Abstract
The identification of Human herpesvirus 6B (HHV-6B) epitopes that are recognized by T-cells could contribute to the development of potential vaccines and immunotherapies. Here, we identified CD4+ and H-2Kd-restricted CD8+ T-cell epitopes on the glycoprotein Q1 of HHV-6B (BgQ1), which is a unique glycoprotein and essential for HHV-6B viral entry, by using in vivo electroporation with a plasmid DNA encoding BgQ1, overlapping peptides spanning the BgQ1 sequence, ELISPOT assay for quantification of gamma interferon (IFN-γ), and computer-based T-cell epitope prediction programs. The CD4+ and CD8+ T-cell epitopes identified in BALB/c mice in this study could be a good animal model system for use in the development of T-cell responses, inducing HHV-6B vaccines or immunotherapies.
Similar content being viewed by others
Introduction
Human herpesvirus 6 (HHV-6), which belongs to the betaherpesvirus subfamily, was first isolated from patients with lymphocytic disorders in 19861. Since 2012, HHV-6 has been classified as two independent virus species, HHV-6A and HHV-6B2, based on genetic and antigenic differences and cell tropism3,4,5. HHV-6B is the causative agent for exanthem subitum6 and is sometimes associated with severe encephalopathy, while the pathogenesis of HHV-6A is still unknown. More than 90% of individuals are infected with HHV-6B during childhood, and the virus remains latent after primary infection7,8. HHV-6B reactivation causes life-threatening encephalitis in immunosuppressed patients9 and is also associated with drug-induced hypersensitivity syndrome10. However, effective immunotherapies based on antibodies, expanded T-cells, or vaccines for controlling HHV-6B infection and reactivation have not been established.
Glycoproteins or their complexes on the surface of enveloped viruses play pivotal roles in the viruses’ infectivity. Glycoprotein Q1 (gQ1) and gQ2 are unique genes that are encoded specifically in HHV-6 and human herpesvirus 7 (HHV-7). Recently, our group reported that human CD134, also called OX40, is a specific cellular receptor for HHV-6B and binds to the HHV-6B gH/gL/gQ1/gQ2 complex11. Moreover, the HHV-6B gQ1 (BgQ1) and gQ2 (BgQ2) subunits are sufficient for CD134 binding, and a region in BgQ1 is critical for the function of the HHV-6B gH/gL/gQ1/gQ2 complex12. When screening for monoclonal antibodies (MAbs) with neutralizing activity against HHV-6B, the neutralizing MAbs obtained were almost all against BgQ113. Thus, the BgQ1 protein, which is unique to HHV-6B, seems to be critical for virus entry and adaptive immunity.
The recognition by CD8+ cytotoxic T lymphocytes (CTL) of antigen peptides presented by class I human leucocyte antigen (HLA) is an essential step in adaptive immunity for virus infection. Identifying immunodominant proteins and epitopes could lead to the design of potential new vaccines and immunotherapies. Recently, some studies have identified HHV-6 antigens that are targeted by the CD4+ and CD8+ T-cell responses14,15. Most of these focused on functional or positional homologs to known T-cell antigens from human cytomegalovirus (HCMV) in chronically infected adults, because HCMV also belongs to the same betaherpesvirus subfamily as HHV-6B. The T-cells responding to the identified HHV-6 antigens are present at low frequency in healthy adults and need to be expanded in vitro for use in autologous immunotherapy16,17,18. In addition, to design vaccines and immunotherapies for HHV-6B infection in humans, a good animal model will be needed to analyze T-cell responses against the HHV-6B antigen.
DNA vaccines can engender not only humoral but also cellular immune responses19. And they offer some advantages including easy construction, preparation, stability. The vaccine immunogenicity and efficacy are significantly enhanced by in vivo electroporation20.
In this report, we attempted to determine T-cell epitopes on BgQ1, which is a unique glycoprotein and essential for HHV-6B viral entry. Due to the similarities in the peptide-binding motifs between H-2Kd and HLA-A24, BALB/c mice were chosen as the animal model hosts21,22,23,24. We identified an H-2Kd-restricted CD8+ T-cell epitope in BALB/c mice by using in vivo electroporation with a plasmid DNA encoding BgQ1, overlapping peptides spanning the BgQ1 sequence, ELISPOT assay for quantification of gamma interferon (IFN-γ), and computer-based T-cell epitope prediction programs.
