were 37 °C and pH 7.5 (Fig. 2b,c). Therefore, subsequent analysis of SyCS was conducted under these conditions. To calculate the kinetic parameters of SyCS, saturation curves of SyCS for both substrates were drawn (Fig. 3). The observed Km value of SyCS for oxaloacetate was 91 ± 11 μM and the kcat value for oxaloacetate was 2.76 ± 0.26 s−1 (Table 1). The kcat/Km value of SyCS for oxaloacetate was 30.50 ± 3.17 s−1mM−1 (Table 1). Similarly, for acetyl-CoA, the Km value was 220 ± 77 μM and the kcat value was 2.51 ± 0.24 s−1 (Table 1). The kcat/Km value of SyCS for acetyl-CoA was 12.07 ± 2.95 s−1mM−1 (Table 1). SyCS showed no catalytic activity for the backward reaction, generating acetyl-CoA and oxaloacetate from citrate and CoA.
Effect of temperature and pH on SyCS activity. (a) SDS-PAGE analysis after affinity purification of GST-tagged SyCS. The purified proteins were electrophoresed by 12% SDS-PAGE and stained with InstantBlue reagent. Arrows (blue) indicate the positions of molecular protein markers. (b) SyCS activity at different temperatures. The experiment was conducted using 150 pmol of SyCS at pH 7.5. The concentrations of oxaloacetate and acetyl-CoA were 1 and 0.5 mM, respectively. SyCS activity was set to 100% at 37 °C. The data represent the mean ± SD from three independent experiments. (c) SyCS activity at different pH values. The experiment was conducted using 150 pmol of SyCS, and the temperature was 37 °C. The concentrations of oxaloacetate and acetyl-CoA were 1 and 0.5 mM, respectively. SyCS activity was set to 100% at pH 7.5. The data represent the mean ± SD from three independent experiments.
Saturation curves of SyCS for oxaloacetate (left) and acetyl-CoA (right). The experiment was conducted using 50 pmol of SyCS at 37 °C, pH 7.5. The concentrations of oxaloacetate and acetyl-CoA were set to 1 and 0.5 mM, respectively. The data represent the mean ± SD from three independent experiments.
Effects of ions on SyCS activity
The effects of monovalent and divalent salts on SyCS activity were examined. SyCS activity was altered by monovalent and divalent salts, which was affected by the concentration of substrates (Fig. 4). At a saturated concentration of both substrates in the absence of effectors, SyCS activity increased to 188% in the presence of 100 mM KCl, 205% in the presence of 100 mM NaCl, 391% in the presence of 100 mM MgCl2, and 251% in the presence of 100 mM CaCl2, while enzyme activity was decreased to 48% in the presence of 100 mM MnCl2 (Fig. 4). At the half-saturated concentration of both substrates in the absence of effectors, SyCS activity increased to 345% in the presence of 100 mM KCl, 336% in the presence of 100 mM NaCl, 1,463% in the presence of 100 mM MgCl2, and 1,050% in the presence of 100 mM CaCl2, while enzyme activity was decreased to 37% in the presence of 100 mM MnCl2 (Fig. 4).
Effect of monovalent and divalent salts on SyCS activity. The experiment was conducted using 50 pmol of SyCS at 37 °C, pH 7.5. The concentrations of monovalent and divalent salts were set to 100 mM. Oxaloacetate and acetyl-CoA were set to the saturated concentrations (left bars; 1 and 0.5 mM, respectively), and the Km values (right bars; 0.091 and 0.220 mM, respectively). SyCS activity in the absence of monovalent and divalent salts was set to 100%. The data represent the mean ± SD from three independent experiments. The asterisks indicate significant differences between the absence and presence of the salt (Student’s t test; *P < 0.05, **P < 0.005).
The activation of SyCS by monovalent and divalent salts was affected by the pH (Fig. 5). In KCl, SyCS activation was concentration-dependent at both pH 7.0 and pH 8.5 (Fig. 5a). The activity of SyCS increased linearly with increasing concentrations of NaCl at pH 7.0 (Fig. 5b). At pH 8.5, SyCS activity did not increase up to 50 mM NaCl; however, it increased by 2.4-fold in the presence of 100 mM NaCl (Fig. 5b). The activation of SyCS by 100 mM monovalent salts at pH 8.5 was greater than that at pH 7.0 (Fig. 5a,b). The activation of SyCS peaked at 20 mM MgCl2 at both pH 7.0 and pH 8.5 (Fig. 5c). The activation of SyCS peaked at 20 mM CaCl2 at pH 8.5 and at 50 mM CaCl2 at pH 7.0 (Fig. 5d). In contrast to monovalent salts, the activation of SyCS by 100 mM divalent salts at pH 7.0 was greater than that at pH 8.5 (Fig. 5c,d).
