Abstract
The lysine-specific demethylase 2A gene (KDM2A) is ubiquitously expressed and its transcripts consist of several alternatively spliced forms, including KDM2A and the shorter form N782 that lacks the 3′ end encoding F-box and LRR. KDM2A binds to numerous CpG-rich genomic loci and regulates various cellular activities; however, the mechanism of the pleiotropic function is unknown. Here, we identify the mechanism of KDM2A played by its CXXC-PHD domain. KDM2A is necessary for a rapid proliferation of post-natal keratinocytes while its 3′ end eclipses the stimulatory effect. EGFP-N782 binds to chromatin together with the XRCC5/6 complex, and the CXXC-PHD domain regulates the CpG-rich IGFBPL1 promoter. In vitro, CXXC-PHD enhances binding of nuclear extract ORC3 to the CpG-rich promoter, but not to the AT-rich DIP2B promoter to which ORC3 binds constitutively. Furthermore, CXXC-PHD recruits 94 nuclear factors involved in replication, ribosome synthesis, and mitosis, including POLR1A to the IGFBPL1 promoter. This recruitment is unprecedented; however, the result suggests that these nuclear factors bind to their cognate loci, as substantiated by the result that CXXC-PHD recruits POLR1A to the rDNA promoter. We propose that CXXC-PHD promotes permissiveness for nuclear factors to interact, but involvement of the XRCC5/6 complex in the recruitment is undetermined.
Similar content being viewed by others
Introduction
Dozens of protein demethylases are found in human and mammalian cells, and the demethylases are currently classified into 9 groups (KDM1 through KDM9) in the UniProt database, based on the type of both demethylase domain and other domains present in the molecule1,2. KDM2A is a lysine-specific demethylase that removes methyl groups from H3K36me2 and H3K36me1 coupling with cofactors Fe (II) and α-ketoglutarate3. It consists of JmjC, CXXC, PHD, F-box, and LRR domains (Fig. 1a). Many reports of the cellular function of KDM2A have been published since it was discovered a decade ago as the H3K36me specific demethylase. However, the fundamental role of KDM2A remains puzzling. The slow progress of investigation on KDM2A is in sharp contrast to the solid progress of that on KDM2B, which is highly similar to KDM2A and found to recruit a variant Polycomb Repressive Complex 1 (PRC1) to certain CpG Islands (CGIs) to silence the gene expression4.
Requirement of N782 for proliferation of keratinocytes. (a) Illustration of KDM2A and N782. C, P, N, and LRR indicate CXXC, PHD, NLS, and leucine repeat region, respectively. (b) Expression of N782 and KDM2A in 7-day cultivated keratinocytes at different densities of 3T3-J2. Blue and red indicate N782 and KDM2A, respectively, throughout Fig. 1. p = 0.0095. (c) Keratinocyte proliferation under the same conditions. (d) Proportional relationship of N782 to the cell proliferation rate. Mean ± SD. (e) Keratinocyte proliferation transduced with N782, KDM2A, and the empty vector (green). p = 0.0043 (N782 vs. the control) and 0.0249 (KDM2A vs the control). (f) Expression of N782 and KDM2A in the presence and absence of 3T3-J2 in EpiLife medium. p = 0.0128. (g) Expression of N782 and KDM2A in hESC. *In Fig. 1 shows a significant difference (P < 0.05) by either ANOVA or t-test. n = 3 for (b–f), 2 for (e), and 6 for (g).
Results of KDM2A in previous reports can be summarized in two parts: first, KDM2A binds via the CXXC domain to thousands of non-methylated CGIs that compose the majority of promoters5,6,7; second, KDM2A is involved in a variety of cellular activities such as cell division, proliferation, development, DNA repair, circadian rhythm, and cancer8,9,10,11,12,13,14,15,16,17,18. However, underlying mechanisms of KDM2A for these cellular activities are virtually unknown as the results from different groups are often conflicting. The discrepancy is especially explicit in the role on cell proliferation: some reports show that KDM2A promotes cell proliferation, while others show that it suppresses proliferation. The discrepancy appears to be derived from both multiple KDM2A isomers and various cells used by different groups such as normal somatic, cancer, and manipulated cells8,12,19,20. Cell cycle arrest at G1 phase by KDM2A knockdown of human apical papilla stem cells10 suggests that KDM2A is essential for human cell proliferation, and at the same time the results reveal that investigation of KDM2A is neither trivial nor simple.
We first encountered KDM2A during complementation experiments of poorly growing keratinocytes derived from human embryonic stem cells (hESC) H98, and subsequently found that N782, which is a shorter isomer of KDM2A, consisting of the first 782 amino acid sequence (Fig. 1a), improves the poor proliferation, but the full-length KDM2A does not. This finding indicated that N782 has domains to stimulate cell proliferation and KDM2A has additional domains antagonizing N782 function. Current research demonstrates that N782 is necessary for rapid cell proliferation and the CXXC-PHD domain enhances recruitment of nuclear factors necessary for proliferation. We begin the investigation with characterizing N782 as a transcript necessary for rapid proliferation of post-natal keratinocytes.
Results
N782 is required for rapid proliferation of post-natal keratinocytes
The N782 mRNA transduced with the aid of a retrovirus vector improved the poor proliferation of hESC-derived keratinocytes but the full length KDM2A mRNA did not8. This result demonstrated that N782 stimulates keratinocyte proliferation but did not prove that the transcript is required for the proliferation. If N782 were necessary for proliferation of keratinocytes, low expression of the transcript would cause slow proliferation of cells. Therefore, we attempted to find culture conditions to lower N782 in post-natal keratinocytes (YF29). Subsequently, we found that lower densities of lethally irradiated 3T3-J2 feeders compared to the standard density decrease N782. As shown in Fig. 1b, the N782 was reduced by 60% at a density of 0.2 × 104/cm2 of 3T3-J2 than 5 × 104/cm2, whereas the total KDM2A was unaffected. Simultaneously, proliferation of keratinocytes was reduced (Fig. 1c); as a result, the proliferation rate was proportional to the expression of N782 (Fig. 1d). Moreover, ectopic expression of N782 improved the slow proliferation to a greater extent in the cultures containing lower density of 3T3-J2 (Fig. 1e). On the other hand, KDM2A’s impact was more modest under the culture conditions. Thus, expression of the KDM2A gene is necessary for rapid proliferation of keratinocytes, and N782 plays the major role for the gene function. EpiLife medium is a feeder-free medium, but 3T3-J2 still increased N782 in the medium (Fig. 1f), thereby providing further evidence that 3T3-J2 enhances the N782 level. Importantly, hESC expressed N782 no less than keratinocytes (Fig. 1g), suggesting that the expression mechanism and the role of N782 are epigenetically transmitted to adult stem cells.