Results
Identification of CD4+ or CD8+ T-cell stimulating peptides by using the synthetic overlapping peptides from BgQ1
As a first preliminary ELISPOT assay, overlapping peptides from BgQ1 (P1–P48) were used for stimulation. Peptides were divided into two groups: the first group contained the first 24 peptides (P1–P24) and the second group contained the other 24 peptides (P25–P48). Splenocytes from BALB/c mice immunized with a plasmid DNA encoding BgQ1 were stimulated with the individual peptides for 40 h, and IFN-γ spots were counted. Independent experiments were carried out from independent mice. As shown in Fig. 1a,b, splenocytes in the presence of 12 peptides (P7, P8, P11, P16, P17, P18 from the first half, and P33, P36, P37, P38, P43, P44 from the second half) showed more than 50 SFC/1 × 106 splenocytes.
IFN-γ ELISPOT assay of splenocytes from BALB/c mice immunized with pCAGGS-MCS BgQ1m. Overlapping peptides (5 µg/ml for each peptide) from glycoprotein Q1 of human herpesvirus 6B (BgQ1) were used for stimulation. Peptides were divided into two groups: the first group contained the first 24 peptides (P1-P24) (a) and the second group contained the other 24 peptides (P25-P48) (b). (c) Splenocytes from immunized mice were stimulated with the selected 12 peptides (5 µg/ml for each peptide). The results were expressed as spot forming cells (SFC)/1 × 106 splenocytes and the wells of “not reliably countable high signals” were defined as 300 SFC/1 × 106 splenocytes. Pep(-) indicates medium alone. The data are the mean ± SD of duplicate (a,b) and triplicated (c) wells. And the data are representative of three independent experiments. Independent experiments were carried out from independent mice.
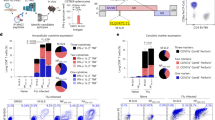
In a second preliminary ELISPOT assay, splenocytes from immunized mice were stimulated with the 12 peptides that showed more than 50 SFC/1 × 106 splenocytes in the first ELISPOT. As shown in Fig. 1c, substantial IFN-γ production was confirmed in splenocytes after stimulation with 5 peptides (P11, P16, P17, P18, and P43). The remaining 7 peptides were considered to have lower immunogenicity. These preliminary results were further confirmed by an independently performed three-color flow cytometric analysis. To reveal the responsive T-cell subset, the findings of intracellular IFN-γ staining after stimulation with these 5 peptides were examined. The results demonstrated that CD8+ T-cells mainly produced IFN-γ in response to P17 and P18 (Fig. 2). On the other hand, CD4+ T-cells mainly produced IFN-γ in the presence of P16 and P43. CD4+ T-cells also produced IFN-γ in the presence of P11, but the IFN-γ signals were lower than the other 4 peptides.
Identification of peptides inducing CD8+ T-cell responses. IFN-γ-producing T-cells in the spleens of BALB/c mice immunized with pCAGGS-MCS BgQ1m. (a) Three-color flow cytometric analysis was performed for the detection of intracellular IFN-γ and cell surface CD4 and CD8 molecules after immune splenocytes were cultured in the presence of the 5 candidate peptides. The data are the percentages of IFN-γ-producing CD4+ or CD8+ cells in lymphocyte after 4 h of stimulation with peptides (mean ± SD of duplicates). Peptide(-) indicates medium alone. The data are representative of three independent experiments with similar results. (b) A representative flow cytometry plot of intracellular IFN-γ and CD8 and CD4 in response to P17 peptide. The lymphocytes were gated by forward scatter (FSC) and side scatter (SSC) and then intracellular IFN-γ levels were detected on CD4+ and CD8+ cells in the lymphocytes. The data are shown as the percentages of IFN-γ-producing cells in CD4+ and CD8+ lymphocyte gate.
