Abstract
Centrioles are conserved organelles fundamental for the organisation of microtubules in animal cells. Oligomerisation of the spindle assembly abnormal protein 6 (SAS-6) is an essential step in the centriole assembly process and may act as trigger for the formation of these organelles. SAS-6 oligomerisation is driven by two independent interfaces, comprising an extended coiled coil and a dimeric N-terminal globular domain. However, how SAS-6 oligomerisation is controlled remains unclear. Here, we show that in the Caenorhabditis elegans SAS-6, a segment of the N-terminal globular domain, unresolved in crystallographic structures, comprises a flexible loop that assists SAS-6 oligomerisation. Atomistic molecular dynamics simulations and nuclear magnetic resonance experiments suggest that transient interactions of this loop across the N-terminal dimerisation interface stabilise the SAS-6 oligomer. We discuss the possibilities presented by such flexible SAS-6 segments for the control of centriole formation.
Similar content being viewed by others
Introduction
Centrioles are conserved organelles widespread in the eukaryotic kingdom1,2,3. In animals, a pair of centrioles comprise the structured core of centrosomes, which direct formation of the microtubule network and the mitotic spindle during cell division4,5. In this capacity, centrioles are crucial for controlling the overall cell architecture, facilitating intracellular cargo transport, anchoring the endoplasmic reticulum and the Golgi apparatus, and ensuring the equitable segregation of genetic material during mitosis. Furthermore, centrioles in all eukaryotic lineages except fungi and higher plants also act close to the membrane, where, as basal bodies, they template formation of microtubule-based cilia and flagella6. In this manner, centrioles are essential for diverse aspects of cellular behaviour including locomotion via flagellar and cillial beating, and sensing, via the antena-like primary cillium. Unsurprisingly, given the wide swath of cellular processes dependant on centrioles, mutations in genes coding for essential components of these organelles are linked to major human genetic disorders and diseases, including male sterility, ectopic pregnancies, multisystemic ciliopathies, primary microcephaly and potentially cancer7,8,9,10,11.
The formation of new centrioles is a highly regulated process which occurs once per cycle in dividing cells12,13,14,15. The main molecular features of the centriole assembly pathway are conserved13,14, and involve the initial localisation at the site of assembly of the coiled coil protein SPD-2 in Caenorhabditis elegans via interactions with the protein SAS-716, followed by the kinase ZYG-1 and SAS-6. Structural and functional studies of SAS-6 have revealed that this protein assists in establishing the canonical radial symmetry of centrioles17, thereby influencing a key element of the overall organelle architecture. SAS-6 forms large 9-fold symmetric oligomers in vitro18,19,20,21 that bear striking resemblance to scaffold-like assemblies observed at the centre of centrioles, the ‘cartwheels’, which are believed to seed formation of these organelles17,22. Disruption of SAS-6 oligomerisation directly abrogates the canonical pathway of centriole formation18,20,23, while SAS-6 variants engineered to form oligomers with symmetry other than 9-fold were seen to influence the organelle radial symmetry24. Thus, a broad consensus has emerged placing SAS-6 oligomerisation as a crucial molecular event at the onset of centriole assembly.
The mechanisms by which SAS-6 oligomerisation is controlled in cells remain, however, poorly understood. At the molecular level, oligomerisation is driven by two independent dimerisation interfaces on SAS-6, comprising a long, parallel, dimeric coiled-coil (the CC interface) and a dimeric globular domain at the protein N-terminus (the NN interface)18,19,20,23,25,26. Interactions across both of these interfaces are essential for SAS-6 oligomer formation; however, whereas the CC interface is relatively stable (Kd ~1 μM)18 and readily forms SAS-6 dimers in the cell cytoplasm27, the N-terminal dimer is significantly weaker (Kd ~50–100 μM in most systems)18,20, thereby presenting a challenge for the assembly of stable SAS-6 oligomers in cells28. SAS-6 is co-recruited to the site of centriole assembly and interacts with the protein SAS-5 in C. elegans25,26,29, while in insects and vertebrates binding to SAS-6 is similarly reported for the proteins Ana230,31 and STIL32,33, respectively. SAS-5, Ana2 and STIL self-associate into hexameric (SAS-5) 34,35 or tetrameric (Ana2/STIL)23,36 complexes, and these complexes have been suggested to assist SAS-6 oligomerisation in cells via an avidity mechanism, whereby multiple weak interactions act cooperatively23,34,35,37,38.