Effect of pH on activation of SyCS by (a) KCl, (b) NaCl, (c) MgCl2, and (d) CaCl2. The experiment was conducted using 50 pmol of SyCS at 37 °C, pH 7.0 or 8.5. The concentrations of oxaloacetate and acetyl-CoA were 1 and 0.5 mM. SyCS activity in the absence of monovalent and divalent salts was set to 100%. The data represent the mean ± SD from three independent experiments.
Regulation of SyCS activity by metabolites
SyCS activity was also altered by different metabolites (Fig. 6). In the presence of 5 mM phosphoenolpyruvate (PEP), citrate (CIT), and 2-oxoglutarate (2OG), SyCS activity markedly decreased to 14%, 1%, and 2%, respectively (Fig. 6). Although PEP is a metabolite in glycolysis, it inhibited SyCS activity (Fig. 6). To examine the changes in the kinetic parameters of SyCS, saturation curves of SyCS for both substrates in the presence of 3 mM PEP were drawn (Fig. 3). In the presence of 3 mM PEP, the Km value of SyCS for oxaloacetate was 108 ± 4 μM, the kcat value of SyCS for oxaloacetate decreased to 1.53 ± 0.06 s−1, and the kcat/Km value of SyCS for oxaloacetate decreased to 14.19 ± 0.99 s−1mM−1 (Table 1). In the presence of 3 mM PEP, the Km value of SyCS for acetyl-CoA was 194 ± 36 μM, the kcat value of SyCS for acetyl-CoA decreased to 1.42 ± 0.18 s−1, and the kcat/Km value of SyCS for acetyl-CoA was 7.39 ± 0.70 s−1mM−1 (Table 1). In contrast, SyCS activity significantly increased to 369% in the presence of 5 mM ADP (Fig. 6). To examine changes in the kinetic parameters of SyCS, the saturation curves of SyCS for both substrates in the presence of 5 mM ADP were drawn (Fig. 3). In the presence of 5 mM ADP, the Km value of SyCS for oxaloacetate was 73 ± 11 μM, the kcat value of SyCS for oxaloacetate was 3.35 ± 0.54 s−1, and the kcat/Km value of SyCS for oxaloacetate increased to 45.65 ± 0.84 s−1mM−1 (Table 1). In the presence of 5 mM ADP, the Km value of SyCS for acetyl-CoA was 153 ± 24 μM, the kcat value of SyCS for acetyl-CoA was 2.99 ± 0.12 s−1, and the kcat/Km value of SyCS for acetyl-CoA increased to 19.90 ± 2.82 s−1mM−1 (Table 1). Compounds related to the TCA cycle and amino acid metabolism [pyruvate (PYR), succinate (SUC), fumarate (FUM), l-malate (MAL), and l-aspartate (ASP)] and nucleotides (ATP, AMP, NADH, NAD+, NADPH, and NADP+) did not significantly alter SyCS activity (Fig. 6).
Effect of metabolites on SyCS activity. The experiment was conducted using 50 pmol of SyCS at 37 °C, pH 7.5. The concentrations of oxaloacetate and acetyl-CoA were the Km values, 0.091 and 0.220 mM, respectively. Concentrations of the metabolites were 5 mM. SyCS activity in the absence of metabolites was set to 100%. The data represent the mean ± SD from three independent experiments. PEP: Phosphoenolpyruvate, PYR: Pyruvate, CIT: Citrate, 2OG: 2-Oxoglutarate, SUC: Succinate, FUM: Fumarate, MAL: l-Malate, ASP: l-Aspartate. The asterisks indicate significant differences between the absence and presence of the metabolite (Student’s t test; *P < 0.05).
