Abstract
Circulating cell-free microRNAs are promising candidates for minimally invasive clinical biomarkers for the diagnosis, prognosis and monitoring of many human diseases. Despite substantial efforts invested in the field, the research so far has failed to deliver expected results. One of the contributing factors is general lack of agreement between various studies, partly due to the considerable technical challenges accompanying the workflow. Pre-analytical variables including sample collection, RNA isolation, and quantification are sources of bias that may hamper biological interpretation of the results. Here, we present a Two-tailed RT-qPCR panel for quality control, monitoring of technical performance, and optimization of microRNA profiling experiments from biofluid samples. The Two-tailed QC (quality control) panel is based on two sets of synthetic spike-in molecules and three endogenous microRNAs that are quantified with the highly specific Two-tailed RT-qPCR technology. The QC panel is a cost-effective way to assess quality of isolated microRNA, degree of inhibition, and erythrocyte contamination to ensure technical soundness of the obtained results. We provide assay sequences, detailed experimental protocol and guide to data interpretation. The application of the QC panel is demonstrated on the optimization of RNA isolation from biofluids with the miRNeasy Serum/Plasma Advanced Kit (Qiagen).
Similar content being viewed by others
Introduction
Circulating cell-free microRNAs have emerged in recent years as promising candidates for minimally invasive clinical biomarkers for diagnosis, prognosis, and monitoring of a multitude of human pathologies1,2,3,4,5,6. After this recognition, a massive wave of research aiming at identifying disease-associated microRNAs followed. A search for the keywords “microRNA”, “biomarker” and “blood” returns over 5000 hits in the PubMed database (September 2018) with the number of studies increasing every year. Despite promising advances in the field (www.clinicaltrials.gov), there is still no microRNA test in clinical practice7,8. There are many reasons behind the current unsatisfactory state and their comprehensive discussion is beyond the scope of this article (for reviews see8,9). One issue is the poor agreement between studies, which may in part be attributed to the lack of standardization10 and technical difficulties associated with the workflow11,12. Protocols for blood collection, sample processing, storage, RNA isolation, and microRNA quantification often vary across laboratories leading to discordant results11,13,14. Efforts are ongoing to standardize the blood sampling and processing steps to improve the reproducibility of microRNA analyses15,16,17,18 (www.cancer-id.eu; www.spidia.eu). However, notable sources of variation remain. These include the RNA isolation, co-purification of inhibitors of enzymatic reactions, and cellular contamination of the biofluid samples19,20,21. These factors may bias the measured microRNA profiles leading to false-positive discoveries of disease-associated biomarkers. Rigorous control of sample quality and technical workflow is therefore of highest importance.
An efficient way to monitor technical variation is the addition of exogenous spike-in molecules prior to RNA isolation22,23,24. The signal from the spike-ins reflect yields and extraction efficiency, which identifies abnormal samples that should be reanalysed or disqualified. A second set of exogenous spike-ins can be added before the microRNA quantification to control for bias introduced downstream of this step, such as the inhibition of enzymatic reactions. This concept has been described previously20,22,23 and is also available as a commercial product (e.g. RNA Spike-In Kit, for RT marketed by Qiagen). Yet, to our knowledge, there is no tool to perform such extended quality control on challenging experimental samples described in detail in literature.
Here, we present the Two-tailed quality control (QC) panel; a tool to assess the technical quality of RNA isolation, degree of inhibition and erythrocyte contamination, primarily in liquid biopsy samples, such as serum and plasma. The panel is based on Two-tailed RT-qPCR; a highly specific method for microRNA quantification25. We provide detailed experimental protocol, guide to data interpretation, and sequences of the RNA oligonucleotides and RT-qPCR assays used (Supplementary file) that can be ordered from any licensed oligo manufacturer. The QC panel is based on standard reagents and is intended to provide researchers a convenient tool to assess technical performance and quality of the samples before investing resources into extensive quantification experiment, such as small-RNA sequencing or high-throughput RT-qPCR. We demonstrate its utility by optimizing an RNA isolation protocol, screening for haemolysis, and testing for outliers with compromised quality. We also report data obtained with the recently launched miRNeasy Serum/Plasma Advanced Kit (Qiagen) for two biofluids collected from human and rat.