SAS-6 binding to Ana2/STIL depends on phosphorylation of these proteins by the Plk4 kinase, the vertebrate and insect analogue of ZYG-1. This dependence offers a putative mechanism for control of SAS-6 oligomerisation in insects and vertebrates via modulation of the Ana2/STIL–SAS-6 interaction affinity, and hence ‘fine tuning’ of the aforementioned avidity effect. In C. elegans, however, a model system for centriole cell biology, no such direct modulation of the SAS-5–SAS-6 interaction has been observed, although SAS-5 protein levels and targeting to the site of centriole assembly are reportedly controlled by the PP2A phosphatase39,40,41. Instead, earlier studies suggested that direct SAS-6 phosphorylation by the ZYG-1 kinase at a specific amino acid, S123, triggers centriole formation and ensures that SAS-6 is stably incorporated in the organelle42. Interestingly, S123 locates at the SAS-6 N-terminal domain, and its phosphorylation was proposed to affect the NN interface dimerisation affinity and, thus, the propensity of SAS-6 to oligomerise26. However, a later study convincingly demonstrated using S123 substitutions that phosphorylation of this SAS-6 residue is not required for C. elegans centriole formation43.
Nevertheless, the molecular logic of modulating the SAS-6 NN dimerisation affinity in order to control oligomer formation remains a strong one. Compared to the SAS-6 coiled-coil dimer, which spans hundreds of amino acids18, the N-terminal dimer principally depends on the interaction of a single amino acid, I154 in C. elegans, with a hydrophobic cavity across the NN dimerisation interface18,20. In this manner, it offers an attractive target for a relatively small, trigger-like molecular event to exert maximum influence on the oligomerisation propensity of SAS-6. Furthermore, we noted that a substantial segment of the C. elegans SAS-6 N-terminal domain, which includes S123, remained unresolved in all crystallographic structures of this domain to date. Thus, we set out to explore the effect of this C. elegans SAS-6 segment on the protein properties.
Here, we report that C. elegans SAS-6 features a ~30-amino acid flexible loop that does not have a counterpart in the algal, insect or vertebrate SAS-6 variants studied to date. The location and length of this loop allow it to transiently interact with multiple amino acids across the NN dimerisation interface, and these transient but frequent interactions cumulatively stabilise formation of SAS-6 oligomers. We note that many SAS-6 variants, including those from several species of human-infective parasites, feature similar, presumed flexible, insertions, and we discuss their possible role as elements controlling the trigger of centriole assembly.
Results
C. elegans SAS-6 features a long, flexible loop in its N-terminal domain
The C. elegans SAS-6 N-terminal domain (henceforth, CeSAS-6N) has been the subject of previous X-ray crystallographic studies that resolved the structures of the wild-type (WT) protein18 as well as derivatives25. In both cases an extended segment of this domain connecting α-helix 2 (α2) and β-strand 5 (β5), spanning amino acids 105–128 of CeSAS-6N, was absent from the structures as no electron density could be observed for the corresponding residues. The α2-β5-connecting segment was, thus, presumed disordered and, in the interest of efficient protein crystallisation, was removed from subsequent CeSAS-6 crystallographic efforts by excising residues I103 to P130 from the protein expression constructs. The resulting Δ103–130 variant of CeSAS-6N remained folded and showed only minimal structural changes compared to the WT protein as judged by X-ray crystallography (Cα RMSD of 0.3 Å)25; nuclear magnetic resonance (NMR) 15N heteronuclear single quantum coherence (HSQC) spectra of CeSAS-6N variants also showed a very high degree of similarity, suggesting limited long-range structural changes to this domain as a result of modifications (Supplemental Fig. 1).
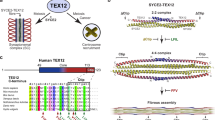
A similarly extended amino acid segment between α2 and β5 was not observed in the structures of Chlamydomonas reinhardtii (green algae)18, fruit fly23 or zebrafish20 SAS-6 N-terminal domains, and sequence alignments suggest it is similarly absent from the human and frog variants (Fig. 1). However, we noted that SAS-6 proteins from the Sar eukaryotic supergroup44, which includes several animal and plant pathogen species such as the malaria parasite Plasmodium falciparum, feature an extended segment connecting α2 and β5 that, in the case of P. falciparum, spans approximately 90 amino acids. Thus, this feature of SAS-6 is not restricted to nematode sequences but is likely also present in other branches of the eukaryotic kingdom.