The N782 protein binds to chromatin with the XRCC5/6 complex
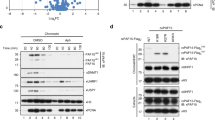
To shed light on mechanisms involving N782, we carried out co-immunoprecipitations using an antibody to GFP and DNase-untreated cell lysates of keratinocytes transfected with pEGFP-N782. The immunoprecipitated sample was resolved by SDS-PAGE and stained with Coomassie blue (Fig. 2a left panel). It contained 2 protein bands in the range of 70–90 kDa besides EGFP-N782. The same protein bands were obtained from lysates of keratinocytes/pEGFP-KDM2A. However, the bands were weaker than those from keratinocytes/pEGFP-N782; and the band of EGFP-KDM2A itself was also weaker than that of EGFP-N782. Western blotting analysis of the similarly prepared sample also presented unbalanced-amount bands (EGFP-KDM2A < EGFP-N782) (Fig. 2a right panel). Therefore, we conjectured that lower yield of EGFP-KDM2 was derived from lower expression of the protein in keratinocytes, and turned our attention to KDM2A C-terminus that is not present in N782. Fusion of either the C-terminus (C387) with F-box and LRR or the C-terminus (C233) with LRR to EGFP strongly reduced green fluorescence in cells and also reduced immunoprecipitation yield of EGFP (Fig. 2a right panel). As F-box and LRR form the SCF (Skp1, Cullin, and F-box) ubiquitin-ligase complex21, the C-termini seem to have targeted either its own N-terminus (EGFP) or an unknown, but KDM2A-associated, activator. This result suggests that instability of KDM2A or lowered activators makes KDM2A ineffective in stimulating keratinocyte proliferation. Note that immunoprecipitates from keratinocytes/EGFP-N782 contained genomic DNA fragments (Fig. 2b). This means that immunoprecipitates contained proteins that interacted with N782 either directly or indirectly. Mass spectrometry of the precipitates from keratinocytes/pEGFP-N782 revealed that 70–90 kDa bands contained a pair of the X-ray repair cross-complementing protein 5/6 (XRCC5/6) complex (Fig. 2c red circle), various chaperones including HSPA5, HSPA8, and HSP90AA1, nuclear-envelope proteins LMNA (blue circle), and TMPO (blue circle) as the major proteins. The last 2 proteins indicate that N782 can bind to CGIs on boarders of lamina-associated domains (LAD)22. Although the immuneprecipitate also contained bands in the range of 12–14 kDa, we did not analyze them further to avoid complications resulting from small molecular mass chromatin proteins. Unlike previous reports14,23, neither E2F1 nor SUZ12 was found in the immunoprecipitate.
Interaction of N782 with the XRCC5/6 complex. * Indicates EGFP tag. (a) Left panel shows proteins immunoprecipitated with a rabbit anti-GFP antibody (black circle) and co-immunoprecipitated proteins (arrow). It contains both rabbit IgG heavy chain (50 kD wide band) and the light chain (25 kD wide band) derived from the anti-GFP antibody. Right panel shows Western blots of immunoprecipitated EGFP-KDM2A, EGFP-N782, and EGFP-KDM2A C-termini. The proteins were detected by a combination of mouse anti-GFP (primary) and goat anti-mouse IgG. (b) Keratinocyte genomic DNA fragments co-immunoprecipitated with EGFP-N782 and EGFP. A marker (250 or 500 ng) or sample (5 μl) was applied to each well, respectively. (c) Mass spectrometry of proteins included in the 70–90 kDa protein bands [proteins from top to bottom, (XRCC5 and XRCC6 in red) and (TMPO and LMNA in blue)]. (d) Western blotting analysis of XRCC5 (arrow) co-immunoprecipitated with N782 or the mutants (circle): Δ = deletion, C = CXXC, J = JmjC, and P = PHD. (e) Binding of GST-CN, GST-PN, and GST-NLSR to 3-length IGFBPL1 baits. Full-length blots are presented in Supplementary Fig. S1. (f) Far Western blotting analysis of the interaction of KMD2A domains with XRCC5 and 6. Probe and protein dye are on the left; cross-reaction of antibody to an E. coli protein, > and <. Full-length blots are presented in Supplementary Fig. S2. (g) Amino acid sequence of CPN domains expressed in cells (CPN, 561V - R733; CN, 561V- T619 plus 676Y - R733; PN, 619T - R733; NLSR, 676Y - R733). Typically defined domains C (red) and P (violet), and the N (blue). We have defined N by deletion analysis (unpublished).
As the XRCC5/6 complex is recruited together with several other proteins to the pS2 promoter of breast cancer MCF-7 cells treated with 17β-estadiol24, co-recruitment of the XRCC5/6 complex does not seem to be unique to our result and rather the complex seems to be important for various regulatory elements. Accordingly, the N782-XRCC5/6 interaction was investigated further by deletion of the domains. Expression of mutant proteins was consistently lower than that of EGFP-N782 and therefore the yield of their immunoprecipitation was also lower, especially lower in JmjC deletion mutant proteins. Nonetheless, EGFP-N782ΔJmjC exhibited molecular binding activity for XRCC5 (XRCC5/EGFP-N782ΔJmjC) at a similar level as the parent protein, whereas EGFP-N782ΔCXXC completely lost the binding activity (Fig. 2d). Thus, CXXC is responsible for the interaction with the XRCC5/6 complex. Taking into account that CXXC primarily binds to CpG-rich DNA (Fig. 2e), the results suggest that N782 indirectly binds to the XRCC5/6 complex via the CXXC-CpG interaction. However, the possibility that N782 directly interacts with the XRCC5/6 complex on the chromatin still remains.
To investigate the direct interaction, size-exclusion chromatography of proteins was carried out with the purified XRCC5/6 complex alone, purified GST-CXXC-PHD-NLSR (GST-CPN) alone (NLSR, nuclear localization signal region), and a mixture of both. The result was inconclusive owing to unusual exclusion of the XRCC5/6 proteins (N. Sekiyama, personal communication). Accordingly, we took a Far Western blotting approach for the investigation by using a high concentration of XRCC5 and XRCC6 immobilized on a nitrocellulose membrane (Fig. 2f; see Fig. 2g for domains of N782). GST-domain fusion proteins bound to XRCC6 as long as the proteins contained the CXXC domain. On the other hand, both GST-CXXC-NLSR (GST-CN) and GST-PHD-NLSR (GST-PN) weakly bound to XRCC5, and GST-CPN further weakly bound to it as if CP tertiary structure inhibited the binding. These results suggest that CP can interact with the XRCC5/6 complex by alternative ways, formation of a heterotrimer with the XRCC5/6 complex via association with XRCC6 and expulsion of the complex via repelling of XRCC5.
CP regulates gene expression at a CpG-rich promoter but not at a CpG-poor promoter
To pursue the role of N782 and the CP domain at genomic CpG loci, we unbiasedly isolated pieces of genomic DNA by binding activity of immobilized KDM2A CXXC domain, and subsequently isolated the 312-bp promoter of the insulin-like growth factor-binding protein-like 1 (IGFBPL1) gene as a paradigm of CpG sequence. Selection criteria for the sequence were: (1) it is a highly CpG-rich sequence; (2) it is a short promoter sequence allowing us to easily monitor binding of the CP domain and its outcome; and (3) it is bound by CP both in vivo and in vitro. The promoter of the IGFBPL1 gene best fitted the criteria (Fig. 3a) among the sequences that we obtained by the enrichment. Therefore, we cloned the promoter region (312 bp) from the IGFBPL1 gene, and fused it to the reporter plasmid pGluc Basic 2 (Fig. 3a). The resulting construct, pIGFBPL1-312-Gluc, was transfected into HEK293T cells (instead of inefficiently transfectable keratinocytes) and the luciferase activity was assessed. The construct induced 2.5–4 fold more luciferase activity than the empty vector, confirming that the 312-bp DNA includes the promoter although the expression was not especially high. This low expression may be associated with the character of CGIs genes proximal to LAD22 or with competition of various nuclear proteins for the promoter that we describe later. By using pIGFBPL1–312-Gluc, we then investigated which domains interact with the 312 bp sequence. As shown in Fig. 3c, co-transfected pEGFP-NLSR, which additionally codes the KDM2A/N-782 N-terminal 80 amino acid sequence as a linker between EGFP and NLSR, slightly reduced the luciferase activity. However, pEGFP-KDM2A substantially reduced it and furthermore pEGFP-N782 did so even more strongly. Subsequently, we found that pEGFP-CPN could severely repress the CpG-rich promoter. Although pEGFP-CN also strongly repressed it, the repression was alleviated by removal of the N-terminal EGFP (Fig. 3e). Thus, CP binds and regulates the IGFBPL1 promoter. pEGFP-PN alone had no effect on the expression as expected from the result that PN did not bind to the GFBPL1 promoter (Figs 2e, 3c).