Results
Design of the QC panel
The Two-tailed QC panel is composed of five synthetic spike-in microRNAs and eight Two-tailed assays targeting these synthetic spike-ins, and three endogenous microRNAs (Fig. 1). The spike-ins are based on C. elegans microRNAs and artificial sequences and have no significant homology to any known human, mouse or rat microRNA (Table 1). All spike-ins have 5′ terminal phosphate to mimic endogenous microRNAs, and to allow incorporation into microRNA libraries for Next Generation Sequencing (NGS).
QC workflow with the Two-tailed QC panel. (A) A mix of three synthetic RNA spike-ins (cel-miR-54, spike-A, spike-B) is added prior to RNA isolation from the biofluid sample. A second mix of two spike-ins (cel-miR-76, cel-miR-2) is added before cDNA synthesis step. Optionally, a diluted isolation spike-in mix is used as a template in a “spike-only” control reaction to determine spike-in baseline signal (for details see Supplementary file section 3.2.2). Two-tailed RT-qPCR is used to quantify the spike-ins along with three endogenous microRNAs (let-7a, miR-23a and miR-451a) to evaluate the technical quality of RNA isolation, effect of inhibition and the level of haemolysis. (B) Decision chart for data interpretation and troubleshooting (see also Supplementary file section 4).
Three spike-in RNAs (cel-miR-54, spike-A and spike-B) comprise the isolation spike-in mix and are added to the samples at a known constant amount prior to RNA isolation, serving as control for the technical performance of the RNA isolation protocol (Fig. 1A). The three spike-ins have varying GC content (41.7–63.6%) and are present at concentrations reflecting high (cel-miR-54, 1e + 7 copies/μl), moderate (spike-A, 2e + 5 copies/μl), and low (spike-B, 4e + 3 copies/μl) abundant microRNAs (Supplementary file). The ΔCq’s between the isolation spike-ins should, in absence of inhibition, be in the range 3.5–5.5 cycles (accounting for differences in RT-PCR efficiencies of the Two-tailed assays), however, these values may be influenced differently by individual isolation protocols due to various biases26,27.
Two RNA spike-ins (cel-miR-76 and cel-miR-2) comprise the reverse transcription (RT) spike-in mix and are added to the RT reaction serving as controls for cDNA synthesis, PCR amplification and as general controls for the presence of inhibitors in RNA eluates (Fig. 1A). Cel-miR-76 (1e + 7 copies/μl) is added at 100x higher concentration than cel-miR-2 (1e + 5 copies/μl) and their ΔCq should be 5.5–6.5 cycles (accounting for differences in PCR efficiency of the Two-tailed assays).
The QC panel also contains assays for the three endogenous microRNAs: let-7a, miR-23a and miR-451a. Let-7a is abundant in plasma and serum20,28,29 and serves as positive control. Mir-23a is also abundant in plasma/serum and its level is independent of haemolysis, while miR-451a is highly abundant in erythrocytes and its level increases dramatically upon haemolysis20,30. The ΔCq (mir-23a – mir-451a) indicates degree of haemolysis in the samples20.
Optimization of sample input volume
A factor that is often neglected, but can have major impact on the quality of microRNA quantification data, is the initial input volume used for the RNA isolation13,26. Liquid biopsy samples contain very low amounts of microRNAs and researchers may be tempted to use as much sample material as possible for RNA isolation. However, with increasing amount of starting material risk of carryover of contaminating substances and saturation of the purification column increases31,32. Most commercial RNA isolation kit manufacturers recommend 200 μl starting serum/plasma volume, however, optimum volume depends on the isolation protocol, sample type and also organism26. Optimizing the sample volume is therefore recommended when setting up a new isolation protocol or extracting a new type of sample. For such optimization the Two-tailed QC panel is a tool to assess relative isolation efficiency, absolute yield, and test for the presence of inhibitors to decide the optimal input volume. With this strategy we optimized protocol based on the miRNeasy Serum/Plasma Advanced Kit (Qiagen) for RT-qPCR analysis of human plasma, human serum, and rat serum (Fig. 2).
