Abstract
Resistance-associated substitutions (RASs) of hepatitis C virus (HCV) in the NS5A region impair the efficacy of NS5A inhibitors. In this study, we evaluated the characteristics of the novel RASs observed in treatment-failure patients, A92K and a deletion at P32 (P32del), and the susceptibility of viruses with these RASs to various anti-HCV reagents by using JFH-1 based recombinant HCV with NS5A from a genotype 1b Con1 strain (JFH1/5ACon1). We introduced A92K or P32del solely or in combination with Q24K, L28M, R30Q or L31F into the NS5A of JFH1/5ACon1. Viruses harboring R30Q/A92K showed high extracellular core antigens and infectivity titers, whereas the other viruses with RASs showed low replication levels and infectivity titers. All the viruses with A92K or P32del were markedly resistant to ledipasvir, velpatasvir and elbasvir. Interestingly, viruses with R30Q/A92K were more susceptible to grazoprevir than viruses without RAS. All the viruses had a similar susceptibility to ribavirin and sofosbuvir. In conclusion, combination RASs R30Q/A92K enhanced virus production whereas other RASs impaired virus replication. Both A92K and P32del conferred severe resistance even to second generation NS5A inhibitors. However, these viruses were susceptible to grazoprevir, ribavirin and sofosbuvir. Thus, combination regimens with these reagents may eradicate viruses harboring A92K or P32del.
Similar content being viewed by others
Introduction
Hepatitis C virus (HCV) infection is a global health problem. The number of people infected with HCV is approximately 80 million worldwide1,2. Of note, HCV infection leads to chronic hepatitis, cirrhosis, end-stage liver dysfunction and hepatocellular carcinoma. The global eradication of this virus is the ultimate goal to solve HCV-associated health problems. For this purpose, many anti-viral reagents have been investigated, and novel reagents designated direct-acting antiviral agents (DAAs) that inhibit viral proteins directly have been established3. The use of DAAs has markedly improved the sustained virological response (SVR) rate of anti-HCV treatments1,4. Among currently available DAAs, NS5A inhibitors have potent anti-HCV effects and are described as key reagents that are included in many recommended DAA regimens in the treatment guidelines for hepatitis C5,6,7. However, the potency of NS5A inhibitors has been impaired by the emergence of specific amino acid substitutions designated resistance-associated substitutions (RASs)8,9,10. RASs of L31F/M/V/I and Y93H in the NS5A region are frequently observed in both DAA naïve and experienced patients infected with HCV genotype 1b strains9,11,12,13. These RASs were reported to be associated with virologic failure of the DAA therapies including first generation NS5A inhibitors, such as daclatasvir (DCV), ledipasvir (LDV) and ombitasvir9,14. To achieve a high SVR rate, even in patients with these RASs, second generation NS5A inhibitors were developed and their superior efficacy was reported15,16,17. However, in recent studies, novel RASs in the NS5A region, A92K or a deletion at P32 (P32del), were identified18,19,20. These novel RASs were not detected in patients before treatment but emerged after DAA treatment alone or in combination. These RASs are considered resistant to DAAs, but the resistance levels, especially in combination with other RASs, have not been investigated. Therefore, the evaluation of these RASs in terms of resistance to DAAs and the identification of effective compounds for these RASs are urgent issues.
HCV subgenomic replicon system have been used in many studies to investigate RASs. However, this system is not able to produce infectious virus and is not suitable for evaluating the HCV whole life cycle. Whereas, HCV cell culture system enables to assess the virus whole life cycle. Previously, we used a recombinant virus of JFH-1 to assess the effects of the RASs, L31M/V/I and Y93H, on the replication and infectious virus production of HCV and to identify effective DAAs to these RASs21. The used recombinant virus has NS5A from the genotype 1b strain and enabled the assessment for the effects of RASs on the HCV life cycle and susceptibility to anti-HCV reagents against genotype 1b strains. The current study used this system to evaluate the effects of the novel RASs A92K and P32del on the HCV life cycle and susceptibility to various anti-HCV reagents.
Results
Effects of A92K Substitution on the HCV Life Cycle
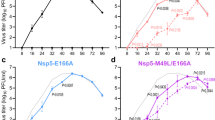
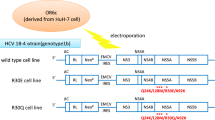
To assess the effect of A92K on HCV propagation, we used the JFH-1 based recombinant virus with the NS5A of genotype 1b strain (JFH1/5AC-wt)21,22. A92K was introduced in this virus alone (JFH1/5AC-A92K) or in combination with R30Q (JFH1/5AC-R30Q/A92K) or Q24K, L28M and R30Q (JFH1/5AC-Q24K/L28M/R30Q/A92K) (Fig. 1a). We transfected the full-length RNAs of these viruses into Huh7.5.1 cells and measured the HCV core antigen (Ag) levels on days 1, 2 and 3 post-transfection. Time-dependent increases of extracellular HCV core Ag levels were observed in these cells, but the levels were different between recombinant viruses (Fig. 1b). The extracellular core Ag levels of JFH1/5AC-wt and -R30Q/A92K were higher than that of the other recombinant viruses. In JFH1/5AC-R30Q/A92K transfected cells, the extracellular core Ag level was higher, but the intracellular core Ag level was slightly lower than that of JFH1/5AC-wt at day 3 post-transfection (Fig. 1c). In addition, both the extra- and intracellular HCV core Ag levels of JFH1/5AC-A92K and -Q24K/L28M/R30Q/A92K were substantially lower than that of JFH1/5AC-wt. To explore the steps in the HCV life cycle affected by the RASs, we performed a single cycle virus production assay with Huh7-25 cells. Huh7-25 cell line supports HCV replication and virus production after the transfection of full-length HCV RNA, but it is not re-infected with progeny viruses because it lacks the expression of CD81 on the cell surface23. Therefore, it allows to assess which steps were affected in a single cycle of virus production. The results of this experiment were similar with that using Huh-7.5.1 cells; the HCV core Ag levels of JFH1/5AC-R30Q/A92K were approximately 2.5-fold higher in culture medium and were comparable in cell lysates compared with JFH1/5AC-wt (Fig. 1d). Both the extra- and intracellular HCV core Ag levels of other viruses were less than 0.5-fold that of JFH1/5AC-wt.
