Abstract
Mitochondrial genomes exhibit diverse features among eukaryotes in the aspect of gene content, genome structure, and the mobile genetic elements such as introns and plasmids. Although the number of published mitochondrial genomes is increasing at tremendous speed, those of several lineages remain unexplored. Here, we sequenced the complete mitochondrial genome of a unicellular heterotrophic eukaryote, Marophrys sp. SRT127 belonging to the Centroheliozoa, as the first report on this lineage. The circular-mapped mitochondrial genome, which is 113,062 bp in length, encodes 69 genes typically found in mitochondrial genomes. In addition, the Marophrys mitochondrial genome contains 19 group I introns. Of these, 11 introns have genes for homing endonuclease (HE) and phylogenetic analyses of HEs have shown that at least five Marophrys HEs are related to those in green algal plastid genomes, suggesting intron transfer between the Marophrys mitochondrion and green algal plastids. We also discovered a putative mitochondrial plasmid in linear form. Two genes encoded in the circular-mapped mitochondrial genome were found to share significant similarities to those in the linear plasmid, suggesting that the plasmid was integrated into the mitochondrial genome. These findings expand our knowledge on the diversity and evolution of the mobile genetic elements in mitochondrial genomes.
Similar content being viewed by others
Introduction
Mitochondria have emerged by the endosymbiosis between the last common ancestor of eukaryotes and an α-proteobacterium. All extent eukaryotes have mitochondria or mitochondrial remnant organelles, with only one exception of Monocercomonoides sp. PA2031 (now classified as Monocercomonoides exilis2). Although most of the genes in the α-proteobacterium that gave rise to the ancestral mitochondrion have been lost or transferred to the host nucleus during organellogenesis, typical mitochondria, which can carry out aerobic respiration, still retain their genomes3 (mitochondrial genomes, mtDNAs). The gene repertories in mtDNAs have differently reduced in individual lineages in eukaryotes from the ancestral mtDNA that encoded at least about 100 genes for oxidative phosphorylation, translation, transcription, protein transport, protein maturation and RNA processing as found in mtDNA of jakobids4,5. In contrast, apicomplexan parasites and their relatives have only three to five genes in their mtDNAs6. mtDNAs also vary with regard to genome architecture, from a simple monocircular molecule (e.g., human mtDNA) to a complex network comprising thousands of chromosomes (e.g., mtDNAs in kinetoplastids). Monolinear mtDNAs were reported from separate branches of eukaryotes as well as mtDNAs consisting of multiple linear chromosomes7,8,9,10,11. Likewise, the size range of mtDNA is broad among eukaryotes. Apicomplexan parasites are known to have the smallest mtDNAs, approximately 6 Kb in size8. On the other hand, land plants tend to contain large-sized mtDNAs, up to 11 Mb in Silene conica12.
mtDNAs of extant species vary markedly in terms of their gene repertories, genome structure, and genome size, as described above. On top of that, mobile genetic elements confer additional layers of mtDNA diversity. Group I introns and group II introns are the examples of the mobile genetic elements found in bacterial, mtDNA and plastid genomes13. Group I and group II introns are also known to be able to catalyze self-splicing by forming distinct structures, which enable us to distinguish two types of introns from DNA sequences: Group I introns form the ribozyme structure consisting of 10 stem-loops14, while group II introns require a characteristic secondary structure, such as a central wheel with six stems, for the self-splicing reaction15. Both of group I and group II introns show patchy distributions, which are considered to be formed by intron invasion from an intron-containing locus to a homologous, but intron-less locus in the same species and/or a distantly related organisms16,17. Group I intron transfers are facilitated by homing endonucleases (HEs) encoded by introns themselves (Note that HEs can act as maturases that facilitate intron splicing as well13). The HEs introduce a double-strand break in the recipient (intron-less) allele that leads to the homologous recombination between intron-containing and intron-less alleles. Individual HEs possess distinct specificities for the nucleotide sequences to recognize and digest. Therefore, a particular HE can introduce the corresponding intron only into a certain position in a genome16,18. Conversely, the group I introns (and the corresponding HEs) found in the homologous position in different genomes are expected to share their evolutionary origin19,20,21. The mechanism of group II intron transfer is different from that of group I introns, but intron-encoded proteins and their sequence recognition specificities play a central role in invasion for group II intron, as in group I intron. Thus, the evolutionary history of group II introns can be retraced by analyzing intron positions and intron-encoded proteins22,23.
Another type of mobile genetic elements in mitochondria is the linear plasmids24. Typical linear plasmids in mitochondria contain terminal inverted repeats and one or some open reading frames (ORFs) usually encoding virus-type DNA and/or RNA polymerases, designated as dpo and rpo, respectively, suggesting their independent replication and transcription25. Linear plasmids are extrachromosomal elements in mitochondria, but can occasionally integrate into mtDNA through recombination26. To date, linear plasmids in mitochondria or plasmid-derived dpo/rpo sequences in mtDNA have been reported from fungi27, a slime mold28, plants24 and a ciliate29. Consistent with the genetic mobility proposed for the linear plasmids, the phylogenies of dpo/rpo were found to be inconsistent with the organismal phylogeny30.
Heliozoa was traditionally defined as a taxonomic group of the axopodium-bearing, heterotrophic unicellular eukaryotes mainly living in freshwater environments31. At present, Heliozoa comprises two classes, namely Endohelea and Centrohelea32,33. The phylogenetic position of Heliozoa in the eukaryotes has been controversial for a long time, but recent phylogenomic study suggests that Heliozoa and haptophytes form a monophyletic clade, Haptista, as a sister to the supergroup consisting of stramenopiles, alveolates, and Rhizaria34,35,36. Although data on the nuclear-encoded genes have been accumulated for heliozoans, as far as we know, no complete mtDNA sequence has been obtained from any member belonging to this group.