A α2-β5 insertion is common in nematode and pathogen SAS-6 proteins. Shown here is a sequence alignment focused on the α2-β6 region of SAS-6 proteins. The secondary structure of CeSAS-6 is represented schematically at the top. The α2-β5 segment shown to be flexible in C. elegans (K101-T131) is highlighted in red. Double slash (‘//’) marks denote areas were 5 or more amino acids have been removed for clarity. Sequences were aligned manually using crystallographic (C. elegans, C. reinhardtii, D. melanogaster, D. rerio)18,20,23 or predicted SAS-6 structures as guides. Structure predictions were performed by Phyre268. Sequences derive from UniProt accession numbers C6KSS4 (P. falciparum), A0A0F7V199 (T. gondii), Q5CPW9 (C. parvum), A0A0G4ERZ9 (V. brassicaformis), M4B318 (H. arabidopsis), W2PGE7 (P. parasitica), D7FSC1 (E. siliculosus), A9CQL4 (C. reinhardtii), Q9VAC8 (D. melanogaster), Q7ZVT3 (D. rerio), Q6NRG6 (X. laevis), Q6UVJ0 (H. sapiens), O62479 (C. elegans), G0N6C0 (C. brenneri), E3NH99 (C. remanei), Q60P76 (C. briggsae) and A0A2H2I8W7 (C. japonica).
To characterise the structural state of the CeSAS-6 α2-β5-connecting segment we employed NMR, which can provide residue-specific information on amino acid properties in solution. In particular, the NMR chemical shifts of amino acid Cα and Cβ atoms are sensitive to the protein secondary structure, and display characteristic patterns of deviation from random coil chemical shift values depending on the type of secondary structure present in a given protein sequence45. We observed that the Cα and Cβ chemical shifts of CeSAS-6N amino acids showed patterns consistent with the secondary structure elements revealed by X-ray crystallography of this domain (Fig. 2A,B); however, chemical shift deviations from random coil were small at the α2-β5 segment, suggesting that this region of the protein lacks stable secondary structure elements.
The CeSAS-6 α2-β5 loop is flexible and unstructured. Shown here are per amino acid NMR chemical shift differences from random coil of CeSAS-6N Cα (top panel) and Cβ (middle panel) atoms, as well as {1H}-15N NOE ratios from the same amino acids (bottom panel). The CeSAS-6N secondary structure elements inferred from the crystallographic structure of this domain18 are represented schematically at the top. Black dashed lines (top and middle panels) denote thresholds over which the chemical shift differences are considered to support the presence of stable secondary structure elements45. A red dashed line (bottom panel) denotes a threshold below which {1H}-15N NOE ratios suggest that amino acids have substantial high-frequency (sub-ns timescale) motions46.
Furthermore, we analysed the CeSAS-6N amino acid mobility using heteronuclear {1H}-15N NOE NMR experiments, which are sensitive to motions in the picosecond to nanosecond time scale. Values of {1H}-15N NOE ratios over 0.6 are considered as indicative of structured protein segments, whereas NOE ratio values lower than that correspond to protein regions of increasing mobility46. As seen in Fig. 2C, {1H}-15N NOE ratios in CeSAS-6N support the rigid state of secondary structure elements observed by crystallography, whereas loops connecting secondary structure elements, such as between β1–β2, β2–β3 and β6–β7, are more mobile. Strikingly, the CeSAS-6N α2-β5 segment shows evidence of very high mobility, with {1H}-15N NOE ratios lower than 0.6 for a continuous span of amino acids between K101 and T131. We conclude that CeSAS-6N features a ~30 amino acid-long, flexible loop connecting α2 with β5, removal of which does not compromise the folded state of CeSAS-6N.
The α2-β5 loop is necessary for CeSAS-6 oligomerisation
SAS-6 oligomerisation is a defining property of this protein that is essential for canonical centriole assembly18,20,23; thus, we assessed the impact of the α2-β5 loop in the ability of CeSAS-6 to form oligomers. We performed analytical size exclusion chromatography (SEC) experiments using a CeSAS-6 construct that included both the N-terminal domain as well as a short stretch of the coiled coil (CeSAS-6N-CC). In previous studies WT and variants of CeSAS-6N-CC were observed to form stable dimers mediated by the CC interface, which then assembled into large oligomers in a concentration-dependent manner via the NN interaction25. Consistent with these previous results, SEC experiments showed increased apparent molecular size of CeSAS-6N-CC WT as function of protein concentration as judged by the reduction in elution volume from the SEC column (Fig. 3A). In contrast, a similar protein construct lacking the α2-β5 loop (CeSAS-6N-CC Δ103–130) showed no increase in apparent molecular size beyond the CC interface-mediated CeSAS-6N-CC dimer, even in concentrations as high as 10 mg/ml (Fig. 3B). We surmised that the α2-β5 loop contributes to the formation of large CeSAS-6 oligomers by strengthening the NN dimerisation of this protein.
