Abstract
Human ileal bile acid-binding protein (hI-BABP) has a key role in the intracellular transport of bile salts. To explore the role of histidine protonation in the binding process, the pH-dependence of bile salt binding and internal dynamics in hI-BABP was investigated using NMR spectroscopy and biophysical tools. Thermodynamic and kinetic measurements show an increase in the overall binding affinity and the association rate constant of the first binding step below the pKa of the histidines, suggesting that ligand binding is favoured by the protonated state. The overlap between residues exhibiting a high sensitivity to pH in their backbone amide chemical shifts and protein regions undergoing a global ms conformational exchange indicate a connection between the two processes. According to 15N NMR relaxation dispersion analysis, the slow motion is most pronounced at and above the pKa of the histidines. In agreement with the NMR measurements, MD simulations show a stabilization of the protein by histidine protonation. Hydrogen-bonding and van der Waals interactions mediating the flow of information between the C/D- and G/H-turn regions hosting the three histidines, suggest a complex way of pH-governed allosteric regulation of ligand entry involving a transition between a closed and a more open protein state.
Similar content being viewed by others
Introduction
Intracellular lipid binding proteins (iLBPs) are small, 14–15 kDa polypeptide chains that are thought to function in the transcellular trafficking of fatty acids, retinoids, and bile salts1,2. Additionally, there is increasing evidence that they have a role in stimulating the transcriptional activities of nuclear receptors initiated by a ligand-induced nuclear translocation3,4,5. The molecular mechanism by which iLBPs regulate the transport of lipid-like compounds within the cell is not yet fully understood. One of the fundamental questions is the way of ligand entry and exit and how ligand transfer is accomplished between the protein and the cell membrane.
Despite the variability of their lipophilic ligands and the divergence in their amino acid sequence, iLBPs share a common topology comprised of two orthogonal antiparallel beta-sheets and a helix-loop-helix motif covering the beta-barrel (Fig. 1A)1,6. The binding cavity of ~1000Å3 is located inside of the beta-barrel. Internal dynamics appear to have a major role in iLBP-ligand interactions7. Stopped-flow kinetic studies of both fatty acid binding proteins (FABP)8 and bile acid binding proteins (BABP)9 indicate a rate-limiting conformational change preceding the binding step. Moreover, NMR relaxation measurements show evidence of the presence of µs-ms conformational fluctuations with a possible role in mediating ligand entry in FABPs10, cellular retinol binding proteins (CRBPs)11, and BABPs12,13. Despite the common topology, there are substantial differences between the three subfamilies regarding the protein regions involved in the fluctuations. In FABPs, the helical segments together with the proximal C/D-turn are thought to be part of a dynamic portal region mediating access to the enclosed binding cavity10. Similarly, in some of the reported CRBP structures, the helical region exhibiting considerable disorder in the apo state has been found to undergo a millisecond timescale fluctuation in the absence of ligand11,14. More recent MD simulation studies of CRBP 1 and 2 indicate two different entry sites for the ligand, both involving the helical cap and proximal loop regions15. In unligated BABPs, both in the human and the chicken analogues, the E/F and the G/H regions of the C-terminal half exhibit the most intense slow internal motion and are hypothesized to form an alternate portal region13,16. Additionally, in human ileal bile acid-binding protein (hI-BABP), there is a second cluster of residues involving the C/D-turn and the B and D beta-strands undergoing an exchange process with a slightly different exchange rate17, the functional relevance of which is yet unclear. Temperature-dependent relaxation dispersion (Rex) measurements show a strong enthalpy-entropy compensation for both clusters between the ground and a sparsely populated higher energy state characteristic of order-disorder transitions17. More recently, a joint analysis of Rex measurements and NMR thermal melting data has indicated a structural and thermodynamic connection between ms timescale conformational fluctuations at and below room temperature (10–25 °C) and thermal unfolding (60 °C)18. Supporting the NMR relaxation data, MD simulations show evidence of correlated motions involving the helical cap, the E-F strands, and the bottom of the β-barrel. Additionally, the calculations capture a partially unfolded state with an open EF-region and an increased hydrophobic surface, posing an aggregation risk for the protein in the absence of ligands18.
(A) Ribbon diagram of the heterotypic human I-BABP:GCDA:GCA complex determined by solution NMR (PDB entry 2MM36). Bound bile salts are shown in a ball-and-stick representation. (B) Chemical structure of glycocholic acid (GCA) and glycochenodeoxycholic acid (GCDA), the two most abundant bile salts in humans.
Besides mediating ligand entry, conformational flexibility and slow internal fluctuations have also been suggested to have a role in transmitting information between the two binding sites of human I-BABP, a protein displaying a moderate-to-high degree of positive binding cooperativity in its interactions with di- and trihydroxy bile salts (Fig. 1B)19,20,21. Specifically, stopped-flow kinetic studies of human I-BABP indicate that positive cooperativity is related to a slow conformational change of the protein (~30–50 s−1 at 25 °C) occurring after the second binding step9.
Proton transfer mechanisms and tautomer equilibria near physiological pH may have a role in conformational transitions associated with ligand entry and/or a subsequent communication between the binding sites, in particular that hydrogen bond networks have been shown to be key determinants of positive binding cooperativity in human I-BABP21. Due to their multiple tautomeric and charged states, histidines play an important role in binding processes in proteins, in particular that the pKa of their imidazole ring is usually close to physiological pH allowing them to act as either acids or bases22,23. In human I-BABP, there are three histidine residues, two in the C/D-turn (H52, H57) and one in the H beta strand (H98) that is within regions undergoing slow conformational fluctuations in the apo state. Both in the human ileal and the chicken liver analogues, slow motions cease upon bile salt binding suggesting that ligation stabilizes one of the two conformations12,13,17. Moreover, in chicken liver bile acid-binding protein (cL-BABP), a buried histidine in the G/H-region has been indicated in an opening/closing motion12 and a possible role in the cooperativity between the two binding sites as well24. In human I-BABP, with additional histidines in the C/D-turn region, the exchange dynamics appears to be more complex with two subsets of residues exhibiting different exchange kinetics17. To explore the role of histidine protonation in human I-BABP-bile salt recognition and gain more insight into the functional role of conformational fluctuations in the protein, we performed a pH-dependent study of the thermodynamics and kinetics of bile salt binding in conjunction with NMR relaxation measurements and MD simulations. Our results unveil an intimate relation between histidine protonation and an opening/closing motion mediating ligand entry in the apo form of hI-BABP raising the possibility of a pH-dependent mechanism pf bile salt uptake in the enterocytes of the distal small intestine.
Results
Titration calorimetry experiments to characterize the thermodynamics of bile salt binding
Injection profiles and binding isotherms for the interactions of human I-BABP with glycocholic acid (GCA) and glycochenodeoxycholic acid (GCDA), the two most abundant bile salts in humans, at pH = 5.8 and pH = 7.2 are shown in Fig. 2. Similar to previous findings, a biphasic binding profile was obtained for both interactions. To quantitate the thermodynamic parameters of ligand binding, isothermal titration calorimetry (ITC) data sets were fit to the stepwise binding model shown in eq. 1 by Bayesian analysis. Initial values for the parameter search were obtained from nonlinear least-squares fitting. As it was described earlier19, Bayesian analysis overcomes the problem of underestimating the parameter space consistent with the data, a commonly encountered difficulty in nonlinear least-squares analysis of systems with positive cooperativity. The dissociation constants and stepwise binding enthalpies together with the calculated Hill coefficients of positive cooperativity, are listed in Table 1. Bayesian analysis of the probability distribution for the fitted stepwise dissociation constants for the binding of GCA and GCDA at the two different values of pH is shown in Fig. 2E,F. The obtained stepwise binding parameters at pH = 7.2 are in reasonable agreement with previously reported values20,21 taken into consideration the difficulty of the determination of ΔH1 in highly cooperative systems even with Bayesian analysis19.
