Abstract
Mitochondrial (mt) DNA encodes factors essential for cellular respiration, therefore its level and integrity are crucial. ABF2 encodes a mitochondrial DNA-binding protein and its null mutation (Δabf2) induces mtDNA instability in Saccharomyces cerevisiae. Mhr1 is a mitochondrial recombinase that mediates the predominant form of mtDNA replication and acts in mtDNA segregation and the repair of mtDNA double-stranded breaks (DSBs). However, the involvement of Mhr1 in prevention of mtDNA deletion mutagenesis is unknown. In this study we used Δabf2 mhr1-1 double-mutant cells, which lose mitochondrial function in media containing fermentable carbon sources, to investigate whether Mhr1 is a suppressor of mtDNA deletion mutagenesis. We used a suppresivity assay and Southern blot analysis to reveal that the Δabf2 mutation causes mtDNA deletions rather than an mtDNA-lacking (ρ0) phenotype, and observed that mtDNA deletions are exacerbated by an additional mhr1-1 mutation. Loss of respiratory function due to mtDNA fragmentation occurred in ∆mhr1 and ∆abf2 mhr1-1 cells. However, exogenous introduction of Mhr1 into Δabf2 mhr1-1 cells significantly rescued respiratory growth, suggesting that Mhr1-driven homologous mtDNA recombination prevents mtDNA instability.
Similar content being viewed by others
Introduction
The mitochondrial genome (mtDNA) encodes rRNAs, tRNAs and electron transport chain subunits essential for cellular respiration. Thus, maintenance of mtDNA level and integrity are crucial for healthy respiratory function1,2,3,4. In human cells, high mtDNA deletion levels can cause mitochondrial dysfunction and have been linked to neuromuscular disorders5,6.
MtDNA is packaged into a nucleoprotein complex termed the mitochondrial nucleoid, which is regarded as the unit of mtDNA inheritance7. Abf2 is a key component of the nucleoid with a histone-like role8,9, and contains two high mobility group (HMG) domains for the binding and efficient packaging of linear double-stranded DNA without supercoiling10. Abf2 wraps and bends mtDNA, but is not required for the activity of promoters at ori sequences and has no transcriptional role in yeast11,12,13. Mutants lacking ABF2 (∆abf2) display a loss-of-mtDNA phenotype8,14,15,16 when utilizing fermentable carbon sources for growth, but are able to maintain wild-type mtDNA in non-fermentable media, indicating that the requirement for Abf2 in ρ+ cells is conditional. The ∆abf2 phenotype is considered typical of nuclear gene mutations that affect mtDNA maintenance, since more than 100 nuclear genes that influence mtDNA integrity in yeast have been identified1,2.
Mhr1-1 is a temperature-sensitive point mutation in the nuclear gene MHR1, which causes deficiency in mtDNA homologous recombination17. Mhr1 is a mitochondrial recombinase18,19 that acts in double-stranded break (DSB) repair, mediates the predominant form of mtDNA replication in ρ+ cells15,17,20,21,22,23 and increases mtDNA content without additional Abf221. DSBs can be created at replication origin (ori) sequences by excision repair enzymes such as Ntg121,24. Following procession of DSBs by the 5′-3′exonuclease activity of Din722, 3′-single stranded DNA can be used by Mhr1 to form a heteroduplex joint, in which the 3′-single stranded DNA tail serves as a primer to initiate rolling circle DNA replication, which produces linear multiple-unit-sized mtDNA molecules, termed concatemers, that promote segregation of heteroplasmy towards homoplasmy20.
In this study, we investigate whether Mhr1-driven mtDNA replication and homologous recombination contributes to the maintenance of mtDNA content and genomic integrity in mhr1-1 cells with a null abf2 genetic background, which show a loss-of-mtDNA phenotype in fermentable media due to deletion mutagenesis.
Results
Double-mutant ∆abf2 mhr1-1 cells rapidly lose respiratory function in fermentable media
In order to examine severely compromised mtDNA maintenance, we used ∆abf2 cells (Table 1), which display a well-documented loss-of-mtDNA phenotype upon cultivation in fermentable media8,14,15. In order to compare the extent of respiratory function loss in these backgrounds, we first selectively pre-cultivated wild-type (WT), single-mutant ∆abf2, mhr1-1, or double-mutant ∆abf2 mhr1-1 cells in glycerol medium, a carbon source requiring mitochondrial respiration for its utilization. We then transferred the cells to synthetic complete fermentable media containing glucose (Glu) or raffinose and galactose (RGal) as carbon sources. While both Glu and RGal media are fermentable, the use of RGal allows for distinction from the transcriptional effects of glucose, which changes the global gene expression pattern25. We cultivated cells for nearly eight generations at 30 °C or 34 °C (Fig. 1a) and then spread equal amounts of dilute culture of each strain onto rich glucose (YPD) and rich glycerol (YPGly) plates (Fig. 1b). The proportion of colony-forming units (CFUs) that retained mitochondrial respiratory activity and were thus able to grow on YPGly, compared to the total number of CFUs on YPD, was quantified (Fig. 1c).
Loss of respiratory function in ∆abf2, mhr1-1 and ∆abf2 mhr1-1 cells. (a) Scheme of respiratory function assay. Cells were selectively pre-cultured in YPGly medium, then 106 cells were transferred to Glu or RGal media and cultivated for <8 generations at 30 °C or 34 °C. Equal volumes of dilute culture were then spread onto YPD and YPGly plates to measure the proportion of CFUs retaining respiratory function. (b) Representative plate images of wild-type, ∆abf2, mhr1-1, and ∆abf2 mhr1-1 CFU formation on YPD and YPGly plates following cultivation in Glu or RGal media at 30 °C or 34 °C. (c) ρ+ CFU formation rate based on n = 3 independent experiments described in (a). (d) Scheme of extended respiratory function assay. Cells were selectively grown in YPGly media, then 105 cells were transferred to RGal media and cultivated for two consecutive 48-hour rounds (approximately 20 generations) at 30 °C. Equal volumes of dilute culture were then: (e) Spread onto YPD and YPGly plates, or (f) spread onto YPD plates, grown for four days, and then replica-plated onto YPGly plates. (g) ρ+ CFU formation rate from cells simultaneously spread onto YPD and YPGly, based on n = 3 independent experiments. All error bars represent ± SD.