Discussion
We purified and biochemically characterised CS from the cyanobacterium Synechocystis sp. PCC 6803 in this study. The kcat value of SyCS was lower than those of reported CSs from heterotrophic bacteria (8–263 s−1)23,24,25,26 (Table 1). The Km value of SyCS for acetyl-CoA was similar to that of CS from Arthrobacter strain DS2-3R (200 μM)26 but higher than those of reported CSs from heterotrophic bacteria (14–200 μM)23,24,25,26,27,28,29,30 (Table 1). Similarly, the Km value of SyCS for oxaloacetate was higher than those of CSs from heterotrophic bacteria (4–20 μM)23,24,25,26,27,28,29,30 (Table 1). These observations indicate that SyCS is an inefficient enzyme, similar to malate dehydrogenase from Synechocystis 6803 (encoded by citH, sll0891), which is involved in the oxidative TCA cycle (Fig. 1)31. In Synechocystis 6803, carbon flux through the TCA cycle is low under heterotrophic, photoheterotrophic, photomixotrophic, and photoautotrophic conditions in contrast to other metabolic pathways such as the glycolysis and oxidative pentose phosphate pathways22,32,33,34,35,36. The low activity of SyCS may be a factor in the observed low carbon flux through the oxidative TCA cycle in Synechocystis 6803. Additionally, CSs from Thermosulfidibacter takaii ABI70S6T37 and Desulfurella acetivorans38 catalyse the backward reaction, the cleavage of citrate. SyCS did not catalyse the backward reaction. This result suggests that SyCS is not involved in the reductive TCA cycle (Fig. 1).
Monovalent and divalent salts and cellular metabolites altered SyCS activity (Figs 4–6). Monovalent salts activate numerous bacterial CSs2, while divalent salts inhibit many bacterial CSs39,40. In contrast to other bacterial CSs, SyCS was markedly activated by the divalent salts MgCl2 and CaCl2 rather than monovalent salts (Fig. 4). Among divalent salts, MnCl2 inhibited SyCS (Fig. 4). In CS from E. coli, a higher pH leads to greater activation by 0.1 M KCl39. In SyCS, a higher pH led to greater activation by 0.1 M KCl and NaCl (Fig. 5a,b). In contrast, a lower pH led to greater activation by 0.1 M MgCl2 and CaCl2 (Fig. 5c,d). In monovalent and divalent salts, MgCl2 was the strongest activator of SyCS (Figs 4 and 5). In the stroma in spinach chloroplasts, free Mg2+ concentration is affected by light-dark transition41. It is expected that free Mg2+ concentration in Synechocystis 6803 is similarly affected by light-dark transition42. Therefore, SyCS activity may also be affected by light-dark transition. NADH is a feedback inhibitor of CSs in most Gram-negative bacteria1. However, SyCS was not inhibited by NADH (Fig. 6). Phylogenetic analysis revealed that SyCS belongs to a cyanobacterial clade being different from clades of Gram-positive bacteria and Gram-negative bacteria (Fig. 7). Unlike heterotrophic bacteria, cyanobacteria do not possess 2-oxoglutarate dehydrogenase in the TCA cycle, and Synechocystis 6803 is rerouted to produce succinate by two alternate pathways (Fig. 1)43,44. Isocitrate dehydrogenase from Synechocystis 6803 (encoded by icd, slr1289) (Fig. 1) is specific to NADP+ and generates NADPH but not NADH45. Additionally, malate dehydrogenase from Synechocystis 6803 (encoded by citH, sll0891) (Fig. 1) specifically catalyses the reductive reaction generating NAD+ but not NADH31. These previous studies explain why NADH does not function as a feedback inhibitor of SyCS. CIT and 2OG are feedback inhibitors of SyCS and several bacterial CSs1,2,46 (Fig. 6). We demonstrated that PEP is a unique inhibitor of SyCS (Fig. 6). In the presence of PEP, the Km values of SyCS for both substrates were unchanged and the kcat values of SyCS for both substrates were decreased (Table 1). These results indicate that PEP is a non-competitive inhibitor of SyCS. Phosphoenolpyruvate carboxylase from Synechocystis 6803 (encoded by pps, sll0920) catalyses the carboxylation of PEP to generate oxaloacetate, a substrate of SyCS (encoded by gltA, sll0401) and malate dehydrogenase (encoded by citH, sll0891) (Fig. 1)11. The reaction catalysed by phosphoenolpyruvate carboxylase (encoded by pps, sll0920) (Fig. 1) is the rate-limiting step in the production of succinate via the reductive TCA cycle in Synechocystis 68039,10. PEP has been suggested to regulate both the oxidative and reductive TCA cycle in Synechocystis 6803. The intracellular concentrations of CIT and PEP are much higher than that of 2OG (CIT: 7.3 times, PEP: 8.5 times)13. In inhibitors of metabolites, CIT and PEP may have large inhibitory effects on SyCS under in vivo conditions. While ATP did not alter SyCS activity (Fig. 6), it acts as a feedback inhibitor for other CSs because of its structural similarity with acetyl-CoA1,2. In terms of activators, AMP upregulates the activity of CSs from Azotobacter vinelandii47 and Streptomyces diastaticus No. 7 strain M103324. However, SyCS was activated by ADP rather than AMP (Fig. 6). ADP enhanced the catalytic efficiency (kcat/Km) of SyCS (Table 1). Generally, ADP inhibits CS for the same reason as ATP1,2. Our study indicates that ADP is a unique activator of SyCS. Synechocystis 6803 has more ADP than ATP in the cells13 in contrast to E. coli48 whose CS is inhibited by ATP and hardly affected by ADP49. This may be a reason that SyCS is affected by not ATP but ADP.