ITC analysis of the pH-dependence of bile salt binding to human I-BABP. Injection profiles for (A) GCA and (B) GCDA at pH = 7.2. The discontinuity at an x axis value of approximately 17 min represents a change in the injection volume from 0.4 to 0.8 µL. Least-squares fitted binding isotherms for (C) GCA and (D) GCDA at pH = 5.8 (red) and 7.2 (black). The curve through the points of the isotherms corresponds to a least-squares fit of the raw data using the stepwise binding model defined in eq. 1 with parameters listed in Table 1. Scatter plot of Kd2obs versus Kd1obs for the binding of (E) GCA and (F) GCDA to human I-BABP at the two investigated values of pH as obtained from the Bayesian analysis of the probability distribution for the fitted stepwise binding parameters. A Markov chain Monte Carlo method was used to sample 10000 points from the posterior probability distribution of Kd1obs, Kd2obs, ΔH1obs, and ΔH2obs given the calorimetry data. The last 9000 points plotted here for each dataset were used to obtain the average values of the parameters listed in Table 1.
As it is revealed by the products of Kd1 and Kd2, a slight increase in the overall binding affinity is observed for both bile salts by lowering the pH, which arises from both enthalpic (ΔH1 = 0.86 ± 0.03 kcal/mol at pH = 5.8 vs. ΔH1 = 2.0 ± 0.1 kcal/mol at pH = 7.2 for GCDA and ΔH1 = −1.3 ± 0.7 kcal/mol at pH = 5.8 vs. ΔH1 = −0.3 ± 0.9 kcal/mol at pH = 7.2 for GCA) and entropic (TΔS2 = −3.24 ± 0.01 kcal/mol at pH = 5.8 vs. TΔS2 = −5.72 ± 0.03 kcal/mol at pH = 7.2 for GCDA and TΔS2 = −1.6 ± 0.1 kcal/mol at pH = 5.8 vs. TΔS2 = −2.7 ± 0.1 kcal/mol at pH = 7.2 for GCA) effects. The large negative enthalpic contribution of the second binding step reflecting bile salt-amino acid interactions as well as, most likely, interactions of bile salts with bound water molecules within the binding cavity, shows a slightly more pH sensitivity for the dihydroxy bile salt (ΔH2 = −9.9 ± 0.1 kcal/mol at pH = 5.8 vs. ΔH2 = −12.1 ± 0.1 kcal/mol at pH = 7.2) than for GCA (ΔH2 = −9.3 ± 0.7 kcal/mol at pH = 5.8 vs. ΔH2 = −10.0 ± 0.1 kcal/mol at pH = 7.2).
Regarding positive cooperativity, the difference between the Hill coefficients at pH = 5.8 and 7.2 observed for GCA (trihydroxy bile salt with more pronounced positive binding cooperativity) appears to be within experimental error, whereas in the case of GCDA, there is no detectable change at all in binding cooperativity in the investigated pH range. This is likely a manifestation of a strong entropy-enthalpy compensation in the system, which appears to be more pronounced for the binding of the dihydroxy bile salt.
Ligand binding kinetics
Stopped-flow fluorescence experiments were used to reveal the effect of pH on the kinetics of human I-BABP-bile salt interaction. The experiments have been performed under pseudo-first-order conditions by mixing the protein with an excess of ligand. Stopped-flow traces showing fluorescence changes upon GCA and GCDA binding to human I-BABP are shown in Fig. 3. As it has been shown previously9, the kinetic mechanism of bile salt binding can be described in a four-step sequential model according to eq. 3 (Materials and Methods). Accordingly, a rate-limiting conformational transition is followed by two sequential binding steps, and an additional slow conformational change. Stopped-flow traces shown in Fig. 3A,B can most adequately be fitted by a biexponential (eq. 2), with the exponents corresponding to the two observed rates (eigenvalues). Among the two, kapp,1 can be given in a polynomial function of the ligand concentration, which at high values of [L] becomes nearly linear. Under these conditions, the rate constant of the association (k2 in eq. 3) can be estimated from the slope of the curve. The other observed rate, kapp,2 (k−4 in eq. 3) shows no change (within experimental error) with bile salt concentration and as it has been shown earlier corresponds to a slow conformational change following the second binding step associated with positive binding cooperativity.
Stopped-flow kinetic analysis of the pH-dependence of bile salt binding to human I-BABP. (A,B) Representative stopped-flow traces showing fluorescence changes upon mixing human I-BABP with GCA in buffer containing 20 mM potassium phosphate, 135 mM KCl, 10 mM NaCl, and 0.05% NaN3 at (A) pH = 7.2 and (B) pH = 5.8 at 15 °C (λex = 290 nm, λem > 320 nm). Final protein concentration was 1 µM. Final bile salt concentrations were 20, 30, 40, 50, 100, 150, 300 µM. The curve through the data points corresponds to a least-squares fit using a double exponential with observed rates depicted in (C). (C,D) Observed rates for the binding of (C) GCA and (D) GCDA to human I-BABP as a function of ligand concentration as determined from the stopped-flow experiments performed at pH = 5.8 (orange) and 7.2 (green). The solid lines are nonlinear least-squares fits according to the four-step sequential binding mechanism defined in eq. 3 with rate constants listed in Table 2.
The observed rates obtained from the biexponential fits for GCA and GCDA at pH = 5.8 and 7.2 (15 °C) are shown in Fig. 3C,D. Shifting the pH toward more acidic conditions markedly increases the association rate for both bile salts, whereas the rate of the unimolecular step remains unchanged. The solid lines are least-square fits of the observed rates according to the mechanism shown in eq. 3 with parameters listed in Table 2. Observed rates were calculated as eigenvalues of coefficient matrices as a function of ligand concentration using Mathematica. Initial values for the fits were obtained from our previous analysis at 25 °C and pH = 7.29. Similarly to our previous findings at room temperature, the association rate constant of the first binding step (k2) is substantially faster for GCDA than for GCA and the difference between the two bile salts is maintained at both values of pH. For GCA, the increased value of k2 at more acidic conditions parallels with a faster dissociation rate constant (k−2) yielding a similar ratio of k−2/k2 at the two investigated pHs at this temperature. For GCDA, k−2 is significantly smaller at lower pH. Regarding the second binding step, the decrease in pH appears to have a significantly larger effect on the binding kinetics of the trihydroxy bile salt with slower association (k3) and dissociation rates (k−3) under more acidic conditions. Finally, the interconversion between PL2 and PL2* shows nearly the same kinetics at pH = 7.2 and pH = 5.8 for both bile salts. Similar to our previous observation at room temperature, incorporation of an initial unimolecular step (P’ ↔ P) slightly improved the quality of the fits but had no effect on the value of the rate constants for the subsequent steps. We note that the time scale of the P’ ↔ P transition matches well the value of kex inferred from relaxation dispersion NMR measurements in the apo form17 between pH = 6.3–8.0 (cf below).
pH response of backbone amide chemical shifts
To explore the long-range effects of histidine protonation, chemical shifts have been monitored in apo human I-BABP and the doubly-ligated human I-BABP:GCDA:GCA (1.0:1.5:1.5) complex in the range of pH = 4.5–9.0. Superimposed partial 1H−15N-HSQC spectra of apo human I-BABP showing the pH-dependence of the three histidines together with some of the residues in their vicinity exhibiting a similar pH-response are depicted in Fig. 4A. Quantifiable chemical shift changes as a function of pH have been fitted using eq. 4 (Fig. 4B), yielding apparent local pKa values listed in Table 3. In the apo state, H52, H57, and H98 titrate with a pKa = 6.7, 6.4, and 6.6, respectively, which is in reasonable agreement with theoretical pKa values (6.2, 6.3, and 6.3, respectively) calculated using a Gaussian-based dielectric function25. Upon ligand binding, the pKa of H57 increases markedly to 7.9 due the presence of the negatively charged bile salt side chains in the upper segment of the binding cavity and favourable electrostatic interactions with the closing helical cap. A smaller but significant increase in pKa of ~0.4 unit occurs for H98 as well upon bile salt binding, whereas the pKa of H52 remains nearly unaffected. Long range effects of histidine protonation are detected in two main regions of human I-BABP (Fig. 5). pH-induced chemical shift changes in βB (T38-V40) are dominated by the effect of the electrostatic interaction between the carboxylate group of E39 and the imidazole ring of H52, whereas perturbations in βG (G88-N93), the G/H-turn (F94, N96), and part of the I/J-region (T113, Y119) reflect changes in the protonation state of H98. The latter is indicated by highly similar values of pKa in the apo state (pKa = 6.6–6.8) and a similar shift of pKa upon ligation (pKa = 7.0–7.2) for the affected residues (Table 3). Importantly, for most of this protein region, pronounced 15N chemical shift changes (0.2–0.4 ppm) suggest significant structural perturbations upon changes in H98 protonation. An H-bonding network (H98-T113, S112-Y119, Y97-F94, Q99-V92) involving the neighboring GHIJ strands appears to have an important role in the transmission of pH-coupled structural rearrangements. Long-range structural effects seem to be associated with H52 protonation as well manifested in significantly above average pH-induced 15N chemical shift changes nearly the entire length of βB. Additionally, as indicated by their pKa in the holo state, residues at the beginning of βD (T58-T60), exhibit a pH-response similar to H52 rather than the more solvent-exposed H57.