Here, we first present the complete mtDNA of a member of the Centrohelea, Marophrys sp. strain SRT127. The circular-mapped mtDNA of Marophrys contains 69 typical mtDNA-encoded genes. Up to 20 group I introns were found in six out of the 69 genes, and phylogenetic analyses of the corresponding HEs suggested that at least five introns share their origins with those in green algal plastid genomes. We also identified a linear plasmid carrying genes encoding dpo and rpo. The plasmid is likely to localize in mitochondrion because both mtDNA and plasmid share a deviant genetic code in which UGA codon assigns tryptophan. We further provide the evidence for the linear plasmid being integrated into the mtDNA. These findings expand our knowledge of the diversity of mobile genetic elements in mtDNAs.
Results and Discussion
Mitochondrial genome overview
The mtDNA of a centrohelid Marophrys sp. SRT127 is 113,062 bp in length and mapped as circular (Fig. 1). The G + C contents of the mtDNA is 44.7%. It contains 42 kinds of protein-coding genes, which have been vertically inherited from the ancestor of all mitochondria. These 42 protein-coding genes include those for translation elongation factor (tufA) and the subunit of cytochrome c oxidase assembly (cox11), which are notable as most of the mtDNAs do not retain these genes except those in a limited number of eukaryotes4,37,38,39,40,41,42. In addition, the Marophrys mtDNA includes genetically mobile genes, namely those of dpo, rpo and 11 intronic HEs (see below). The Marophrys mtDNA also contains 12 functionally unidentified open reading frames (ORFs) longer than 100 amino acid residues. Large and small subunits of rRNA genes plus 5S rRNA gene were detected. We identified twenty kinds of genes for tRNAs that could translate 50 codons that cover 17 amino acids in total. During our BLAST analyses of the Marophrys mtDNA sequence against NCBI nr database, we observed UGA at the conserved tryptophan positions in the putative amino acid sequences. Thus, we concluded that UGA in the Marophrys mtDNA is used as tryptophan codon, instead of the termination signal for translation. Genes of the tRNAs that bind the codons UUA for leucine, UGA for tryptophan, ACN for threonine, AAR for lysine and CGN for arginine (R = A or G; N = A, C, G or T) were not found (Table S1). Genes coding for the RNA components for tmRNA and RNAase P could not be detected.
Mitochondrial genome of Marophrys sp. SRT127. Protein-coding genes and rRNA genes are shown in the cyan and orange boxes, respectively. Introns are indicated by light green and genes for homing endonuclease (HE) in introns are shown in the gray boxes. The genes for tRNA are depicted as black lines. Two red arrows indicate inverted repeat regions.
Overview of the introns in the Marophrys mtDNA
In the Marophrys mtDNA, the genes for cytochrome oxidase c subunit 1 (cox1), apocytochrome b (cob), ATP synthetase subunit 1 (atp1), and large and small subunits of rRNA (rnl and rns) appeared to be split by a single or up to nine introns. The sequence motif shared among the canonical group I introns was detected from all of the introns in the Marophrys mtDNA (Table 1). A series of RT-PCR experiments confirmed that all of the introns were removed from the mature mRNAs (Fig. S1A). Henceforth here, we designate the introns found in the Marophrys mtDNA and those of the HEs harbored in the introns as follows. Introns are designated by adding ‘_i’ (intron) to their host gene names with the ascending numbers from the 5′ terminus (e.g., rnl_i1 and rnl_i2 are the first and second introns in the rnl gene, respectively). The HEs are distinct from each other by adding the corresponding superscripted host intron names (e.g., HEs harbored in rnl_i3 and rnl_i4 are designated as HErnl_i3 and HErnl_i4, respectively).
The gene for NADH dehydrogenase subunit 5 (nad5) appeared to be split into two distant loci in the mtDNA, which were designated as nad5_a and nad5_b encoding the N- and C-terminal halves of the protein, respectively (Fig. 1). According to the organization of the two loci in the mtDNA, they are most likely transcribed independently. We successfully obtained the evidence for a single, contentious mRNA molecule comprising the transcripts from nad5_a and nad5_b by performing reverse transcription (RT)-PCR using cDNA as template and two primers—one specific to the nad5_a nucleotide sequence and the other to the nad5_b nucleotide sequence (Figs 2A and S1B). Thus, the expression of nad5 most likely requires trans-splicing. Likewise, we identified separate loci encoding the N- and C-terminal of Cox1, termed cox1_a and cox1_b, respectively (Fig. 1). Again, the RT-PCR provided evidence for the transcripts from cox1_a and cox1_b being trans-spliced into a single mRNA molecule encoding the entire Cox1 (Figs 2A and S1B).