The CeSAS-6 α2-β5 loop supports NN protein dimerisation. (A,B) Size-exclusion chromatography traces showing the elution profiles of CeSAS-6N-CC WT (A) or Δ103–130 (B) variants at different protein concentrations. WT CeSAS-6N-CC elutes at smaller retention volumes as protein concentration increases, indicating formation of larger oligomers. In contrast, CeSAS-6N-CC Δ103–130 does not form large oligomers under the same conditions. The apparent molecular masses of standard samples passed through the same size-exclusion column are shown as dashed lines. For reference, the calculated molecular weight of the CeSAS-6N-CC WT dimer is 48.8 kDa. (C) Fluorescence polarisation-monitored titrations of 1,5-IAEDANS-conjugated CeSAS-6N WT and variants as function of protein concentration. Points and error bars represent means and standard deviations, respectively, derived from three independent experiments. Solid lines denote fits of ideal self-association models to the data, with the estimated dissociation constants (Kd) shown. Raw fluorescence polarisation data were converted to fractions of CeSAS-6 NN dimers formed using the maximum polarisation change estimated from the fits.
To confirm this observation, we examined the dimerisation propensity of the CeSAS-6 N-terminal domain in isolation. Quantitative interaction assays using CeSAS-6N WT site-specifically labelled with 1,5-IAEDANS showed increased fluorescence polarisation as function of protein concentration, consistent with the formation of CeSAS-6N dimers mediated by the NN interface with a Kd of approximately 100 μM (Fig. 3C). In contrast, a CeSAS-6N I154E variant, which lacks the hydrophobic residue critical for NN dimerisation18, showed no increase in fluorescence polarisation in these assays. Similarly, CeSAS-6N Δ103–130 did not produce changes in fluorescence polarisation under the same conditions, suggesting that CeSAS-6 NN dimerisation affinity is greatly weakened in the absence of the α2-β5 loop. Thus, both SEC and fluorescence polarisation experiments independently support the role of loop α2-β5 in strengthening NN dimerisation and, hence CeSAS-6 oligomerisation.
Simulations and NMR reveal transient interactions formed by the α2-β5 loop
We proceeded to analyse how the α2-β5 loop strengthens CeSAS-6 NN dimerisation using atomistic molecular dynamics (MD) simulations. The NN-mediated dimer is well defined in the CeSAS-6N crystallographic structures; however, these structures did not resolve the α2-β5 loop and, thus, cannot provide starting positions for the loop amino acids for computational simulations. For that reason, we constructed models of CeSAS-6N dimers where the α2-β5 loop residues were placed in energetically favourable but variable arrangements. We derived three different models of CeSAS-6N dimers with distinct α2-β5 loop conformations for each monomeric subunit, and performed nine, 50 ns-long MD simulations (three simulations starting from each CeSAS-6N dimer model) to explore the available structural landscape. The starting and end points of one MD simulation for each CeSAS-6N dimer are shown in Fig. 4A. We observed that in all cases the α2-β5 loop of CeSAS-6N rearranges to form interactions with helices α1-α2 of the opposing CeSAS-6N monomer. Notably, these interactions were not stable; rather the α2-β5 loops continuously repositioned over α1-α2 during the course of simulations, breaking and reforming interactions with several residues therein. The α2-β5 loop conformations did not stabilise even when simulations were extended to 100 ns length, suggesting that loop mobility observed in simulations reflects the flexibility of this protein segment shown by NMR experiments.
The CeSAS-6 α2-β5 loop forms transient contacts across the NN dimerisation interface. (A) Snapshots of three structural models of CeSAS-6N that include the α2-β5 loop in different starting conformations, at the beginning (0 ns) and end (50 ns) of representative atomistic MD simulations. The α2-β5 loop is coloured red at the starting and blue at the end point of simulations; the α1-α2 helices are shown in gold. Note that in all cases the α2-β5 loop forms extensive contacts with the structured core of CeSAS-6N during the MD simulations. (B,C) Quantitative analysis of contacts between α1-α2 residues and the α2-β5 loop (B), and vice versa (C), in nine MD simulations (three simulations per starting CeSAS-6N dimer model). Contacts are expressed as fraction of simulation time during which residues are in close proximity (distance <3.5 Å) to α1-α2 helices (C) or the α2-β5 loop (B). The amino acid sequences are shown, as is the position (B) of α1-α2 helices within the sequence. (D–J) Sections of NMR 1H-15N HSQC spectra of (D) the monomeric CeSAS-6N I154E variant at 500 μM protein concentration, (E) CeSAS-6N WT at 25 (light blue) or 500 μM (purple) protein concentration, (F) CeSAS-6N S123E at 500 μM protein concentration and (G) the CeSAS-6N Δ103–130 variant at 500 μM protein concentration. (H–J) NMR spectra of CeSAS-6N at 500 μM concentration and different amounts of NaCl as shown. The resonances of G111, G120, G128 and G157 amino acids are indicated. Note that all four glycine resonances are strong in the monomeric CeSAS-6N I154E variant (D) as is also the case for G157 in the CeSAS-6N Δ103–130 variant (F). In contrast, in CeSAS-6N WT or S123E the G120 and G157 resonances disappear as function of protein concentration, indicating the formation of μs-ms time scale contacts by these residues. At high ionic strength conditions the G120 resonance increases in intensity, suggesting that loop α2-β5 forms fewer contacts. (K) Fractional intensities of the G120 and G157 resonances as function of NaCl concentration. Intensities were normalised to those of the G111 and G128 resonances in the same spectra. Error bars derive from the spectral signal-to-noise ratios. The G120 resonance intensity is fit to a linear regression model with the indicated R2.