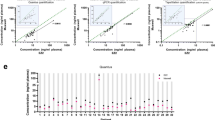
Optimizing input volumes of (A) human plasma, (B) human serum, and (C) rat serum for RNA isolation. Data are presented as ΔCq between Cqs obtained with the tested volume and an input volume of 200 μl (human) or 100 μl (rat). Each dot is one isolation replicate. Optimum starting serum/plasma volumes based on absolute endogenous microRNA yields are 250 μl for human plasma, 300–500 μl for human serum, and 150 μl for rat serum (blue mean profiles). Error bars on mean profiles panels indicate standard deviation (SD).
We found a non-linear relation between the input sample volume and cDNA yield as reflected by RT-qPCR signal of endogenous microRNAs (Fig. 2). The non-linearity is caused neither by RT nor PCR inhibition, as the signals from the RT spike-ins were independent of volume. Rather the non-linear response is due to variations in RNA isolation efficiency, as reflected by the RT-qPCR response of the isolation spike-ins (Fig. 2). We observed poor isolation efficiency with low input volumes (<200 μl for human, <100 μl for rat), but also with higher input volumes (≥300 μl for human, ≥200 μl for rat), where the response was also more variable (Fig. 2). Based on our results, optimum starting sample volumes with our workflow are: 250 μl for human plasma, 300–500 μl for human serum, and 150 μl for rat serum.
Assessing the effect of co-precipitants in the isolation procedure
Since biofluids like serum and plasma contain very low amounts of RNA, significant portion may be lost during the isolation procedure due to adsorption to the pipette tips, tube walls etc. Losses can be reduced by adding carriers such as MS2 phage RNA or yeast tRNA to the samples before RNA isolation33,34. However, RNA-based carriers are less suited when NGS is used for downstream analysis as the exogenous RNAs may consume sequencing reads. Other carriers, such as linear acrylamide, BSA or glycogen may then be used instead24. Using the Two-tailed QC panel we tested the impact of using glycogen as carrier in our isolation procedure (Fig. 3). In accordance with previous observations33,34, we found that addition of glycogen significantly improved the reproducibility of isolation (F-test, p < 0.001) and significantly increased the yield (average Cq difference 1.25; paired T-test p = 0.011) with no negative effects on the downstream RT-qPCR analysis (Fig. 3). Based on these findings, we recommend addition of glycogen to increase the robustness and efficiency of microRNA isolation with the miRNeasy Serum/Plasma Advanced Kit (Qiagen).
Effect of glycogen carrier on microRNA quantification in human plasma. Identical sample aliquots were isolated with (n = 5) or without (n = 6) addition of glycogen carrier starting from 200 μl, and quantified with the Two-tailed QC panel. Extractions with glycogen had significantly higher yields (average difference between Cq means: 1.25 cycles; paired T-test p = 0.011) and higher reproducibility (F-test, p < 0.001).
Assessing the level of haemolysis in serum/plasma samples
A major complication in microRNA analysis of serum/plasma samples is contamination with microRNAs derived from lysed blood cells20,30,35 and in particular haemolysed erythrocytes. Plasma and serum samples should therefore be assessed for haemolysis. Standard method is to measure absorption at 414 nm, 540 nm and 578 nm, which are the absorption peaks of free oxyhemoglobin36. An alternative approach, which is applicable also when the original sample is no longer available, is to measure the ratio of miR-23a, which is insensitive to haemolysis, and miR-451a, which is highly enriched in erythrocytes20. Blondal et al.20 established threshold ΔCq (miR-23a–miR-451a) values as quality indicators for human samples: ΔCq > 5 indicates there may be erythrocyte contamination, and ΔCq > 7 indicates high risk of haemolysis. A complication, however, is that the indicator is sensitive to the relative isolation yields of miR-23a and miR-451, as well as their relative RT yields and PCR efficiencies of the assays used to quantify them. Hence, the threshold ΔCq’s reported by Blondal et al.20 are valid only for their particular workflow and protocol, and should not be used as general indicators. Here, we establish threshold ΔCq values for the Two-tailed QC panel and our recommended workflows (Supplementary file).