Characteristics of the recombinant virus JFH1/5AC with A92K. (a) Schematic representation of the recombinant virus (JFH1/5AC), subgenomic replicon (JFH1/5ACRep) and their derivatives with NS5A resistance-associated substitutions. (b) Time course of extracellular core Ag levels of HCV RNA-transfected Huh7.5.1 cells. (c,d) In vitro transcribed HCV RNAs were electroporated into Huh 7.5.1 (c) or Huh7-25 (d) cells. On day 3 post-transfection, extra- and intracellular core Ag levels were quantified. (e) Extra- and intracellular infectivity titers of HCV RNA-transfected Huh7-25 cells. (f) Luciferase activity of HCV-subgenomic replicon RNA-transfected Huh-7.5.1 cells on day 3 post-electroporation. Replication levels were calculated as fold increases at 4 hours. (c–f) Data are expressed as the percentages of JFH1/5AC-wt. *p < 0.05.
To assess the infectivity of the viruses with these RASs, we next evaluated the extra- and intracellular infectivity titers of these recombinant viruses. Extracellular infectivity titers of JFH1/5AC-R30Q/A92K transfected cells were more than 7-fold higher than that of JFH1/5AC-wt, whereas few HCV infected cells were observed in the infection of culture medium and cell lysates from JFH1/5AC-A92K and -Q24K/L28M/R30Q/A92K transfected cells (Fig. 1e and Supplementary Fig. S1). These results suggest that substitutions of R30Q/A92K enhanced the production levels of infectious viruses, and that substitutions of A92K and Q24K/L28M/R30Q/A92K attenuated the virus replication.
To confirm the attenuated replication efficiencies of JFH1/5AC-A92K and -Q24K/L28M/R30Q/A92K, we used the JFH-1 based recombinant subgenomic replicon system (JFH1/5ACRep). Similar to the full-length recombinant viruses, we introduced the RASs of A92K, R30Q/A92K and Q24K/L28M/R30Q/A92K and generated JFH1/5ACRep-A92K, -R30Q/A92K and -Q24K/L28M/R30Q/A92K, respectively (Fig. 1a). In vitro transcribed replicon RNAs were electroporated into Huh7.5.1 cells and luciferase activities were measured on day 3 after electroporation (Fig. 1f). The luciferase activities of JFH1/5ACRep-A92K and -Q24K/L28M/R30Q/A92K were significantly lower than those of JFH1/5ACRep-wt and those of JFH1/5ACRep-R30Q/A92K were slightly higher than -wt. These results were consistent with the results of the intracellular HCV core Ag levels of full-length HCV transfected Huh7-25 cells. These results indicate that A92K reduced virus replication but that R30Q compensated for it mainly by enhancing infectious virus production and partially by enhancing replication.
Effects of P32del Substitution on the HCV Life Cycle
To evaluate the effect of P32del on the virus life cycle, we used JFH1/5AC-wt by introducing P32del alone (JFH1/5AC-P32del) or in combination with L31F (JFH1/5AC-L31F/P32del) (Fig. 2a). When Huh7.5.1 cells were transfected with the full-length RNAs of these viruses, the extracellular HCV core Ag levels of JFH1/5AC-P32del and -L31F/P32del increased in a time-dependent manner although the levels were lower than that of JFH1/5AC-wt (Fig. 2b). At day 3 post-transfection, both the extra- and intracellular core Ag levels of JFH1/5AC-P32del and -L31F/P32del were substantially lower than that of JFH1/5AC-wt (Fig. 2c). The core Ag levels of JFH1/5AC-L31F/P32del were slightly higher than that of JFH1/5AC-P32del. To evaluate which steps in the HCV life cycle were affected by these RASs, we performed the single cycle virus production assay (Fig. 2d). In this experiment, the results were similar to that with Huh-7.5.1 cells. Both the extra- and intracellular HCV core Ag levels of JFH1/5AC-P32del and -L31F/P32del were less than approximately 0.2-fold that of JFH1/5AC-wt and the HCV core Ag levels of JFH1/5AC-L31F/P32del were more than 3-fold higher than JFH1/5AC-P32del. The results suggest the attenuated replication efficiencies of these viruses and complementary effect of L31F on the reduced replication of JFH1/5AC-P32del. We next assessed the extra- and intracellular infectivity titers of these recombinant viruses. Few HCV infected cells were detected by infection with culture medium and cell lysates from JFH1/5AC-P32del and -L31F/P32del transfected cells (Fig. 2e and Supplementary Fig. S1). To confirm the attenuated replication efficiencies of JFH1/5AC-P32del and -L31F/P32del, we generated the subgenomic replicons corresponding to these viruses (JFH1/5ACRep-P32del and -L31F/P32del) and performed the luciferase assay (Fig. 2a and 2f). The luciferase activities of JFH1/5ACRep-P32del and -L31F/P32del were significantly lower than that of JFH1/5ACRep-wt. These results were consistent with the data of the single cycle virus production assay.