To quantitatively compare the α2-β5 loop conformations across different simulations we evaluated the length of time during which loop amino acids are in close proximity (<3.5 Å distance) to residues of the α1-α2 region as proportion of the total MD simulation time. As shown in Fig. 4B, in most simulations one or more α2-β5 loop amino acids contact the C-terminus of helix α1 and the α1-α2 linker for over 50% of MD time; residues of helix α2 are also contacted in a minority of cases. Similar analysis showed that the α2-β5 loop region primarily involved in α1-α2 contacts spans amino acids R116-A125, which locate approximately at the middle of the α2-β5 loop (Fig. 4C). We note that the majority of contact residues in both the α1-α2 region and the α2-β5 loop are hydrophilic in nature, including a large number of charged amino acids; indeed, these residues primarily form hydrogen bond and electrostatic interactions in the MD simulations.
Analysis of 1H-15N HSQC NMR spectra supports the formation of transient contacts by residues at the middle of the α2-β5 loop. Specifically, we used the glycine amino acids of the α2-β5 loop, which are easily distinguished in NMR spectra (Fig. 4D–G), as probes to quickly ascertain the structural state of loop residues. In the monomeric CeSAS-6N I154E protein variant (Fig. 4D) these glycine residues (G111, G120 and G128) always give rise to strong resonances, as is also the case for WT CeSAS-6N at low concentrations (25 μM) when the protein is mostly monomeric (Fig. 4E, light blue). In contrast, at high (500 μM) protein concentrations, when WT CeSAS-6N forms NN-mediated dimers, the NMR resonance of G120 nearly disappears (Fig. 4E, purple), indicating that this residue at the middle of the α2-β5 loop is involved in μs-ms timescale interactions. A similar effect is seen for the resonance of G157 at high CeSAS-6N concentrations, as this amino acid is located directly at the CeSAS-6 NN dimerisation interface. In contrast, we observed no perturbation of the G111 or G128 resonances regardless of protein concentration. To assess whether the transient contacts of the α2-β5 loop are electrostatic in nature we titrated NaCl to samples of WT CeSAS-6N at high protein concentration (500 μM). Under these conditions, the G120 increases in intensity as function of ionic strength (Fig. 4H–K), suggesting that the α2-β5 loop forms fewer μs-ms timescale interactions. This is consistent with electrostatic contacts of the α2-β5 loop being masked by increased NaCl amounts. In contrast, the G157 resonance intensity is virtually unchanged upon NaCl titration, suggesting that CeSAS-6N remains dimeric. We conclude that formation of the CeSAS-6 NN dimer causes residues at the middle of the α2-β5 loop, including G120, to engage in intermediate timescale electrostatic interactions, as suggested by the MD simulations.
Interactions of the α2-β5 loop stabilise the CeSAS-6 NN dimer
We proceeded to examine using steered MD simulations whether the transient, interchangeable interactions formed between the α2-β5 loop and the structured core of CeSAS-6N may cumulatively stabilise formation of the CeSAS-6 NN dimer. Pulling forces in opposing directions were applied in silico on the monomeric subunits of the CeSAS-6 NN dimer, and the work required to pull the dimer apart was measured during the course of 14 independent simulations for each of CeSAS-6N WT and Δ103–130 variants. We observed that the work necessary for disruption of CeSAS-6 NN dimers varied substantially across different simulations, reflecting the non-equilibrium nature of these experiments; however, in all cases CeSAS-6N Δ103–130 dimers were pulled apart faster and with greater ease compared to CeSAS-6N WT dimers (Fig. 5). A representative example of this is shown in Fig. 5A, where a dimer of CeSAS-6N Δ103–130 has lost all amino acid contacts across the NN interface after 15 ns of simulation time, whereas a dimer of CeSAS-6N WT maintains contacts at the same time point partly through the α2-β5 loop. We surmise that contacts between the α2-β5 loop and the structured core of CeSAS-6N can indeed stabilise the NN dimerisation interface in MD simulations.