We prepared duplicate haemolysis dilution series for each sample type and constructed standard curves to correlate ΔCq (miR-23a – miR-451a) values to absorbance at 414 nm (Fig. 4B). To increase the number of data points, samples screened in other experiments with the same workflow were also included. Correlation between A414 and linear transformation of ΔCq (2ΔCq) is significant for all three biofluids (Pearson r ≥ 0.80, p < 0.0001). A414 for plasma sample 1 was outside the linear range of the absorption spectrophotometer and was estimated by interpolation. A540 and A578 nm dependences show the same trend, although those peaks are considerably less significant in the absorbance spectrum (Fig. 4A). A414 ≤ 0.2–0.25 has previously been recommended as threshold for non-haemolysed samples13,20,35. Based on our calibration this corresponds to a ΔCq of 15 cycles for human plasma, 11 cycles for human serum, and 6 cycles for rat serum for our workflow (Fig. 4B; Supplementary file).
Assessing haemolysis in serum/plasma samples. (A) Human plasma samples with varying degree of haemolysis, corresponding A414, ΔCq (miR-23a–miR-451a), and selected UV-Vis spectra. (B) Correlation between ΔCq (miR-23a–miR-451a) and A414, A540 and A578, respectively. Exponential regression line with 95% confidence interval is shown. Dashed red line indicates A414 = 0.25 as threshold for non-haemolysed samples. Corresponding ΔCq thresholds are ~15 (human plasma), ~11 (human serum), and ~6 (rat serum).
Discussion
We present an RT-qPCR based protocol to assess the technical performance of workflows for analysis of microRNAs in body fluids such as serum and plasma. The QC panel developed is based on two sets of synthetic spike-in molecules and three endogenous microRNAs to assess RNA isolation yield, RT yield, PCR efficiency, and haemolysis (Fig. 1).
A highly error-prone step in microRNA analysis workflow is RNA isolation. Several studies have studied the effect of input volume on microRNA recovery, reporting varying results26,37,38. However, consistent observations are substantial variability between replicate isolations and non-linear dependence of the input volume on the amount of microRNAs detected26,31. Here, we studied the effect of input sample volume when extracting with the recently launched miRNeasy Serum/Plasma Advanced Kit from Qiagen and found that optimal input volume is different for the three sample types: human plasma, human serum, and rat serum (Fig. 2). We also found that higher input volumes (>300 μl for human, >150 μl for rat), although still in the range recommended by the manufacturer, lead to less reproducible Cq values compared to moderate input volumes (200–300 μl for human, 100–150 μl for rat). Using spike-in controls we showed this is due to inhibition of neither cDNA synthesis nor PCR, as suggested previously20, but rather to impaired isolation efficiency, possibly because of saturation of the purification column. We confirm previous observations that adding a carrier improves extraction yield and reproducibility32,33,34,39. We also show glycogen is a suitable alternative to RNA-based carriers when using the miRNeasy Serum/Plasma Advanced Kit (Qiagen) conferring advantage when samples shall be analysed with NGS (Fig. 3).
Another contribution to bias is microRNAs from leaking blood cells30,35. While cellular contamination can be minimized by careful removal of the plasma fraction and dual centrifugation to efficiently remove platelets, haemolysis remains a problem. Haemolysis can occur during sampling and handling procedures and the released cellular microRNAs distort the measured microRNA profiles, which no longer reflect exclusively cell free microRNA30,35,36. This not only hampers biological interpretation of the results, but can distort normalization or RT-qPCR data. For example, miR-16-5p is widely used as reference microRNA9, but it is also one of the most abundant microRNAs in erythrocytes30 and its level is therefore perturbed even at low level of haemolysis35,40.
Haemolysis can be assessed by visual inspection of the samples or, more precisely, spectroscopically. An alternative approach is to compare the levels of the erythrocyte-enriched miR-451a and the haemolysis-insensitive miR-23a20. While visual inspection is rather subjective and not particularly sensitive, spectroscopic assessment and RT-qPCR quantification of miR-451a and miR-23a levels reveal even low degree of haemolysis35. Shah et al.41, compared several methods to assess the level of haemolysis in human serum samples and found ΔCq (miR-23a – miR-451a) to be the most sensitive indicator41. In contrast, Vliet et al.13 reported that absorption measurement is more sensitive for rat plasma samples. Our results show that the approaches are comparable and correlate well even at very low levels of haemolysis for all three sample types we tested (Fig. 4). An advantage of the qPCR approach is that haemolysis can be assessed even when the original sample is no longer available. The same strategy can be used to assess contamination with other cell-types when needed. For example, miR-425 level may reflect contamination of platelets13. It is important to be aware that the ΔCq (miR-23a–miR-451) indicator must be calibrated for every new biofluid, isolation procedure and RT-qPCR method, as the ratio of the measured levels of miR-23a and miR-451 depend on the relative bias introduced by the methods used26,27,42, but also the particular species and biofluids analysed. Indeed, in our study we concluded different threshold ΔCq values for the three sample types analysed. The ΔCq indicator should therefore be established for every workflow. Once calibrated the ΔCq indicator can be used to compare processed samples to identify outliers that should be reanalysed or discarded (see Supplementary file).