(A) Selected region of the 15N-HSQC spectrum of apo human I-BABP at 10 °C and pH 4.7 (dark green), 5.1 (pink), 5.9 (blue), 6.4 (orange), 7.0 (purple), 7.5 (grey), 8.0 (light green). (B) Titration curves for the backbone 15N resonance of the histidines in the apo (○) and the holo (●) states. Dashed lines show the estimated value of pKa as determined from fitting the observed 15N chemical shifts to eq. 4. (C) 1H-15N long-range HSQC spectra of apo human I-BABP at pH 3.7 (left), 6.4 (middle), and 7.0 (right) (10 °C).
Backbone amide chemical shift differences for apo human I-BABP between pH = 5.9 and 7.0 (top left). Combined (1HN, 15N) chemical shifts were calculated using the equation of Δδ1HN, 15N =√[(ΔδHN)2 + (w1*ΔδN)2], where w1 (=0.154) is a weight factor determined using the BioMagResBank chemical shift database61. The dashed line corresponds to the mean value. Residues with substantially above average (mean + 0.5 SD and mean + SD in light and dark pink, respectively) are mapped on the ribbon diagram of hI-BABP (PDB entry 1O1U56). The position of the histidines is indicated in a ball-and-stick representation. The effect of ligand binding on pH-induced chemical shift changes is depicted at the bottom.
Changes in histidine protonation states as a function of pH have also been monitored by long-range 15N-HSQC experiments that correlate histidine 15Nδ1 and 15Nε2 chemical shifts with nonexchangeable C-H ring protons (Fig. 4C). Based on studies of free histidine and model compounds, the protonated and deprotonated nitrogen of the neutral tautomer in either the δ1 or the ε2 position is expected to resonate at 167.5 and 249.5 ppm, respectively26, whereas in the cationic form the resonance frequency of 15Nδ1 and 15Nε2 are at ~178 and 174 ppm, respectively27,28. As expected, at low pH (3.7), the protonated form dominates for each of the three histidines in apo human I-BABP. At pH = 6.4, H52 and H98 exist in a mixture of different forms, including the neutral Nδ1-H/Nε2 and Nδ1/Nε2-H tautomers with the most abundant Nδ1/Nε2-H form appearing to be more prevalent. Based on the 15N chemical shifts, a substantial population of the cationic form exists as well. The proton of H57 at this pH seems to be in fast exchange between Nδ1 and Nε2. Further increasing the pH to 7.0 results in the typical Γ pattern of cross-peaks29 characteristic of the Nδ1/Nε2-H neutral tautomer for H98 whereas for H52 the proton is exchanging between Nδ1 and Nε2 in a fast tautomerization process. The cationic form of H52 and H98 is still present. The connectivity pattern of H57 is not detectable due to high mobility. Similar connectivity patterns were obtained for the holo state as well with slight perturbations in chemical shifts (data not shown).
pH dependence of slow conformational fluctuations
To relate the macroscopic thermodynamic and kinetic parameters of ligand binding to slow internal dynamics, relaxation dispersion (Rex) NMR measurements30,31 were performed on apo human I-BABP and the doubly-ligated human I-BABP:GCDA:GCA (1:1.5:1.5) complex at different values of pH at a static magnetic field strength of 14.1 T. Specifically, our previously obtained dataset at pH = 6.317 was complemented by three additional measurement series at pH = 5.4, pH = 6.8, and pH = 8.0. As judged by the number of residues exhibiting values of Rex > 2 Hz in the apo form, conformational fluctuations on the Rex-sensitive 0.3–10 ms timescale are most pronounced between pH = 6.3–6.8 (Fig. 6). Under more basic conditions (pH = 8.0), slow motions remain prevalent in the C-terminal half but become diminished in the BCD-region. The most dramatic change occurs under more acidic conditions (pH = 5.4), where the number of residues sensing a contribution to transverse relaxation from ms timescale fluctuations drops to less than 10%. Representative transverse relaxation dispersions as a function of Carr-Purcell-Meiboom-Gill (CPMG) field strength as determined for apo human I-BABP at the four investigated values of pH are plotted in Fig. 7. As we have described it earlier17, at pH = 6.3, dispersion profiles can be grouped into two clusters of residues and fit adequately with the assumption of two separate two-state exchange processes. On the basis of the slightly different exchange rate constants (kA↔B = 836 ± 59 s−1 and kA↔C = 294 ± 40 s−1) for the two subsets of residues, they were termed as fast and slow clusters, respectively. Importantly, a fairly good spatial separation has been observed for the two clusters with residues undergoing the faster A ↔ B process are primarily located in the E-F and G-H regions (cluster I), whereas residues involved in the slower A ↔ C process are located in β-strands B and D, the C/D-turn, and the helical cap (cluster II) (Fig. 8). Upon increasing the pH to 6.8, the exchange kinetics becomes faster and the slower cluster catches up with the faster one merging into a single fluctuating network of residues with values of kex averaging around 1316 ± 174 s−1. Similarly, at pH = 8.0, residues undergoing a ms timescale conformational exchange in the C-terminal half fit well to a two-state exchange model with average values of kex = 1472 ± 233 s−1, justifying a global fit. We note that a similar merging of distinct ms fluctuations have been observed for ribonuclease A at the pH of optimal catalytic activity32. As opposed, under more acidic conditions (pH = 5.4), residues with non-flat Rex profile display a more heterogeneous exchange kinetics in human I-BABP with kex ranging between 12–1850 s−1 accompanied by large differences in the excited state populations (0.3–23%). Kinetic and thermodynamic parameters determined from the global (pH = 6.3–8.0) and individual (pH = 5.4) fit analysis are listed in Tables 4 and S1–S4 of the Supplementary Information. Besides the kinetics becoming faster, the population of the higher energy state (pE) increases slightly upon increasing the pH, suggesting that the conformational exchange is triggered by the formation of the deprotonated state. The majority of Rex-sensing residues can be characterized with a 15N chemical shift difference between the ground and higher energy states of |Δδ| = 0.5–2.2 ppm in the investigated pH range (Fig. 8). The exceptions include V91 exhibiting |Δδ| = 3.5 ppm at pH = 6.3 and some of the residues at pH = 5.4 (Table S1).
Values of 15N Rex (at a static magnetic field strength of 14.1 T) of 2H/15N-labeled apo human I-BABP at 10 °C and pH = 5.4 (green), 6.3 (grey), 6.8 (blue), and 8.0 (magenta) as a function of amino acid sequence. Rex was estimated from the difference in R2eff at the lowest and highest νCPMG values. Secondary structural elements are indicated at the top. For representative residues, 15N Rex at the four investigated values of pH are shown in large at the top.
Transverse relaxation dispersions of the backbone 15N nuclei of selected residues in apo human I-BABP as a function of CPMG B1 field strength at 10 °C and pH = 5.4 (green), 6.3 (grey), 6.8 (blue), and 8.0 (magenta). At pH = 6.3, 6.8, and 8.0, solid black lines correspond to global two-state exchange models with parameters listed in Table 4A. Parameters providing an adequate fit for the relaxation dispersion of T73, V91, and Q99 at pH = 5.4 are listed in Table 4B. Dashed lines indicate the flatness of the dispersion profile of T60 at pH = 5.4 and 8.0.