We observed that WT cells retained almost all respiratory function under each condition, while ∆abf2 cells showed remarkable loss of respiratory activity in Glu media, to 59.1 ± 5.9% ρ+ at 30 °C and 14.0 ± 5.3% ρ+ at 34 °C. On the other hand, cultivation of ∆abf2 cells in RGal medium resulted in a large proportion of cells retaining respiratory activity, forming ρ+ CFUs at rates of 103.9 ± 14.3% at 30 °C and 83.9 ± 20.7% at 34 °C. These observations appear consistent with the previously reported mtDNA instability phenotype of ∆abf2 cells in glucose14. In the mhr1-1 background, a large proportion of CFUs retained respiratory activity, with ρ+ CFU formation rates of 103.9 ± 0.7% and 77.6 ± 6.0% ρ+ in Glu, and 96.5 ± 13.5% and 83.7 ± 10.6% ρ+ in RGal, at 30 °C and 34 °C, respectively. The slight decreases in respiratory activity of mhr1-1 cells grown at 34 °C was previously observed as mhr1-1 temperature sensitivity17,20. Furthermore, ∆abf2 mhr1-1 double-mutant cells displayed ρ+ CFU formation rates of 68.3 ± 10.4% and 15.4 ± 7.3% ρ+ in Glu, and 64.3 ± 8.8% and 37.2 ± 3.7% ρ+ in RGal, at 30 °C and 34 °C, respectively. Severe, temperature-dependent loss of respiratory function occurred in the ∆abf2 single-mutant in Glu but not RGal, while ∆abf2 mhr1-1 double mutant cells displayed an additive increase in temperature sensitivity in RGal. These results indicate that loss of mitochondrial function in this background occurs independently of glucose repression.
We next explored the effect of extended cultivation time on loss of cellular respiratory function. We pre-cultivated the four strains in YPGly medium, then transferred the cells to RGal medium and cultivated for approximately 20 generations (Fig. 1d). Finally, we spread the cells on YPD and subsequently replicated the colonies onto YPGly plates (Fig. 1e), or simultaneously spread equal amounts of dilute culture onto YPD and YPGly plates (Fig. 1f). In contrast to WT and mhr1-1 single-mutant cells, which remained almost entirely ρ+, only 27.0 ± 18.4% of the ∆abf2 cells were able to form ρ+ colonies on glycerol plates following simultaneous spreading. Remarkably, none of the ∆abf2 mhr1-1 double-mutant cells were able to grow on YPGly plates after growth for approximately 20 generations in RGal (Fig. 1g). Therefore, extended cultivation of the ∆abf2 single-mutant in fermentable RGal media increases loss of cellular respiratory function, while the additional loss of Mhr1 function causes rapid and complete loss of cellular respiratory function1,14.
Nucleoid numbers are significantly reduced in ∆abf2 mhr1-1 cells
Next, we investigated the relative abundance of mtDNA nucleoids in WT, Δabf2 or mhr1-1 single-mutant, or Δabf2 mhr1-1 double-mutant cells selectively grown in YPGly media, or grown in YPD media at 30 °C or 34 °C for more than 20 generations (Fig. 2). In Δabf2 single-mutants, approximately 27% of CFUs remained ρ+ after cultivation in glucose (Fig. 1g), yet all ∆abf2 mother cells we observed displayed mtDNA-derived nucleoid signals after cultivation in YPD at 30 °C (Fig. 2b), suggesting that loss of respiratory function in ∆abf2 cells may be due to mtDNA deletion mutagenesis. In addition, double-mutant ∆abf2 mhr1-1 cells showed the lowest number of mitochondrial nucleoid signals after cultivation in YPD media (Fig. 2a,b), and nucleoid signals were absent from 44.4% of ∆abf2 mhr1-1 mother cells following cultivation at 34 °C, compared to 0% of WT and mhr1-1 cells and only 10.0% of ∆abf2 mother cells. These results indicate that mtDNA maintenance is defective without Abf2 and fully functional Mhr1. Since we observed that respiratory function and nucleoid abundance generally decrease in ∆abf2 mhr1-1 cells relative to ∆abf2 cells, it is likely Mhr1 plays a role in protecting mtDNA genomic integrity.
Reduction in mtDNA-derived DAPI signals in ∆abf2, mhr1-1 and ∆abf2 mhr1-1 mutant cells. (a) Mitochondrial nucleoid signals in wild-type, ∆abf2, mhr1-1 and ∆abf2 mhr1-1 cells cultivated in YPGly media, or in YPD media to log-phase at 30 °C or 34 °C. Scale bar = 2 µm. (b) Numbers of mtDNA-DAPI foci in individual mother cells. The number of individual cells measured for WT, ∆abf2, mhr1-1 and ∆abf2 mhr1-1 mother cells cultivated in YPGly at 30 °C was n = 14, 8, 7 and 7, respectively. For cells grown in YPD at 30 °C, n = 13, 12, 14 and 17, respectively. For cells grown in YPD at 34 °C, n = 28, 30, 38 and 27, respectively. Horizontal lines represent the mean number of DAPI foci.