Promoters of the IGFBPL1 and the DIP2B genes and their characterization. Promoters of both IGFBPL1 (a) and DIP2B (b) are underlined with broken line for DNase I hypersensitive site according to ENCODE data of UCSC browser. The 5′ 67-bp fragment of the DIP2B consisting of a few CpGs that were bound by CXXC during the affinity purification of genomic fragments. CpG, red); and A, T, and AT stretches, blue. (c) Luciferase activity expressed from pIGFBPL1-312-GLuc (blue) and the empty vector pGLuc Basic 2 (red) in HEK293T cells that were co-transfected with effector plasmids encoding N782 and the domains. The activity of the culture co-transfected with pIGFBPL1-312-GLuc and pEGFP-NLSR is defined as 1.0. Small * shows EGFP. (d) Luciferase activity expressed from pDIP2B-557-Gluc and the empty vector. (e) The same-type experiment as in (c) but co-transfected effector plasmids were free of the EGFP tag sequence. Large * shows a significant difference by t-test (p < 0.05, n = 4).
To compare behavior of CP at the CpG-rich promoter with that at a CpG-poor promoter, we selected the disco interacting protein2 homolog B (DIP2B) gene from the collection made by affinity chromatography (see Fig. 3b legend) and fused the 557-bp promoter to the Gluc reporter gene (Fig. 3b). The first 312-bp region of the promoter contained only 11 CpG and had many A, T and AT stretches. The resulting construct, pDIP2B-Gluc-557, gave rise to 10-fold higher luciferase activity than the empty vector (Fig. 3d). The activity was strongly decreased by pEGFP-NLSR presumably via involvement of KDM2A/N782 N-terminal 80 sequence, but the residual activity was not reduced further by pEGFP-CPN. Thus, the CP domain does not have a role in regulation of the CpG-poor DIP2B promoter, but it has separate regulation.
CP stimulates binding of ORC3 to a specific site of the IGFBPL1 promoter
CpG-rich promoters provide not only gene-specific transcription factors, but also Origin Recognition Complex (ORC) with binding sites25,26. In fact, ORC binds to as many as 13,500 human CpG-rich promoters27. On the other hand, KDM2A binds to roughly equivalent number of CGIs around promoters in mouse cells5. Therefore, the IGFBPL1 promoter gave us an excellent opportunity to investigate pleiotropic regulatory feature of KDM2A with ORC. To pursue this objective, we prepared nuclear extracts from HEK293T transfected with either pEGFP-CPN or the control plasmid pEGFP-NLSR, and mixed them with immobilized IGFBPL1-312 bait, respectively, and then quantified bound ORC3 by Western blotting. ORC3 bound to the bait at a slow time-dependent rate in the control extracts (Fig. 4a). However, ORC3 bound to the bait faster in the EGFP-CPN-containing extracts. In contrast, ORC3 very rapidly bound to AT-rich promoter of the DIP2B (Fig. 4b) and the binding was not enhanced by CP. Interestingly, EGFP-CPN also enhanced binding of ORC3 to a hybrid 312-bp DNA containing 5′ 69-bp of the IGFBPL1 bait (Fig. 4c), but not to the bait without the 69-bp sequence (Fig. 4d). Thus, the enhancement element was present in the 69-bp DNA of the IGFBPL1 or a sequence slightly longer than the 69-bp segment. Binding of EGFP-CPN to the baits (Fig. 4e and 4g) suggests that direct interaction of CP with the element enabled ORC3 to more efficiently bind.
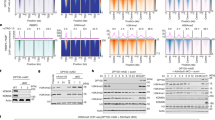
Binding kinetics of nuclear proteins to 4 different DNA baits in the presence and absence of EGFP-CPN. Investigated proteins: ORC3 in (a–d); EGFP-CPN and EGFP-NLSR in (e–h); XRCC5 (solid) and XRCC6 (dotted) in (i–l); H2 (dotted) and H3 (solid) in (m–p). Baits used for the binding are shown above the kinetics graphs along with the number of CpG (blue) counted from 5′ to 3′. *Indicates biotinylated 5′ end of baits. Green and orange kinetics indicate data obtained with EGFP-CPN-containing and EGFP-NLSR-containing nuclear extracts, respectively. Each point on the graphs represents an average of 2 independent kinetics data, but p-values of ANOVA test are calculated with all the independent points (not from the averages). Inserted cartoon suggests protein binding to the bait. Full-length blots for Fig. 4. (a through l) are presented in Supplementary Fig. S3.
Our earlier result hinted that the XRCC5/6 complex has an affinity for CpG-rich sequences (Fig. 2b,c). The notion was indeed demonstrated by the kinetics result that the XRCC5/6 complex bound to IGFBPL1-312 bait twice as much as to DIP2B-312 bait (Fig. 4i,j). Note that the XRCC5/6 complex is well established to bind to double-stranded DNA broken ends, regardless of the nucleotide composition (see inserted illustrations)28,29. Both H2 and H3 more slowly bound to the IGFBPL1-312 in the presence of EGFP-CPN than ORC3 did, suggesting that binding of histones does not interfere with binding of ORC3 to the IGFBPL1 promoter (Fig. 4a,m).
CP recruits various protein complexes essential for cell proliferation to CpG-rich promoters
During the investigation of ORC3 binding, we found that the protein-band profile of the IGFBPL1-312-interacting proteins in the presence of EGFP-CPN differed from that in the control extract on SDS-PAGE gel, and the altered profile similarly occurred with the shorter bait, IGFBPL1-171 (Fig. 5a). This shorter bait does not include the region corresponding to the POLRII initiation complex binding site (Fig. 3a)30. Accordingly, by using it, we could investigate the recruited protein species without involvement of the major transcription complex. Following analysis of the proteins by mass spectrometry and manual data curation from the protein group, we described the recruitment of each protein by the ratio of protein amounts bound in the presence and absence of EGFP-CPN to dissociate recruited proteins from the rest. The majority of ratios for all analyzed proteins (1017 proteins) was closed to or smaller than 1, suggesting that binding of most proteins was not enhanced or excluded by CP (Fig. 5b). However, a considerable number of proteins had this ratio greater than 1. We took a ratio of 2 as indication of binding enhancement as 1.8 was the ratio for ORC3 in the same reaction period (Fig. 4a), as such, we obtained the statistic values 0.92 (mean) ± 0.33 (SD) for a group of the proteins with less than ratio 2. With this value, we subsequently calculated that 127 of 1017 proteins have ratios greater than 1.58 (0.92 + 0.33 × 2) and 94 proteins have ratios greater than 2. Thus, at least 94 proteins bound more abundantly to the CpG-rich DNA by CP. The ratio of ORC2 was slightly lower than expected in this experiment, but the mean of 2 independent experiments was 1.75 and that for the ORC-stabilizing protein LRWD31 was 2.1. Thus, the result was consistent even if the enhancement was not substantial. The ratio of ORC4 and ORC5 was 2 and 1.3, respectively. CDC45, required for initiation of replication and interacting with ssDNA32, was 4-fold more highly bound to the bait by CP. However, ORC1 was not identified even in the nuclear extracts (Fig. 5c). It seems that the cultures did not sufficiently contain G1 phase cells that are known to express ORC133. MCM2-7 was virtually not recovered from the nuclear extracts (the ratio of MCM2, 3, 4, 5, 6, and 7 was 0, 0.75, 0, 0, 0, 0.41, respectively) even though all the MCM were present in the nuclear extracts. Likewise, GINS, CDC6, and CDT1 did not bind although these proteins were also present in the nuclear extracts. Taken together, our results show that ORC and the relevant proteins bind to cis elements by the action of CP during S phase without ORC1. The CP domain may replace MCM2-7 to build up the replication initiation complex and prevent second initiation in S phase34.