Backbone 15N chemical shift differences between the ground and higher energy state (|Δδi|) as derived from CPMG relaxation dispersion measurements on apo human I-BABP at different values of pH with parameters listed in Table 4A. Values of |Δδi| are mapped on the ribbon representation of the mean structure of the protein (PDB entry 1O1U56 in pink-to-red (pH = 6.3 ‘fast’ cluster, pH = 6.8, pH = 8.0) and yellow-to-orange (pH = 6.3 ‘slow’ cluster) gradients. Residues with non-flat Rex profile at pH = 5.4 are indicated in green. Residues exhibiting a flat dispersion profile or with no available data are coloured in grey.
Although the changes in |Δδ| vs. pH (Tables S2–S4) are small and the number of data points where comparisons can be made are rather limited, a subtle difference seems to exist between the two protein regions hosting the three histidines. Specifically, a decrease in |Δδ| vs. pH is indicated at T60 of βD (1.5 ± 0.1 ppm at pH = 6.3 vs. 0.8 ± 0.1 ppm at pH = 6.8), T73 of the proximate E/F-turn (2.8 ± 0.1 ppm at pH = 6.3 vs. 1.7 ± 0.2 ppm at pH = 6.8 vs. 1.9 ± 0.2 ppm at pH = 8.0), and to a smaller extent, at E16 of the nearby α-I (0.83 ± 0.05 ppm at pH = 6.8 vs. 0.6 ± 0.1 ppm at pH = 8.0). This trend appears to be different from what we observe in the I/J-region of the C-terminal half, where a slight increase in |Δδ| is found at T113 (0.57 ± 0.03 ppm at pH = 6.3 vs. 0.9 ± 0.1 ppm at pH = 8.0) and Y119 (0.75 ± 0.03 ppm at pH = 6.3 vs. 1.1 ± 0.1 ppm at pH = 8.0). In the G/H-region, the observed differences are either within experimental error or the change of |Δδ| with pH appears to have a maximum (Y97, T100, S101) or a minimum (V91) at pH = 6.8, that is near the pKa of H98. To confirm these trends, additional experiments are required, but the observed differences may indicate that while in the vicinity of the C/D-turn (hosting H52 and H57), the excited state may be reminiscent of a non-protonated form, in the I/J-region (proximate to H98) it may carry features of a protonated state.
In contrast to the unligated state, in holo human I-BABP the vast majority (>95%) of relaxation dispersion profiles remain flat in the investigated 5.4–8.0 pH range. Exceptions occur sporadically throughout the sequence and are limited to F2, Y14, S25, R32, T78, and G115 located in turn or linker regions or at the termini of secondary structural elements. Importantly, ceasing of slow motions upon ligand binding is observed in all regions exhibiting an exchange in the apo state. This is in agreement with our previous findings13,17 and indicates that ms conformational fluctuations giving rise to Rex in the unligated form become abolished upon bile salt binding in a wide pH range.
Molecular dynamics simulations
We tested the effect of protonation levels of the three His residues on the flexibility and stability of human I-BABP by MD simulations. First, simulations with uncharged, Nε2 protonated His residues (designated as 52ε57ε98ε) and with the fully protonated 52p+57p+98p+ models were carried out at 300, 350, 370, and 400 K. At 300 K and 350 K, on the short, μs time scale of the MD simulations there were no significant differences observable between the variants. At 400 K, both variants exhibited a complete unfolding during the simulation, however, the 52ε57ε98ε variant unfolded significantly faster (Fig. 9A). We found that on the time scale of these MD simulations, 370 K is the temperature where the stability of the I-BABP molecule is challenged and differences are well expressed between His protonation variants. We have found that at this temperature protonation decreases the flexibility in most regions and stabilizes the secondary structural elements in general. In the case of 52ε57ε98ε I-BABP, starting with the flexible EF strands, the C-terminal half of the molecule became unfolded after 350 ns followed by a complete unfolding in the second half of the simulation. The fully His-protonated variant 52p+57p+98p+ was proved to be substantially more stable, however, this variant also lost its structure completely at the end of a 2 μs simulation (Fig. 9B).
The effect of His protonation on the stability and flexibility of human I-BABP as revealed by MD simulations. (A,B) I-BABP with uncharged (left) and all cationic (right) H52, H57, and H98 residues simulated at 400 K (A) and 370 K (B). (C) MD at 370 K of single charged variants, with cationic H52 (top), H57 (middle) and H98 (bottom). (D) MD at 370 K of double charged variants, 52ε57p+98p+ (top), 52p+57ε98p+ (middle) and 52p+57p+98ε (bottom). Secondary structure composition is presented along the sequence as a function of time (colour coding: red: α-helix; purple: 310-helix, yellow: β-strand, green: turn, grey: irregular). 3D structures corresponding to every 200 ns are also presented.
In an attempt to distinguish the role of the individual histidines in the stability and dynamics of hI-BABP, we carried out 2 μs simulations with one (Fig. 9C) and two (Fig. 9D) charged histidines in all variations. Uncharged histidines were protonated at Nε2. Comparing the α-helical region (Tyr14-Phe34) in different human I-BABP His protonation variants, it was found that the positive charge at His98 exhibited a stabilization effect increasing the average α-helix content in all but one cases (Table 5). The exception was the variant charged only at His57, which exhibited the highest of all helix content and has proven to be the most stable variant (cf below).
Compared to the molecule with uncharged histidines, all single charged variants proved to be more stable, however, to a different extent. The C terminal β-sheet of 52p+57ε98ε hI-BABP unfolded after 1 μs, while the N-terminal part was basically intact until the 2 μs end of the simulation (Fig. 9C, top). The 52ε57p+98ε variant showed the highest stability not only in single charged but among all variants with low fluctuations in the β-barrel structure (Fig. 9C, middle). The 52ε57ε98p+ variant was stable during the 2 μs simulation, however, its DEF strands unfolded at ~550 ns, and part of the former EF region made a new strand pairing to the C strand forming a misfolded intermediate with an open, exposed substrate binding site which was stable for the rest of the simulation (Fig. 9C, bottom).
Double charged human I-BABP variants were proved to be stable in the first 1 μs of the simulations with showing the previously described flexibility18 in the region of the EF strands (Fig. 9D). However, the 52ε57p+98p+ variant became unfolded around 1200 ns. The 52p+57ε98p+ variant was more stable but its C-terminal unfolded around 1300 ns followed by a complete unfolding around 1800 ns. The most stable double charged variant was 52p+57p+98ε with uncharged His98 showing flexible α-helices (Fig. 9D, bottom) and EF region with a partial C-terminal unfolding at the end of the 2 μs simulation.
In summary, compared to the molecule with all neutral histidines, the cationic His residues increased the stability of human I-BABP, however, in a non-additive manner. The most stable variant was proved to be the one with single cationic His57. Protonation of His98 increased the stability of the N-terminal α-helices. With all three histidines charged, the stability of hI-BABP was in the middle among the variants. The unfolding of hI-BABP usually started with unfolding of the flexible EF strands and the C-terminal β-sheet, indicating the vulnerability of the C-terminal half and higher stability and rigidity of the N-terminal β-sheet (A-D strands) of the molecule.
Discussion
Bile salts are amphipathic molecules synthesized from cholesterol in the liver, which in the small intestine facilitate the absorption of dietary lipids, cholesterol, and fat-soluble vitamins. Besides their role in digestion, they activate signaling pathways33,34, nuclear hormone receptors35,36, and have been shown to interact with the G-protein coupled receptor TGR537,38. Participating in a diverse set of activation processes, bile acid-controlled signaling pathways are important new targets for the treatment of metabolic disorders39. Human I-BABP binds and carries bile salts in the epithelial cells of the distal small intestine and has a key role in the enterohepatic circulation of bile salts40. Additionally, BABPs are thought to have a role in gene regulation by controlling the presentation of bile salts to the farnesoid X receptor (FXR)35.
Positive binding cooperativity between the two binding sites in combination with their relatively low intrinsic affinity enables human I-BABP to act as a buffering agent, which at low bile salt concentration allows a sizeable fraction of bile salts to remain unbound (passing through the enterocytes as monomers) whereas at high bile salt concentrations it protects cells from toxicity20. The extensive hydrogen bond network mediating the communication between the two binding sites21 in conjunction with large conformational rearrangements observed upon ligand binding in the G/H- and C/D-turn regions6 hosting the three histidines of the protein prompted us to investigate the effect of pH on the binding process and internal dynamics of human I-BABP.