Despite several advances, circulating microRNA research has been hampered by inconsistency and poor reproducibility1. The Two-tailed quality control panel developed here is a simple yet powerful tool for researchers to optimize new workflows, assess the technical performance of an analysis, identify outlier samples, and generally improve the reliability of circulating microRNA data.
Methods
Oligonucleotides
Sequences of mature microRNAs were obtained from the miRBase release 22 (www.mirbase.org). RNA oligonucleotides with 5′-phosphate were synthesized and quantified by Integrated DNA Technologies. Spike-in miRNA sequences were screened in silico for homology against human, mouse and rat miRBase records (Release 22) with the following parameters - search sequences: mature miRNAs, search method: SSEARCH, e-value cut-off: 100, max. no. of hits: 100. No significant homology was found. DNA oligonucleotides were synthesized and quantified by Invitrogen. Sequences are available in Supplementary file.
Samples
For the preparation of human serum, blood was collected from two healthy volunteers into 8.5 ml BD Vacutainer SST II Advance tubes (Beckman Dickinson) and allowed to clot for at least 30 min before centrifugation at 1500 g for 10 min at room temperature. The serum was then transferred to 2 ml tubes (Eppendorf) and stored at −80 °C. For the preparation of human plasma, blood was collected from four healthy volunteers into K2EDTA BD Vacutainer tubes (Beckman Dickinson) and centrifuged within 30 min at 1500 g for 15 min at room temperature. The plasma fraction was aspirated and transferred to 2 ml tubes (Eppendorf) and centrifuged again for 15 min at 3000 g. The supernatant was transferred to new 2 ml tubes and stored at −80 °C until analysis. Informed consent was obtained from all volunteers participating in the study. All procedures involving the use of human samples were performed in accordance with the ethical standards of Institute of Experimental Medicine, Academy of Sciences of the Czech Republic and with the Declaration of Helsinki. All methods were approved by the Ethical committee of the Institute of Experimental Medicine (decision on 22 June 2018, approval number 04/2018). For the preparation of rat serum, animals were anesthetized using 2–4% isoflurane. One millilitre of blood was collected from orbital plexus into 2 ml tubes (Eppendorf) using glass capillary. Blood was allowed to clot for 1 hour at room temperature and then centrifuged at 1000 g for 10 min. The clot was mechanically retracted from the tube wall before the centrifugation. Serum was transferred to another 2 ml tube and centrifuged a second time at 3000 g for 10 min. The supernatant was then transferred to cryovials (Biologix) and stored at −80 °C until analysis. All procedures involving the use of laboratory animals were performed in concordance with the European Community Council Directive of 24 November 1986 (86/609/EEC) and animal care guidelines approved by the Institute of Experimental Medicine, Academy of Sciences of the Czech Republic (Animal Care Committee decision on 17 April 2009; approval number 85/2009).
Haemolysis dilution series
After whole-blood centrifugation, erythrocytes from the lower phase were collected into a separate tube and subjected to a freeze-thaw cycle followed by vigorous vortexing for at least 90 seconds to lyse the erythrocytes. The haemolysed test sample was prepared by adding 1% (v/v) of lysed erythrocytes into a non-haemolysed sample. A two-fold haemolysis dilution series was prepared by diluting the haemolysed sample sequentially with non-haemolysed sample. Dilution series from two subjects were prepared for each biofluid type (human serum, human plasma, and rat serum). Absorbance of free haemoglobin was measured at 414 nm, 540 nm, and 578 nm with a NanoDrop 2000 spectrophotometer (ThermoFisher) in duplicates. RNA was isolated from the serum and plasma samples as described below, starting with either 200 μl (human) or 150 μl (rat) input volume.