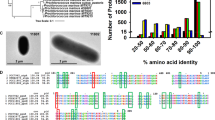
Phylogenetic analysis of bacterial CS. Sequences of 23 bacterial CS obtained from GenBank were aligned using CLC Sequence Viewer ver. 8.0. A maximum-likelihood tree based on 357 preserved amino acid residues was designed using PhyML online (http://www.atgc-montpellier.fr/phyml/). Bootstrap values were calculated by 500 replications.
Taken together, we found that the characteristics of SyCS significantly differ from those of other bacterial CSs.
Methods
Construction of the recombinant SyCS expression vector
The region containing gltA (sll0401, encoding SyCS) in the Synechocystis 6803 genome was commercially synthesized by Eurofin Genomics Japan (Tokyo, Japan) and cloned into the BamHI-XhoI site of the pGEX5X-1 vector (GE Healthcare, Little Chalfont, UK).
Affinity purification of recombinant proteins
The expression vector was transformed into E. coli DH5α cells (TakaraBio, Shiga, Japan). A 2-L culture of transformed DH5α cells in LB media was prepared by shaking at 125 rpm at 30 °C overnight. Expression of the recombinant proteins was induced by 0.01 mM isopropyl β-d-1-thiogalactopyranoside (Wako Chemicals, Osaka, Japan). Affinity purification of the recombinant proteins was performed as described previously12. The DH5α cells were lysed by sonication (VC-750, EYELA, Tokyo, Japan) for 200 s at 20% intensity. The supernatant was collected after centrifugation (5,800 × g for 2 min at 4 °C) and 560 μL of Glutathione-Sepharose 4B resin (GE Healthcare) was added to the supernatant. The mixture was incubated on ice for 30 min with constant shaking. After 30 min, 1 mM ATP and 1 mM MgSO4·7H2O were added to the mixture and shaken for 30 min at 37 °C. After centrifugation (5,800 × g for 2 min at 4 °C) to remove the supernatant, the resin was re-suspended in 700 μL of PBS-T (1.37 M NaCl, 27 mM KCl, 81 mM Na2HPO4·12H2O, 14.7 mM KH2PO4, and 0.05% Tween-20) with 1 mM ATP/1 mM MgSO4·7H2O. The resin was washed 10 times using PBS-T, and the recombinant proteins were eluted with 700 μL of GST elution buffer (50 mM Tris-HCl, pH 8.0, and 10 mM reduced glutathione) four times. The eluted protein fractions were concentrated using a VivaSpin 500 MWCO 50000 device (Sartorius, Göttingen, Germany). The protein concentration was measured using a Pierce BCA Protein Assay Kit (Rockford, IL, USA). To evaluate protein purity, SDS-PAGE followed by staining with InstantBlue (Expedion Protein Solutions, San Diego, CA, USA) was performed.