The α2-β5 loop stabilises CeSAS-6N dimerisation in steered MD simulations. (A) Snapshots of CeSAS-6N WT and Δ103–130 dimers at different time points during representative steered MD simulations. The α2-β5 loop is coloured red and amino acid I154, which directly mediates CeSAS-6 NN dimerisation, is shown as sticks. The forces applied on the CeSAS-6N monomeric subunits are represented by black arrows as reference. (B) Graphic representation of work applied to separate the CeSAS-6N monomeric subunits versus distance pulled in steered MD simulations. Shown here are 14 independent simulations performed on CeSAS-6N WT (red graphs) and the same number of simulations for CeSAS-6N Δ103–130 (blue graphs).
To validate the effect of interactions formed by the α2-β5 loop on CeSAS-6 NN dimerisation, we sought to examine whether changes within the loop modulate the NN dimerisation affinity. We chose to modify S123, which locates at the middle section of this loop and forms transient interactions in MD simulations (Fig. 4C). Although S123 phosphorylation has been shown not to have an effect on C. elegans centriole assembly43, we reasoned that changes at this site might provide an informative in vitro tool. Thus, we analysed the effect of a S123E substitution on CeSAS-6N dimerisation using quantitative fluorescence polarisation experiments. Although this substitution does not induce local structural rearrangements or stabilisation of the α2-β5 loop, as judged by heteronuclear {1H}-15N NOE NMR experiments (Supplemental Fig. 3), we observed that S123E increases NN affinity by approximately 2-fold (Fig. 3C). It is likely that this small but notable change in CeSAS-6 NN affinity upon substituting S123 may have contributed to the lack of clarity on the functional role of this residue in the literature. However, for the purpose of our analysis, this substitution provides evidence that changes in the α2-β5 loop can indeed modulate CeSAS-6 properties.
Discussion
SAS-6 oligomerisation is a key property for the function of this protein, not least as it assists the establishment of centriolar 9-fold radial symmetry17,24. The weakest molecular ‘link’ in SAS-6 oligomerisation is dimerisation of this protein’s N-terminal domain via the NN interface18,19,20,23,25. With this in mind, we examined the properties of a previously unresolved sequence element in the N-terminal domain of C. elegans SAS-6. We found that this element, which spans approximately 30 amino acids and connects α2 and β5 of the CeSAS-6 N-terminal domain, is unstructured and highly dynamic in solution (Fig. 2). The flexible nature of this α2-β5 loop is consistent with the lack of electron density for this region in crystallographic structures of the N-terminal domain. Nevertheless, despite the apparent lack of structure, we noted that the α2-β5 loop has a stabilising role in CeSAS-6 NN dimerisation, to the extent that removing this loop abrogated formation of CeSAS-6 oligomers in vitro (Fig. 3). The α2-β5 loop enhances CeSAS-6 N-terminal domain dimerisation by forming transient interactions, evident by both NMR and computation (Fig. 4), with the structured core of this domain. Crucially, substituting a single amino acid in the α2-β5 loop further enhanced CeSAS-6 NN dimerisation by approximately 2-fold (Fig. 3C).
The concept of disordered protein segments engaging in, and being important for, protein interactions is widely accepted47, and such disordered segments are believed to confer enhanced interaction specificity as well as plasticity. In most cases, disordered segments fold into stable structures upon binding their physiological partner. However, in a subset of protein interactions disordered segments remain unfolded, which has given rise to the notion of ‘fuzzy complexes’ during the last decade48,49. Such complexes comprise conformational ensembles even in their functional state, with the relative populations of discrete states within these ensembles subject to change according to the cellular context in order to fine-tune activity. Our work strongly suggests that the CeSAS-6 N-terminal domain forms a fuzzy complex, at least in part, as the α2-β5 loop remains disordered even at the physiologically relevant dimeric state of this domain.
A long α2-β5 loop with high levels of sequence conservation is found in many species of nematode worms, such as throughout the Caenorhabditis genus (Fig. 1), despite these species diverging over 30 million years ago50. Although our analysis shows that this loop serves to stabilise the CeSAS-6 NN dimer, it is clear from the vertebrate, insect and algal SAS-6 variants lacking this loop that such stabilisation could be achieved more simply by a handful of amino acid changes, not least by replacing I154 with an aromatic amino acid24,25. If correct, this raises the question of what is the true purpose of the long α2-β5 loop so that it is maintained across millions of years. We can only speculate the answer to this question; however, a likely clue is offered by the observation that amino acid changes within the α2-β5 loop directly affect the CeSAS-6 NN dimerisation affinity (Fig. 3C). Although phosphorylation of the specific amino acid substituted in our study, S123, is not physiologically relevant43, our work demonstrated the principle, common to fuzzy complexes, that even small changes in the disordered segment can affect complex formation. Thus, we propose that the physiological role of the SAS-6 α2-β5 loop in nematode species may be as a modulator of NN dimerisation, and consequently as a molecular control mechanism for regulating SAS-6 oligomerisation. It should be noted that in a multimeric system such as SAS-6 oligomers, where a complex of at least 9 protein dimers is necessary to define centriolar 9-fold symmetry, even small changes in self-association affinity can exert a powerful effect. Indeed, simple simulations suggest that increasing the CeSAS-6 NN dimerisation affinity from 100 μM to 50 μM Kd leads to a ~150-fold increase in the likelihood 9 CeSAS-6 dimers associate into an oligomer, and, hence, in the probability that a core structural element of centrioles forms.