Characteristics of the recombinant virus JFH1/5AC with P32del. (a) Schematic representation of the recombinant virus (JFH1/5AC), subgenomic replicon (JFH1/5ACRep) and their derivatives with NS5A resistance-associated substitutions. (b) Time course of extracellular core Ag levels of HCV RNA-transfected Huh7.5.1 cells. (c,d) In vitro transcribed HCV RNAs were electroporated into Huh7.5.1 (c) or Huh7–25 (d) cells. On day 3 post-transfection, extra- and intracellular core Ag levels were quantified. (e) Extra- and intracellular infectivity titers of HCV RNA-transfected Huh7-25 cells. (f) Luciferase activity of HCV-subgenomic replicon RNA-transfected Huh-7.5.1 cells on day 3 post-electroporation. Replication levels were calculated as fold increases at 4 hours. (c–f) Data are expressed as the percentages of JFH1/5AC-wt. *p < 0.05.
Susceptibility of JFH1/5AC with RASs to NS5A inhibitors
To verify the conferred resistance of A92K, P32del and associated RASs to NS5A inhibitors, we assessed the susceptibility of JFH1/5AC-A92K, -R30Q/A92K, -Q24K/L28M/R30Q/A92K, -P32del and -L31F/P32del to LDV, VEL or EBR. The resistance levels of these recombinant viruses were compared with JFH1/5AC-wt, Y93H introduced JFH1/5AC (JFH1/5AC-Y93H) or L31V/Y93H introduced (JFH1/5AC-L31V/Y93H). We treated the HCV RNA-transfected cells with LDV, EBR or VEL for 72 hours and measured the extracellular HCV core Ag levels to determine the EC50 values. Treatment with NS5A inhibitors inhibited the HCV core Ag level of JFH1/5AC-wt at a low concentration. The EC50 values of JFH1/5AC-wt to these NS5A inhibitors were comparable and were 1.1–3.9 pM (Fig. 3 and Table 1). However, all recombinant viruses with A92K or P32del exhibited marked resistance to LDV, and the EC50 values were higher than 20 μM (Fig. 3a and Table 1). For the treatment with VEL or EBR, JFH1/5AC-R30Q/A92K and -Q24K/L28M/R30Q/A92K showed marked resistance comparable to or higher than for JFH1/5AC-L31V/Y93H. The EC50 values of JFH1/5AC-R30Q/A92K, -Q24K/L28M/R30Q/A92K and - L31V/Y93H for VEL were 2.4 × 103, 7.5 × 104, and 8.1 × 103 pM, respectively, and for EBR they were 3.4 × 104, 6.8 × 104 and 3.2 × 104 pM, respectively (Fig. 3b,c and Table 1). Interestingly, JFH1/5AC-A92K exhibited a greater resistance to LDV and EBR compared with VEL, and JFH1/5AC-P32del had a marked resistance to all NS5A inhibitors. Moreover, JFH1/5AC-L31F/P32del had a greater resistance to VEL and EBR compared with JFH1/5AC-P32del, and the EC50 values were 6.3 × 107 pM for LDV, 2.3 × 108 pM for VEL and 1.0 × 107 pM for EBR (Fig. 3a–c and Table 1).
Susceptibility of JFH1/5AC and its derivatives with NS5A RAS to NS5A inhibitors. In vitro transcribed HCV RNAs of JFH1/5AC-wt and indicated derivatives were electroporated into Huh-7.5.1 cells. Four hours after transfection, the cells were treated with LDV (a) VEL (b) or EBR (c). After incubation for 72 hours, the supernatants were harvested and extracellular HCV core Ag levels were quantified. The data are expressed as the percentages of the DMSO-treated control.
These results suggest that A92K substitution was responsible for the resistance of JFH1/5AC-R30Q/A92K and -Q24K/L28M/R30Q/A92K to NS5A inhibitors and that P32del conferred a marked resistance to NS5A inhibitors.
Susceptibility of JFH1/5AC with RASs to the other anti-HCV reagents
To evaluate the susceptibilities of recombinant viruses with RASs to the other anti-HCV reagents, we measured the extracellular HCV core Ag levels after treatment with SOF, RBV or GZR. The core Ag levels of viruses with RASs were reduced by treatment with SOF, RBV or GZR in a concentration-dependent manner (Fig. 4a–c). When treated with SOF or RBV, the dose-response curves of viruses with RASs were similar to that of JFH1/5AC-wt (Fig. 4a,b), and the EC50 values of these strains were comparable (Table 2). These data suggest that the inhibitory effect of SOF and RBV on these strains with NS5A RASs were comparable to JFH1/5AC-wt.
Susceptibility of JFH1/5AC and its derivatives with NS5A RAS to SOF, RBV and GZR. In vitro transcribed HCV RNAs of JFH1/5AC-wt and indicated derivatives were electroporated into Huh-7.5.1 cells. Four hours after transfection, the cells were treated with SOF (a) RBV (b) or GZR (c). After incubation for 72 hours, the supernatants were harvested and extracellular HCV core Ag levels were quantified. Data are expressed as the percentages of the DMSO-treated control.