Trans-splicing for nad5 and cox1 gene expressions in the Marophrys mitochondrial genome. (A) Reverse transcription PCR using a set of primers specific to nad5_a and nad5_b loci (left) and that specific to cox1_a and cox1_b loci (right). The DNA fragment was amplified from the cDNA template which most likely contained the spliced product connecting the two RNA fragments transcribed from the two separate loci together (lanes labelled with “cDNA”). On the other hand, no specific amplification was observed in the PCR using the genomic DNA template due to the configuration of the two separate loci in the mtDNA (lanes labelled with “gDNA”). (B-C) Model for nad5 mRNA trans-splicing. (B) Primary structures of nad5_a and nad5_b loci. mtDNA, exons, and introns are shown in thin black lines, boxed, and thick lines, respectively. The 5′ exon and subsequent intronic region are colored in red, while the 3′ exon and its preceding intronic region are indicated in blue. The two loci are located on the different strands, and thus transcribed independently from each other. (C) Putative group I intron-like secondary structure of nad5_a and nad5_b transcripts. Five stem-loop structures conserved among group I intron ribozymes (P3, P4, P6, P7, and P8) can be formed within the nad5_a transcript and between the nad5_a and nad5_b transcripts. This secondary structure was predicted by RNAweasel followed by manual inspection and modification. Watson–Crick base pairings and a wobble bond in the five stem-loop structures are indicated by the black lines and a circle, respectively. The typical secondary structure of group I intron ribozymes is schematically shown as an inset. (D) Mature nad5 mRNA.
RNAweasel suggested that the 3′ and 5′ flanking region of the transcript from nad5_a and nad5_b loci, respectively, can form the group I-specific secondary structure together (Fig. 2B–D). Interestingly, two of the predicted stem structures, P7 and P8, can be folded by a combination of the transcripts from nad5_a and nad5_b loci (Fig. 2C). Therefore, the trans-splicing between nad5_a and nad5_b transcripts is most likely mediated by a group I intron (Fig. 2B–D). Prior to this study, the group I intron-mediated trans-splicing was found in the mtDNAs of a placozoan43, green algae44,45 and an arbuscular mycorrhizal fungus46. However, these examples were limited to the cox1 and rnl genes. Hence, the Marophrys mtDNA is the first example of group I intron-mediated trans-splicing in nad5.
Neither RNAweasel nor Infernal detected any conserved motif of group I (or group II) introns in the nucleotide sequences flanking the cox1_a or cox1_b locus. Thus, we currently have no insight into the mechanism mediating the trans-splicing between the transcripts from the two cox1 loci.
Origins of group I introns harboring HEs
Overall, 11 of the 19 introns identified in the Marophrys mtDNA harbor intronic ORFs encoding LAGLIDADG motif-containing HEs, which have been typically been found in group I introns16. All of the HEs in the Marophrys mtDNA belong to either LAGLIDADG_1 (pfam00961) or LAGLIDADG_2 (pfam031611). Here, we explored the evolutionary origins of the 11 group I introns harboring HEs by combining phylogenetic affinities of the HEs and the insertion positions of the introns. We are aware of the cases in which the evolution of a group I intron and that of the corresponding intron-encoded protein disagreed to one another47. Unfortunately, we could not exam whether introns and their intron-encoded proteins coevolved, as the intron (nucleotide) sequences dealt in this study are too diverged for phylogenetic analyses. Thus, we assumed the coevolution of each pair of a group I introns and its HE in the following sections. Table 1 summarizes the group I introns found in the Marophyrs mtDNA and their putative origins.
Introns sharing the origins with green algal organellar genomes
Marophrys atp1 intron (atp1_i1) harbors an HE (HEatp1_i1). The HE phylogeny grouped HEatp1_i1 with those harbored in atpA introns, which were found in the plastid genomes (plDNAs) of green algae belonging to core Chlorophyta, with an ML bootstrap value (MLBP) of 97% (node A in Figs 3 and S2). The atpA introns in green algal plDNA and Marophrys atp1_i1 appeared to be inserted in the homologous positions, namely, phase 0 of the 164th codon for glutamine in atp1 and the 165th codon for arginine in atpA (Figs 3 and S2). The results described above consistently and strongly suggest that lateral transfer of an intron took place between a centrohelid mtDNA and a green algal plDNA. Nevertheless, the data presented above are insufficient to draw definitive conclusions about whether the ancestral intron emerged in an mtDNA or a plDNA. Interestingly, DNA transfer from an mtDNA to a plDNA has been considered to occur rarely48,49, and many cases of DNA transfer with the opposite direction have been documented50,51. Thus, we favor the intron transfer from a green alga to a centrohelid over that in the opposite direction. Moreover, centrohelids probably encounter opportunities to uptake the genetic materials of green algae in the natural environments. Marophrys sp. SRT127 and other centrohelids prey on green algae, and some centrohelids are capable of sequestering the plastids of their prey algae for a certain period, which is known as kleptoplasty52. Taking these findings together, we propose that atp1_i1 in the Marophrys mtDNA was laterally transferred from a green algal plDNA.
Maximum likelihood (ML) analysis of the homing endonucleases harbored in the first intron in atp1 (HEatp1_i1) in the Marophrys mitochondrial genome. OTU names consist of species name, intron-hosting gene (bold) and accession number in brackets. Green algal sequences are shown in green. Gene names are colored in green when they reside in plastid genomes. The inserted positions of atpA/atp1 introns are presented the numbers of the amino acid residues correspond to the Marophrys sequence. Asterisks indicate insertion site in triplet codon. For instance, “*Gln164” means that an intron was found between the codon for Gln164 and that for the 163th amino acid (phase class = 0). Ultrafast bootstrap values higher than 70% are shown. Collapsed clades are indicated by triangles and the linages included in the clade are shown with the number of genes in parentheses. The detailed ML tree is given in Fig. S2.