The mhr1-null mutation (Δmhr1) causes mtDNA fragmentation
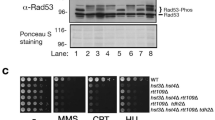
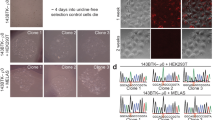
In order to demonstrate that Mhr1 is required to maintain mtDNA integrity, we introduced mhr1::LEU DNA fragments into WT/WT diploid cells to disrupt one of the two MHR1 alleles, thereby creating WT/Δmhr1 haploinsufficient diploid cells (see: Table 1). We then conducted tetrad analysis to determine whether haploid spores with the nuclear genotype mhr1::LEU (Δmhr1) retain respiratory function. All leucine prototrophic spores were unable to grow on YPGly plates, confirming that the Δmhr1 mutation causes complete loss of respiratory function (Fig. 3a). We further cultivated WT cells and seven Δmhr1 spores (Fig. 3b,a–g) in glucose medium, after selecting cells that still displayed mtDNA signals upon DAPI-staining. ApaI-digests of mtDNA from wild-type cells gave rise to many discrete bands (Fig. 3b), while ApaI-digests of mtDNA from Δmhr1 cells gave only a few discrete bands, indicating mtDNA deletions occur in Δmhr1 cells with some observable amounts of mtDNA remaining.
Tetrad analysis of respiratory function and mtDNA deletions in ∆mhr1 cells. (a) Respiratory function of ∆mhr1 spores was analyzed by replica plating colonies derived from spores (top) onto synthetic media lacking leucine (middle) and YPGly media (bottom). (b) ApaI digests of purified mtDNA molecules derived from wild-type and ∆mhr1 spores.
MtDNA deletion mutagenesis in ∆abf2 mhr1-1 cells
To determine whether mtDNA deletion mutagenesis or the complete loss of mtDNA occurred in ∆abf2 single- or ∆abf2 mhr1-1 double-mutant cells, we analyzed suppressiveness according to previously established methods24. The degree of suppressiveness is determined by: (1) The length of the remnant mtDNA molecule after undergoing a deletion and (2) the preservation of an active ori sequence (Fig. 4a). Small mtDNA deletions result in relatively large remnant molecules. Therefore, when crossed with ρ+ haploid cells, small mtDNA deletion-bearing cells give rise to diploid populations that are in a range of 10% to 90% ρ+. In contrast, when mitochondrial genomes undergo a large deletion event but retain at least one active replication origin, crosses of these haploid cells with ρ+ haploid cells of the opposite mating type will give rise to <5% ρ+ diploid progeny, a phenotype termed “hypersuppressive”26. Finally, crossing haploid cells without mtDNA (ρ0) with ρ+ haploid cells will result in 100% ρ+ diploid progeny, a phenotype termed “non-suppressive” (Fig. 4a).
Degree of mtDNA suppressivity in ∆abf2 and ∆abf2 mhr1-1 mutant cells. (a) Illustration of the effect of mtDNA deletion size on the suppressive phenotype. (b) Frequency distribution of the ρ+ phenotype in crosses of ρ+, ρ−, HS ρ−, ρ0, ∆abf2 or ∆abf2 mhr1-1 cells (top to bottom, respectively) with ρ+ haploid cells (see Table 1). The numbers of independent crosses conducted for each result (shown top to bottom) was, n = 5, 10, 5, 5, 32 and 20, respectively.
We crossed a ρ+ strain with haploid WT ρ+, ρ−, HS ρ− or ρ0 cells as controls (Fig. 4b, top four panels) and ∆abf2 or ∆abf2 mhr1-1 cells, and analyzed the respiratory phenotypes of the resulting diploid progeny. In crosses with the ∆abf2 background, 20~80% of diploid colonies were ρ+, with a single distribution centered at around 55% ρ+ (Fig. 4b, second panel from bottom). In contrast to this moderately suppressive phenotype, crossing ∆abf2 mhr1-1 cells yielded a bimodal distribution in which resulting diploid cells either displayed a hypersuppressive, or moderate to non-suppressive phenotype, ranging from 0~20% and 50~100% ρ+, respectively (Fig. 4b, bottom panel). These results indicate that large-scale mtDNA deletions or the complete loss of mtDNA occurs in ∆abf2 mhr1-1 cells, suggesting that the increased production of ρ− progeny (Fig. 1c,g) is due to a deficiency of Mhr1-driven mtDNA recombination.
Next, we analyzed mtDNA from ρ− ∆abf2 and ∆abf2 mhr1-1 colonies by Southern blot analysis. Compared to ρ+ WT and ∆abf2 controls (Fig. 5a), we observed that ρ− ∆abf2 mtDNA generally lacked several mtDNA-specific bands that were present in ρ+ mtDNA (Fig. 5b). Similarly, ∆abf2 mhr1-1 double-mutant cells generally lacked many mtDNA-specific bands and some samples lacked mtDNA signals altogether, compared to the mtDNA signals derived from ρ+ WT and mhr1-1 cells (Fig. 5a,b). Also, to our surprise we observed several more small, mtDNA-specific bands in the mhr1-1 control compared to the ρ+ WT and ∆abf2 controls. One explanation is that mtDNA deletions caused by mhr1-1 mutation result in heteroplasmy, which would be stable since the mhr1-1 mutation delays mitochondrial allele segregation20. Collectively, these results indicate that mtDNA deletions occur in ρ− ∆abf2 cells and that large-scale mtDNA deletion or the complete loss of mtDNA occurs in ρ− or ρ0 ∆abf2 mhr1-1 cells.