Identification of an MHC class Ia restriction molecule for P17 and P18 of BgQ1
Since P17 and P18 were found to stimulate CD8+ T-cells, we next tried to determine which MHC class Ia molecule was involved in the presentation of peptides to CD8+ T-cells. BW5147 (H2k) lymphoma cell lines expressing either H-2Kd, H-2Dd, H-2Ld, or just the wild type molecule (H2k) were co-cultured with P17 or P18, and used for stimulation of splenocytes from BgQ1-immunized mice. IFN-γ production for both P17 and P18 was only observed in H-2Kd expressing BW5147 (Fig. 3), indicating P17 and P18 contained the H-2Kd-restricted CD8+ T-cell epitope.
Identification of MHC class Ia restriction molecules for P17 and P18 of BgQ1. BW5147 (H2k) lymphoma cell lines expressing either H-2Kd, H-2Dd, H-2Ld, or just the wild type were co-cultured with P17 or P18, and used for stimulation of splenocytes from BALB/c mice immunized with pCAGGS-MCS BgQ1m. IFN-γ productions were measured by ELISPOT assay. Results are expressed as spot forming cells (SFC)/1 × 106 splenocytes. The data are the mean ± SD of three independent experiments of one mouse.
Identification of a 9-mer CD8+ T-cell epitope on P17 and P18 of BgQ1
Several CD8+ T-cell epitope candidates within the P17 and P18 were predicted by using two computer-based programs BIMAS HLA Peptide Binding Prediction (http://bimas.dcrt.nih.gov/cgi-bin/molbio/ken_parker_comboform) and SYFPEITHI Epitope Prediction (http://www.syfpeithi.de/). Table 1 shows the results of these programs. Since P17 and P18 are both presented by H-2Kd, 4 peptides which had high scores were synthesized as epitope candidates: an 8-mer (FCPMTSKL), a 9-mer (AFCPMTSKL), and a 10-mer (IAFCPMTSKL) peptide from the overlapping region of P17 and P18; and a 9-mer peptide (KPLTAMTAI) from the front half of P17.
As shown in Fig. 4a,b, the 9-mer peptide (AFCPMTSKL) from the overlapping region of P17 and P18 induced the strongest intracellular IFN-γ signals in the CD8+ T-cells by flow cytometric analysis, followed by the 10-mer (IAFCPMTSKL) and 8-mer (FCPMTSKL) peptides, indicating that this 9mer peptide is the optimal H-2Kd-restricted CD8+ T-cell epitope on BgQ1, in BALB/c mice. Another 9-mer peptide (KPLTAMTAI) from the front half of P17 induced only a background level of intracellular IFN-γ synthesis.
Determination of an optimal T-cell epitope on the P17 and P18 peptides and a T-cell subset recognizing the epitope in BALB/c mice. (a) IFN-γ-producing T-cell subsets in the spleens of BALB/c mice immunized with pCAGGS-MCS BgQ1m. Three-color immunofluorescence analysis was performed on flow cytometer to detect intracellular IFN-γ, cell surface CD4, and CD8 molecules after immune splenocytes were cultured in the presence of the peptides. The data are the percentages of IFN-γ-producing CD4+ or CD8+ cells in lymphocyte after 4 h of stimulation with peptides (mean ± SD of duplicates). Peptide(-) indicates medium alone. Representative data from three independent experiments with similar results are shown. (b) A representative flow cytometry plot of intracellular IFN-γ and CD8 (left) and CD4 (right) in response to the 9-mer peptide (AFCPMTSKL) from an overlapping region of P17 and P18. Very few intracellular IFN-γ-positive CD4+ T-cells were observed after stimulation with the AFCPMTSKL peptide (right). The data are shown as the percentages of IFN-γ-producing cells in lymphocyte gate.
Discussion
In this study, we identified an H-2Kd-restricted CD8+ T-cell epitope on the BgQ1 molecule. In addition, we found that P43 and P16 of the BgQ1 contain CD4+ T-cell epitopes in BALB/c mice.