Sequence analysis suggests that a large α2-β5 loop is not restricted to nematode SAS-6 proteins, but also found throughout the Sar eukaryotic supergroup (Fig. 1). Of particular interest there are apicomplexan parasites, including Plasmodium, Cryptosporidium and Toxoplasma, that are responsible for widespread and severe human diseases. It will be interesting to examine whether the α2-β5 loop in apicomplexan SAS-6 acts in a similar capacity as in nematode SAS-6 to modulate NN dimerisation. If so, such a behaviour would represent a distinct departure from vertebrate SAS-6 oligomerisation and, thus, may be a mechanism open to exploitation by putative therapeutic agents. Although SAS-6 has only just begun to be studied in these parasites, we note that Plasmodium SAS-6 appears essential for malaria transmission51.
In conclusion, we report here that a previously uncharacterised loop in the CeSAS-6 N-terminal domain reinforces the self-association interactions of this protein, and that changes in this loop can modulate the formation of large SAS-6 oligomers. As formation of such SAS-6 oligomers is an essential step for the initiation of centriole formation, we postulate that changes in the α2-β5 loop, putatively through yet uncharacterised amino acid modifications, may act as molecular switches that assist in triggering centriole assembly.
Materials and Methods
Protein production and purification
C. elegans SAS-6 (Uniprot ID 062479) fragments were prepared as described earlier18,25; briefly, fragments comprising the protein N-terminal domain (CeSAS-6N, amino acids 1–168) or the N-terminal domain plus a short stretch of the coiled-coil interface (CeSAS-6N-CC, amino acids 1–215) were cloned in a modified pET15b vector containing an N-terminal His6-tag, transformed into Escherichia coli BL21 (DE3) cells grown in Luria-Bertani (LB) media, and protein expression was induced for 16 h with 0.25 mM final concentration of isopropylb-D-1-thiogalactopyranoside at 18 °C. Cell pellets were resuspended in lysis buffer containing 20 mM Tris HCl buffer pH 7.5, 500 mM NaCl, 0.5% v/v Triton X-100 and Complete protease inhibitor tablets (Roche), and sonicated for cell lysis. Metal affinity purification of clarified lysates was performed using His-Trap HP columns (GE LifeSciences), followed by His6-tag cleavage using thrombin protease (Sigma-Aldrich) and size exclusion chromatography on Sephadex G75 columns (GE LifeSciences) equilibrated in PBS (20 mM sodium phosphate buffer pH 7.0, 150 mM NaCl and 2 mM DTT). For the production of isotopically labelled protein samples E. coli cells were grown in M9 minimal media supplemented with 15NH4Cl and 13C6-glucose (Isotech) as necessary.
NMR experiments
Sequence-specific NMR resonance assignments were performed as described previously52. Briefly, NMR experiments were performed using Bruker Avance II and Avance III spectrometers with cryogenic TCI probeheads, and 11.7–14.1 T magnetic field strengths. Samples of 13C/15N-enriched CeSAS-6N S123E I154E variant at 1 mM concentration in PBS buffer were supplemented with 5% v/v D2O, 0.02% w/v NaN3 and 50 μM 4,4-dimethyl-4-silapentane-1-sulfonic acid. Assignment experiments were performed at 20 °C using 3D CBCA(CO)NH, CBCANH and HNCA pulse sequences. NMR data were processed using NMRpipe53 and analysed using PIPP54. Assignments were deposited in BioMagResBank under accession number 27607. Chemical shift assignments were transferred to CeSAS-6N WT by overlaying spectra. Spectra overlays were prepared with Sparky55. Comparisons of 13Cα and 13Cβ chemical shifts to those of random coil were performed using the Chemical Shift Index method56. Heteronuclear {1H}-15N NOE experiments were performed in a manner analogous to that described previously57.
Fluorescence polarisation and size exclusion chromatography
Protein samples for fluorescence polarisation were disolved in PBS and featured 1,5-IAEDANS (Invitrogen) fluorescence labels conjugated to C100 of CeSAS-6N using the manusfacturer’s recommended protocol. Measurements were recorded using a PHERASTAR FS fluorimeter (BMG Labtech, λex = 340 nm, λem = 520 nm). Analytical size exclusion chromatography assays were performed using protein samples in PBS and Superdex 75 10/300 GL columns (GE LifeSciences).