EGFP-CPN enhances nuclear protein binding to the IGFBPL1-171 bait. (a) SDS-PAGE analysis of proteins that bound to the bait with EGFP-CPN-containing (*CPN) and EGFP-NLSR-containing (*NLSR) nuclear extracts. Slice identification number is shown on the right. (b) Mass spectrometry analysis of the bound proteins. Y-axis is ratio of bound-protein between the presence and absence of EGFP-CPN, and X-axis is the slice number. Two stars (**) show the ratio for ORC3 (2.0). (c) Mass spectrometry analysis of the EGFP-CPN-containing and EGFP-NLSR-containing nuclear extracts. Y-axis is the same as in (b), and X-axis is the protein identification number. Two stars (**) show the ratio for ORC3 (1.03). (d) Alignment of promoters of the IGFBPL1 and the rRNA genes. Nucleotides are numbered from the 5′ end of our clone for the IGFBPL, while TSS is shown by 1 in the rRNA promoter. (e) Time course of POLR1A binding (arrowed) to rDNA-260 promoter bait in the presence and absence of EGFP-CPN. 800 channel intensity, 6.5, was used to scan the image. Full-length blots are presented in Supplementary Fig. S4. (f) Binding of XRCC5/6, ORC3, and EGFP-CPN to the rDNA bait with the same extracts. (g) Quantification data of (f). 700 and 800 channel intensities were set to 4.5 and 6.5 for the scanning, respectively. * Shows a significant difference by t-test. p = 0.04 for ORC3, 0.01 for XRCC6, and <0.001 for *CPN vs *NLSR, n = 4. (h) Roles of CP on proliferation of keratinocytes. YF29 cultures were transduced with CPN (blue), NLSR (red), N782 (green), and the empty vector (violet) and cultivated for up to 20 days on 0.2 × 104/cm2 3T3-J2 (duplicates). * Shows a significant difference by ANOVA test (p = 0.0030 for CPN vs NLSR, and 0.0002 for N782 vs the control. Cell density of each measuring point was presented by fold over that of cultures transduced with the empty vector.
Proteins most strongly recruited by CP included POLR1A, POLR1B (Table 1), and the activator proteins (TAF1A, TAF1B, and TAF1C), as well as nucleolus proteins involved in processing of the transcripts and ribosome synthesis35. In addition, GNL3L required for ribosome synthesis36 was also recruited. The TOP3B/TDRD3 complex, which processes pre-ribosomal RNA and also activates gene expression and protects against DNA damage by breaking down the R-loop37, was also recruited. Notably, both UBTF (ratio, 1.0) that interacts with POLR1 via TAF1B and NUCL (ratio, 0.65) that is required for the pre-RNA processing were not recruited by CP, suggesting that the recruited proteins of this group have either a strong affinity for POLR1A/POLR1B or a direct affinity for the IGFBPL1-171 bait by other means.
Proteins engaging in chromosome maintenance, congression, segregation, and progression of cell cycle were also highly recruited by CP. Both KIF18A and KIF2C work together to achieve the proper attachment of microtubles to the kinetochore, congression of chromosome on the spindle equator, and segregation of chromosome by depolymerizing microtubules and suppressing the process38,39,40,41. CLASP1 is a member of outer kinetochore proteins and important in the proper chromosome segregation processes along with TPX2 and KNSTRN42,43,44. Proteins in this class recruited by CP include CEP170, KIF18A, KIF23, KIF2C, KIFC1, and MARK2. The recruitment of all these factors explains the mechanisms how abnormal congression and segregation of chromosomes happened by mutations or knockdowns of KDM2A15.
Unexpectedly, CP reduced binding of the XRCC5/6 complex by half (XRCC5 was reduced to 0.52 and XRCC6 to 0.5). This result was reproduced with IGFBPL1-171 bait, but this result was not consistent with that of the IGFBPL1-312 bait (Fig. 4i). The difference between the two results can be reconciled by the observation that CP can either bind or expel the XRCC5/6 complex (Fig. 2f).
To ensure that CP enhanced the protein-DNA interaction but not nuclear protein concentration, we analyzed proteins present in the both nuclear extracts by tandem mass tag based mass spectrometry45. We confirmed that the recruited proteins, for example, in Table 1, were evenly present between the EGFP-CPN- and EGFP-NLSR-containing extracts and less in the former extracts (Fig. 5c). Exceptionally, however, a very small fraction of a total of 7811 proteins contained proteins that were more abundant in the EGFP-CNP-containing extracts. The fraction included STMN1 (Stathmin) and STMN2 (Stathmin 2) as well as MYCBP. Stathmin, known as a tubulin depolymerization protein, is required for proliferation and is co-related to cancer cells and the invasion46. This protein was 2.8-fold increased in the presence of EGFP-CPN.
An obvious skepticism on this recruitment would be why the IGFBPL1-171 was capable of binding various nuclear proteins by the CP enhancement despite this sequence being known to interact with a small number of specific proteins. We ascribe this capability to the nature of CGIs. As the islands literally have frequent repeats of CpG and high GC content in the sequences, a given CGI is highly similar to other CGIs. For example, IGFBPL1-171 was found to be 49% identical to the core element of the rRNA promoter without gap (Fig. 5d) to which KDM2A and the CXXC domain bind19,20. To prove our argument, we performed binding assay of POLR1A with rDNA-260 bait followed by Western blotting. As shown in Fig. 5e, POLR1A time-dependently bound to rDNA-260 bait in the presence of CP, whereas the binding never took place without it. We also performed binding assays of ORC3 using the same bait. ORC3 bound less to rDNA-260 bait in 10 min than to IGFBPL1-312 in terms of % input, but CP still significantly enhanced ORC3 binding (Fig. 5f,g). This result firmly supports the suggestion that CP recruits various nuclear factors to the cognate binding sites. Perhaps, different nuclear factors bound to distinct CpG sites within IGFBPL1-171 as the bait has many CpGs.
The concern of nuclear factor recruitment to IGFBPL1-171 was further addressed at cellular level. If CP enhanced only the crosstalk between genome and nuclear proteins in cells, the domain would disturb natural partnerships and result in further deterioration of the N782-deficient cell proliferation. Therefore, we approached such an investigation by transducing YF29 with CPN and the control transcripts and by allowing the cells to grow at a low density of 3T3-J2 (0.2 × 104/cm). As we expected, the control keratinocytes proliferated slowly, but the cells transduced with CPN improved proliferation (Fig. 5h). Thus, we conclude that CP recruits the various important proteins to their natural partner CpG-rich sites in cells. The effect on cell proliferation by CPN was to a lesser extent than that of N782, suggesting that other domains of N782, including JmjC, play a supporting role for the recruitment.