We previously demonstrated that viruses with Y93H were more susceptible to the protease inhibitors, simeprevir and asunaprevir than viruses without RASs21. Therefore, we evaluated the susceptibility of RAS introduced viruses including JFH1/5AC-Y93H and -L31V/Y93H to GZR, a novel protease inhibitor. As expected, treatment with GZR reduced the extracellular HCV core Ag levels of JFH1/5AC-Y93H and -L31V/Y93H more efficiently than that of JFH1/5AC-wt (Fig. 4c). Interestingly, the reduction of HCV core Ag levels of JFH1/5AC-R30Q/A92K was also more efficient in comparison with JFH1/5AC-wt and the dose-response curve of this strain shifted to the left suggesting a higher susceptibility of this virus to GZR. The EC50 value of JFH1/5AC-R30Q/A92K was 7.4 nM and was significantly lower than that of JFH1/5AC-wt; however, its susceptibility level was not pronounced compared with the levels of JFH1/5AC-Y93H and -L31V/Y93H. The EC50 values of these viruses to GZR were both 1.9 nM and were 0.1-fold that of JFH1/5AC-wt (Table 2). The EC50 values of the other viruses with RASs were similar or slightly, but not significantly, higher than that of JFH1/5AC-wt.
Discussion
NS5A inhibitors play a pivotal role in the treatment of chronic hepatitis C. NS5A RASs reduced the treatment efficacy and had a great impact on the selection of therapeutic regimens when the 1st generation NS5A inhibitors mainly used9,14,24. Although the recently developed DAA combination therapies including 2nd generation NS5A inhibitors have been reported to eradicate the viruses with NS5A RASs and to have high efficacies, specific or multiple RASs is still associated with the treatment failure especially for cirrhosis patients25. The most described NS5A RASs of L31F/M/V/I and Y93H in genotype 1b are often detected in treatment naïve patients at ~6% and ~10%, respectively9,11,12,13. On the other hand, the prevalence of these RASs extremely increases in DAA-experienced patients up to 40% and 60%, respectively18,26. RAS at amino acid (aa) 24, aa 28 and aa 30 in the NS5A region detected in DAA-experienced patients also increase ~4-folds compared with treatment-naïve patients. In addition, recent studies have reported the emergence of novel RASs: A92K and P32del. These RASs were not detected in treatment naïve patients but emerged at the time of the treatment failure of DAA therapy at ~5% treatment with ASV and DCV and also LDV and SOF19,20,27,28,29,30. Of note, virus harboring P32del have been reported to difficult to eliminate by retreatment with LDV and SOF or glecaprevir and pibrentasvir31,32. Furthermore, in many cases, these NS5A RASs emerge in combination with multiple RASs in DAA-experienced patients18. Based on these backgrounds, we reasoned that the information of NS5A RASs should not be ignored when considering the treatment strategy. Thus, the present study evaluated the effects of these novel RASs on the HCV life cycle by exploiting the HCV cell culture system with JFH1 based NS5A chimeric viruses and assessed the susceptibility to various anti-HCV reagents.
The assessment of effects on the viral life cycle indicated that chimeric viruses harboring A92K and P32del had attenuated fitness. Viruses with these RASs had a lower replication efficiency compared with JFH1/5AC-wt. This finding may explain why these RASs are rarely detected in treatment naïve patients at baseline9,12,18,29,33. In the chimeric virus harboring A92K, the addition of R30Q compensated for the virus propagation by enhancing infectious virus production and may partially by enhancing virus replication (Fig. 1). A similar effect was observed by introducing mutations at the same positions in the genotype 2a JFH-1 strain34. The mutation of K30E/Q compensated for the impaired replication efficiency induced by the C92R mutation in the JFH-1 strain. Therefore, amino acids at these positions appear to coordinate with each other, and this effect is maintained beyond genotypes. The enhancement of infectious virus production by R30Q resembled the effect of Y93H reported in our previous study21. Therefore, similar to HCV with Y93H, HCV with R30Q/A92K may persist for a long period after treatment failure21,35,36. Notably, the observed compensation by R30Q disappeared by the addition of the Q24K/L28M mutations. JFH1/5AC-Q24K/L28M/R30Q/A92K exhibited lower replication and virus production efficiencies.
In the chimeric virus with P32del, the addition of L31F conferred a minor effect on the HCV life cycle. The replication efficiency of JFH1/5AC-L31F/P32del was slightly higher than that of -P32del and the production of infectious viruses were similar (Fig. 2). From these observations, we hypothesized that a virus with P32del will decrease and disappear soon after treatment failure. However, a recent report indicated that a virus with P32del persisted for at least one year after treatment similar to a virus with Y93H20,37. Thus, some unknown mutations might compensate for the attenuated viral fitness induced by P32del and enable the virus to persist.