In general, CTL plays a crucial role in protective immunity against infection with intracellular pathogens, such as certain bacteria and viruses25,26. Although many things are still unknown about the mechanisms of T-cell mediated protection against HHV-6B infection, the necessity of T-cells to control HHV-6B replication is suggested by the higher incidence of persistent HHV-6B viremia in patients without proliferative T-cell responses27. To better understand CD8+ T-cell responses against HHV-6B infection, identifying the epitopes which are recognized by CD8+ T-cells is important. However, the HHV-6B genome encodes hundreds of proteins, making the identification of immunodominant epitopes complicated. Recent reports which defined CD4+ and CD8+ T-cell epitopes by using expanded T-cells in vitro focused on the HHV-6 proteins present at high levels in virus preparations18, or on the HHV-6 homologies of antigens defined for HCMV16,17. In this study, we focused on the BgQ1 protein, which plays an essential role in HHV-6B virus entry and is a promising candidate for antiviral therapy13.
In the present mouse model, we identified the H-2Kd-restricted CTL epitope on the BgQ1 molecule. This epitope is not conserved in HHV-6A gQ1 sequence (strain U1102; PubMed accession NC 001664), but similar sequence (RFCPMTTKL) which has same amino acids in the main anchor positions of nonameric Kd epitope is present (position 2 and 9).
In this study, mice were immunized with a codon-optimized plasmid DNA expressing the BgQ1 by in vivo electroporation to identify the CTL epitopes. DNA vaccination is a powerful tool for identifying T-cell epitopes. However, optimization of codon usage is an important consideration in constructing DNA vaccines26. DNA immunization with an optimized codon has been reported to result in enhanced CTL reactivity by increasing the translational efficiency of plasmid DNA28,29. Moreover, an in vivo electroporation technique to induce transient and reversible permeabilization of the cell membrane improves the efficiency of plasmid DNA transfection30. Electroporation enhances the immunogenicity and effectiveness of DNA vaccines by increasing antigen delivery up to 1000-fold over naked DNA delivery alone20.
IFN-γ producing cells were detected by three-color flow cytometric analysis. It might have been better if we had used Live/Dead and other markers of CTL activity, because the percentages of IFN-γ-producing cells in this study were relatively small. These markers may influence our results. However, similar number of IFN-γ producing cells with similar system were reported in another study31,32 and IFN-γ production was consistently detected over the background level.
We also used the computer-based algorithm programs BIMAS and SYFPEITHI to predict the epitopes33,34,35. The strategy including DNA vaccines, overlapping peptide and these computer-based programs is an effective methods for narrowing down the amino acid region of the T-cell epitope31. Prediction in these programs is restricted to for 8–11mer length peptides. It would be possible CTL can recognize much longer peptides up to 14mer36, however we did not pursue this long CTL peptide in this study. We also found that P43 and P16 of the BgQ1 contain CD4+ T-cell epitopes in BALB/c mice. Especially, P43 contains 15mer peptide which showed high score (VNNIFTVQARYSKQN, score 31) for binding to the H-2Ad molecule by SYFPEITHI Epitope Prediction. But MHC class II prediction tools do not perform as well as class I predictions37, there is room for further research.
In conclusion, we identified an H-2Kd-restricted CD8+ T-cell epitope on the BgQ1 molecule and two peptides which contain CD4+ T-cell epitopes in BALB/c mice. These results warrant further study to examine whether the epitopes are applicable to animal models or humans and could be utilized in vaccines or immunotherapies.
Materials and Methods
Mice
Six-week-old female BALB/c mice (16–20 g) were obtained from Japan SLC (Shizuoka, Japan). The mice were housed in-house under specific pathogen-free conditions maintained at 22 ± 2 °C and 55 ± 5% relative humidity, in a 12-hour light/dark cycle environment, and provided with food and water ad libitum. The health condition of the mice was monitored daily. A total of 25 mice were used including preliminary experiments. All mice were immunized with plasmid DNA and each experiment was independently repeated three times. All of the animal experimental procedures were approved by the Kobe University Institutional Animal Care and Use Committee (Permission number: P131101-R1) and carried out according to the Animal Experimentation Regulations of Kobe University.