Mhr1 overproduction prevents mtDNA deletion mutagenesis
To verify that Abf2 and Mhr1 are required for mtDNA maintenance, we analyzed mtDNA levels relative to nuclear DNA using quantitative PCR27. We observed that the mtDNA level in ∆abf2 cells was less than half (46.3 ± 8.6%) that of WT cells grown in YPGly medium (Fig. 6a). Consistent with our previous results17, a large proportion (83.9 ± 15.3%) of mtDNA content was retained in mhr1-1 cells grown in glycerol medium, while we observed no additive effect on the depletion of mtDNA in ∆abf2 mhr1-1 double-mutant cells (51.4 ± 8.8%), suggesting Abf2 is dispensable for Mhr1-driven mtDNA replication (Fig. 6a).
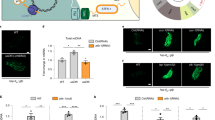
Effects of Mhr1 overproduction on mtDNA content and respiratory function. (a) Relative mtDNA level in wild-type, ∆abf2, mhr1-1 or ∆abf2 mhr1-1 cells cultivated in YPGly media. (b) Immunoblot analysis of Mhr1 protein content in cells containing the pVT or pVT-MHR1 plasmids. (c) Relative mtDNA levels in wild-type or ∆abf2 mhr1-1 cells containing empty or MHR1-overexpressing plasmids, after cultivation in Gly-U media. (d) Scheme of respiratory function assay. Cells were selectively pre-cultured in Gly-U media, then 106 cells were transferred to Glu-U or RGal-U media and cultivated for <8 generations at 30 °C or 34 °C. Cells were then spread onto YPD and YPGly plates. (e) Representative plate images of ∆abf2 and ∆abf2 mhr1-1 CFUs harboring empty or MHR1-overexpressing plasmids, following growth in Glu-U or RGal-U media at 30 °C or 34 °C for <8 generations. (f) ρ+ CFU formation rate based on n = 3 independent experiments described in (d). (g) Scheme of extended respiratory function assay. Cells were selectively grown in Gly-U media, then 105 cells were transferred to RGal-U media and cultivated for two consecutive 48-hour rounds (approximately 20 generations) at 30 °C. Cells were then spread onto YPD and YPGly plates. (h) Representative plate images and (i) ρ+ CFU formation rate based on n = 3 independent experiments described in (g). (j) Scheme of extended respiratory function assay. Cells were selectively grown in Gly-U media, then 105 cells were transferred to RGal-U media and cultivated for two consecutive 48-hour rounds (approximately 20 generations) at 30 °C. Cells were then spread onto RGal-U plates, grown for four to seven days, and then replica-plated onto YPGly plates. (k) Representative plate images for n = 3 independent experiments as described in (j). (l) Scheme of extended respiratory function assay. Cells were selectively grown in Gly-U media, then 105 cells were transferred to Glu-U media and cultivated for two consecutive 48-hour rounds (approximately 20 generations) at 30 °C. Cells were then spread onto RGal-U plates, grown for four to seven days, and then replica-plated onto YPGly plates. (m) Representative plate images for n = 2 independent experiments as described in (l). All error bars indicate ± SD.
To investigate the effects of increasing the amount of Mhr1 on mtDNA content and cellular respiratory function, we constitutively overexpressed MHR1 under the ADH promoter via plasmid in ∆abf2 single-mutant and ∆abf2 mhr1-1 double-mutant cells, and confirmed Mhr1 overproduction by immunoblot analysis (Fig. 6b; Supplementary Fig. 1). qPCR analysis revealed that ∆abf2 mhr1-1 double-mutant cells harboring an empty vector had 68.4 ± 11.9% of the mtDNA level of wild-type cells. In contrast, Mhr1 overexpression resulted in an mtDNA level of 97.5 ± 26.5% (Fig. 6c).
To examine whether exogenous MHR1 expression rescues respiratory function, we compared ∆abf2 single-mutant and ∆abf2 mhr1-1 double-mutant cells harboring empty (pVT) or MHR1-overexpressing plasmids (pVT-MHR1) after 48 hours of growth (equivalent to less than eight generations) in fermentable glucose minus uracil (Glu-U) or raffinose-galactose minus uracil (RGal-U) media at 30 °C or 34 °C (Fig. 6d). We then spread equal amounts of dilute culture on YPD and YPGly plates, as described in Fig. 1. Glucose strongly reduced respiratory growth levels in ∆abf2/pVT cells, a result closely matching that in ∆abf2 cells without plasmid DNA (Fig. 1c). ∆abf2/pVT cells grown in Glu-U medium produced 67.0 ± 6.9% ρ+ colonies at 30 °C and only 16.1 ± 9.4% at 34 °C. In contrast, ∆abf2/pVT cells yielded 92.9 ± 13.3% and 94.6 ± 10.8% ρ+ colonies when grown in RGal-U at 30 °C or 34 °C, respectively. In agreement with our result from ∆abf2 cells, cellular respiratory function is significantly reduced following cultivation of ∆abf2/pVT cells at 34 °C in Glu-U medium, but not in RGal-U. There was a small decrease in respiratory growth upon MHR1-overexpression in the ∆abf2/pVT-MHR1 cells in Glu-U or RGal-U media at either 30 °C or 34 °C, indicating that additional Mhr1 is not sufficient to offset a lack of Abf2. Although nucleoid formation defects in the ∆abf2 background cause respiratory defects28,29, it is very likely such defects are unable to be prevented by MHR1-overexpression. On the other hand, ρ+ CFU formation rates were only 50.3 ± 14.8% and 3.3 ± 4.2% in ∆abf2 mhr1-1/pVT cells in Glu-U medium at 30 °C, and 34 °C, respectively. Importantly, ∆abf2 mhr1-1/pVT cells also displayed highly temperature-sensitive respiratory function after cultivation in RGal-U, giving ρ+ CFU formation rates of 81.0 ± 6.6% and 12.4 ± 7.4% at 30 °C and 34 °C, respectively. Addition of MHR1 in ∆abf2 mhr1-1/pVT-MHR1 cells significantly rescued the respiratory function of these cells in RGal-U at 34 °C, to a ρ+ CFU formation rate of 69.1 ± 27.5%. On the other hand, ∆abf2 mhr1-1/pVT-MHR1 cells cultivated in Glu-U showed no rescue effect, with ρ+ CFU formation rates of 54.5 ± 3.2% and 6.3 ± 6.8% at 30 °C and 34 °C, respectively (Fig. 6e,f).