We also detected four rnl introns in Marophrys mtDNA that shared origins with those in organellar genomes (mtDNAs or plDNAs) in green algae. The HE harbored in Marophrys rnl_i3 (HErnl_i3) grouped together with the HEs in two mitochondrial and one plastid rnl genes in green algae with an MLBP of 97% (node A in Fig. S3). Among the introns harboring the HEs united by node A in Fig. S3, Marophrys rnl_i3 and two other introns appeared to be inserted in the homologous position (i.e., the introns were found between G1809 and A1810; nucleotide numbering is based on the Marophrys rnl gene).
The HE harbored in Marophrys rnl_i4 (HErnl_i4) branched at the base of the clade of nucleus-encoded HEs in land plants with an MLBP of 87% (node A in Fig. S4). The nucleus-encoded HEs in land plants are not encoded by intronic ORFs (designated as “stand-alone” in Fig. S4). Marophrys HErnl_i4 and the nucleus-encoded HEs were then connected with the HEs harbored in eight plastid and one mitochondrial rnl introns in green algae with an MLBP of 85% (node B in Fig. S4). In the clade united by node B, HErnl_i4 was excluded from both stand-alone HEs in land plants, and HEs in organellar rnl introns in green algae. It is difficult to clarify the origin of HErnl_i4 with a confidence based on the HE phylogeny (Fig. S4). However, the particular HE in the Marophrys mtDNA is encoded by an intronic ORF (not by a stand-alone), and Marophrys rnl_i4 and eight plastid rnl introns in green algae appeared to be inserted between T2151 and C2152 (nucleotide numbering is based on the Marophrys rnl gene). Taking these findings together, we propose that Marophrys rnl_i4 and the eight rnl introns in green algal plDNA are derived from a single rnl intron.
The HE harbored in Marophrys rnl_i6 (HErnl_i6) and those found in four plastid and four mitochondrial rnl introns in green algae formed a clade with an MLBP of 91% (node C in Fig. S4). Among the introns harboring the HEs united by node C in Fig. S4, the Marophrys intron was found to share the insertion position with three plastid introns (i.e., the introns were found between C2394 and A2395; nucleotide numbering is based on the Marophrys rnl gene). We concluded that Marophrys rnl_i6 and the rnl introns found in green algal organellar genomes share the common ancestor.
The HE harbored in Marophrys rnl_i8 (HErnl_i8) and that in the rnl intron in the Acanthamoeba castellanii mtDNA grouped together with an MLBP of 94%, and the two introns appeared to be inserted between G2482 and A2483 (nucleotide numbering is based on the Marophrys rnl gene; node A in Fig. S5). Moreover, the clade of HErnl_i8 and the Acanthamoeba HE clustered with two stand-alone HEs in bacterial genomes, HEs found in 24 rnl introns in green algal mtDNAs/plDNAs and a single rnl intron in a diatom mtDNA (MLBP = 86%, node B in Fig. S5). The vast majority of the rnl introns described here were found in a homologous position. Altogether, we suspect that the ancestral intron, which gave rise to the ones in the Marophrys and Acanthamoeba mt rnl genes, resided in the rnl gene in a green algal mtDNA or plDNA.
By combining the phylogenetic affinities of HEs and intron positions, we propose that the four rnl HEs (and their host introns) found in the Marophrys mtDNA share origins with those in organellar genomes in green algae as well as those of atp1. The atp1 and rnl introns described above were likely transmitted from green algae to Marophrys by considering the predator–prey relationship between centrohelids and green algae in the wild (see above). We presume that, once mobile genetic elements resided in a centrohelid mtDNA, these elements may have persisted in the descendant genomes, as centrohelids are unicellular and asexual eukaryotic lineage53. Unfortunately, the data obtained in this study failed to (i) pinpoint the green algal species that donated the five introns or (ii) clarify whether mtDNA or plDNA was the origin of each of the four rnl introns. It is also important to re-examine the putative green algal origins of the five introns (and their HEs) in future studies incorporating organellar genome data sampled from much broader eukaryotes (green algae and centrohelids in particular).
It should be noted that, in the strict sense, we cannot rule out the possibility of intron-HE coevolution being violated in the Marophyrs mtDNA, as the intron phylogenies were not examined due to their extremely divergent natures. Nevertheless, the HE phylogenies (Figs 3 and S2–5) firmly suggest that the HE-coding DNA fragments were transferred from green algal organellar genomes to centrohelid mtDNAs.
Other introns
The HE found in Marophrys rns_i1 was placed within a radiation of the HEs in rns genes in green algal plastid and bacterial genomes, and this clade received an MLBP of 96% (node D in Fig. S4). Within this clade, the affinity between the Marophrys and bacterial HEs could not be excluded, leaving the origin of this intron inconclusive.
The HE phylogeny tied together the HE found in Marophrys cob_i2 and that in a fungal cob intron with an MLBP of 86% (node E in Fig. S4). However, the inserted positions of the two cob introns appeared to be distant from each other. Thus, it remains unclear whether the two cob introns genuinely share the same origin.
The HE phylogeny united the HE found in Marophrys cox1_i1 (HEcox1_i1) and those in two green algal and three fungal cox1 genes with an MLBP of 78% (node A in Fig. S6). However, HEcox1_i1 showed no special affinity to any of the five HEs in this clade. The inserted positions of Marophrys cox1_i1 and those harboring the five HEs grouped with HEcox1_i1 appeared to vary. We consider that the results described above are insufficient to infer the origin of Marophrys cox1_i1 with confidence.
As we yielded no insight for the origin of cox1_i3, cox1_i6 or rnl_i5, as none of their HEs showed any particular affinities to other HEs considered in the analyses (Figs S4 and S7).