Enzyme assays
The enzyme activity of SyCS for the forward reaction, generating citrate and CoA from acetyl-CoA and oxaloacetate, was assessed as described previously50. Enzyme activity was calculated by measuring the change in A412 using a Hitachi U-3310 spectrophotometer (Tokyo, Japan). The purified SyCS proteins were added to 1 mL assay solution [50 mM Tris-HCl containing various concentrations of oxaloacetate, various concentrations of acetyl-CoA, and 0.2 mM DTNB]. The Km and Vmax values of SyCS were calculated using a Lineweaver-Burk double reciprocal plot. The kcat values of SyCS were determined from the Vmax values of SyCS. The enzyme activity of SyCS for the backward reaction, generating acetyl-CoA and oxaloacetate from citrate and CoA, was assessed as described previously38 by measuring the change in A365 using a Hitachi U-3310 spectrophotometer. The purified SyCS proteins were added to 1 mL assay solution [50 mM Tris-HCl containing 1 mM CoA, 1 mM citrate, 5 mM MgCl2, 0.5 mM NADH, and 20 U malate dehydrogenase (Oriental Yeast, Tokyo, Japan)].
References
Weitzman, P. D. Unity and diversity in some bacterial citric acid-cycle enzymes. Adv. Microb. Physiol. 22, 185–244 (1981).
Weitzman, P. D. & Danson, M. J. Citrate synthase. Curr. Top. Cell. Regul. 10, 161–204 (1976).
Katayama, N., Iijima, H. & Osanai, T. Production of bioplastic compounds by genetically manipulated and metabolic engineered cyanobacteria. Adv. Exp. Med. Biol. 1080, 155–169 (2018).
Branco Dos Santos, F., Du, W. & Hellingwerf, K. J. Synechocystis: Not Just a Plug-Bug for CO2, but a Green E. coli. Front Bioeng Biotechnol. 2, 36 (2014).
Lea-Smith, D. J., Vasudevan, R. & Howe, C. J. Generation of Marked and Markerless Mutants in Model Cyanobacterial Species. J Vis Exp (2016).
Wang, B., Eckert, C., Maness, P. C. & Yu, J. P. A Genetic Toolbox for Modulating the Expression of Heterologous Genes in the Cyanobacterium Synechocystis sp PCC 6803. Acs Synthetic Biology. 7, 276–286 (2018).
Iijima, H., Nakaya, Y., Kuwahara, A., Hirai, M. Y. & Osanai, T. Seawater cultivation of freshwater cyanobacterium Synechocystis sp. PCC 6803 drastically alters amino acid composition and glycogen metabolism. Front. Microbiol. 6, 326 (2015).
Osanai, T. et al. Genetic manipulation of a metabolic enzyme and a transcriptional regulator increasing succinate excretion from unicellular cyanobacterium. Front. Microbiol. 6, 1064 (2015).
Hasunuma, T., Matsuda, M. & Kondo, A. Improved sugar-free succinate production by Synechocystis sp. PCC 6803 following identification of the limiting steps in glycogen catabolism. Metab. Eng. Commun. 3, 130–141 (2016).
Hasunuma, T., Matsuda, M., Kato, Y., Vavricka, C. J. & Kondo, A. Temperature enhanced succinate production concurrent with increased central metabolism turnover in the cyanobacterium Synechocystis sp. PCC 6803. Metab. Eng. 48, 109–120 (2018).
Takeya, M., Hirai, M. Y. & Osanai, T. Allosteric inhibition of phosphoenolpyruvate carboxylases is determined by a single amino acid residue in cyanobacteria. Sci. Rep. 7, 41080 (2017).
Ito, S., Takeya, M. & Osanai, T. Substrate specificity and allosteric regulation of a d-lactate dehydrogenase from a unicellular cyanobacterium are altered by an amino acid substitution. Sci. Rep. 7, 15052 (2017).
Dempo, Y., Ohta, E., Nakayama, Y., Bamba, T. & Fukusaki, E. Molar-based targeted metabolic profiling of cyanobacterial strains with potential for biological production. Metabolites. 4, 499–516 (2014).
Osanai, T. et al. Increased bioplastic production with an RNA polymerase sigma factor sige during nitrogen starvation in Synechocystis sp. PCC 6803. DNA Res. 20, 525–535 (2013).
Osanai, T. et al. Pathway-level acceleration of glycogen catabolism by a response regulator in the cyanobacterium Synechocystis species PCC 6803. Plant Physiol. 164, 1831–1841 (2014).