One of the most important findings of our work is the overlap between residues exhibiting an above average sensitivity to pH change in their backbone 1H/15N chemical shifts and protein regions undergoing a conformational exchange on the fast end of the millisecond timescale in apo hI-BABP. As indicated by our NMR relaxation measurements, the contribution of the motion to transverse relaxation is most pronounced between pH = 6.3–6.8, that is near the pKa of the histidines of hI-BABP (Table 3). Under more basic conditions (pH = 8.0), while ms timescale fluctuations in the C-terminal half still persist (with exchange rate constants and populations similar to those observed at pH = 6.8 but with somewhat smaller values of Rex), conformational exchange in the vicinity of the C/D-turn becomes abolished. Regarding the opposite, i. e. under conditions corresponding to each of the histidines being protonated (pH = 5.4), fluctuations on the ms timescale become markedly diminished and more heterogeneous by means of both the kinetics and local thermodynamics of the exchange. The cessation of motion below the pKa of histidines is in agreement with MD simulations showing a decrease in flexibility and enhanced stability of the 52p+57p+98p+ state in comparison to 52ε57ε98ε.
When comparing the 15N chemical shift differences between the ground and the excited states inferred from the Rex measurements (|Δδ|) with differences between chemical shifts obtained under more acidic and more basic conditions, most Rex-sensing residues exhibit a value of |Δδ| exceeding the pH-induced shift change. Accordingly, while the observed conformational fluctuation appears to be associated with the protonation/deprotonation equilibrium of histidines, the resulting conformational change is more complex manifesting in larger changes in the chemical environment of the affected residues. As we have shown previously, the slow conformational fluctuations in the apo state near neutral pH cease upon bile salt binding and the Rex-detected higher energy state in the free form shows similarities with the holo state17. Specifically, the correlation between the Rex-derived values of |Δδ| and 15N chemical shift differences between the apo and holo states for a subset of residues in the C-terminal half (e.g. I71, M74, T78, G88 in the EF- and V91, Q99, S101 in the GH-region) suggests the presence of an equilibrium between a closed and a more open holo-like state in the absence of ligands. According to the conformational selection mechanism we proposed earlier6, the closed state is thermodynamically favoured in the absence of ligands, whereas upon bile salt binding the equilibrium is shifted toward the open state possessing an enlarged gap between the E/F- and G/H-turns. More recently, as revealed by temperature dependent NMR measurements and MD simulations18, ms timescale fluctuations in the C-terminal half are related to a partial unfolding of hI-BABP in the DEF protein region. Specifically, a joint analysis of thermal melting and relaxation dispersion data at pH = 6.3 reveals a correlation between Rex-derived Φ values and the squared chemical shift difference between the folded and a partially unfolded hI-BABP state deconvoluted from NMR thermal melting curves indicating a direct connection between the two processes in the apo form18. This raises a possibility that the excited state carries partially unfolded characteristics in the C-terminal half with an opening in the EFGH region, which becomes stabilized upon ligand binding. We note that the opening in the EFGH region is accompanied by a closure of the helical cap on the β-barrel by both NMR structural studies (holo vs. apo form)6 and MD simulations18. A dynamic connection between the G/H-region and the helical cap is further indicated by MD simulations in the current study suggesting that the protonation state of H98 has a role in the stability of the helical region (Table 5). The MD simulations also highlight the role of H57 in the stability of the protein suggesting that favourable interactions of the protonated H57 with negatively charged side chains of the helical cap (i. e. D26) may trigger a chain of stabilizing interactions propagating from the C/D-turn and the helices to the C-terminal half of the protein with the capability of enhancing the stability of even the loose and vulnerable DEF-strands as well (Fig. 9C, middle). Intriguingly, while protonation of H98 in itself has a long-range stabilizing effect on the helical region, protonation of H98 together with either one or both of the histidines in the C/D-turn appears to be less favourable for both the helical content (Table 5) and the overall stability of the protein (Fig. 9D). One possible explanation for this is that positively charged histidines in both regions interfere with some of the hydrophobic interactions stabilizing the protein scaffold.
Cessation of the slow fluctuation upon either bile salt binding or lowering the pH in the apo form indicates similarities in the dynamic behaviour of the complexed and the histidine-protonated apo forms of the protein. One possibility is that similarly to ligand binding, histidine protonation stabilizes a more open arrangement of the EFGH β-strands in the putative portal region. As we have shown previously for the holo protein6, a more open conformation in the EFGH region can be stabilized by the formation of new hydrogen-bonds with the involvement of H98 and its vicinity (e.g. Nδ1 of H98 and Oδ1 of N93, OH of Y97 and Oε1 of Q99, NH of N96 and CO of G115, OH of T113 and OH of T118, NH of T113 and CO of H98) as well as the rearrangement of hydrophobic interactions between the D and E β-strands (e.g. F63-I71, M59-M74) (Fig. 10). It is also apparent that the accompanying closure of the helical cap results in contacts between the I-J region (I114, Y119) and α-I (F17, M18) as well as between the helical segment (M18, D26, V27) and the C/D- (Y53, H57, M59) and E/F-turn regions (M74). Accordingly, an extensive network of interactions can form in the ‘EFGH-open/helix-closed’ state enabling the transmission of information from the segments hosting the histidines to the IJ-, EF-, and the helical regions (Fig. 10). In agreement with this, MD simulations show the involvement of the C/D-turn and D beta-strand in the correlated ‘breathing’ motion of human I-BABP indicating that the motions in the C-terminal half (EFGH-region) and the CD-region with the together moving helical cap are closely related18. Noteworthy, while the protonation equilibrium of H98 appears to have a dominating role in triggering the fluctuation between the closed and more open states throughout the protein, there is a pH (6.3) where the motion of the EFGH and BCD-αIαII protein regions is decoupled from each other giving rise to a faster and slower cluster of residues in the C- and N-terminal half, respectively. This suggests that the protonation of histidine(s) in the C/D-turn may contribute and fine tune the allosteric regulation of ligand entry in hI-BABP.
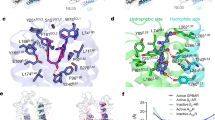
Stabilizing interactions in the doubly-ligated hI-BABP:GCDA:GCA complex (PDB entry 2MM36) corresponding to the ‘EFGH-open/helix-closed’ state of the protein. Hydrogen-bonding and hydrophobic interactions mediate the flow of information between distant sites including βH and the C/D-turn hosting the three histidines and segments in the EF-, IJ-, and helical-regions. Residues with non-flat Rex profile at pH = 6.3 (10–18 °C) are underlined.
We attribute the differences in the dynamic behaviour between the two protein regions (i. e. ceasing of motion in the C-D region at pH = 8.0) to differences between the H98 vs. H52/H57 interaction networks. For instance, an H-bond/salt bridge with the proximate side chain of E39 can have an important role in shifting the protonation equilibrium of H52. Importantly, an additional H-bond with V37 further ties H52 to the neighboring B strand. Furthermore, the vicinity of H52 is tied strongly to βD by multiple H-bonds (Q51-M59, Y53-H57). This may explain the similar dynamic behaviour of βB, βD, and the C/D-turn as revealed by Rex-measurements in a wide temperature range17. Unlike the two histidines in the C/D-turn, H98 in βH forms backbone H-bonds with N93 and T113 contributing to the stabilization of the protein scaffold in the GHI-region by strand-strand interactions.
The shift of the pKa of H98 and residues interacting with it to larger values upon ligand binding is consistent with the notion that bile salt binding is favoured by H98 protonation. This may provide an explanation for the faster association rate constant of bile salt binding below the pKa of H98 as a shift in the conformational equilibrium toward the state with a more open conformation in the EFGH-region is expected to aid the entry of bile salts. We note that as the pKa of bile salts (at least in the unbound state) is ~4, a fraction of them (<10%) may be in the protonated neutral state at the lowest investigated values of pH, which may slightly influence the binding affinity. However, as revealed by structural studies of hI-BABP (PDB: 2MM3), the carboxyl group of GCDA is in favourable electrostatic interaction with the positively charged side chain of K77 and in the vicinity of a number of polar side chains. Therefore, it is unlikely that a protonated GCDA would bind with a higher affinity. In the case of GCA, its carboxyl group is in the vicinity of K30 and H57 (the latter showing a dramatic increase in its pKa upon ligand binding from pKa = 6.4 (apo) to 7.9 (holo), that is favouring a protonated side chain in the presence of bile salts), therefore we find it unlikely that an uncharged GCA with a protonated carboxyl group would bind with higher affinity.