Plasmid and plasmid-derived genes
In addition to the circular mtDNA, we identified a linear plasmid of 5,877 bp in length with inverted repeats at both ends (Fig. S8). This plasmid contains genes encoding virus-type dpo and rpo (Fig. S8). The two genes in the Marophrys plasmid most likely use UGA for tryptophan codon instead of translation termination signal. Although no experimental evidence is available, we consider that the linear plasmid localizes in the Marophrys mitochondrion for two reasons described below. First, the structure and gene content of the Marophrys plasmid resemble those of mitochondrial plasmids found in land plants, fungi, amoeba and a ciliate Oxytricha trifallax24,25,26,27,28,29. Second, the circular mtDNA and linear plasmid share the same deviant genetic code. Given the patchy distribution in the tree of eukaryote, the linear plasmids are regarded as one of the mobile genetic elements23,29,54. Altogether, we here propose that the Marophrys plasmid was laterally acquired from an as-yet-unknown eukaryote.
The Marophrys plasmid showed clear similarity at the nucleotide level to the region containing dpo and rpo in the circular mtDNA (Fig. S8). As observed in other eukaryotes25,54, we suspect that the linear plasmid was integrated into the circular mtDNA in the Marophrys mitochondrion55. The inverted repeats were proposed to facilitate the linear plasmid to integrate into an mtDNA, and, in some cases, the entire plasmid including the repeats were found in the mtDNA56. In the Marophrys mitochondrion, the inverted repeats in the linear plasmid (102 bp in length) were found to be totally different from those surrounding dpo and rpo in the mtDNA (the regions highlighted by red arrows in Fig. 1; 7,659 bp), indicating that only two genes in the plasmid were integrated into the mtDNA. Nevertheless, it is attractive to hypothesize that plasmid integration (more precisely integration of dpo and rpo) triggered the duplication of a ~7.6 Kb region containing nad9, cob, cox1_a, two ORF and three tRNA genes observed in the current Marophrys mtDNA (red arrows in Fig. 1), as Sakurai et al.57 reported the structural change in an mtDNA led by plasmid integration. Unfortunately, the mtDNA from a single centrohelid species is insufficient to examine whether the inverted repeats in the Marophrys mtDNA arose with the plasmid integration. Relevant to the above issue, we need to clarify the mechanism by which plasmid integration introduced the inverted repeats in the recipient DNA molecule. As the first step toward resolving the issues mentioned above, we require additional mtDNA data from multiple centrohelids, particularly close relatives of Marophrys.
Methods
Isolation and culturing
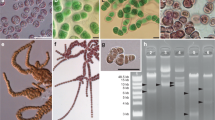
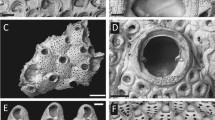
A clonal culture of a centrohelid (strain SRT127) was established using micropipetting method from seawater sample that had been collected from Tokyo Bay, Tokyo, Japan (35.6281°N, 139.7713°E), on July 30, 2011. Based on the morphological characteristics, strain SRT127 was identified as a member of the genus Marophyrs (Fig. S9).
Marophrys sp. strain SRT127 was maintained in MNK medium (http://mcc.nies.go.jp/02medium.html#mnk) with a green alga Pyramimonas sp. as prey at 20 °C under a 14-h light / 10-h dark cycle until it died in March 2016.
DNA/RNA extraction and cDNA synthesis
The centrohelid cells were collected by centrifugation once they had reached confluence and the algal cells were hardly observed in the culture medium under a light microscope. Genomic DNA was extracted with 25:24:1 of phenol:chloroform:isoamyl alcohol and purified by ethanol precipitation. Total RNA was extracted using TRIzol (Thermo Fisher Scientific), following manufacturer’s instructions. The extracted RNA was used to synthesize random hexamer-primed cDNA with SuperScript II reverse transcriptase (Thermo Fisher Scientific). The cDNA was used for the PCR experiments to confirm intron splicing (see detail in Genome annotation section).
Sequencing of the Marophrys sp. mitochondrial genome
Approximately 10 μg of the extracted DNA was submitted to Illumina Sequencing technology (HiSeq 2000 at Eurofins Genomics). A total of 125,733,479 paired-end reads of 100 bp were generated (Approx. 25 Gb in total) and assembled by SPAdes58. To identify mitochondrial sequences from the assembly data, a TBLASTN59 search was performed using the putative amino acid sequences of mitochondrion-encoded proteins in a jakobid Andalucia godoyi4 as queries. A total of 228 contigs were recovered as candidates for Marophrys mtDNA fragments at the threshold E-value of <1e−10. Then, the candidate contigs were used as a queries for a BLASTN59 search against the NCBI non-redundant nucleotides database. The contigs hit to bacterial sequences with ≥ 95% nucleotide identity in the second BLAST analysis were discarded as the genome fragments originated from bacteria contaminating in the Marophrys culture. After filtering bacterial sequences, two contigs, which were 90,947 and 7,652 bp in length, were recovered as mtDNA fragments of Marophrys. By mapping the paired-end reads on the two putative mtDNA fragments using Bowtie260, we additionally identified a contig of 6,811 bp in length as a mtDNA fragment. The third contig was overlooked by the first TBLASTN59 analysis, as this region only carries dpo and rpo genes that encode non-typical mitochondrion-encoded proteins. The physical continuity of three candidate mtDNA fragments was confirmed by PCR. Finally, the Marophrys mtDNA was reconstructed as a circular molecule of 113,062 bp in length.