Although CP enhances the binding of a variety of nuclear proteins, ectopic expression of N782 did not fully recover cell proliferation at low densities of 3T3-J2. This suggests that keratinocytes has another pathway under control of 3T3-J2 and independent of N782. Factors would operate out of nucleus so that those cannot be recruited to genomic sites. Integrins are a good candidate for the factors, as they are a group of transmenbrane proteins but nonetheless stimulate proliferation of keratinocytes and inhibit terminal differentiation of the cells by direction of 3T3-J247,48.
Discussion
Here, we demonstrate that the CP domain of KDM2A enhances binding of 94 nuclear factors including ORC3 to the IGFBPL1 promoter and further that it enhances binding of POLR1A and ORC3 to the rDNA promoter. This result strongly suggests that CP makes loci permissive for cognate nuclear factors to approach and bind to the loci. In other words, it is almost impossible for CP to fetch numerous nuclear factors from nucleosol to the cognate sites. Although 94 is an unusually large number, we think that this number is far below the actual number, and predict that CP enables as many nuclear proteins as the number of its binding sites. Thus, KDM2A should be able to regulate a wide variety of cellular activities even those nuclear factors that are unrelated to cell proliferation, such as circadian rhythm18. The CP domain seizes the basic feature of KDM2A by its broad binding to CpG-rich loci5 and by the permissive effect on the sites for various cognate nuclear factors.
We have not dissected the mechanisms of how CP makes the binding loci permissive, but we speculate that the domain bends or twists CpG-rich DNA and subsequently makes the loci permissive. Two scenarios can be conjectured for this process: (1) only CP is required, (2) besides CP, the XRCC5/6 complex is required. If the first were the case, then CP would eliminate adjacently sitting XRCC5/6 and promote binding of other nuclear proteins. However, if the second were the case, CP would make the sites more permissive with the XRCC5/6 complex especially with the XRCC6 subunit than CP alone. However, the XRCC5/6 complex may be expelled in the end of the reaction. Association of the XRCC5/6 complex with CGIs promoters has been recognized since the 1990’s49, but the role of XRCC5/6 at CpG-rich sites remains undetermined.
KDM2A was originally found to remove methyl groups from H3K36me2 and to a lesser extent from H3K36me1 by the JmjC domain3. However, the cellular role of this domain has not been elucidated. One report shows that the enzyme activity is involved in normal chromosome segregation16. The wild-type KDM2A, but not 2 mutants, the latter of which does not recognize H3K36me2 and thereby does not demethylate it in cells, recovers normal chromosome segregation in KDM2A knockdown cells. This result is consistent with our result that the CP domain of N782 recruits a number of proteins involved in chromosome segregation (Table 1) and that N782, which contains both JmjC and CP domains, more strongly enhances cell proliferation than the CP domain alone (Fig. 5h). It appears therefore that JmjC coordinates with CXXC-PHD domains to stimulate DNA-protein interactions for cell proliferation. Recently, KDM2A was found to bind HP1 through its 779LTVTL803 motif 50. Consequently, KDM2A enables HP1 to bind K3H9me3, and vice versa. The authors suggest that by this mechanism, KDM2A and HP1 are able to establish heterochromatin and expand the heterochromatin loci so as to silence gene expression. This pathway does not require the JmjC domain. However, repression of the rRNA gene by KDM2A, which occurs under starvation conditions, requires JmjC demethylation activity in addition to CXXC binding to the promoter7,19. The shift of KDM2A from activator to repressor states by starvation seems to be caused by modification of KDM2A via AMPK51. The role of KDM2A demetylase remains unresolved.
In this report, we show that KDM2As enhance the proliferation of human post-natal keratinocytes and that the short isomer N782 is virtually responsible for the effect. Further, we suggest that N782 has the same role in other human cells. The CP domain of the protein has the key role to recruit at least 94 proteins with functions ranging from replication to mitosis. Expression of N782 is regulated. In the case of keratinocytes, expression of N782 is turned off accompanying poor proliferation. However, 3T3-J2 feeders activate the expression and allow keratinocytes to proliferate at a rapid rate. Our findings help us understand both the basic mechanisms of human cell proliferation and feeding mechanism of 3T3-J2. Feeding of 3T3-J2 is essential for medical teams to prepare autologous skin grafts for treatment of life-threatening skin afflictions52,53.
Methods
Cell culture, transfection, and transduction
Post-natal keratinocytes (YF29) and hESC (H9) were routinely cultivated on feeder cells as described54,55. HEK293T cells were cultivated in DMEM supplemented with 10% fetal bovine serum. Cells used in this study, the cultivation, and the handling procedures have been approved by Harvard University Committee on Microbiological Safety and Embryonic Stem Cell Research Oversight Committee. We carried out experiments under the guidelines: NIH rDNA Guidelines; and the Federal Occupational Safety and Health Administration Bloodborne Pathogen Standard. Our experiments neither involved humans nor animals as the cultures had been established by other scientists before we started this investigation. For transfection, YF29 was transferred to a dish with feeder-free EpiLife (Invitrogen) one day before transfection, and HEK293T was similarly transferred to a dish with fresh DMEM/FBS. Ten-μg plasmid/10-cm dish and 5-μg plasmid/6-cm dish) were transfected with X-tremeGENE 9 DNA transfection reagent (Roche). Transfected keratinocyte and HEK293T cultures were incubated for 19 to 20 h and for 2 days, respectively. Transfection efficacy of YF29 was as poor as 5% and highly variable from one to another, while that of HEK293T was about 30%. Longer incubation did not improve transfection efficacy of keratinocytes. For measurement of promoter activity, HEK293 cells were inoculated in wells of a 96- well plate; and after one day, the cultures were transfected with 0.1-μg pIGFBPL1-312-GLuc and 0.2-μg plasmid encoding KDM2A or its derivatives. pIGFBPL1-312-GLuc was replaced with the empty vector, pGLuc-Basic 2, as the control. Plasmid pDIP2B-557-Gluc was used for estimation of the DIB2B promoter activity. Supernatant of the cultures were harvested and used for the luciferase activity assay with the BioLuc Gaussia Luciferase Assay kit (NEB). Keratinocytes was transduced with virions prepared by the method8. However, in this study selection for puromycin was skipped to avoid puromycin-induced terminal differentiation of post-natal keratinocytes.
Polymerase chain reaction
DNA sequences of interest were amplified using the Q5 Hot Start High-Fidelity 2x Master Mix (NEB). Primers are described in Tables 2 and 3. In general, template was initially denatured for 30 s at 98 °C and for 10 s before each cycle, annealing was performed at 0–2 °C higher temperature than the Tm of the primers, and polymerization was performed for 20 s per kbp at 72 °C. qPCR was carried out with AB-4167/A (Thermo Scientific). Ten-μl reaction mixture was initially heated for 15 s at 95 °C and 15 s for each cycle, annealing was performed for 30 s at 67 °C, and polymerization was performed for 30 s at 68 °C. Forty cycles were performed. KRT14 and G3PDH mRNA were amplified for normalization of N782 and KDM2A mRNA within keratinocytes and between keratinocytes and hESC, respectively.