The relation of the protonation equilibrium of the conserved H98 to an open/closed conformational change has been indicated for the chicken liver BABP analogue as well12 suggesting that it may be a common means of an allosterically regulated binding mechanism in the iLBP subfamily. Importantly, in cL-BABP there is no histidine in the C/D-region and the interaction network appears to be more limited. (We note that there is a second more exposed histidine in cL-BABP at the bottom of the beta-barrel, which reportedly is not involved in the conformational fluctuation and forms no contacts with the bound bile salts.) Strikingly, there is a significant difference between the two proteins in terms of both the pKa of H98 and its change upon bile salt binding. Unlike in the human form (Table 3), the conserved H98 in the chicken liver analogue is more buried and exhibits a pKa of 4.7 in the apo state, which upon bile salt binding shifts further down41. Accordingly, while in the chicken liver analogue the contribution of H98 toward substrate binding is favoured by an increase in pH, in the human ileal form it appears to be the opposite. The difference in the values of pKa indicates a large difference in the fraction of protonated H98 at physiological pH between cL-BABP and hI-BABP. Given the hypothesized role of the protonation/deprotonation triggered conformational transition in ligand entry in both analogues, the difference in histidine pKa values should have implications for the regulation of free and bound bile salt pool in (human) enterocytes and (chicken) hepatocytes. Noteworthy, in chicken ileal bile acid-binding protein (cI-BABP), there is an additional histidine in the C/D-turn similar to the human form42. Also, unlike in the liver analogue, in cI-BABP, H98 is solvent exposed similar to hI-BABP.
Importantly, in cL-BABP, H98 has also been suggested to be involved in a network of communication between the two binding sites. Specifically, a hydrophobic bridge between the deprotonated H98 and the nearby I111 has been indicated to provide a connection between the two bound bile salts and contribute to binding cooperativity24. As noted above, in the human ileal form, the side chain of H98 is facing the solvent and does not form a contact with the bound bile salts. In agreement with this, our ITC analysis have shown no significant effect of pH on binding cooperativity in human I-BABP between pH = 5.8 and 7.2. Comparison of the bound state of cL-41 and hI-BABP6 suggests that the concerted fluctuation of the CD-turn with the EFGH-region in the human form may have a role in enabling one of the bile salts to enter more deeply into the binding cavity instead of being trapped near the helical cap. Interestingly, in the chicken ileal form, an additional histidine next to H98 has been suggested to exert too much rigidity on the system and thought to be associated with the lack of positive binding cooperativity in the protein42.
The pH-triggered conformational fluctuation between the closed and more open EFGH protein region in human I-BABP has implications for bile salt uptake and targeted delivery in the epithelial cells. Specifically, enterohepatic circulation facilitated by human I-BABP involves a vectorial transport of bile salts between the apical and basolateral membrane of enterocytes40,43. While the mechanism of bile salt uptake through the membrane is not yet clear, human I-BABP has been shown to be colocalized and functionally associated with the apical sodium dependent bile acid transporter (ASBT) on the apical membrane of enterocytes5. Intriguingly, VPP-c, a subunit of a vacuolar proton pump, has been identified as an interacting partner of ASBT44. If bile salt entry to the enclosed binding cavity of human I-BABP is regulated by a protonation/deprotonation triggered mechanism communicated via the histidines, fluctuations in pH brought about by the vacuolar H+-ATPase at the apical membrane of enterocytes may have a mediatory role.
In conclusion, while histidine protonation appears to be a common means of allosteric regulation in BABP proteins, with histidines in both the C/D and G/H turn regions, human I-BABP exhibits a complex regulation of the opening/closing equilibrium mediating bile salt entry. Considering the differences in the surface accessibility and interaction network of histidines among different BABP analogues suggests that fine tuning of conformational fluctuations by protonation equilibria in different organisms and tissues may have changed in accordance with the evolving pattern of H-bonding and hydrophobic interactions dictated by the local bile salt pool.
Materials and Methods
Sample preparation
The methods used for the expression and purification of unenriched and 15N- as well as 2H/15N-labeled human I-BABP used in the experiments are detailed elsewhere13. In the case of the NMR spectroscopic analysis of the holo sample, protein was complexed with an equimolar mixture of GCA and GCDA at a molar ratio of I-BABP:GCDA:GCA = 1.0:1.5:1.5, ensuring that over 99.9% of the protein was in its bound state20. Protein concentration was determined by absorbance at 280 nm using an extinction coefficient of 12930 M−1 cm−1 obtained by composition analysis45.
Isothermal titration calorimetry
Bile salts were obtained from Sigma and were dissolved in a buffer containing 20 mM potassium phosphate, 135 mM KCl, 10 mM NaCl, 0.05% NaN3 at pH = 5.8 or 7.2. Protein was dialyzed into the same buffer at each pH. The calorimetry experiments were performed using a MicroCal iTC200 isothermal titration calorimeter at 25 °C. Eight injections of 0.4 μL aliquots were followed by 37 injections of 0.8 μL aliquots of 5.1 mM bile salt into the reaction cell containing 0.2019 mL of 0.2 mM human I-BABP, unless noted otherwise. Each titration series was repeated three or four times. The heats of injection were corrected for the heat of dilution of the ligand into buffer and normalized to the amount of bile salt injected. The integrated peak intensities were fit to a stepwise binding model shown schematically as
using Bayesian analysis as described elsewhere46.
Stopped-flow fluorescence spectroscopy
Kinetic measurements of bile salt binding to human I-BABP were performed using a BioLogic SFM 300 stopped-flow spectrophotometer. Ligand binding was monitored by observing the change in fluorescence using an excitation wavelength of 290 nm (4-nm bandwidth) and a 320-nm long-pass filter. Experiments were performed at 15 °C in 20 mM potassium phosphate, 135 mM KCl, 10 mM NaCl, 0.05% NaN3, at pH = 5.8 and 7.2. The protein, at a concentration of 2 μM, was mixed with an equal volume of GCA or GCDA in the same buffer so that the final bile salt concentrations ranged from 20 μM to 500 μM. At and below 500 μM, no self-association of bile salts was observed on the basis of light scattering. A total of 8000 points were collected in each trace. Usually, 12–14 individual traces were averaged at each set of conditions. The kinetic curves were fitted to extract the apparent rates and amplitudes using a nonlinear least-squares algorithm, with a single or a sum of exponential functions defined as
where F(t) is the fluorescence intensity at time t, F(∞) is the fluorescence intensity at t = ∞, ζi is the apparent rate of the ith kinetic process occurring with an amplitude Ai. Kinetic simulations and all further analysis of the data were performed with Dynafit47 and Mathematica (Wolfram, Urbana, IL) assuming a four-step mechanism of binding according to
as described in detail previously9.
NMR data collection and analysis
Multidimensional NMR experiments were carried out at 283 K on 600 MHz Varian NMR SYSTEM™ spectrometer equipped with a 5-mm indirect detection triple 1H13C15N resonance z-axis gradient probe. The backbone resonance assignment of apo human I-BABP and the doubly-ligated hI-BABP:GCDA:GCA complex under our experimental conditions at pH = 6.3 has been published earlier13. To monitor the effect of pH on backbone amide chemical shifts, a series of 1H-15N HSQC spectra48 were acquired on [U-15N]-enriched apo and holo human I-BABP in a buffer containing 20 mM potassium phosphate, 50 mM KCl, and 0.05% NaN3 at pH = 4.7, 5.1, 5.9, 6.4, 7.0, 7.5, 8.0 and pH = 4.6, 5.3, 5.9, 6.5, 7.0, 7.9, 9.1, respectively. Protein concentration was 0.5 mM in all experiments. To evaluate the pKa values of histidines and residues in their vicinity, the observed 15N chemical shifts were fitted to the following equation49
where δp and δd are the chemical shifts of the protonated and deprotonated forms, respectively. The protonation and tautomeric state of the histidines was further monitored by long-range 1H-15N HSQC measurements allowing the correlation of histidine 15Nδ1 and 15Nε2 chemical shifts with nonexchangeable C-H ring protons. The delay to refocus 1H-15N correlations during the formation of 1H-15N antiphase magnetization was set to 23 ms. Experiments were performed at pH = 3.7, 6.4, 7.0 and 4.6, 6.5, 7.9 for apo and holo human I-BABP, respectively. Spectral processing, computer-assisted spin-system analysis, and resonance assignment was carried out using Felix 2004 (Accelrys, Inc.) and CCPNMR.