Linear plasmid carrying dpo and rpo
A contig of 5,877 bp in length was found by a TBLASTN59 search against the assembly data using the dpo and rpo genes identified in the circular-mapped mtDNA as queries. This 5,877 bp-contig appeared to bear inverted repeats of 102 bp in length at both 5′ and 3′ ends, and to carry only dpo and rpo. To determine whether the contig is a linear or circular molecule, we mapped the paired-end reads to the contig using Bowtie260. We also conducted a PCR experiment to connect both ends of the contig as done to the circular mtDNA (see above). As neither of the aforementioned experiments positively supported the circular structure of the contig (data not shown), we concluded that the 5,877-bp contig represents a linear DNA molecule.
Genome annotation
The Marophrys mtDNA was annotated by Prokka61, followed by manual curation. NCBI translational Table 4 (https://www.ncbi.nlm.nih.gov/Taxonomy/Utils/wprintgc.cgi#SG4) was applied during the aforementioned annotation, because UGA codons are most likely assigned as tryptophan, not as the termination signal for translation in the Marophrys mtDNA. When a gene appeared to contain one or more introns, the corresponding cDNA was amplified by PCR with the primers listed in Table S2, and the amplicons (Fig. S1A,B) were sequenced by the Sanger method. The precise intron–exon boundaries were determined by comparing the corresponding cDNA and genome sequences. The type of introns was predicted by RNAweasel (http://megasun.bch.umontreal.ca/cgi-bin/RNAweasel/RNAweaselInterface.pl). The secondary structural motif conserved among group I introns was also searched by Infernal62 with covariance models built by Nawrocki et al.63.
Phylogenetic analyses of HEs
The conceptual amino acid sequences of the HEs identified in the Marophrys mtDNA were used as queries for a BLASTP59 search against the NCBI nr database (E-values threshold < 1e−10) to generate phylogenetic datasets of HEs. The HE sequences retrieved from the NCBI nr database, which shared >90% identity at the amino acid sequence level, were clustered by CD-HIT64,65. To reduce the redundancy, the representative sequences from each cluster were selected to build the phylogenetic datasets analyzed in this study. The HE sequences harbored in the third and sixth introns of cox1 gene appeared to share the same evolutionary origin, and we prepared a single dataset including both HEs and their related sequences. For a similar reason, we prepared a single dataset containing HEs harbored in three of the introns found in rnl gene, a single intron in rns gene and one of the introns found in cob gene in the Marophrys mtDNA. Individual datasets were subjected to MAFFT66 with the LINSI option for calculating the alignments. The ambiguously aligned sites in the alignments were discarded manually. Maximum likelihood (ML) analyses using IQ-TREE67 were conducted on each alignment with 1,000 ultrafast bootstrap replicates68. The substitution models were selected by ModelFinder69 implemented in IQ-TREE.
Data Availability
The Marophrys mtDNA sequences with annotation are available at GenBank/EMBL/DDBJ (accession nos AP019310 and AP019311). The alignments of HEs used to estimate phylogenetic tree are available from the corresponding author on request.
References
Karnkowska, A. et al. A Eukaryote without a mitochondrial organelle. Curr. Biol. 26, 1274–84 (2016).
Treitli, S. C. et al. Molecular and Morphological Diversity of the Oxymonad Genera Monocercomonoides and Blattamonas gen. nov. Protist 169(5), 744–783 (2018).
Gray, M. W., Lang, B. F. & Burger, G. Mitochondria of protists. Annu. Rev. Genet. 38, 477–524 (2004).
Burger, G., Gray, M. W., Forget, L. & Lang, B. F. Strikingly bacteria-like and gene-rich mitochondrial genomes throughout jakobid protists. Genome Biol. Evol. 5, 418–38 (2013).
Roger, A. J., Muñoz-Gómez, S. A. & Kamikawa, R. The origin and diversification of mitochondria. Curr. Biol. 27, R1177–92 (2017).
Flegentov, P. et al. Divergent mitochondrial respiratory chains in phototrophic relatives of apicomplexan parasites. Mol. Biol. Evol. 32, 115–31 (2015).
Vahrenholz, C., Riemen, G., Pratje, E., Dujon, B. & Michaelis, G. Mitochondrial DNA of Chlamydomonas reinhardtii: the structure of the ends of the linear 15.8-kb genome suggests mechanisms for DNA replication. Curr. Genet. 24, 241–7 (1993).
Hikosaka, K., Kita, K. & Tanabe, K. Diversity of mitochondrial genome structure in the phylum Apicomplexa. Mol. Biochem. Parasitol. 188, 26–33 (2013).
Nishimura, Y. et al. Mitochondrial genome of Palpitomonas bilix: Derived genome structure and ancestral system for cytochrome c maturation. Genome Biol. Evol. 8(3090-98), 2016 (2016).
Burger, G., Forget, L., Zhu, Y., Gray, M. Q. & Lang, F. Unique mitochondrial genome architecture in unicellular relatives of animals. Proc. Natl. Acad. Sci. USA 100, 892–7 (2003).
Smith, D. R. et al. First complete mitochondrial genome sequence from a box jellyfish reveals a highly fragmented linear architecture and insights into telomere evolution. Genome Biol. Evol. 4, 52–8 (2012).
Sloan, D. B. et al. Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol. 10, e1001241, https://doi.org/10.1371/journal.pbio.1001241 (2012).
Lang, B. F., Laforest, M. J. & Burger, G. Mitochondrial introns: a critical view. Trends Genet. 23, 119–25 (2007).