RNA isolation
Total RNA was isolated from human plasma, and human and rat serum samples using the miRNeasy Serum/Plasma Advanced Kit (Qiagen) according to the manufacturer´s instructions. 1 μl of isolation spike-in mix containing synthetic cel-miR-54 (1e + 7 copies/μl), spike-A (2e + 5 copies/μl), spike-B (4e + 3 copies copies/μl) and, when appropriate 1 μl of GlycoBlue Coprecipitant (15 mg/mL) (Invitrogen), per sample was added at the lysis step. RNA was eluted into 20 μl nuclease-free water and stored at −80 °C.
Reverse transcription and quantitative PCR
Reverse transcription (RT) reactions were performed with the qScript flex cDNA kit (Quantabio) in a total reaction volume of 10 μl. One reaction contained 2 μl of template RNA, 1x buffer, mix of 0.05 μM Two-tailed RT primers, 1 μl of GSP enhancer and 0.5 μl of RT enzyme, and nuclease-free water up to 10 μl. RT reactions were incubated in a CFX 1000 thermocycler (Bio-Rad) for 45 min at 25 °C, 5 min at 85 °C and then held at 4 °C. Immediately after incubation, cDNA was diluted by addition of 50 μl nuclease-free water. Quantitative PCR (qPCR) was performed in a total volume of 10 μl. One reaction contained 1x SYBR Grandmaster Mix (Tataa Biocenter), forward and reverse primer (final concentration 0.4 μM), and 2 μl of diluted cDNA template (resulting in a final cDNA dilution of 15x). qPCR was performed in duplicates and incubated in a 384-well plate in a CFX 384 Real Time Detection System (Bio-Rad) at 95 °C for 30 s, 45 cycles of 95 °C for 5 s, and 60 °C for 15 s followed by melting-curve analysis.
Data Availability
Cq values were pre-processed with CFX Manager 3.1 (Bio-Rad). Missing values were replaced with maximum Cq per assay + 1 (Cqmax + 1). Paired two-tailed T-test was used to calculate significance of difference of mean Cq values between extractions with and without glycogen and F-test was used to calculate significance of difference of spread of replicates (Fig. 3). For the calculation of F-test, Cq values were transformed to achieve normal distribution as: 2^ΔCq (Cq − Cqmean), where Cqmean represents mean Cq of particular assay.
References
He, Y. et al. Current State of Circulating MicroRNAs as Cancer Biomarkers. Clin Chem 61, 1138–1155, https://doi.org/10.1373/clinchem.2015.241190 (2015).
Anfossi, S., Babayan, A., Pantel, K. & Calin, G. A. Clinical utility of circulating non-coding RNAs - an update. Nat Rev Clin Oncol 15, 541–563, https://doi.org/10.1038/s41571-018-0035-x (2018).
Ojha, R., Nandani, R., Pandey, R. K., Mishra, A. & Prajapati, V. K. Emerging role of circulating microRNA in the diagnosis of human infectious diseases. J Cell Physiol, https://doi.org/10.1002/jcp.27127 (2018).
Blandford, S. N., Galloway, D. A. & Moore, C. S. The roles of extracellular vesicle microRNAs in the central nervous system. Glia, https://doi.org/10.1002/glia.23445 (2018).
Guay, C. & Regazzi, R. Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol 9, 513–521, https://doi.org/10.1038/nrendo.2013.86 (2013).
Matsuzaki, J. & Ochiya, T. Circulating microRNAs and extracellular vesicles as potential cancer biomarkers: a systematic review. Int J Clin Oncol 22, 413–420, https://doi.org/10.1007/s10147-017-1104-3 (2017).
Kreth, S., Hubner, M. & Hinske, L. C. MicroRNAs as Clinical Biomarkers and Therapeutic Tools in Perioperative Medicine. Anesth Analg 126, 670–681, https://doi.org/10.1213/ANE.0000000000002444 (2018).
Lee, I., Baxter, D., Lee, M. Y., Scherler, K. & Wang, K. The Importance of Standardization on Analyzing Circulating RNA. Mol Diagn Ther 21, 259–268, https://doi.org/10.1007/s40291-016-0251-y (2017).
Witwer, K. W. Circulating microRNA biomarker studies: pitfalls and potential solutions. Clin Chem 61, 56–63, https://doi.org/10.1373/clinchem.2014.221341 (2015).