Relaxation dispersion data were obtained on [80% 2H, 99% 15N]-labeled protein at 283 K and pH = 5.4, 6.3, 6.8, and 8.0, using a relaxation compensated Carr-Purcell-Meiboom-Gill (CPMG) dispersion experiment performed in a constant time manner50,51. The constant time delay (TCP) was set to 40 ms. Spectra were collected as a series of 19 two-dimensional data sets with CPMG field strengths (υCPMG) of 25, 50, 74, 99, 123, 147, 172, 195, 219, 242, 289, 335, 380, 425, 469, 556, 641, 764, and 883 Hz. A reference spectrum was obtained by omitting the CPMG period in the pulse sequence52. Spectra (3 s interscan delay, 24 transients) were acquired in duplicate. Contributions to transverse relaxation rates of conformational exchange were analyzed assuming a two-state exchange process using the all-timescales multiple quantum Carver-Richards-Jones formulation53 implemented in GUARDD54.
MD simulations
Molecular dynamics simulations were carried out as it was described earlier18 using the GROMACS package55 on the solution structure of human I-BABP (PDB: 1O1U56) using the AMBER-ff99SB*-ILDNP force field57 and TIP4P parametrization58. The total charge of the system was neutralized, and the physiological salt concentration was set by placing Na+ and Cl− ions. Energy minimization of starting structures was followed by sequential relaxation of constraints on protein atoms in three steps and an additional NVT step (all of 200 ps) to stabilize pressure. 1 or 2 μs trajectories of NPT simulations at 300, 350, or 370 K at 1 bar were recorded (collecting snapshots at every 20 ps). Secondary structure compositions of the frames of MD trajectories were determined by DSSP algorithm59. As partial protonation of histidines, which is realized in real solution, was not possible in these simulations, we compared neutral (Nε2) and cationic histidines in all combinations. Specifically, a series of simulations were carried out by varying the protonation levels of the three histidine residues, H52, H57, and H98, setting them individually as protonated at Nε2, or Nδ1, or at both nitrogens (positively charged form). Molecular graphics was performed with the UCSF CHIMERA package (University of California, San Francisco)60.
Data Availability
All data generated or analysed during this study are included in the published article and its Supplementary Information File.
References
Banaszak, L. et al. Lipid-binding proteins: A family of fatty acid and retinoid transport proteins. Adv Protein Chem 45, 89–151 (1994).
Veerkamp, J. H. & Maatman, R. G. Cytoplasmic fatty acid-binding proteins: Their structure and genes. Prog Lipid Res 34, 17–52 (1995).
Sessler, R. J. & Noy, N. A ligand-activated nuclear localization signal in cellular retinoic acid binding protein-II. Mol Cell 18, 343–353 (2005).
Ayers, S. D., Nedrow, K. L., Gillilan, R. E. & Noy, N. Continuous nucleocytoplasmic shuttling underlies transcriptional activation of PPARγ by FABP4. Biochemistry 46, 6744–6752 (2007).
Nakahara, M. et al. Ileal bile acid-binding protein, functionally associated with the farnesoid X receptor or the ileal bile acid transporter, regulates bile acid activity in the small intestine. J Biol Chem 280, 42283–42289 (2005).
Horváth, G. et al. Structural determinants of ligand binding in the ternary complex of human ileal bile acid binding protein with glycocholate and glycochenodeoxycholate obtained from solution NMR. FEBS J 283, 541–555 (2016).
Ragona, L. et al. The role of dynamics in modulating ligand exchange in intracellular lipid binding proteins. Biochim Biophys Acta Proteins Proteomics 1844, 1268–1278 (2014).
Kim, K., Cistola, D. P. & Frieden, C. F. Intestinal fatty acid-binding protein: The structure and stability of a helix-less variant. Biochemistry 35, 7553–7558 (1996).
Toke, O., Monsey, J. D. & Cistola, D. P. Kinetic mechanism of ligand binding in human ileal bile acid binding protein as determined by stopped-flow fluorescence analysis. Biochemistry 46, 5427–5436 (2007).
Hodsdon, M. E. & Cistola, D. P. Discrete backbone disorder in the nuclear magnetic resonance structure of apo intestinal fatty acid-binding protein: Implications for the mechanism of ligand entry. Biochemistry 36, 1450–1460 (1997).
Lu, J., Cistola, D. P. & Li, E. Two homologous rat cellular retinol-binding proteins differ in local conformational flexibility. J Mol Biol 330, 799–812 (2003).
Ragona, L. et al. NMR dynamic studies suggest that allosteric activation regulates ligand binding in chicken liver bile acid-binding protein. J Biol Chem 281, 9697–9709 (2006).
Horváth, G., Király, P., Tárkányi, G. & Toke, O. Internal motions and exchange processes in human ileal bile acid binding protein as studied by backbone 15N nuclear magnetic resonance spectroscopy. Biochemistry 51, 1848–1861 (erratum appears in Biochemistry 51, 10119), (2012).
Lu, J. et al. The structure and dynamics of rat apo-cellular retinol-binding protein II in solution: comparison with the X-ray structure. J Mol Biol 286, 1179–1195 (1999).
Menozzi, I., Polverini, E. & Berni, R. Deciphering protein dynamics changes along the pathway of retinol by cellular retinol-binding proteins 1 and 2. Archives Biochem Biphys 645, 107–116 (2018).
Cogliati, C. et al. Site-specific investigation of the steady-state kinetics and dynamics of the multistep binding of bile acid molecules to a lipid carrier protein. Chem Eur J 16, 11300–11310 (2010).
Horváth, G., Egyed, O. & Toke, O. Temperature dependence of backbone dynamics in human ileal bile acid-binding protein: Implications for the mechanism of ligand binding. Biochemistry 53, 5186–5198 (2014).
Horváth, G. et al. Structural insight into a partially unfolded state preceding aggregation in an intracellular lipid-binding protein. FEBS Journal 284, 3637–3661 (2017).
Tochtrop, G. P. et al. Energetics by NMR: Site-specific binding in a positively cooperative system. Proc Natl Acad Sci USA 99, 1847–1852 (2002).
Tochtrop, G. P., Bruns, J. M., Tang, C., Covey, D. F. & Cistola, D. P. Steroid ring hydroxylation patterns govern cooperativity in human bile acid binding protein. Biochemistry 42, 11561–11567 (2003).
Toke, O. et al. Determinants of cooperativity and site-selectivity in human ileal bile acid-binding protein. Biochemistry 45, 727–737 (2006).
Shimba, N. et al. Quantitative identification of the protonation state of histidines in vitro and in vivo. Biochemistry 42, 9227–9234 (2003).
Hass, M. A. S., Hansen, D. F., Christensen, H. E. M., Led, J. J. & Kay, L. E. Characterization of conformational exchange of a histidine side chain: Protonation, rotamerization, and tautomerization of His61 in plastocyanin from Anabaena variabilis. J Am Chem Soc 130, 8460–8470 (2008).
Pedo, M., D’Onofrio, M., Ferranti, P., Molinari, H. & Assfalg, M. Towards the elucidation of molecular determinants of cooperativity in the liver bile acid binding protein. Proteins 77, 718–731 (2009).
Wang, L., Li, L. & Alexov, E. pKa predictions for proteins, RNAs and DNAs with the Gaussian dielectric function using DelPhiPKa. Proteins. 83, 2186–2197 (2015).