Construction of the plasmid DNAs, pCAAGGS-MCS BgQ1m
To optimize the gene expression in mammalian cells, the BgQ1 original cDNA (GenBank accession no. MK388090) codon was converted to more common in human genes and cloned into the pMX plasmid by Invitrogen (Carlsbad, CA). The amplified BgQ1 DNA fragments were digested with SacI and KpnI and ligated into the digested pCAGGS-MCS vector, yielding the plasmid pCAGGS-MCS BgQ1m. The plasmid was amplified in DH5α Escherichia coli and purified using QIAGEN Plasmid Mega Kit (QIAGEN, Hilden, Germany) following the manufacturer’s instructions.
Peptides
BgQ1 of the HHV-6B HST strain has 516-amino-acid (aa). Except for the signal sequence (aa 1 to 25), peptides spanning the total 491 aa BgQ1 sequence were prepared as 20-mers overlapping by 10 residues (Supplementary Fig. S1). Forty-eight lyophilized peptides in total (P1–P48) were purchased from Eurofins Genomics (Tokyo, Japan). The purity of the peptides was 50% or more, confirmed by high-performance liquid chromatography (HPLC) and verified for correct sequence by mass spectroscopy (MS). All peptides were dissolved in dimethyl sulfoxide (DMSO) at a concentration of 10 mg/ml and stored at −80 °C until use.
Immunization of mice
Mice were immunized with a plasmid DNA expressing the BgQ1 by in vivo electroporation. After anesthesia by isoflurane, 100 µg of pCAGGS-MCS BgQ1m was injected twice into both femoral muscles of the mice, and an NEPA21 electroporator (NepaGene, Tokyo, Japan) was used for transfection. Immunizations were performed three times at 2-week intervals and the spleens were removed one week after the third immunization.
Cell lines
BW5147 (H2k) lymphoma cell line (JCRB9002) was obtained from JCRB Cell Bank (Japan). BW5147 cells retrovirally transduced with one of the genes encoding H-2Kd, H-2Dd, or H-2Ld were made as previously described32, and used to determine the CD8+ T-cell epitope-presenting MHC Ia molecule. The cells were cultured in RPMI 1640 medium with 10% heat-inactivated fetal bovine serum (RPMI/10FBS) in an incubator with a humidified atmosphere containing 5% CO2. FBS, which originate from Canada (endotoxin level ≤ 50 EU/mL, hemoglobin level ≤ 25 mg/dL), was purchased from Gibco (cat. # 12483020, Thermo Fisher Scientific, Waltham, MA).
Preparation of splenocytes from an immunized mouse
Immunized mice were euthanized by an intraperitoneal injection of sodium pentobarbital and cervical dislocation. The spleens were aseptically removed and single cell suspensions were prepared after dissociation of the spleen through a cell strainer. Splenocytes were isolated by Ficoll density gradient centrifugation, suspended in RPMI/10FBS at a concentration of 1 × 107 cells/ml, and used for assay.
Quantification of IFN-γ by ELISPOT assay
The wells of a 96-well MultiScreen HA plate (Millipore, Billerica, MA) were precoated and blocked with RPMI/10FBS according to the manufacturer’s instructions (Ready-Set-Go Mouse IFN-γ ELISPOT kit; eBioscience, San Diego, CA). After blocking, 50 µl of RPMI/10FBS was added to each well followed by 100 µl of splenocytes (1 × 106 cells) from immunized mice. All dissolved peptides in DMSO were diluted in RPMI/10FBS to a concentration of 20 µg/ml, and 50 µl of the resulting solution was added to each well. The total volume was 200 µl/well and the final concentration of each peptide was 5 µg/ml. Measurements were made in duplicate for each peptide in the first preliminary ELISPOT, and in triplicate in the second preliminary ELISPOT.
The plates were incubated for 40 h at 37 °C in a 5% CO2 humidified incubator. IFN-γ spot-forming cells (SFC) were detected and developed according to the manufacturer’s instructions (Ready-Set-Go Mouse IFN-γ ELISPOT kit; eBioscience). As a minor modification, 100 µl of 3,3′,5,5′-tetramethylbenzidine-H substrate (Moss, Pasadena, MD) was used for development of spots, and the spots were developed for 3 min at room temperature. After stopping the substrate reaction by washing wells with water, the plates were dried. The spots were then enumerated using the KS ELISPOT system (Carl Zeiss Microscopy GmbH, Jena, Germany) for automated spot counting. Because too many spots can cause signal overlapping and a reduction in counting accuracy, the wells of undercounting high signals were defined as 300 SFC per 1 × 106 splenocytes.