Hausner, G., Hafez, M. & Edgell, R. Bacterial group I introns: mobile RNA catalysts. Mob. DNA 5, 8, https://doi.org/10.1186/1759-8753-5-8. (2014).
Toro, N. Bacterial and Archaea group II introns: additional mobile genetic elements in the environment. Environ. Microbiol. 5, 143–51 (2003).
Haugen, P., Simon, D. M. & Bhattacharya, D. The natural history of group I introns. Trends Genet. 21, 111–9 (2005).
Zimmerly, S. & Semper, C. Evolution of group II introns, Mob. DNA 6, 7, https://doi.org/10.1186/s13100-015-0037-5 (2014).
Belfort, M. & Roberts, R. J. Homing endonucleases: keeping the house in order. Nucleic Acids Res. 25, 3379–88 (1997).
Cho, Y., Qiu, Y. L., Kuhlman, P. & Palmer, J. D. Explosive invasion of plant mitochondria by a group I intron. Proc. Natl. Acad. Sci. USA 95, 14244–9 (1998).
Sanchez-Puerta, M. V. et al. phylogenetic local horizontal transfer of the cox1 group I intron in flowering plant mitochondria. Mol. Biol. Evol. 25, 1762–77 (2008).
Nishimura, Y., Kamikawa, R., Hashimoto, T. & Inagaki, Y. Separate origins of group I intron in two mitochondrial genes of the katablepharid Leucocryptos marina. PloS One 7, e37307, https://doi.org/10.1371/journal.pone.0037307 (2012).
Bonen, L. & Vogel, J. The ins and outs of group II introns. Trends Genets. 17, 322–31 (2001).
Toor, N., Hausner, G. & Zimmerly, S. Coevolution of group II intron RNA structures with their intron-encoded reverse transcriptases. RNA 7, 1142–52 (2001).
Handa, H. Linear plasmid in plant mitochondria: peaceful coexistences or malicious invasions? Mitochondrion 8, 15–25 (2008).
Kuzmin, E. V. & Levchenko, I. V. S1 plasmid from cms-S-maize mitochondria encodes a viral type DNA-polymerase. Nucleic Acids Res. 15, 6758 (1987).
Bertrand, H., Griffiths, A. J., Court, D. A. & Cheng, C. K. An extrachromosomal plasmid is the etiological precursor of kalDNA insertion sequences in the mitochondrial chromosome of senescent Neurospora. Cell 47, 829–37 (1986).
Chan, B. S., Court, D. A., Vierula, P. J. & Bertrand, H. The kalilo linear senescence-inducing plasmid of Neurospora is an invertron and encodes DNA and RNA polymerases. Curr. Genet. 20, 225–37 (1991).
Takano, H., Kawano, S. & Kuroiwa, T. Genetic organization of a linear mitochondrial plasmid (mF) that promotes mitochondrial fusion in Physarum polycephalum. Curr. Genet. 26, 506–11 (1994).
Swart, E. C. et al. The Oxytricha trifallax mitochondrial genome. Genome Biol. Evol. 4, 136–54 (2012).
Warren, J. M., Simmons, M. K., Wu, Z. & Sloan, D. B. Linear plasmids and the rate of sequence evolution in plant mitochondrial genomes. Genome Biol. Evol. 8, 364–74 (2016).
Mikrjukov, K. A. System and phylogeny of Heliozoa: should this taxon exist in modern systems in protists? Zool Z. 79, 883–97 (2000).
Yabuki, A., Chao, E. E., Ishida, K. & Cavalier-Smith, T. Microheliella maris (Microhelida ord. n.), an ultrastructurally highly distinctive new axopodial protist species and genus, and the unity of phylum Heliozoa. Protist 163, 356–88 (2012).
Cavalier-Smith, T., Chao, E. E. & Lewis, R. Multiple origins of Heliozoa from flagellate ancestors: New cryptist subphylum Corbihelia, superclass Corbistoma, and monophyly of Haptista, Cryptista, Hacrobia and Chromista. Mol. Phylogenet. Evol. 93, 331–62 (2015).
Sakaguchi, M., Nakayama, T., Hashimoto, T. & Inouye, I. Phylogeny of the Centrohelida inferred from SSU rRNA, tubulins, and actin genes. J. Mol. Evol. 61, 765–75 (2005).
Burki, F., Okamoto, N., Pombert, J. F. & Keeling, P. J. The evolutionary history of haptophytes and cryptophytes: phylogenomic evidence for separate origins. Proc. Biol. Sci. 279, 2246–54 (2012).
Burki, F. et al. Untangling the early diversification of eukaryotes: a phylogenomic study of the evolutionary origins of Centrohelida, Haptophyta and Cryptista. Proc. Biol. Sci. 283, 20152802, https://doi.org/10.1098/rspb.2015.2802 (2016).
Lang, B. F. et al. An ancestral mitochondrial DNA resembling a eubacterial genome in miniature. Nature 387, 493–7 (1997).
Herman, E. K. et al. The mitochondrial genome and a 60-kb nuclear DNA segment from Naegleria fowleri, the causative agent of primary amoebic meningoencephalitis. J. Eukaryot. Microbiol. 60, 179–91 (2013).
Fučíková, K. & Lahr, D. J. Uncovering cryptic diversity in two Amoebozoan species using complete mitochondrial genome sequences. J. Eukaryot. Microbiol. 63, 112–22 (2016).
Kamikawa, R., Shiratori, T., Ishida, K., Miyashita, H. & Roger, A. J. Group II intron-mediated trans-splicing in gene-rich mitochondrial genome of an enigmatic eukaryote. Diphylleia rotans, Genome Biol. Evol. 3, 458–66 (2016).