To further advance the notion that Mhr1 can protect mtDNA integrity, we examined the effect of an extended cultivation time in fermentable media of approximately 20 generations. Simultaneous spreading of ∆abf2 mhr1-1/pVT and ∆abf2 mhr1-1/pVT-MHR1 cells onto YPD and YPGly plates (Fig. 6g) yielded 4.5 ± 4.1% and 54.0 ± 3.6% ρ+ CFUs, respectively (Fig. 6h,i), reinforcing the notion of a rescue effect for additional Mhr1 in RGal-U media. Similarly, replica-plating ∆abf2 mhr1-1/pVT and ∆abf2 mhr1-1/pVT-MHR1 colonies from RGal-U to YPGly plates showed a clear increase in the proportion of ρ+ CFUs upon MHR1 overexpression (Fig. 6j,k), while only a small proportion of CFUs remained ρ+ after cultivation in Glu-U medium (Fig. 6l,m). These results indicate that glucose impairs Mhr1-mediated action, which protects against mtDNA deletions. In summary, Mhr1 functions to prevent loss of respiratory function due to mtDNA deletion mutagenesis, although deletions are not completely prevented by overproduced Mhr1 (Fig. 7).
Model for the prevention of mtDNA deletion mutagenesis by Mhr1-driven recombination and mtDNA replication. MtDNA deletions threatening cellular respiratory function arising from genetic backgrounds such as ∆abf2 mhr1-1 can be prevented by increased homologous recombination via overproduction of Mhr1 recombinase.
Discussion
In this study, we found that increasing Mhr1 protein level prevents loss of respiratory function in cells lacking Abf2 and functional Mhr1, which display an mtDNA-instability phenotype similar to several other nuclear mutations in yeast. Our results provide further support for the notion that Mhr1 has a pivotal role in the maintenance of mitochondrial genomic integrity30, and that DSB-induced mtDNA replication by Mhr1 is the predominant form of mtDNA replication in ρ+ yeast cells23. We previously reported that Mhr1-dependent mtDNA replication and homologous recombination are crucial for repair of mtDNA DSBs22. Collectively, our results here suggest that mitochondrial homologous DNA recombination may have utility in preventing the spontaneous generation of mtDNA deletions in a variety of circumstances (Fig. 7).
The requirement for Abf2 and Mhr1 in mtDNA stability raises the question of how increasing the amount of Mhr1 alone prevents generation of deleted mtDNA and helps to sustain cellular respiratory function. The answer likely comes from one of the mechanisms of mtDNA deletion formation proposed by Krishnan et al., in which mtDNA deletions occur during repair of DSB-induced mtDNA damage, rather than during replication31. We inferred that a lack of Abf2 might weaken DSB repair by homologous recombination and increase the accumulation of deleted mtDNA molecules. Since DSB repair can be accomplished by homologous DNA recombination32,33,34, an increased amount of Mhr1 may enhance the number of homologous DNA recombination events to repair DSBs22. This interpretation implies that mtDNA deletions are prevented, mtDNA integrity is maintained, and a large proportion of cells with the ∆abf2 mutation sustain respiratory function upon overexpression of MHR1 (Fig. 6). On the other hand, since mtDNA deletion mutagenesis still occurs upon Mhr1 overproduction in ∆abf2 cells, the stabilization of mtDNA recombination intermediates by Abf214 is important. In addition, we found that a fraction of the population of ∆abf2 mhr1-1 cells contains HS ρ− mtDNA molecules (less than 20%; Fig. 4b), however this proportion is likely large enough to reduce the beneficial effect of Mhr1 overexpression on preventing deleterious mtDNA mutations. Due to the presence of HS ρ− mtDNA mutant molecules, Mhr1 overexpression likely increases the amounts of both wild type and HS ρ− mtDNA. Thus, it is very unlikely that the observed rescue of ∆abf2 mhr1-1 cells upon MHR1 overexpression (Fig. 6) is due only to increased replication and selection for wild-type mtDNA over deleterious mtDNA mutations. These results therefore indicate that increased amounts of Mhr1 can prevent mitochondrial genomic instability.
Mhr1 promotes homologous DNA pairing18,19, while Abf2 packages mtDNA and is the main component of the nucleoid7,35. Both of these molecular functions resemble that of RecA in Escherichia coli. RecA forms a nucleoprotein complex that both ensures the formation of heteroduplex DNA and protects DNA from nuclease degradation36. Since these roles are performed by discrete proteins in S. cerevisiae mitochondria, whether mtDNA-Mhr1 nucleoprotein formation occurs prior to packaging of concatemeric mtDNA molecules by Abf2 or by a contrary process, remains for further investigation.