Intracellular IFN-γ staining and flow cytometry
Splenocytes from the immunized mice were treated with ammonium chloride-tris (ACT) buffer for 3 min at room temperature to remove red blood cells. The cells were then washed twice with RPMI 1640 medium and resuspended in RPMI/10FBS at a concentration of 1 × 107 cells/ml. The cells (2 × 106 cells) were incubated for 4 h at 37 °C in the presence or absence of peptide at a concentration of 10 µg/ml with Golgistop stock solution (monensin solution; BD Biosciences, San Jose, CA) diluted 1:1,500. The cells were then washed twice with 2% bovine serum albumin (BSA) in phosphate buffered saline (PBS) followed by staining with fluorescein isothiocyanate (FITC)-conjugated anti-CD8 (clone KT15; MBL, Nagoya, Japan) and phycoerythrin-indotricarbocyanine (PE/Cy7)-conjugated anti-CD4 (clone GK1.5; BioLegend, San Diego, CA) monoclonal antibodies (mAbs) for 30 min at 4 °C. The cells were washed twice and then intracellular cytokine staining (ICS) was performed by using a BD Cytofix/Cytoperm kit (BD Biosciences) according to the manufacturer’s protocol. ICS for IFN-γ was performed with PE-conjugated anti-IFN-γ (clone XMG1.2; BD Biosciences) mAbs for 30 min at 4 °C. The cells were washed twice, resuspended in PBS with 2% BSA, and then analyzed by flow cytometry with a SA3800 spectral analyzer (Sony Corporation, Tokyo, Japan).
Determination of the restricted MHC Ia molecule
The CD8+ T-cell epitope-presenting H-2d molecules, i.e., H-2Kd, H-2Dd, and H-2Ld, were identified as previously reported32. Briefly, BW5147-Kd, BW5147-Dd, BW5147-Ld, or BW5147 wild type cells (4 × 106 cells) were co-cultured with each peptide (10 µg/ml) at 37 °C for 1 h. The cells were washed three times with RPMI 1640 medium and resuspended in RPMI/10FBS at a concentration of 4 × 106 cells/ml. Splenocytes (1 × 106 cells) from the immunized mice were stimulated with 2 × 105 of each type of peptide-pulsed BW5147 cells in 200 µl of RPMI/10FBS for 40 h at 37 °C, and the levels of IFN-γ production were determined by ELISPOT assay.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Salahuddin, S. Z. et al. Isolation of a new virus, HBLV, in patients with lymphoproliferative disorders. Science 234, 596–601 (1986).
Adams, M. J. & Carstens, E. B. Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2012). Arch Virol 157, 1411–1422, https://doi.org/10.1007/s00705-012-1299-6 (2012).
Wyatt, L. S., Balachandran, N. & Frenkel, N. Variations in the replication and antigenic properties of human herpesvirus 6 strains. J Infect Dis 162, 852–857 (1990).
Aubin, J. T. et al. Several groups among human herpesvirus 6 strains can be distinguished by Southern blotting and polymerase chain reaction. J Clin Microbiol 29, 367–372 (1991).
Campadelli-Fiume, G., Guerrini, S., Liu, X. & Foa-Tomasi, L. Monoclonal antibodies to glycoprotein B differentiate human herpesvirus 6 into two clusters, variants A and B. J Gen Virol 74(Pt 10), 2257–2262, https://doi.org/10.1099/0022-1317-74-10-2257 (1993).
Yamanishi, K. et al. Identification of human herpesvirus-6 as a causal agent for exanthem subitum. Lancet 1, 1065–1067 (1988).
Saxinger, C. et al. Antibody reactivity with HBLV (HHV-6) in U.S. populations. J Virol Methods 21, 199–208 (1988).