Construction of plasmids
Coding regions of the KDM2A and N782 genes were amplified by PCR and ligated to the BglII/SalI sites of pEGFP C2, so that the N-termini of the proteins were fused to the C-terminus of EGFP (pEGFP-KDM2A and pEGFP-N782). From these, additional plasmids were constructed by deletion and insertion by using restriction enzymes or PCR. DNA encoding KDM2A domains was cloned into pEGFP-N80 (a pEGFP derivative with the sequence for the first 80 amino acids of KDM2A as a linker) or pGEX-5X-3 and pET-28C(+) for production of the domains in human cells or E. coli. The coding regions of the XRCC5 and XRCC6 genes were amplified by PCR using a YF29 cDNA pool as templates that were prepared using the ProtoScript first strand cDNA synthesis kit (NEB) and cloned into pET-28C(+) at the HindIII restriction site. XRCC5 and XRCC6 expressed from the resulting plasmids in E. coli specifically reacted to mouse-monoclonal antibodies to XRCC5 and XRCC6, respectively.
Cloning of genomic DNAs
We unbiasedly cloned human CpG-rich loci56. Briefly, Sau3AI digest fragments of a YF29 genomic DNA were collected by affinity chromatography with purified and immobilized 6x his-tagged KDM2A CXXC, and ligated to BamHI site of pBluescriptKS (+) and amplified by PCR. The product was ligated back to the empty vector. This amplification procedure was carried out only once not to amplify CpG-rich sequences methylated in cells. The prepared plasmids were transformed and amplified in E. coli, and the randomly isolated. Subsequently the inserts were identified by sequencing. Based on the identification, the promoter regions of the IGFBPL1 and the DIP2B genes were cloned, and their validity was confirmed by sequencing. The 455-bp sequence of the IGFBPL1 has been deposited (GenBank accession number, KX688890). The sequence of the DIP2B promoter was identical to that from 50504472 through 50504994 of GenBank ID NC_000012.12. The rDNA promoter (−179 through + 81) was cloned by PCR, whose sequence was the same as that in GenBank ID MF164270.1
Immunoprecipitation
Keratinocyte cultures were lysed in 50 mM Tris-HCl (pH 7.4) with 1% Triton X-100, 10% glycerol, 150 mM NaCl, 1 mM EDTA, 0.2 mM sodium orthovanadate, 1× EDTA-free Complete Protease Inhibitor Cocktail (Roche), and 1 mM PMSF in a cold room. The lysates were pre-cleared for 1 h with protein A magnetic beads (NEB) that had been charged with normal-rabbit-IgG (Santa Cruz). The cleared supernatant was then mixed with protein A magnetic beads that had been charged with rabbit anti-GFP (Acris, SP3005P) and rotated for 3 h. Then, beads bearing protein complexes were collected and washed twice with the lysis buffer. The immunoprecipitated complexes were released from the beads by heating in a 3× loading buffer.
Preparation of extracts
We prepared nuclear extracts, slightly modifying the reported procedure57. The nuclear fraction was suspended in nuclear extraction buffer consisting of 50 mM Tris-HCl pH 7.4, 1% Triton X-100, 10% glycerol, 150 mM NaCl, 0.2 mM sodium orthovandate, 1x EDTA- free Complete Protease Inhibitor Cocktail, 1 mM PMSF, 5 μl/ml RNase (Roche), and 25 μM ZnSO4. The suspension was sonicated 3 times for 3 s each time. The lysates were centrifuged after addition of EDTA to 1 mM and the supernatants were used as nuclear extracts. Cell extracts containing XRCC5, XRCC6, and GST-fusion probes were prepared from E. coli BL21 (DE3) or a derivative in which each protein were overexpressed. Collected bacterial cell pellets were subjected to freezing-thawing 3 times, then suspended in the nuclear extraction buffer containing 1 mg/ml lysozyme. The lysates were sonicated and the supernatants were saved. When necessary, the pellets were suspended in the same buffer and homogenized by brief sonication.
DNA-protein interaction
Bait used in this study was prepared by PCR including a 5′ end biotinylated forward primer and a backward primer. Four pmol bait was bound to hydrophilic streptavidin magnetic beads (NEB, S1421S), and incubated with 200 µg nuclear extracts in 50 µl up to 30 min at room temperature. The beads were washed twice with 100 µl extraction buffer. The bound proteins were released in 20-µl 1.5X sample buffer by heating for 5 min at 100 °C, and collected immediately for subsequent SDS-PAGE. For mass spectrometry, the IGFBPL1-171 bait (4 pmol) was incubated with 500 µg nuclear extracts in 72 µl for 10 min. Binding of GST-CN to IGFBLL1 baits similarly performed in the presence of 10 units of poly[d(A-T)] per reaction by 10-min incubation. Protein samples interacted with DNA were separated on 4–20% Gold Precast Gels by SDS-PAGE. Then, proteins on the gel were stained with ProtoBlue Safe Colloidal Coomassie G-250 (National Diagnostics) or transferred to a nitrocellulose membrane (Bio-Rad) for Western blotting. Both stained gel and nitrocellulose membrane were scanned using a LI-COR Odyssey Imager. Resolution and Quality were set to 169 μm and medium, respectively. Seven hundred and 800 channel intensities were set to 3.5 and 5.0, respectively, unless otherwise indicated. ImageStudioLite2 was used to collect images and quantitate data. The following antibodies were used for Western blotting: rabbit anti-ORC3 (Thermofisher, PA5-28083); rabbit anti-POLR1A (Thermofisher, PA5-37156); mouse anti-XRCC5 (Thermo Scientific, MA5-15873); mouse anti-XRCC6 (Abnova, H00002547-M01), rabbit polyclonal anti-H3 (Thermofisher Scientific, PA-16183), mouse anti-H2B (Biolegend, 688702), anti-GFP monoclonal antibody (Clontech), rabbit anti-GFP (Acris, SP3005P); goat anti-rabbit IgG (H + L), IRDy 800 conjugated (Rockland, 611-132-122); and IRDye 680RD goat anti-mouse IgG (H + L) (LI-COR, 926-68170).
Protein-protein interaction
Far Western blotting was used to investigate interaction of KDM2A domains with XRCC5 and XRCC6. Both XRCC5 and XRCC6 were prepared from E. coli particulate fractions and immobilized onto nitrocellulose membrane after SDS-PAGE, and then renatured by the method of Einarson58. The proteins on the membrane were incubated overnight at 4 °C in the HEPES buffer (without guanidine hydrochloride) with 150 mM NaCl and 5% non-fat dry milk for blocking, and then incubated for 2 h at room temperature in the HEPES buffer consisting of 150 mM NaCl/1% non-fat milk with bacterial extracts (soluble fractions) containing each probe: GST-CN, GST-PN, GST-CPN, or GST-NLSR. The membrane was washed 4 times in the HEPES buffer/NaCl. Eventually, the probes on the membrane were immunologically identified with goat-anti GST (Pharmacia).
Mass spectrometry
For identification of DNA bait- and N782-binding proteins, mass spectrometry was performed by the previously described procedure59. For identification of total nuclear proteins, tandem mass tag-based mass spectrometry was performed45. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE1 partner repository with the dataset identifier PXD012941.
Analytical software
BLAST, Cn3D, and GenBank database at National Center for Biotechnology Information and USCS Genome Browser at University of California, Santa Cruz were used for various computational analysis of proteins and nucleic acids. The human Uniprot database was used for identification of bait-CPN-binding proteins. Lasergene (DNASTAR) was used for sequence alignment. Microsoft Excel software and GraphPad Prism were used for statistical analysis. t- and ANOVA-test: p < 0.05 indicates a significant difference.