TFAM, the mammalian ortholog of Abf2, contains an additional C-terminal domain that has been predicted to be essential for transcription13. TFAM can complement Abf2 in yeast by rescuing the loss-of-mtDNA phenotype of yeast ∆abf2 cells37. In contrast to Abf2/TFAM, the existence of a metazoan ortholog of Mhr1 remains an open issue38. To date, there have been several lines of evidence to indicate that human mtDNA recombination may occur39,40,41. For example, we demonstrated that ROS stimulate mitochondrial allele segregation from heteroplasmy towards homoplasmy in human fibroblasts42, a result consistent with the stimulatory effect of ROS and mechanism of recombination-driven mtDNA replication in yeast mitochondria21,43. Consistent with the results reported here, a method to stimulate recombination function in human mitochondria may similarly prevent human mtDNA instability and present numerous other health benefits.
Materials and Methods
Yeast strains and media
Yeast strains used in this study are listed in Table 1. General genetic techniques used in this study are described in44. Yeast transformation was carried out using the lithium-acetate method45. The assay for hypersuppressiveness was carried out according to a procedure previously described24,46. Overexpression of MHR1 was achieved using a yeast multiple-copy plasmid with a constitutive promoter (pVT100U)47. The ORF (open reading frame) of MHR1 was amplified by PCR with addition of Sac1 and Xba1 restriction sites at the 5′ and 3′ ends respectively, and inserted into pVT100U to generate pVT100U-MHR1. Immunoblot analysis for Mhr1 detection was performed using rabbit antiserum against Mhr1, according a previously established procedure22.
Media were prepared as previously described17,18,24. Selective pre-cultivation of cells for respiratory function assays was conducted either in rich glycerol (YPGly), synthetic glycerol (Gly) or synthetic glycerol minus uracil (Gly-U) media. Fermentable media, used to promote loss of respiratory function, was rich glucose (YPD), synthetic glucose (Glu), synthetic glucose minus uracil (Glu-U), synthetic raffinose-galactose (RGal), or synthetic raffinose-galactose minus uracil (RGal-U). Glycerol, glucose, raffinose and galactose concentrations used were 3% v/v, 2%, 2% and 2%, respectively. Selection for diploid cells following crossing with ∆abf2 or ∆abf2 mhr1-1 cells was carried out by cultivating cells in liquid synthetic minimal (SD) or synthetic minimal plus tryptophan (SD + W) medium containing 2% glucose at 30 °C overnight and then spreading cells on agar plates containing the same nutrients. For control crosses, diploids were selected by spreading mated cells directly onto synthetic minimal media agar plates containing leucine plus uracil (SD + LU) or tryptophan (SD + W) and 2% glucose.
Mitochondrial nucleoid analysis
WT, mhr1-1, ∆abf2 and ∆abf2 mhr1-1 cells were cultivated in YPGly media to early log-phase at 30 °C and stained with DAPI. Cells were subsequently transferred to YPD media and cultivated for two consecutive rounds at 30 °C or 34 °C, stained with DAPI and observed again. DAPI staining was performed by transferring aliquots to fresh media containing 1 µg/ml DAPI and incubating for 15 min. Cells were then washed and resuspended in fresh media and 1% low-melting point agarose at a 1:1 ratio. Cells were then mounted on glass slides and analyzed with a Deltavision fluorescence microscopy system (Applied Precision, Inc.) equipped with an Olympus IX71 microscope. DAPI foci in mother cells were counted using ImageJ software.
Tetrad analysis
We used an Olympus micromanipulation system for tetrad dissection to separate the four ascus-encapsulated spores derived from diploid WT/Δmhr1 cells. Spores were separated and placed on synthetic defined (SD) complete plates supplemented with adenine (A), leucine (L), uracil (U), histidine (H) and tryptophan (W), and cultivated at 30 °C for 7 days. Colonies were then replica-plated onto SD complete plates lacking L and YPGly plates.
Purification of yeast mtDNA and analysis by restriction enzyme digestion
MtDNA was purified from Δmhr1 spore-derived yeast cells using cesium chloride density-gradient centrifugation48. The purified mtDNA was digested with ApaI and run on a 1% agarose gel alongside DNA size markers. The DNA fragments were photographed under ultraviolet irradiation after staining the gel with 0.5 μg/ml ethidium bromide.
Southern blot analysis
∆abf2 and ∆abf2 mhr1-1 cells were cultivated in YPGly media, then transferred to RGal complete media and cultivated at 30 °C for two days. Total cellular DNA was then prepared and digested by ApaI. Approximately 15 µg of total DNA was separated by electrophoresis on a 1.0% agarose gel, run at 24 °C for 80 h at 5 V/cm, and transferred to a nylon membrane (Amersham Hybond N Plus; GE Healthcare). Signals for mtDNA were detected using 32P-labeled full-length mtDNA from budding yeast as a probe. Signals were analyzed using a Typhoon FLA 7000 biomolecular imager (GE Healthcare).
Analysis of mtDNA level by quantitative real-time PCR
Primers used for real-time PCR were as follows: COX3-Forward, 5′-TCCATTCAGCTATGAGTCCTGATG-3′; COX3-Reverse, 5′-AATTCGGTAGGTTGTACAGCTTCAA-3′; NUC1-Forward, 5′-TTTAGGTCGGGCTATGATCGA-3′; NUC1-Reverse, 5′-TCCATGGCCTGTTGAGAAAAT-3′. The 20 µl reaction volumes contained 10 µl SYBR Premix Ex TaqTM II (Takara), 0.4 µM of each primer, and 100 ng of template genomic DNA. A LightCycler 480 (Roche) was used for real-time PCR analysis27.
References
Contamine, V. & Picard, M. Maintenance and integrity of the mitochondrial genome: a plethora of nuclear genes in the budding yeast. Microbiol Mol Biol Rev 64, 281–315 (2000).
Chen, X. J. & Clark-Walker, G. D. The petite mutation in yeasts: 50 years on. Int Rev Cytol 194, 197–238 (2000).