Pelton, J. G., Torchia, D. A., Meadow, N. D. & Roseman, S. Tautomeric states of the activesite histidines of phosphorylated and unphosphorylated IIIGlc, a signal-transducing protein from Escherichia coli, using two-dimensional heteronuclear NMR techniques. Protein Sci 2, 543–558 (1993).
Singer, A. U. & Forman-Kay, J. D. pH titration studies of an SH2 domain-phosphopeptide complex: Unusual histidine and phosphate pKa values. Protein Sci 6, 1910–1919 (1997).
Shimahara, H. et al. Tautomerism of histidine 64 associated with proton transfer in catalysis of carbonic anhydrase. J Biol Chem 282, 9646–9656 (2007).
Van Dijk, A. A., Scheek, R. M., Dijkstra, K., Wolters, G. K. & Robillard, G. T. Characterization of the protonation and hydrogen bonding state of the histidine residues in IIAmtl, a domain of the phosphoenolpyruvate-dependent mannitol-specific transport protein. Biochemistry 31, 9063–9072 (1992).
Palmer, A. G. III, Kroenke, C. D. & Loria, J. P. Nuclear magnetic resonane methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339, 204–238 (2001).
Vallurupalli, P., Hansen, D. F. & Kay, L. E. Structures of invisible, excited states by relaxation dispersion NMR spectroscopy. Proc Natl Acad Sci USA 105, 11766–11771 (2008).
Khirich, G. & Loria, J. P. The complexity of protein energy landscapes studied by solution NMR relaxation dispersion experiments. J Phys Chem B 119, 3743–3754 (2015).
Gupta, S., Stravitz, R. T., Dent, P. & Hylemon, P. B. Down-regulation of cholesterol 7alpha-hydroxylase (CYP7A1) gene expression by bile acids in primary rat hepatocytes is mediated by the c-Jun N-terminal kinase pathway. J Biol Chem 276, 15816–15822 (2001).
Qiao, L. et al. Bile acid regulation of C/EBPbeta, CREB, and c-Jun function, via the extracellular signal-regulated kinase and c-Jun NH2-terminal kinase pathways, modulates the apoptotic response of hepatocytes. Mol Cell Biol 23, 3052–3066 (2003).
Makishima, M. et al. Identification of a nuclear receptor for bile acids. Science 284, 1362–1365 (1999).
Parks, D. J. et al. Bile acids: natural ligands for an orphan nuclear receptor. Science 284, 1365–1368 (1999).
Maruyama, T. et al. Identification of membrane-type receptor for bile acids (M-BAR). Biochem Biophys Res Comm 298, 714–719 (2002).
Kawamata, Y. et al. A G protein-coupled receptor responsive to bile acids. J Biol Chem 278, 9435–9440 (2003).
Houten, S. M., Watanabe, M. & Auwerx, J. Endocrine functions of bile acids. The EMBO Journal 25, 1419–1425 (2006).
Small, D. M., Dowling, R. H. & Redinger, R. N. The enterohepatic circulation of bile salts. Arch Intern Med 130, 552–573 (1972).
Eliseo, T. et al. Structural and dynamic determinants of ligand binding in the ternary complex of chicken liver bile acid binding protein with two bile salts revealed by NMR. Biochemistry 46, 12557–12567 (2007).
Zanzoni, S., Assfalg, M., Giorgetti, A., D’Onofrio, M. & Molinari, H. Structural requirements for cooperativity in ileal bile acid-binding proteins. J Biol Chem 286, 39307–39317 (2011).
Alrefai, W. A. & Gill, R. K. Bile acid transporters: Structure, Function. Regulation, and Patophysiological implications. Pharmac Res 24, 1803–1823 (2007).
Sun, A. Q., Balasubramaniyan, N., Liu, C. J., Shahid, M. & Suchy, F. J. Association of the 16-kDa subunit c of vacuolar proton pump with the ileal Na+-dependent bile acid transporter. J Biol Chem 279, 16295–16300 (2004).
Pace, C. N., Vajdos, F., Fee, L., Grimsley, G. & Gray, T. How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4, 2411–2423 (1995).
Berry, D. A. Statistics: A Bayesian Perspective (Duxbury Press, Belmont, CA), (1996).
Kuzmic, P. Program DYNAFIT for the analysis of enzyme kinetic data: application to HIV proteinase. Anal Biochem 237, 260–273 (1996).
Kay, L. E., Keifer, P. & Saarinen, T. Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114, 10663–10665 (1992).
Hass, M. A. S., Thuesen, M. H., Christensen, H. E. M. & Led, J. J. Characterization of μs-ms dynamics of proteins using a combined analysis of 15N NMR relaxation and chemical shift: Conformational exchange in plastocyanin induced by histidine protonations. J Am Chem Soc 126, 753–765 (2004).
Loria, J. P., Rance, M. & Palmer, A. G. III A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121, 2331–2332 (1999).
Tollinger, M., Skrynnikov, N. R., Mulder, F. A. A., Forman-Kay, J. D. & Kay, L. E. Slow dynamics of folded and unfolded states of an SH3 domain. J Am Chem Soc 123, 11341–11352 (2001).
Mulder, F. A., Skrynnikov, N. R., Hon, B., Dahlquist, F. W. & Kay, L. E. Measurement of slow (micros-ms) time scale dynamics in protein side chains by (15)N relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123, 967–975 (2001).
Korzhnev, D. M., Kloiber, K. & Kay, L. E. Multiple-quantum relaxation dispersion NMR spectroscopy probing millisecond time-scale dynamics in proteins: theory and application. J Am Chem Soc 126, 7320–7329 (2004).
Kleckner, I. R. & Foster, M. P. GUARDD:user-friendly MATLAB software for rigorous analysis of CPMG RD NMR data. J Biomol NMR 52, 11–22 (2012).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Kurz, M. et al. Insights into the bile acid transportation system: The human ileal lipid-binding protein-cholyltaurine complex and its comparison with homologous structures. Proteins 50, 312–328 (2003).
Aliev, A. E. et al. Motional timescale predictions by molecular dynamics simulations: case study using proline and hydroxyproline sidechain dynamics. Proteins 82, 195–215 (2014).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J Chem Phys 79, 926–935 (1983).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogenbonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Pettersen, E. F. et al. UCSF Chimera – a visualization system for exploratory research and analysis. J Comput Chem 25, 1605–1612 (2004).
Evenäs, J. et al. Ligand-induced structural changes to maltodextrin-binding protein as studied by solution NMR spectroscopy. J Mol Biol 309, 961–74 (2001).
Acknowledgements
This work was supported by National Research, Development and Innovation Office, Hungary (NKFIH) Grants K-109035 (to O.T.), K-117062 and K-123989 (to M.K.), K-120391 (to J.K. and A.M.), KTIA_NAP_13-2-2014-0017 (to J.K. and A.M.), ERC_HU 117680 (to M.K.), the “Momentum” Program of the Hungarian Academy of Sciences (LP2011-006/2011 to M.K.), the ‘János Bolyai Research Scholarship’ program of the Hungarian Academy of Sciences (to O.T. and A.M.), the New National Excellence Program ÚNKP-18-4-ELTE-108 (to A.M.), and a MedinProt grant (to M.K. and O.T.). The authors thank Prof. David Cistola for the pMON5840 plasmid, Ádám Fizil for the ITC measurements, and Dr. Nikolett T. Nagy for assistance with the stopped-flow experiments.
Author information
Authors and Affiliations
Contributions
G.H. produced the isotopically labelled proteins and analysed some of the relaxation NMR experiments. E.O. carried out the pH titration NMR measurements. C.T. carried out the Bayesian analysis of the ITC data. M.K. performed the fluorescence experiments. A.M. and J.K. performed and analysed the MD simulations and wrote part of the manuscript. O.T. carried out and analysed the vast majority of the NMR experiments, designed the research, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Horváth, G., Egyed, O., Tang, C. et al. Ligand entry in human ileal bile acid-binding protein is mediated by histidine protonation. Sci Rep 9, 4825 (2019). https://doi.org/10.1038/s41598-019-41180-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-41180-7
This article is cited by
-
Mapping the electrostatic potential of the nucleosome acidic patch
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.