Kirschner, M. B., van Zandwijk, N. & Reid, G. Cell-free microRNAs: potential biomarkers in need of standardized reporting. Front Genet 4, 56, https://doi.org/10.3389/fgene.2013.00056 (2013).
Moldovan, L. et al. Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med 18, 371–390, https://doi.org/10.1111/jcmm.12236 (2014).
Chugh, P. & Dittmer, D. P. Potential pitfalls in microRNA profiling. Wiley Interdiscip Rev RNA 3, 601–616, https://doi.org/10.1002/wrna.1120 (2012).
van Vliet, E. A. et al. Standardization procedure for plasma biomarker analysis in rat models of epileptogenesis: Focus on circulating microRNAs. Epilepsia 58, 2013–2024, https://doi.org/10.1111/epi.13915 (2017).
Watson, A. K. & Witwer, K. W. Do platform-specific factors explain microRNA profiling disparities? Clin Chem 58, 472–474; author reply 474–475, https://doi.org/10.1373/clinchem.2011.175281 (2012).
Tuck, M. K. et al. Standard Operating Procedures for Serum and Plasma Collection: Early Detection Research Network Consensus Statement Standard Operating Procedure Integration Working Group. J Proteome Res 8, 113–117, https://doi.org/10.1021/pr800545q (2009).
Ainsztein, A. M. et al. The NIH Extracellular RNA Communication Consortium. J Extracell Vesicles 4, 27493, https://doi.org/10.3402/jev.v4.27493 (2015).
Laurent, L. C. & Alexander, R. Serum Collection Procedure (Small Scale) for the analysis of extracellular RNA (2015).
Laurent, L. C. & Alexander, R. Plasma Collection Procedure (Small Scale) for the analysis of extracellular RNA (2015).
McDonald, J. S., Milosevic, D., Reddi, H. V., Grebe, S. K. & Algeciras-Schimnich, A. Analysis of circulating microRNA: preanalytical and analytical challenges. Clin Chem 57, 833–840, https://doi.org/10.1373/clinchem.2010.157198 (2011).
Blondal, T. et al. Assessing sample and miRNA profile quality in serum and plasma or other biofluids. Methods 59, S1–6, https://doi.org/10.1016/j.ymeth.2012.09.015 (2013).
Marzi, M. J. et al. Optimization and Standardization of Circulating MicroRNA Detection for Clinical Application: The miR-Test Case. Clin Chem 62, 743–754, https://doi.org/10.1373/clinchem.2015.251942 (2016).
Li, Y. & Kowdley, K. V. Method for microRNA isolation from clinical serum samples. Anal Biochem 431, 69–75, https://doi.org/10.1016/j.ab.2012.09.007 (2012).
Mitchell, P. S. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 105, 10513–10518, https://doi.org/10.1073/pnas.0804549105 (2008).
Buschmann, D. et al. Toward reliable biomarker signatures in the age of liquid biopsies - how to standardize the small RNA-Seq workflow. Nucleic Acids Res 44, 5995–6018, https://doi.org/10.1093/nar/gkw545 (2016).
Androvic, P., Valihrach, L., Elling, J., Sjoback, R. & Kubista, M. Two-tailed RT-qPCR: a novel method for highly accurate miRNA quantification. Nucleic Acids Res 45, e144, https://doi.org/10.1093/nar/gkx588 (2017).
El-Khoury, V., Pierson, S., Kaoma, T., Bernardin, F. & Berchem, G. Assessing cellular and circulating miRNA recovery: the impact of the RNA isolation method and the quantity of input material. Sci Rep 6, 19529, https://doi.org/10.1038/srep19529 (2016).
Li, X., Mauro, M. & Williams, Z. Comparison of plasma extracellular RNA isolation kits reveals kit-dependent biases. Biotechniques 59, 13–17, https://doi.org/10.2144/000114306 (2015).
Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin Chem 56, 1733–1741, https://doi.org/10.1373/clinchem.2010.147405 (2010).
Max, K. E. A. et al. Human plasma and serum extracellular small RNA reference profiles and their clinical utility. Proc Natl Acad Sci USA 115, E5334–E5343, https://doi.org/10.1073/pnas.1714397115 (2018).