Regarding the drug susceptibility analysis, we evaluated the resistance of these RASs introduced viruses to NS5A inhibitors (LDV, EBR, and VEL) and other anti-HCV reagents (SOF, RBV, and GZR). JFH1/5AC-wt was susceptible to the NS5A inhibitors with EC50 values ranging from 1.1 to 3.9 pM (Fig. 3 and Table 1). The observed values were consistent with previous reports38,39,40,41. However, viruses harboring the novel RASs of A92K or P32del exhibited high resistance to the conventional NS5A inhibitor, LDV. The EC50 values of these viruses to LDV were significantly higher than that of JFH1/5AC-Y93H or -L31V/Y93H reported in our previous paper21. The resistance levels of viruses with these RASs to the second generation NS5A inhibitors VEL and EBR were varied. JFH1/5AC-Y93H exhibited low-level resistance to EBR and no resistance to VEL in comparison with JFH1/5AC-wt. The addition of L31V to Y93H resulted in enhanced resistance and the EC50 value was increased to greater than 103-fold that of JFH1/5AC-wt. The highest EC50 values for these reagents were observed in JFH1/5AC-L31F/P32del followed by -P32del. Resistance due to A92K alone was not potent but higher than that of Y93H, and the EC50 value of this virus was lower than that of JFH1/5AC-P32del. The combination of R30Q with A92K enhanced the resistance levels to NS5A inhibitors, but the EC50 value of this virus was still lower than that of JFH1/5AC-P32del and comparable to JFH1/5AC-L31V/Y93H. In clinical studies, the detection of a single A92K RAS is very rare in both treatment-naïve and treatment-experienced patients19,33. In treatment-experienced patients, A92K is often observed with R30Q or with Q24K/L28M/R30Q, whereas the detection of L31F/M/V/I or Y93H with A92K is very rare19,33. RASs of L28M or R30Q were often detected in approximately 10% of treatment naïve patients9,29,33. Thus, A92K is important for patients infected with viruses harboring L28M and/or R30Q but without L31F/M/V/I and/or Y93H viruses as an additional RAS that confers increased resistance to NS5A inhibitors. In contrast, a single mutation of P32del conferred marked resistance to NS5A inhibitors. As expected, none of the DAA-experienced patients with P32del could be rescued by combination therapy with the potent NS5A inhibitor pibrentasvir and the second generation protease inhibitor glecaprevir31,42.
Regarding anti-HCV reagents other than NS5A inhibitors, no difference was observed in susceptibility to SOF and RBV between viruses with NS5A RASs and without RASs (Fig. 4 and Table 2). Interestingly, we found that JFH1/5AC-R30Q/A92K was more susceptible to GZR than JFH1/5AC-wt as well as JFH1/5AC-Y93H and -L31V/Y93H. This suggests that GZR is a good treatment option for viruses harboring R30Q/A92K. Similarly, we previously demonstrated that viruses with Y93H were more susceptible to the NS3/4A protease inhibitors, simeprevir and asunaprevir21. Intriguingly, both viruses harboring R30Q/A92K and Y93H enhanced the production of infectious virus particles. There may be some association between high virus production and the strong effect of protease inhibitors. HCV NS3 protein was reported to be essential for viral assembly in cooperation with NS5A or other virus proteins43,44,45. Thus, protease inhibitors might have various levels of inhibitory effect on virus assembly processes through the NS3 protein that follows the recruitment of NS5A protein to produce virus particles. Because the recombinant viruses used in this study contained the NS3 of genotype 2a and NS5A of genotype 1b, it is unclear whether this result is applicable to genotype 1b viruses. Confirmation by clinical observation is required to verify these in vitro findings and future studies should determine the mechanisms involved in the anti-viral effects of protease inhibitors to these viruses. Taken together with our susceptibility analysis, treatment with SOF, RBV, or GZR may eradicate viruses harboring P32del or A92K.
In conclusion, our experiments using JFH1 based NS5A chimeric recombinant viruses clearly demonstrated that R30Q/A92K substitution in NS5A contributes to enhanced virus propagation and that this might cause persistent infection similar to the Y93H variant. To treat this virus, our data suggests a DAA regimen containing GZR will be a useful option. Furthermore, viruses with a deletion at P32 were strongly resistant to all NS5A inhibitors including VEL. However, the propagation levels of these variants were very low and they were susceptible to SOF, RBV, and GZR. Combination regimens containing these anti-HCV reagents other than NS5A inhibitors will be better options for treating patients infected with viruses harboring P32del.
Materials and Methods
Cell Culture
The HuH-7-derived cell lines Huh-7.5.1 (provided by Francis Chisari, Scripps Research Institute, La Jolla, CA, USA) and Huh7-25 were cultured in Dulbecco’s Modified Eagle’s Medium supplemented with 10% fetal bovine serum essential medium at 37 °C in a 5% CO2 as described previously46.
Plasmid Construction
We used the JFH-1 based chimeric virus which NS5A was replaced with that from the Con1 strain (JFH1/5AC-wt) as described previously21,22. In this study, we focused on two novel RASs in NS5A, A92K and P32del. These RASs were not detected in DAA naïve patients but emerged at the time of the virologic failure of DAA treatment18. The RAS of A92K has been reported to emerge in combination with R30Q or with Q24K/L28M/R30Q19,33. Therefore, we made three variants of A92K by site-directed mutagenesis with appropriate primers as follows: JFH1/5AC-A92K, -R30Q/A92K and -Q24K/L28M/R30Q/A92K. The emergence of P32del was reported alone or in combination with L31F19. Therefore, we made two variants of P32del as follows: JFH1/5AC-P32del and - L31F/P32del. We also used the JFH-1 based subgenomic firefly luciferase reporter replicon with the NS5A from the Con1 strain (JFH1/5ACRep-wt), and constructed subgenomic reporter replicons with the RASs described above.