Kang, D. & Hamasaki, N. Maintenance of mitochondrial DNA integrity: repair and degradation. Curr Genet 41, 311–322, https://doi.org/10.1007/s00294-002-0312-0 (2002).
Alexeyev, M., Shokolenko, I., Wilson, G. & LeDoux, S. The maintenance of mitochondrial DNA integrity–critical analysis and update. Cold Spring Harb Perspect Biol 5, a012641, https://doi.org/10.1101/cshperspect.a012641 (2013).
Holt, I. J., Harding, A. E. & Morgan-Hughes, J. A. Deletions of muscle mitochondrial DNA in mitochondrial myopathies: sequence analysis and possible mechanisms. Nucleic Acids Res 17, 4465–4469 (1989).
Corral-Debrinski, M. et al. Marked changes in mitochondrial DNA deletion levels in Alzheimer brains. Genomics 23, 471–476, https://doi.org/10.1006/geno.1994.1525 (1994).
Chen, X. J. & Butow, R. A. The organization and inheritance of the mitochondrial genome. Nat Rev Genet 6, 815–825, https://doi.org/10.1038/nrg1708 (2005).
Diffley, J. F. & Stillman, B. DNA binding properties of an HMG1-related protein from yeast mitochondria. J Biol Chem 267, 3368–3374 (1992).
Newman, S. M., Zelenaya-Troitskaya, O., Perlman, P. S. & Butow, R. A. Analysis of mitochondrial DNA nucleoids in wild-type and a mutant strain of Saccharomyces cerevisiae that lacks the mitochondrial HMG box protein Abf2p. Nucleic Acids Res 24, 386–393 (1996).
Brewer, L. R. et al. Packaging of single DNA molecules by the yeast mitochondrial protein Abf2p. Biophys J 85, 2519–2524, https://doi.org/10.1016/S0006-3495(03)74674-8 (2003).
Fisher, R. P., Lisowsky, T., Parisi, M. A. & Clayton, D. A. DNA wrapping and bending by a mitochondrial high mobility group-like transcriptional activator protein. J Biol Chem 267, 3358–3367 (1992).
Van Dyck, E. & Clayton, D. A. Transcription-dependent DNA transactions in the mitochondrial genome of a yeast hypersuppressive petite mutant. Mol Cell Biol 18, 2976–2985 (1998).
Kukat, C. & Larsson, N. G. mtDNA makes a U-turn for the mitochondrial nucleoid. Trends Cell Biol 23, 457–463, https://doi.org/10.1016/j.tcb.2013.04.009 (2013).
MacAlpine, D. M., Perlman, P. S. & Butow, R. A. The high mobility group protein Abf2p influences the level of yeast mitochondrial DNA recombination intermediates in vivo. Proc Natl Acad Sci USA 95, 6739–6743 (1998).
Cho, J. H., Lee, Y. K. & Chae, C. B. The modulation of the biological activities of mitochondrial histone Abf2p by yeast PKA and its possible role in the regulation of mitochondrial DNA content during glucose repression. Biochim Biophys Acta 1522, 175–186 (2001).
Zelenaya-Troitskaya, O., Newman, S. M., Okamoto, K., Perlman, P. S. & Butow, R. A. Functions of the high mobility group protein, Abf2p, in mitochondrial DNA segregation, recombination and copy number in Saccharomyces cerevisiae. Genetics 148, 1763–1776 (1998).
Ling, F., Makishima, F., Morishima, N. & Shibata, T. A nuclear mutation defective in mitochondrial recombination in yeast. EMBO J 14, 4090–4101 (1995).
Ling, F. & Shibata, T. Recombination-dependent mtDNA partitioning: in vivo role of Mhr1p to promote pairing of homologous DNA. EMBO J 21, 4730–4740 (2002).
Ling, F., Yoshida, M. & Shibata, T. Heteroduplex joint formation free of net topological change by Mhr1, a mitochondrial recombinase. J Biol Chem 284, 9341–9353, https://doi.org/10.1074/jbc.M900023200 (2009).
Ling, F. & Shibata, T. Mhr1p-dependent concatemeric mitochondrial DNA formation for generating yeast mitochondrial homoplasmic cells. Mol Biol Cell 15, 310–322, https://doi.org/10.1091/mbc.E03-07-0508 (2004).
Hori, A., Yoshida, M., Shibata, T. & Ling, F. Reactive oxygen species regulate DNA copy number in isolated yeast mitochondria by triggering recombination-mediated replication. Nucleic Acids Res 37, 749–761, https://doi.org/10.1093/nar/gkn993 (2009).
Ling, F. et al. Din7 and Mhr1 expression levels regulate double-strand-break-induced replication and recombination of mtDNA at ori5 in yeast. Nucleic Acids Res 41, 5799–5816, https://doi.org/10.1093/nar/gkt273 (2013).
Prasai, K., Robinson, L. C., Scott, R. S., Tatchell, K. & Harrison, L. Evidence for double-strand break mediated mitochondrial DNA replication in Saccharomyces cerevisiae. Nucleic Acids Res 45, 7760–7773, https://doi.org/10.1093/nar/gkx443 (2017).
Ling, F., Hori, A. & Shibata, T. DNA recombination-initiation plays a role in the extremely biased inheritance of yeast [rho-] mitochondrial DNA that contains the replication origin ori5. Mol Cell Biol 27, 1133–1145, https://doi.org/10.1128/MCB.00770-06 (2007).
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Ephrussi, B., Jakob, H. & Grandchamp, S. Etudes Sur La SuppressivitE Des Mutants a Deficience Respiratoire De La Levure. II. Etapes De La Mutation Grande En Petite Provoquee Par Le Facteur Suppressif. Genetics 54, 1–29 (1966).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55, 611–622, https://doi.org/10.1373/clinchem.2008.112797 (2009).