Sequence and mass spectrometry data obtained during current study are available as stated in the Methods section.
References
Klose, R. J., Kallin, E. M. & Zhang, Y. JmjC-domain-containing proteins and histone demethylation. Nature reviews. Genetics 7, 715–727, https://doi.org/10.1038/nrg1945 (2006).
Pedersen, M. T. & Helin, K. Histone demethylases in development and disease. Trends in cell biology 20, 662–671, https://doi.org/10.1016/j.tcb.2010.08.011 (2010).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816, https://doi.org/10.1038/nature04433 (2006).
Farcas, A. M. et al. KDM2B links the Polycomb Repressive Complex 1 (PRC1) to recognition of CpG islands. eLife 1, e00205, https://doi.org/10.7554/eLife.00205 (2012).
Blackledge, N. P. et al. CpG islands recruit a histone H3 lysine 36 demethylase. Molecular cell 38, 179–190, https://doi.org/10.1016/j.molcel.2010.04.009 (2010).
Zhou, J. C., Blackledge, N. P., Farcas, A. M. & Klose, R. J. Recognition of CpG island chromatin by KDM2A requires direct and specific interaction with linker DNA. Molecular and cellular biology 32, 479–489, https://doi.org/10.1128/MCB.06332-11 (2012).
Tanaka, Y., Umata, T., Okamoto, K., Obuse, C. & Tsuneoka, M. CxxC-ZF domain is needed for KDM2A to demethylate histone in rDNA promoter in response to starvation. Cell structure and function 39, 79–92 (2014).
Iuchi, S. & Green, H. Lysine-specific demethylase 2A (KDM2A) normalizes human embryonic stem cell derived keratinocytes. Proceedings of the National Academy of Sciences of the United States of America 109, 9442–9447, https://doi.org/10.1073/pnas.1206176109 (2012).
Kawakami, E., Tokunaga, A., Ozawa, M., Sakamoto, R. & Yoshida, N. The histone demethylase Fbxl11/Kdm2a plays an essential role in embryonic development by repressing cell-cycle regulators. Mechanisms of development 135, 31–42, https://doi.org/10.1016/j.mod.2014.10.001 (2015).
Gao, R. et al. Depletion of histone demethylase KDM2A inhibited cell proliferation of stem cells from apical papilla by de-repression of p15INK4B and p27Kip1. Molecular and cellular biochemistry 379, 115–122, https://doi.org/10.1007/s11010-013-1633-7 (2013).
Wagner, K. W. et al. KDM2A promotes lung tumorigenesis by epigenetically enhancing ERK1/2 signaling. The Journal of clinical investigation 123, 5231–5246, https://doi.org/10.1172/JCI68642 (2013).
Lu, T. et al. Regulation of NF-kappaB by NSD1/FBXL11-dependent reversible lysine methylation of p65. Proceedings of the National Academy of Sciences of the United States of America 107, 46–51, https://doi.org/10.1073/pnas.0912493107 (2010).
Du, J., Ma, Y., Ma, P., Wang, S. & Fan, Z. Demethylation of epiregulin gene by histone demethylase FBXL11 and BCL6 corepressor inhibits osteo/dentinogenic differentiation. Stem cells 31, 126–136, https://doi.org/10.1002/stem.1255 (2013).
Rizwani, W., Schaal, C., Kunigal, S., Coppola, D. & Chellappan, S. Mammalian lysine histone demethylase KDM2A regulates E2F1-mediated gene transcription in breast cancer cells. PloS one 9, e100888, https://doi.org/10.1371/journal.pone.0100888 (2014).
Frescas, D. et al. KDM2A represses transcription of centromeric satellite repeats and maintains the heterochromatic state. Cell cycle 7, 3539–3547 (2008).
Cheng, Z. et al. A molecular threading mechanism underlies Jumonji lysine demethylase KDM2A regulation of methylated H3K36. Genes & development 28, 1758–1771, https://doi.org/10.1101/gad.246561.114 (2014).
Cao, L. L. et al. ATM-mediated KDM2A phosphorylation is required for the DNA damage repair. Oncogene, https://doi.org/10.1038/onc.2015.81 (2015).
Reischl, S. & Kramer, A. Fbxl11 Is a Novel Negative Element of the Mammalian Circadian Clock. Journal of biological rhythms 30, 291–301, https://doi.org/10.1177/0748730415587407 (2015).
Tanaka, Y. et al. JmjC enzyme KDM2A is a regulator of rRNA transcription in response to starvation. The EMBO journal 29, 1510–1522, https://doi.org/10.1038/emboj.2010.56 (2010).
Okamoto, K., Tanaka, Y. & Tsuneoka, M. SF-KDM2A binds to ribosomal RNA gene promoter, reduces H4K20me3 level, and elevates ribosomal RNA transcription in breast cancer cells. International journal of oncology, https://doi.org/10.3892/ijo.2017.3908 (2017).
Kipreos, E. T. & Pagano, M. The F-box protein family. Genome biology 1, REVIEWS3002, https://doi.org/10.1186/gb-2000-1-5-reviews3002 (2000).
Van Steensel, B. & Belmont, A. S. Lamina-Associated Domains: Links with Chromosome Architecture, Heterochromatin, and Gene Repression. Cell 169, 780–791, https://doi.org/10.1016/j.cell.2017.04.022 (2017).
Gonzalez, I. et al. A lncRNA regulates alternative splicing via establishment of a splicing-specific chromatin signature. Nature structural & molecular biology 22, 370–376, https://doi.org/10.1038/nsmb.3005 (2015).
Ju, B. G. et al. A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription. Science 312, 1798–1802, https://doi.org/10.1126/science.1127196 (2006).
Necsulea, A., Guillet, C., Cadoret, J. C., Prioleau, M. N. & Duret, L. The relationship between DNA replication and human genome organization. Molecular biology and evolution 26, 729–741, https://doi.org/10.1093/molbev/msn303 (2009).
Langley, A. R., Graf, S., Smith, J. C. & Krude, T. Genome-wide identification and characterisation of human DNA replication origins by initiation site sequencing (ini-seq). Nucleic acids research 44, 10230–10247, https://doi.org/10.1093/nar/gkw760 (2016).
Miotto, B., Ji, Z. & Struhl, K. Selectivity of ORC binding sites and the relation to replication timing, fragile sites, and deletions in cancers. Proceedings of the National Academy of Sciences of the United States of America 113, E4810–4819, https://doi.org/10.1073/pnas.1609060113 (2016).
Taccioli, G. E. et al. Ku80: product of the XRCC5 gene and its role in DNA repair and V(D)J recombination. Science 265, 1442–1445 (1994).
Fell, V. L. & Schild-Poulter, C. The Ku heterodimer: function in DNA repair and beyond. Mutation research. Reviews in mutation research 763, 15–29, https://doi.org/10.1016/j.mrrev.2014.06.002 (2015).
Hahn, S. Structure and mechanism of the RNA polymerase II transcription machinery. Nature structural & molecular biology 11, 394–403, https://doi.org/10.1038/nsmb763 (2004).
Shen, Z. et al. A WD-repeat protein stabilizes ORC binding to chromatin. Molecular cell 40, 99–111, https://doi.org/10.1016/j.molcel.2010.09.021 (2010).