RNA Transfection and Quantification of the HCV Core Antigen
RNA transfection and quantification of the HCV core Ag were performed as described previously21. Briefly, in vitro transcribed HCV RNAs were electroporated into cells. The transfected cells and the supernatants were harvested and the concentration of the HCV core Ags were quantified using a Lumipulse Ortho HCV Ag kit (Ortho Clinical Diagnostics, Tokyo, Japan) according to the manufacturer’s instructions. In vitro RNA synthesis and electroporation were performed as described previously21,46.
Susceptibility Analysis for Antiviral Reagents
Susceptibility analysis for antiviral reagents were performed as described previously21. Briefly, in vitro transcribed HCV RNAs were electroporated into Huh-7.5.1 cells. Four hours after electroporation, transfected cells were treated with various concentrations of anti-HCV reagents. After incubation for 72 hours, the supernatants of the cells were harvested and the HCV core Ags were quantified. The anti-HCV reagents used were as follows: ribavirin (RBV) (Sigma-Aldrich, St. Louis, MO, USA), ledipasvir (LDV) (GS5885; AdooQ Bioscience, Irvine, CA, USA), elbasvir (EBR) (Chem Scene, Monmouth Junction, NJ, USA), velpatasvir (VEL) (Chem Scene), grazoprevir (GZR) (MK-5172; Chem Scene), and sofosbuvir (SOF) (PSI-7977; Medchemexpress LLC, NJ, USA). Dose-response curves were fitted using GraphPad Prism 7 software (GraphPad Software, Inc., CA, USA) and the values of the effective concentrations required to inhibit 50% (EC50) were determined with the results of the extracellular HCV core Ag quantification. The fitted model is as follows: Y = 100/(1 + 10((X-LogIC50))), where X is the concentration of the reagent and Y is the response expressed as the percentage of HCV core Ag related to the corresponding control.
Titration of HCV Infectivity and Indirect Immunofluorescence Analysis
The HCV infectivity titers were measured by indirect immunostaining as described previously21,46. For the indirect immunofluorescence analysis, the cells were fixed with cold methanol and incubated with an anti-core antibody (clone C7–50; Abcam, Cambridge, UK) and subsequently with fluorescent labeled secondary antibody. The infectivity titer was calculated the focus-forming units per mL and expressed as relative to JFH1/5AC-wt.
Subgenomic Replicon Analysis
In vitro transcribed HCV RNAs of subgenomic replicons were electroporated into Huh-7.5.1 cells. Four hours after transfection, the culture media were replaced with fresh media. After incubation for 3 days, cells were harvested and a luciferase assay was performed according to the manufacturer’s instructions (Bright-Glo™ Luciferase Assay System, Promega).
Statistical Analysis
Statistical analysis was performed by two-way ANOVA with Dunnett’s multiple comparisons test. The p-values were determined in comparison with the data of JFH1/5AC-wt. P-values less than 0.05 were considered statistically significant.
References
Webster, D. P., Klenerman, P. & Dusheiko, G. M. Hepatitis C. Lancet 385, 1124–1135 (2015).
Gower, E., Estes, C., Blach, S., Razavi-Shearer, K. & Razavi, H. Global epidemiology and genotype distribution of the hepatitis C virus infection. J. Hepatol. 61, S45–S57 (2014).
Pawlotsky, J. M. New Hepatitis C Therapies: The Toolbox, Strategies, and Challenges. Gastroenterology 146, 1176–1192 (2014).
Zoulim, F. et al. Hepatitis C virus treatment in the real world: optimising treatment and access to therapies. Gut 64, 1824–1833 (2015).
AASLD-IDSA HCV Guidance Panel. Hepatitis C Guidance 2018 Update: AASLD-IDSA Recommendations for Testing, Managing, and Treating Hepatitis C Virus. Infection. Clin. Infec. Dis. 67, 1477–1492 (2018).
Asahina, Y. et al. JSH Guidelines for the Management of Hepatitis C Virus Infection: A 2016 update for genotype 1 and 2. Hepatol. Res. 46, 129–165 (2016).
European Association for the Study of the Liver. EASL Recommendations on Treatment of Hepatitis C 2018. J. Hepatol. 69, 461–511 (2018).
Pawlotsky, J. M. NS5A inhibitors in the treatment of hepatitis C. J. Hepatol. 59, 375–382 (2013).
Sarrazin, C. The importance of resistance to direct antiviral drugs in HCV infection in clinical practice. J. Hepatol. 64, 486–504 (2016).
Lontok, E. et al. Hepatitis C virus drug resistance–associated substitutions: State of the art summary. Hepatology 62, 1623–1632 (2015).
Kumada, H. et al. Daclatasvir plus asunaprevir for chronic HCV genotype 1b infection. Hepatology 59, 2083–2091 (2014).
Welzel, T. M. et al. Global epidemiology of HCV subtypes and resistance-associated substitutions evaluated by sequencing-based subtype analyses. J. Hepatol. 67, 224–236 (2017).
Karino, Y. et al. Characterization of virologic escape in hepatitis C virus genotype 1b patients treated with the direct-acting antivirals daclatasvir and asunaprevir. J. Hepatol. 58, 646–654 (2013).