Khetkorn, W., Incharoensakdi, A., Lindblad, P. & Jantaro, S. Enhancement of poly-3-hydroxybutyrate production in Synechocystis sp. PCC 6803 by overexpression of its native biosynthetic genes. Bioresour. Technol. 214, 761–768 (2016).
Wang, B., Pugh, S., Nielsen, D. R., Zhang, W. & Meldrum, D. R. Engineering cyanobacteria for photosynthetic production of 3-hydroxybutyrate directly from CO2. Metab. Eng. 16, 68–77 (2013).
Anfelt, J. et al. Genetic and nutrient modulation of acetyl-CoA levels in Synechocystis for n-butanol production. Microb. Cell Fact. 14, 167 (2015).
Wang, Y. et al. Biosynthesis of platform chemical 3-hydroxypropionic acid (3-HP) directly from CO2 in cyanobacterium Synechocystis sp. PCC 6803. Metab. Eng. 34, 60–70 (2016).
Wang, Y., Chen, L. & Zhang, W. Proteomic and metabolomic analyses reveal metabolic responses to 3-hydroxypropionic acid synthesized internally in cyanobacterium Synechocystis sp. PCC 6803. Biotechnol. Biofuels. 9, 209 (2016).
Veetil, V. P., Angermayr, S. A. & Hellingwerf, K. J. Ethylene production with engineered Synechocystis sp. PCC 6803 strains. Microb. Cell Fact. 16, 34 (2017).
Xiong, W. et al. The plasticity of cyanobacterial metabolism supports direct CO2 conversion to ethylene. Nat. Plants. 1, 15053 (2015).
Bond, D. R. et al. Characterization of citrate synthase from Geobacter sulfurreducens and evidence for a family of citrate synthases similar to those of eukaryotes throughout the Geobacteraceae. Appl. Environ. Microbiol. 71, 3858–3865 (2005).
Ge, Y., Cao, Z., Song, P. & Zhu, G. Identification and characterization of a novel citrate synthase from Streptomyces diastaticus No. 7 strain M1033. Biotechnol. Appl. Biochem. 62, 300–308 (2015).
Francois, J. A. et al. Structure of a NADH-insensitive hexameric citrate synthase that resists acid inactivation. Biochemistry. 45, 13487–13499 (2006).
Gerike, U., Danson, M. J. & Hough, D. W. Cold-active citrate synthase: mutagenesis of active-site residues. Protein Eng. Des. Sel. 14, 655–661 (2001).
Mitchell, C. G. & Weitzman, P. D. J. Proteolysis of Acinetobacter citrate synthase by subtilisin. FEBS Lett. 151, 265–268 (1983).
Robinson, M. S., Danson, M. J. & Weitzman, P. D. Citrate synthase from a Gram-positive bacterium. Purification and characterization of the Bacillus megaterium enzyme. Biochem. J. 213, 53–59 (1983).
Sievers, M., Stöckli, M. & Teuber, M. Purification and properties of citrate synthase from Acetobacter europaeus. FEMS Microbiol. Lett. 146, 53–58 (1997).
Swissa, M. & Benziman, M. Factors affecting the activity of citrate synthase of Acetobacter xylinum and its possible regulatory role. Biochem. J. 153, 173–179 (1976).
Takeya, M., Ito, S., Sukigara, H. & Osanai, T. Purification and characterisation of malate dehydrogenase from Synechocystis sp. PCC 6803: biochemical barrier of the oxidative tricarboxylic acid cycle. Front. Plant Sci. 9, 947 (2018).
Wan, N. et al. Cyanobacterial carbon metabolism: Fluxome plasticity and oxygen dependence. Biotechnol. Bioeng. 114, 1593–1602 (2017).
You, L., He, L. & Tang, Y. J. Photoheterotrophic fluxome in Synechocystis sp. strain PCC 6803 and its implications for cyanobacterial bioenergetics. J. Bacteriol. 197, 943–950 (2015).
Nakajima, T. et al. Integrated metabolic flux and omics analysis of Synechocystis sp. PCC 6803 under mixotrophic and photoheterotrophic conditions. Plant Cell Physiol. 55, 1605–1612 (2014).