Marin-Garcia, J., Pi, Y. & Goldenthal, M. J. Mitochondrial-nuclear cross-talk in the aging and failing heart. Cardiovasc Drugs Ther 20, 477–491, https://doi.org/10.1007/s10557-006-0584-6 (2006).
Lee, S. R. & Han, J. Mitochondrial Nucleoid: Shield and Switch of the Mitochondrial Genome. Oxid Med Cell Longev, https://doi.org/10.1155/2017/8060949 (2017).
Fritsch, E. S., Chabbert, C. D., Klaus, B. & Steinmetz, L. M. A genome-wide map of mitochondrial DNA recombination in yeast. Genetics 198, 755–771, https://doi.org/10.1534/genetics.114.166637 (2014).
Krishnan, K. J. et al. What causes mitochondrial DNA deletions in human cells? Nat Genet 40, 275–279, https://doi.org/10.1038/ng.f.94 (2008).
Mehta, A. & Haber, J. E. Sources of DNA double-strand breaks and models of recombinational DNA repair. Cold Spring Harb Perspect Biol 6, a016428, https://doi.org/10.1101/cshperspect.a016428 (2014).
Osman, F. & Subramani, S. Double-strand break-induced recombination in eukaryotes. Prog Nucleic Acid Res Mol Biol 58, 263–299 (1998).
Jasin, M. & Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb Perspect Biol 5, a012740, https://doi.org/10.1101/cshperspect.a012740 (2013).
Kucej, M., Kucejova, B., Subramanian, R., Chen, X. J. & Butow, R. A. Mitochondrial nucleoids undergo remodeling in response to metabolic cues. J Cell Sci 121, 1861–1868, https://doi.org/10.1242/jcs.028605 (2008).
Cox, M. M., Morrical, S. W. & Neuendorf, S. K. Unidirectional branch migration promoted by nucleoprotein filaments of RecA protein and DNA. Cold Spring Harb Symp Quant Biol 49, 525–533 (1984).
Parisi, M. A., Xu, B. & Clayton, D. A. A human mitochondrial transcriptional activator can functionally replace a yeast mitochondrial HMG-box protein both in vivo and in vitro. Mol Cell Biol 13, 1951–1961 (1993).
Wiuf, C. Recombination in human mitochondrial DNA? Genetics 159, 749–756 (2001).
Hagelberg, E. et al. Evidence for mitochondrial DNA recombination in a human population of island Melanesia. Proc Biol Sci 266, 485–492, https://doi.org/10.1098/rspb.1999.0663 (1999).
Kraytsberg, Y. et al. Recombination of human mitochondrial DNA. Science 304, 981, https://doi.org/10.1126/science.1096342 (2004).
Zsurka, G. et al. Recombination of mitochondrial DNA in skeletal muscle of individuals with multiple mitochondrial DNA heteroplasmy. Nat Genet 37, 873–877, https://doi.org/10.1038/ng1606 (2005).
Ling, F. et al. Reactive oxygen species stimulate mitochondrial allele segregation toward homoplasmy in human cells. Mol Biol Cell 27, 1684–1693, https://doi.org/10.1091/mbc.E15-10-0690 (2016).
Ling, F., Mikawa, T. & Shibata, T. Enlightenment of yeast mitochondrial homoplasmy: diversified roles of gene conversion. Genes (Basel) 2, 169–190, https://doi.org/10.3390/genes2010169 (2011).
Kaiser, C., Michaelis, S. & Mitchell, A. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual. Cold Spring Harbor Laboratory Press, Plainview, New York, NY. (1994).
Ito, H., Fukuda, Y., Murata, K. & Kimura, A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol 153, 163–168 (1983).
Bradshaw, E., Yoshida, M. & Ling, F. Regulation of Small Mitochondrial DNA Replicative Advantage by Ribonucleotide Reductase in Saccharomyces cerevisiae. G3 (Bethesda) 7, 3083–3090, https://doi.org/10.1534/g3.117.043851 (2017).
Westermann, B. & Neupert, W. Mitochondria-targeted green fluorescent proteins: convenient tools for the study of organelle biogenesis in Saccharomyces cerevisiae. Yeast 16, 1421–1427, https://doi.org/10.1002/1097-0061(200011)16:15<1421::AID-YEA624>3.0.CO;2-U (2000).
Hudspeth, M. E., Shumard, D. S., Tatti, K. M. & Grossman, L. I. Rapid purification of yeast mitochondrial DNA in high yield. Biochim Biophys Acta 610, 221–228 (1980).
Acknowledgements
This work was supported in part by a Grant-in-Aid for Scientific Research (C) (No. 23510237 and No. 17K07294) from the Ministry of Education, Culture, Sports, Science and Technology of Japan to F.L.; by an Incentive Research Grant from RIKEN to F.L.; by a grant from the RIKEN Strategic Research Program; and by a grant from JST-CREST to F.L.
Author information
Authors and Affiliations
Contributions
L.F. designed the study. L.F. and E.B. designed and performed experiments and wrote the paper. L.F. and E.B. prepared the figures. M.Y. provided critical feedback and advice for the writing of the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ling, F., Bradshaw, E. & Yoshida, M. Prevention of mitochondrial genomic instability in yeast by the mitochondrial recombinase Mhr1. Sci Rep 9, 5433 (2019). https://doi.org/10.1038/s41598-019-41699-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41699-9
This article is cited by
-
Shared and more specific genetic determinants and pathways underlying yeast tolerance to acetic, butyric, and octanoic acids
Microbial Cell Factories (2024)
-
petiteFinder: an automated computer vision tool to compute Petite colony frequencies in baker’s yeast
BMC Bioinformatics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.