Pawlotsky, J.-M. Hepatitis C Virus Resistance to Direct-Acting Antiviral Drugs in Interferon-Free Regimens. Gastroenterology 151, 70–86 (2016).
Komatsu, T. E. et al. Regulatory Analysis of Effects of Hepatitis C Virus NS5A Polymorphisms on Efficacy of Elbasvir and Grazoprevir. Gastroenterology 152, 586–597 (2017).
Younossi, Z. M. et al. Sofosbuvir/velpatasvir improves patient-reported outcomes in HCV patients: Results from ASTRAL-1 placebo-controlled trial. J. Hepatol. 65, 33–39 (2016).
Feld, J. J. et al. Sofosbuvir and Velpatasvir for HCV Genotype 1, 2, 4, 5, and 6 Infection. N. Engl. J. Med. 373, 2599–2607 (2015).
Krishnan, P. et al. Integrated Resistance Analysis of CERTAIN-1 and CERTAIN-2 Studies in Hepatitis C Virus-Infected Patients Receiving Glecaprevir and Pibrentasvir in Japan. Antimicrob. Agents Chemother. 62, e02217–17, https://doi.org/10.1128/AAC.02217-17 (2018).
Itakura, J. et al. Complex Pattern of Resistance-Associated Substitutions of Hepatitis C Virus after Daclatasvir/Asunaprevir Treatment Failure. PLoS One 11, e0165339, https://doi.org/10.1371/journal.pone.0165339 (2016).
Doi, A. et al. Nonstructural protein 5A/P32 deletion after failure of ledipasvir/sofosbuvir in hepatitis C virus genotype 1b infection. Hepatology 68, 380–383 (2018).
Nitta, S. et al. Effects of Resistance-Associated NS5A Mutations in Hepatitis C Virus on Viral Production and Susceptibility to Antiviral Reagents. Sci. Rep. 6, 34652, https://doi.org/10.1038/srep34652 (2016).
Okamoto, Y. et al. Development of recombinant hepatitis C virus with NS5A from strains of genotypes 1 and 2. Biochem. Biophys. Res. Commun. 410, 404–409 (2011).
Akazawa, D. et al. CD81 Expression Is Important for the Permissiveness of Huh7 Cell Clones for Heterogeneous Hepatitis C Virus Infection. J. Virol. 81, 5036–5045 (2007).
Zeuzem, S. et al. NS5A resistance-associated substitutions in patients with genotype 1 hepatitis C virus: Prevalence and effect on treatment outcome. J. Hepatol. 66, 910–918 (2017).
Harrington, P. R. et al. Impact of hepatitis C virus polymorphisms on direct‐acting antiviral treatment efficacy: Regulatory analyses and perspectives. Hepatology 67, 2430–2448 (2018).
Dietz, J. et al. Patterns of Resistance-Associated Substitutions in Patients With Chronic HCV Infection Following Treatment With Direct-Acting Antivirals. Gastroenterology 154, 976–988 (2018).
Paolucci, S. et al. Baseline and Breakthrough Resistance Mutations in HCV Patients Failing DAAs. Sci. Rep. 7, 16017, https://doi.org/10.1038/s41598-017-15987-1 (2017).
Wyles, D. et al. Post-treatment resistance analysis of hepatitis C virus from phase II and III clinical trials of ledipasvir/sofosbuvir. J. Hepatol. 66, 703–710 (2017).
Sarrazin, C. et al. Prevalence of Resistance-Associated Substitutions in HCV NS5A, NS5B, or NS3 and Outcomes of Treatment With Ledipasvir and Sofosbuvir. Gastroenterology 151, 501–512 (2016).
Iio, E. et al. Clinical evaluation of sofosbuvir/ledipasvir in patients with chronic hepatitis C genotype 1 with and without prior daclatasvir/asunaprevir therapy. Hepatol. Res. 47, 1308–1316 (2017).
Poordad, F. et al. Glecaprevir/Pibrentasvir in patients with hepatitis C virus genotype 1 or 4 and past direct-acting antiviral treatment failure. Hepatology 67, 1253–1260 (2018).
Ng, T. I. et al. In Vitro Antiviral Activity and Resistance Profile of the Next-Generation Hepatitis C Virus NS5A Inhibitor Pibrentasvir. Antimicrob. Agents Chemother. 61, e02558–16, https://doi.org/10.1128/AAC.02558-16 (2017).
Mawatari, S. et al. New resistance-associated substitutions and failure of dual oral therapy with daclatasvir and asunaprevir. J. Gastroenterol., 1–13 (2017).
Fridell, R. A. et al. Distinct functions of NS5A in hepatitis C virus RNA replication uncovered by studies with the NS5A inhibitor BMS-790052. J. Virol. 85, 7312–7320 (2011).
Wang, C. et al. Persistence of resistant variants in hepatitis C virus-infected patients treated with the NS5A replication complex inhibitor daclatasvir. Antimicrob. Agents Chemother. 57, 2054–2065 (2013).
Yoshimi, S. et al. Long term persistence of NS5A inhibitor-resistant hepatitis C virus in patients who failed daclatasvir and asunaprevir therapy. J. Med. Virol. 87, 1913–1920 (2015).
Jeong, Y. et al. Evolution and persistence of resistance-associated substitutions of hepatitis C virus after direct-acting antiviral treatment failures. J. Viral. Hepat. 25, 1251–1259 (2018).