You, L., Berla, B., He, L., Pakrasi, H. B. & Tang, Y. J. 13C-MFA delineates the photomixotrophic metabolism of Synechocystis sp. PCC 6803 under light- and carbon-sufficient conditions. Biotechnol. J. 9, 684–692 (2014).
Young, J. D., Shastri, A. A., Stephanopoulos, G. & Morgan, J. A. Mapping photoautotrophic metabolism with isotopically nonstationary 13C flux analysis. Metab. Eng. 13, 656–665 (2011).
Nunoura, T. et al. A primordial and reversible TCA cycle in a facultatively chemolithoautotrophic thermophile. Science. 359, 559–563 (2018).
Mall, A. et al. Reversibility of citrate synthase allows autotrophic growth of a thermophilic bacterium. Science. 359, 563–567 (2018).
Faloona, G. R. & Srere, P. A. Escherichia coli citrate synthase. Purification and the effect of potassium on some properties. Biochemistry. 8, 4497–4503 (1969).
Flechtner, V. R. & Hanson, R. S. Regulation of the tricarboxylic acid cycle in bacteria. A comparison of citrate synthases from different bacteria. Biochim Biophys Acta. 222, 253–264 (1970).
Ishijima, S., Uchibori, A., Takagi, H., Maki, R. & Ohnishi, M. Light-induced increase in free Mg2+ concentration in spinach chloroplasts: measurement of free Mg2+ by using a fluorescent probe and necessity of stromal alkalinization. Arch Biochem Biophys. 412, 126–132 (2003).
Osanai, T. et al. ChlH, the H subunit of the Mg-chelatase, is an anti-sigma factor for SigE in Synechocystis sp. PCC 6803. Proc Natl Acad Sci USA 106, 6860–6865 (2009).
Zhang, S. & Bryant, D. A. The tricarboxylic acid cycle in cyanobacteria. Science. 334, 1551–1553 (2011).
Xiong, W., Brune, D. & Vermaas, W. F. J. The γ-aminobutyric acid shunt contributes to closing the tricarboxylic acid cycle in Synechocystis sp. PCC 6803. Mol. Microbiol. 93, 786–796 (2014).
Muro-Pastor, M. I. & Florencio, F. J. Purification and properties of NADP-isocitrate dehydrogenase from the unicellular cyanobacterium Synechocystis sp. PCC 6803. Eur. J. Biochem. 203, 99–105 (1992).
Johnson, D. E. & Hanson, R. S. Bacterial citrate synthases: purification, molecular weight and kinetic mechanism. Biochim. Biophys. Acta - Enzymol. 350, 336–353 (1974).
Rault-Leonardon, M., Atkinson, M. A. L., Slaughter, C. A., Moomaw, C. R. & Srere, P. A. Azotobacter vinelandii citrate synthase. Biochemistry. 34, 257–263 (1995).
Bennett, B. D. et al. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat Chem Biol. 5, 593–599 (2009).
Jangaard, N. O., Unkeless, J. & Atkinson, D. E. The inhibition of citrate synthase by adenosine triphosphate. Biochim Biophys Acta. 15, 225–235 (1968).
Srere, P. A., Brazil, H., Gonen, L. & Takahashi, M. The citrate condensing enzyme of pigeon breast muscle and moth flight muscle. Acta Chem. Scand. 17(supl.), 129–134 (1963).
Acknowledgements
This work was supported by the Ministry of Education, Culture, Sports, Science, and Technology, Japan, by a grant to T.O., from JST-ALCA (Japan Science and Technology Agency, Advanced Low Carbon Technology Research and Development Program, the project name is “Production of cyanobacterial succinate by the genetic engineering of transcriptional regulators and circadian clocks”) (Grant Number JPMJAL1306) and from by Japan Society for the Promotion of Science, KAKENHI Grant-in-Aid for Scientific Research on Innovative Areas (Grant Number 16H06559). No conflicts of interest to declare. All authors contributed to the work and approved the submission of the manuscript.
Author information
Authors and Affiliations
Contributions
S.I. designed the research, performed the experiments, analysed the data, and wrote the manuscript. N.K. performed the experiments and analysed the data. T.O. designed the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ito, S., Koyama, N. & Osanai, T. Citrate synthase from Synechocystis is a distinct class of bacterial citrate synthase. Sci Rep 9, 6038 (2019). https://doi.org/10.1038/s41598-019-42659-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42659-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.