Okuno, T. et al. Seroepidemiology of human herpesvirus 6 infection in normal children and adults. J Clin Microbiol 27, 651–653 (1989).
Ogata, M., Fukuda, T. & Teshima, T. Human herpesvirus-6 encephalitis after allogeneic hematopoietic cell transplantation: what we do and do not know. Bone Marrow Transplant 50, 1030–1036, https://doi.org/10.1038/bmt.2015.76 (2015).
Tohyama, M. et al. Association of human herpesvirus 6 reactivation with the flaring and severity of drug-induced hypersensitivity syndrome. Br J Dermatol 157, 934–940, https://doi.org/10.1111/j.1365-2133.2007.08167.x (2007).
Tang, H. et al. CD134 is a cellular receptor specific for human herpesvirus-6B entry. Proc Natl Acad Sci USA 110, 9096–9099, 10.1073/pnas.1305187110 (2013).
Tang, H., Wang, J., Mahmoud, N. F. & Mori, Y. Detailed study of the interaction between human herpesvirus 6B glycoprotein complex and its cellular receptor, human CD134. J Virol 88, 10875–10882, https://doi.org/10.1128/JVI.01447-14 (2014).
Kawabata, A. et al. Analysis of a neutralizing antibody for human herpesvirus 6B reveals a role for glycoprotein Q1 in viral entry. J Virol 85, 12962–12971, 10.1128/JVI.05622-11 (2011).
Becerra, A., Gibson, L., Stern, L. J. & Calvo-Calle, J. M. Immune response to HHV-6 and implications for immunotherapy. Curr Opin Virol 9, 154–161, https://doi.org/10.1016/j.coviro.2014.10.001 (2014).
Papadopoulou, A. et al. Activity of broad-spectrum T cells as treatment for AdV, EBV, CMV, BKV, and HHV6 infections after HSCT. Sci Transl Med 6, 242ra283, https://doi.org/10.1126/scitranslmed.3008825 (2014).
Martin, L. K., Schub, A., Dillinger, S. & Moosmann, A. Specific CD8(+) T cells recognize human herpesvirus 6B. Eur J Immunol 42, 2901–2912, https://doi.org/10.1002/eji.201242439 (2012).
Gerdemann, U. et al. Immunotherapeutic strategies to prevent and treat human herpesvirus 6 reactivation after allogeneic stem cell transplantation. Blood 121, 207–218, https://doi.org/10.1182/blood-2012-05-430413 (2013).
Nastke, M. D. et al. Human CD4+ T cell response to human herpesvirus 6. J Virol 86, 4776–4792, 10.1128/JVI.06573-11 (2012).
Gurunathan, S., Klinman, D. M. & Seder, R. A. DNA vaccines: immunology, application, and optimization*. Annu Rev Immunol 18, 927–974, https://doi.org/10.1146/annurev.immunol.18.1.927 (2000).
Sardesai, N. Y. & Weiner, D. B. Electroporation delivery of DNA vaccines: prospects for success. Curr Opin Immunol 23, 421–429, https://doi.org/10.1016/j.coi.2011.03.008 (2011).
Maier, R. et al. Peptide motifs of HLA-A3, -A24, and -B7 molecules as determined by pool sequencing. Immunogenetics 40, 306–308 (1994).
Okugawa, T. et al. A novel human HER2-derived peptide homologous to the mouse K(d)-restricted tumor rejection antigen can induce HLA-A24-restricted cytotoxic T lymphocytes in ovarian cancer patients and healthy individuals. Eur J Immunol 30, 3338–3346, 10.1002/1521-4141(200011)30:11<3338::AID-IMMU3338>3.0.CO;2-3 (2000).
Falk, K., Rotzschke, O., Stevanovic, S., Jung, G. & Rammensee, H. G. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. 1991. J Immunol 177, 2741–2747 (2006).
Inoue, M. et al. Identification of SPARC as a candidate target antigen for immunotherapy of various cancers. Int J Cancer 127, 1393–1403, https://doi.org/10.1002/ijc.25160 (2010).
Gulzar, N. & Copeland, K. F. CD8+ T-cells: function and response to HIV infection. Curr HIV Res 2, 23–37 (2004).