Szambowska, A. et al. DNA binding properties of human Cdc45 suggest a function as molecular wedge for DNA unwinding. Nucleic acids research 42, 2308–2319, https://doi.org/10.1093/nar/gkt1217 (2014).
Sclafani, R. A. & Holzen, T. M. Cell cycle regulation of DNA replication. Annual review of genetics 41, 237–280, https://doi.org/10.1146/annurev.genet.41.110306.130308 (2007).
Yeeles, J. T., Deegan, T. D., Janska, A., Early, A. & Diffley, J. F. Regulated eukaryotic DNA replication origin firing with purified proteins. Nature 519, 431–435, https://doi.org/10.1038/nature14285 (2015).
Grummt, I. Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus. Genes & development 17, 1691–1702, https://doi.org/10.1101/gad.1098503R (2003).
Lin, T. et al. Nucleostemin and GNL3L exercise distinct functions in genome protection and ribosome synthesis, respectively. Journal of cell science 127, 2302–2312, https://doi.org/10.1242/jcs.143842 (2014).
Yang, Y. et al. Arginine methylation facilitates the recruitment of TOP3B to chromatin to prevent R loop accumulation. Molecular cell 53, 484–497, https://doi.org/10.1016/j.molcel.2014.01.011 (2014).
Cleveland, D. W., Mao, Y. & Sullivan, K. F. Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112, 407–421 (2003).
Walczak, C. E., Gayek, S. & Ohi, R. Microtubule-depolymerizing kinesins. Annual review of cell and developmental biology 29, 417–441, https://doi.org/10.1146/annurev-cellbio-101512-122345 (2013).
Mayr, M. I. et al. The human kinesin Kif18A is a motile microtubule depolymerase essential for chromosome congression. Current biology: CB 17, 488–498, https://doi.org/10.1016/j.cub.2007.02.036 (2007).
Du, Y., English, C. A. & Ohi, R. The kinesin-8 Kif18A dampens microtubule plus-end dynamics. Current biology: CB 20, 374–380, https://doi.org/10.1016/j.cub.2009.12.049 (2010).
Fu, J. et al. TPX2 phosphorylation maintains metaphase spindle length by regulating microtubule flux. The Journal of cell biology 210, 373–383, https://doi.org/10.1083/jcb.201412109 (2015).
Kern, D. M., Nicholls, P. K., Page, D. C. & Cheeseman, I. M. A mitotic SKAP isoform regulates spindle positioning at astral microtubule plus ends. The Journal of cell biology 213, 315–328, https://doi.org/10.1083/jcb.201510117 (2016).
Maiato, H. et al. Human CLASP1 is an outer kinetochore component that regulates spindle microtubule dynamics. Cell 113, 891–904 (2003).
Paulo, J. A., O’Connell, J. D. & Gygi, S. P. A Triple Knockout (TKO) Proteomics Standard for Diagnosing Ion Interference in Isobaric Labeling Experiments. Journal of the American Society for Mass Spectrometry 27, 1620–1625, https://doi.org/10.1007/s13361-016-1434-9 (2016).
Biaoxue, R., Xiguang, C., Hua, L. & Shuanying, Y. Stathmin-dependent molecular targeting therapy for malignant tumor: the latest 5 years’ discoveries and developments. Journal of translational medicine 14, 279, https://doi.org/10.1186/s12967-016-1000-z (2016).
Green, H. Terminal differentiation of cultured human epidermal cells. Cell 11, 405–416 (1977).
Watt, F. M. Role of integrins in regulating epidermal adhesion, growth and differentiation. The EMBO journal 21, 3919–3926, https://doi.org/10.1093/emboj/cdf399 (2002).
Toth, E. C. et al. Interactions of USF and Ku antigen with a human DNA region containing a replication origin. Nucleic acids research 21, 3257–3263 (1993).
Borgel, J. et al. KDM2A integrates DNA and histone modification signals through a CXXC/PHD module and direct interaction with HP1. Nucleic Acids Res 45, 1114–1129, https://doi.org/10.1093/nar/gkw979 (2017).
Tanaka, Y. et al. Mild Glucose Starvation Induces KDM2A-Mediated H3K36me2 Demethylation through AMPK To Reduce rRNA Transcription and Cell Proliferation. Mol Cell Biol. 35, 4170–84, https://doi.org/10.1128/MCB (2015).
Gallico, G. G. 3rd, O’Connor, N. E., Compton, C. C., Kehinde, O. & Green, H. Permanent coverage of large burn wounds with autologous cultured human epithelium. The New England journal of medicine 311, 448–451, https://doi.org/10.1056/NEJM198408163110706 (1984).
Hirsch, T. et al. Regeneration of the entire human epidermis using transgenic stem cells. Nature 551, 327–332, https://doi.org/10.1038/nature24487 (2017).
Allen-Hoffmann, B. L. & Rheinwald, J. G. Polycyclic aromatic hydrocarbon mutagenesis of human epidermal keratinocytes in culture. Proceedings of the National Academy of Sciences of the United States of America 81, 7802–7806 (1984).
Iuchi, S. et al. An immortalized drug-resistant cell line established from 12–13-day mouse embryos for the propagation of human embryonic stem cells. Differentiation; research in biological diversity 74, 160–166, https://doi.org/10.1111/j.1432-0436.2006.00067.x (2006).
Iuchi, S. & Green, H. Basonuclin, a zinc finger protein of keratinocytes and reproductive germ cells, binds to the rRNA gene promoter. Proceedings of the National Academy of Sciences of the United States of America 96, 9628–9632 (1999).
Huang, J. et al. RAD18 transmits DNA damage signalling to elicit homologous recombination repair. Nature cell biology 11, 592–603, https://doi.org/10.1038/ncb1865 (2009).
Einarson, M. B. Detection of protein-protein interactions using Far Western with GST fusion proteins. Molecular cloning, 3rd Ed., (edited by Sambrook, J. and Russel, D. W.) 18.48–18.54 (CSH, 2001).
Paulo, J. A. Sample preparation for proteomic analysis using a GeLC-MS/MS strategy. Journal of biological methods 3, https://doi.org/10.14440/jbm.2016.106 (2016).
Acknowledgements
We dedicate this work to Dr. Howard Green who discovered the 3T3-J2-based keratinocyte cultivation method, saved lives of severely burned patients, and supported this research, but passed away in 2015. We thank Dr. Danesh Moazed for all his support, Dr. Steven Gygi for providing access to mass spectrometry facilities, Dr. Alfred Goldberg for providing access to qPCR machinery and Drs. Dong-Hoon Lee and Zhe Sha for providing introductory lessons to it, ICCB-Longwood Screening Facility for providing access to Molecular Devices SpectraMax M5, Dr. Naotaka Sekiyama of Kyoto University for the communication, Dr. Mark A. Currie for reading the manuscript and commenting on it. We also thank the Japan Tissue Engineering Co., Ltd. (J-TEC) for a research grant.
Author information
Authors and Affiliations
Contributions
J.A.P. performed MS; S.I. performed the other experiments. S.I. designed the study and wrote the manuscript; J.A.P. edited the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Iuchi, S., Paulo, J.A. Lysine-specific demethylase 2A enhances binding of various nuclear factors to CpG-rich genomic DNAs by action of its CXXC-PHD domain. Sci Rep 9, 5496 (2019). https://doi.org/10.1038/s41598-019-41896-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41896-6
This article is cited by
-
RNAmetasome network for macromolecule biogenesis in human cells
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.