Cheng, G. et al. In Vitro Antiviral Activity and Resistance Profile Characterization of the Hepatitis C Virus NS5A Inhibitor Ledipasvir. Antimicrob. Agents Chemother. 60, 1847–1853 (2016).
Liu, R. et al. Susceptibilities of Genotype 1a, 1b, and 3 Hepatitis C Virus Variants to the NS5A Inhibitor Elbasvir. Antimicrob. Agents Chemother. 59, 6922–6929 (2015).
Lawitz, E. J. et al. Clinical Resistance to Velpatasvir (GS-5816), a Novel Pan-Genotypic Inhibitor of the Hepatitis C Virus NS5A Protein. Antimicrob. Agents Chemother. 60, 5368–5378 (2016).
Gottwein, J. M. et al. Efficacy of NS5A Inhibitors Against Hepatitis C Virus Genotypes 1–7 and Escape Variants. Gastroenterology 154, 1435–1448 (2018).
Kumada, H. et al. Efficacy and safety of glecaprevir/pibrentasvir in HCV-infected Japanese patients with prior DAA experience, severe renal impairment, or genotype 3 infection. J. Gastroenterol. 53, 566–575 (2018).
Ma, Y., Yates, J., Liang, Y., Lemon, S. M. & Yi, M. NS3 Helicase Domains Involved in Infectious Intracellular Hepatitis C Virus Particle Assembly. J. Virol. 82, 7624–7639 (2008).
Lai, C.-K., Jeng, K.-S., Machida, K. & Lai, M. M. C. Association of Hepatitis C Virus Replication Complexes with Microtubules and Actin Filaments Is Dependent on the Interaction of NS3 and NS5A. J. Virol. 82, 8838–8848 (2008).
Isken, O. et al. A Conserved NS3 Surface Patch Orchestrates NS2 Protease Stimulation, NS5A Hyperphosphorylation and HCV Genome Replication. PLoS Pathog. 11, e1004736, https://doi.org/10.1371/journal.ppat.1004736 (2015).
Kato, T. et al. Cell culture and infection system for hepatitis C virus. Nat. Protoc. 1, 2334–2339 (2006).
Acknowledgements
This work was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology-Japan, the Japan Society for the Promotion of Science (16H05285, 17K19647), Japan Agency for Medical Research and Development (JP18fk0210002, JP18fk0210025 and JP18fk0210021).
Author information
Authors and Affiliations
Contributions
S. Nitta performed the experiments. S.N. and Y. Asahina planned this study. T. Kato., S. Kakinuma, Y.A. and M. Watanabe supervised various aspects of the study. S.N., T.K., S.K., J. Tsuchiya, A. Sato, M. Miyoshi, E. Inoue-Shinomiya, T. Tsunoda, F.K.-Kitahata, M. Murakawa, Y. Itsui, M. Nakagawa, S. Azuma, H. Hikita and T. Takehara discussed the strategy and the results. S.N., T.K. and Y.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
Dr. Yasuhiro Asahina and Dr. Sei Kakinuma belong to a donation-funded department funded by Toray Industries Inc., Gilead Sciences, AbbVie GK, Chugai Pharmaceutical Co. Ltd., Fujirebio Inc. and Merck Sharp & Dohme. Dr. Mamoru Watanabe received research support from the following, Kyorin Pharmaceutical Co., Ltd., Mitsubishi Tanabe Pharma Co., Ltd., Takeda Pharmaceutical Co., Ltd., Eisai Co., Ltd., Otsuka Pharma Co., Ltd., Kyowa Hakko Kirin Co., Ltd., Zeria Pharmaceutical Co., Ltd., JIMRO Co., Ltd., Asahi Kasei Kuraray Medical Co., Ltd., EA Pharma Co., Ltd., AbbVie GK, Astellas Pharma Inc., Daiichi Sankyo Co., Ltd., Toray Industries, Inc, MSD K.K., Chugai Pharmaceutical Co., Ltd., Gilead Sciences, Nippon Kayaku Co., Ltd., Mochida Pharmaceutical Co., Ltd., Taiho Pharmaceutical Co., Ltd., Miyarisan Pharmaceutical Co., Ltd., Ayumi Pharmaceutical Co., Shionogi & Co., Ltd. Dr. Tetsuo Takehara has received grants from Chugai Pharmaceutical Co. Ltd., Bristol‐Myers Squibb, MSD K.K., AbbVie Inc. and Gilead Sciences. Dr. Tetsuo Takehara is on the speakers’ bureau for Bristol‐Myers Squibb, MSD K.K., AbbVie Inc. and Gilead Sciences. Dr Hayato Hikita has received grants from MSD K.K.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nitta, S., Asahina, Y., Kato, T. et al. Impact of novel NS5A resistance-associated substitutions of hepatitis C virus detected in treatment-experienced patients. Sci Rep 9, 5722 (2019). https://doi.org/10.1038/s41598-019-42114-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42114-z
This article is cited by
-
Analysis of the susceptibility of refractory hepatitis C virus resistant to nonstructural 5A inhibitors
Scientific Reports (2024)
-
Mutational spectrum of hepatitis C virus in patients with chronic hepatitis C determined by single molecule real-time sequencing
Scientific Reports (2022)
-
Real-word efficacy of sofosbuvir, velpatasvir plus ribavirin therapy for chronic hepatitis patients who failed to prior DAA therapy with NS5A-P32 deletion mutated HCV infection
Clinical Journal of Gastroenterology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.