Uchijima, M., Yoshida, A., Nagata, T. & Koide, Y. Optimization of codon usage of plasmid DNA vaccine is required for the effective MHC class I-restricted T cell responses against an intracellular bacterium. J Immunol 161, 5594–5599 (1998).
Wang, F. Z., Linde, A., Dahl, H. & Ljungman, P. Human herpesvirus 6 infection inhibits specific lymphocyte proliferation responses and is related to lymphocytopenia after allogeneic stem cell transplantation. Bone Marrow Transplant 24, 1201–1206, https://doi.org/10.1038/sj.bmt.1702058 (1999).
Andre, S. et al. Increased immune response elicited by DNA vaccination with a synthetic gp120 sequence with optimized codon usage. J Virol 72, 1497–1503 (1998).
Nagata, T., Uchijima, M., Yoshida, A., Kawashima, M. & Koide, Y. Codon optimization effect on translational efficiency of DNA vaccine in mammalian cells: analysis of plasmid DNA encoding a CTL epitope derived from microorganisms. Biochem Biophys Res Commun 261, 445–451, 10.1006/bbrc.1999.1050 (1999).
Mir, L. M., Moller, P. H., Andre, F. & Gehl, J. Electric pulse-mediated gene delivery to various animal tissues. Adv Genet 54, 83–114, https://doi.org/10.1016/S0065-2660(05)54005-7 (2005).
Aoshi, T. et al. Identification of an HLA-A*0201-restricted T-cell epitope on the MPT51 protein, a major secreted protein derived from Mycobacterium tuberculosis, by MPT51 overlapping peptide screening. Infect Immun 76, 1565–1571, 10.1128/IAI.01381-07 (2008).
Aoshi, T., Suzuki, M., Uchijima, M., Nagata, T. & Koide, Y. Expression mapping using a retroviral vector for CD8+ T cell epitopes: definition of a Mycobacterium tuberculosis peptide presented by H2-Dd. J Immunol Methods 298, 21–34, https://doi.org/10.1016/j.jim.2004.12.015 (2005).
Rammensee, H., Bachmann, J., Emmerich, N. P., Bachor, O. A. & Stevanovic, S. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50, 213–219 (1999).
Parker, K. C., Bednarek, M. A. & Coligan, J. E. Scheme for ranking potential HLA-A2 binding peptides based on independent binding of individual peptide side-chains. J Immunol 152, 163–175 (1994).
Chen, A. et al. H-2 Kd-restricted hepatitis B virus-derived epitope whose specific CD8+ T lymphocytes can produce gamma interferon without cytotoxicity. J Virol 79, 5568–5576, https://doi.org/10.1128/JVI.79.9.5568-5576.2005 (2005).
Ekeruche-Makinde, J. et al. Peptide length determines the outcome of TCR/peptide-MHCI engagement. Blood 121, 1112–1123, 10.1182/blood-2012-06-437202 (2013).
Wang, P. et al. A systematic assessment of MHC class II peptide binding predictions and evaluation of a consensus approach. PLoS Comput Biol 4, e1000048, https://doi.org/10.1371/journal.pcbi.1000048 (2008).
Acknowledgements
We thank Dr. Jun-ichi Miyazaki for providing the pCAGGS-MCS plasmid. This work was partially supported by Acceleration Transformative research for Medical innovation (ACT-MS) from Japan Agency for Medical Research and Development (AMED) under Grant Number JP17im0210601.
Author information
Authors and Affiliations
Contributions
Y.M. conceived and supervised the project. T.A. designed the analysis and reviewed the manuscript. S.N. wrote the manuscript. S.N., A.K., Y.Y., M.N., S.K. and K.M. performed the experiments and analysis. H.Y supervised the project. All authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nagamata, S., Aoshi, T., Kawabata, A. et al. Identification of CD4 and H-2Kd-restricted cytotoxic T lymphocyte epitopes on the human herpesvirus 6B glycoprotein Q1 protein. Sci Rep 9, 3911 (2019). https://doi.org/10.1038/s41598-019-40372-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-40372-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.