Janouškovec, J. et al. A new lineage of eukaryotes illuminates early mitochondrial genome reduction. Curr. Biol. 27, 3717–24 (2017).
Yabuki, A., Gyaltshen, Y., Heiss, A. A., Fujikura, K. & Kim, E. Ophirina amphinema n. gen., n. sp., a New Deeply Branching Discobid with Phylogenetic Affinity to Jakobids. Scientific Reports 8(1) (2018).
Burger, G., Yan, Y., Javadi, P. & Lang, B. F. Group I-intron trans-splicing and mRNA editing in the mitochondria of placozoan animals. Trends Genet. 25, 381–6 (2009).
Pombert, J. F. & Keeling, P. J. The mitochondrial genome of the entomoparasitic green alga Helicosporidium. PloS One 5, e8954, https://doi.org/10.1371/journal.pone.0008954 (2010).
Pombert, J. F., Otis, C., Turmel, M. & Lemieux, C. The mitochondrial genome of the prasinophyte Prasinoderma coloniale reveals two trans-spliced group I introns in the large subunit rRNA gene. PloS One 8, e84325, https://doi.org/10.1371/journal.pone.0084325 (2013).
Nadimi, M., Beaudet, D., Forget, L., Hijri, M. & Lang, B. Group I intron-mediated trans-splicing in mitochondria of Gigaspora rosea and a robust phylogenetic affiliation of arbuscular mycorrhizal fungi with Mortierellales. Mol. Biol. Evol. 29, 2199–210 (2012).
Haugen, P. et al. The recent transfer of a homing endonuclease gene. Nucleic Acids Res. 33, 2734–41 (2005).
Bock, R. The give-and-take of DNA: horizontal gene transfer in plants. Trends Plant Sci. 15, 11–22 (2010).
Iorizzo, M. et al. De novo assembly of the carrot mitochondrial genome using next generation sequencing of whole genomic DNA provides first evidence of DNA transfer into an angiosperm plastid genome. BMC Plant Biol. 12, 61, https://doi.org/10.1186/1471-2229-12-61 (2012).
Turmel, M. et al. Evolutionary transfer of ORF-containing group I introns between different subcellular compartments (chloroplast and mitochondrion). Mol. Biol. Evol. 12, 533–45 (1995).
Alverson, A. J. et al. Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol. Biol. Evol. 27, 1436–48 (2010).
Patterson, D. J. & Dürrschmidt, M. Selective retention of chloroplasts by algivorous heliozoa: Fortuitous chloroplast symbiosis? Eur. J. Protistol. 23, 51–5 (1987).
Speijer, D., Lukeš, J. & Eliáš, M. Sex is a ubiquitous, ancient, and inherent attribute of eukaryotic life. Proc. Natl. Acad. Sci. USA 112, 8827–34 (2015).
Robison, M. M. & Wolyn, D. J. A mitochondrial plasmid and plasmid-like RNA and DNA polymerases encoded within the mitochondrial genome of carrot (Daucus carota L.). Curr Genet. 47, 57–66 (2005).
Court, D. A. & Bertrand, H. Genetic organization and structural features of maranhar, a senescence-inducing linear mitochondrial plasmid of Neurospora crassa. Curr Genet. 22, 385–97 (1992).
Sakurai, R., Sasaki, N., Takano, H., Abe, T. & Kawano, S. In vivo conformation of mitochondrial DNA revealed by pulsed-field gel electrophoresis in the true slime mold, Physarum polycephalum. DNA Res. 7, 83–91 (2000).
Hausner, G. Introns, mobile elements and plasmids. In Organelle Genetics: Evolution of Organelle Genomes and Gene Expression. (Ed. Bullerwell, C. E.) 329–358 (Springer Verlag, 2012).
Nurk, S. et al. Assembling single-cell genomes and mini-metagenomes from chimeric MDA products. J. Comput. Biol. 20, 714–37 (2013).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–10 (1990).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9, 357–9 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–9 (2014).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–35 (2013).
Nawrocki, E. P., Jones, T. A. & Eddy, S. R. Group I introns are widespread in archaea. Nucleic Acids Res. 46, 7970–76 (2018).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–9 (2006).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–2 (2012).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–8 (2005).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–74 (2015).
Hoang, H. C., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: Improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–22 (2018).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods. 14, 587–9 (2017).
Acknowledgements
This work was supported in part by grants from the Japanese Society for Promotion of Science awarded to Y.N. (18K14783), Y.I. (23117006, 16H04826 and 18KK0203) and T.H. (23117005 and 15H05231). This work was also supported, in part by the RIKEN Competitive Program for Creative Science and Technology (to M.O.). Y. N. was supported by a Research Fellowships from the JSPS for Young Scientists (no. 25789).
Author information
Authors and Affiliations
Contributions
Y.N. conceived and designed experiments. Y.N. and T.S. performed experiments. Y.N. and Y.I. analyzed the data. Y.N., T.S., K.I., T.H., M.O., and Y.I. discussed results and implications. Y.N. and Y.I. wrote the manuscript and all authors joined in revision. T.H. and Y.I. supervised Y.N. K.I. supervised T.S.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nishimura, Y., Shiratori, T., Ishida, Ki. et al. Horizontally-acquired genetic elements in the mitochondrial genome of a centrohelid Marophrys sp. SRT127. Sci Rep 9, 4850 (2019). https://doi.org/10.1038/s41598-019-41238-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41238-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.