Pritchard, C. C. et al. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev Res (Phila) 5, 492–497, https://doi.org/10.1158/1940-6207.CAPR-11-0370 (2012).
Brunet-Vega, A. et al. Variability in microRNA recovery from plasma: Comparison of five commercial kits. Anal Biochem 488, 28–35, https://doi.org/10.1016/j.ab.2015.07.018 (2015).
McAlexander, M. A., Phillips, M. J. & Witwer, K. W. Comparison of Methods for miRNA Extraction from Plasma and Quantitative Recovery of RNA from Cerebrospinal Fluid. Front Genet 4, 83, https://doi.org/10.3389/fgene.2013.00083 (2013).
Ramon-Nunez, L. A. et al. Comparison of protocols and RNA carriers for plasma miRNA isolation. Unraveling RNA carrier influence on miRNA isolation. Plos One 12, e0187005, https://doi.org/10.1371/journal.pone.0187005 (2017).
Andreasen, D. et al. Improved microRNA quantification in total RNA from clinical samples. Methods 50, S6–9, https://doi.org/10.1016/j.ymeth.2010.01.006 (2010).
Kirschner, M. B. et al. Haemolysis during sample preparation alters microRNA content of plasma. Plos One 6, e24145, https://doi.org/10.1371/journal.pone.0024145 (2011).
Kirschner, M. B. et al. The Impact of Hemolysis on Cell-Free microRNA Biomarkers. Front Genet 4, 94, https://doi.org/10.3389/fgene.2013.00094 (2013).
Kim, D. J. et al. Plasma components affect accuracy of circulating cancer-related microRNA quantitation. J Mol Diagn 14, 71–80, https://doi.org/10.1016/j.jmoldx.2011.09.002 (2012).
Sourvinou, I. S., Markou, A. & Lianidou, E. S. Quantification of circulating miRNAs in plasma: effect of preanalytical and analytical parameters on their isolation and stability. J Mol Diagn 15, 827–834, https://doi.org/10.1016/j.jmoldx.2013.07.005 (2013).
Duy, J., Koehler, J. W., Honko, A. N. & Minogue, T. D. Optimized microRNA purification from TRIzol-treated plasma. BMC Genomics 16, 95, https://doi.org/10.1186/s12864-015-1299-5 (2015).
Pizzamiglio, S. et al. A methodological procedure for evaluating the impact of hemolysis on circulating microRNAs. Oncol Lett 13, 315–320, https://doi.org/10.3892/ol.2016.5452 (2017).
Shah, J. S., Soon, P. S. & Marsh, D. J. Comparison of Methodologies to Detect Low Levels of Hemolysis in Serum for Accurate Assessment of Serum microRNAs. Plos One 11, e0153200, https://doi.org/10.1371/journal.pone.0153200 (2016).
Kim, Y. K., Yeo, J., Kim, B., Ha, M. & Kim, V. N. Short structured RNAs with low GC content are selectively lost during extraction from a small number of cells. Mol Cell 46, 893–895, https://doi.org/10.1016/j.molcel.2012.05.036 (2012).
Acknowledgements
This study was supported by following grants: Czech Science Foundation P303/18/21942S; RVO 86652036; BIOCEV CZ.1.05/1.1.00/02.0109; EU H2020 SC1-HCO-02-2016 “Standardisation of pre-analytical and analytical procedures for in vitro diagnostics in personalised medicine”; SPIDIA4P and IMI call 11 “Blood-based biomarker assays for personalised tumour therapy: value of latest circulating biomarkers”; CANCER-ID.
Author information
Authors and Affiliations
Contributions
P.A. and L.V. conceived and designed the study. P.A. performed experiments, analysed the data, prepared figures and drafted the manuscript. N.R. and L.U.-M. provided and processed animal samples. E.R., L.V. and M.K. commented on the manuscript and produced the final version. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
M.K. has shares in TATAA Biocenter AB. P.A., N.R., L.U.-M., E.R. and L.V. declare no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Androvic, P., Romanyuk, N., Urdzikova-Machova, L. et al. Two-tailed RT-qPCR panel for quality control of circulating microRNA studies. Sci Rep 9, 4255 (2019). https://doi.org/10.1038/s41598-019-40513-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-40513-w
This article is cited by
-
Current challenges and best practices for cell-free long RNA biomarker discovery
Biomarker Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.