Molecular modelling and all-atom simulations
A complete structure of the CeSAS-6N domain, including residues 103–130 of the α2-β5 loop, was built using Modeller58 starting from the crystallographic structure of CeSAS-6N Δ103–130 (RCSB ID 4G79)25. 100 models were created, and models for MD simulations were selected visually preferring those structures that minimised clashes in the α2-β5 loop while also lacking secondary structure elements there. Protein models were placed in a 100 × 100 × 100 Å boxes with periodic boundary conditions, and MD simulations were initiated using the all-atom force field AMBER99SB-ILDN59 with explicit TIP3P60 water molecules and an ionic concentration of 150 mM NaCl. The model was energy minimised using the steepest descent method with a target energy of 100 kJ/(mol nm). For NVT equilibration 200 ps of MD simulations were run with constant temperature at 300 K using a Berendsen thermostat61, while applying position restraints for protein heavy atoms. NPT equilibration was achieved by 200 ps of MD simulations in constant pressure of 1 bar using a Berendsen barostat61. Position restraints on heavy atoms were removed for production runs of 50 ns, which were started from the same equilibrated starting point but using different seed parameters. All trajectories were generated and analysed with GROMACS v5.0262. The distance cut-off for van der Waals and short-range electrostatic interactions was set to 10 Å. Long-range electrostatics were accounted for using the particle mesh Ewald method63,64 and the LINCS65 algorithm was selected to treat all bonds as constraints, allowing a time step of 2 fs. Residue encounters were calculated using a tcl/tk script and VMD66 with a distance cut-off of 3.5 Å.
For steered MD simulations the collective variable (CV) was the distance between the centres of mass of each CeSAS-6N domain monomer. The centres of mass were calculated using the Cα atoms of residues 1–102 and 131–168, thereby excluding residues of the flexible loop. During steered MD simulations the CV was steered towards a distance of 20 Å, which was judged as sufficient to separate the CeSAS-6 NN dimer. Constant velocity of 1 Å/ns and a force constant of 1000 kJ/mol/nm were used. Steered MD simulations were setup and analysed using PLUMED v2.267 and GROMACS v5.0262.
References
Hodges, M. E., Scheumann, N., Wickstead, B., Langdale, J. A. & Gull, K. Reconstructing the evolutionary history of the centriole from protein components. J Cell Sci 123, 1407–1413, https://doi.org/10.1242/jcs.064873 (2010).
Marshall, W. F. Centriole evolution. Curr Opin Cell Biol 21, 14–19, https://doi.org/10.1016/j.ceb.2009.01.008 (2009).
Carvalho-Santos, Z., Azimzadeh, J., Pereira-Leal, J. B. & Bettencourt-Dias, M. Evolution: Tracing the origins of centrioles, cilia, and flagella. J Cell Biol 194, 165–175, https://doi.org/10.1083/jcb.201011152 (2011).
Conduit, P. T., Wainman, A. & Raff, J. W. Centrosome function and assembly in animal cells. Nat Rev Mol Cell Biol 16, 611–624, https://doi.org/10.1038/nrm4062 (2015).
Bornens, M. The centrosome in cells and organisms. Science 335, 422–426, https://doi.org/10.1126/science.1209037 (2012).
Marshall, W. F. Basal bodies platforms for building cilia. Curr Top Dev Biol 85, 1–22, https://doi.org/10.1016/S0070-2153(08)00801-6 (2008).
Bettencourt-Dias, M., Hildebrandt, F., Pellman, D., Woods, G. & Godinho, S. A. Centrosomes and cilia in human disease. Trends Genet 27, 307–315, https://doi.org/10.1016/j.tig.2011.05.004 (2011).
Venghateri, J. B., Jindal, B. & Panda, D. The centrosome: a prospective entrant in cancer therapy. Expert opinion on therapeutic targets 19, 957–972, https://doi.org/10.1517/14728222.2015.1018823 (2015).
Gönczy, P. Centrosomes and cancer: revisiting a long-standing relationship. Nat Rev Cancer 15, 639–652, https://doi.org/10.1038/nrc3995 (2015).
Chavali, P. L., Putz, M. & Gergely, F. Small organelle, big responsibility: the role of centrosomes in development and disease. Philos Trans R Soc Lond B Biol Sci 369, https://doi.org/10.1098/rstb.2013.0468 (2014).
Thornton, G. K. & Woods, C. G. Primary microcephaly: do all roads lead to Rome? Trends Genet 25, 501–510, https://doi.org/10.1016/j.tig.2009.09.011 (2009).
Azimzadeh, J. & Marshall, W. F. Building the centriole. Curr Biol 20, R816–825, https://doi.org/10.1016/j.cub.2010.08.010 (2010).
Gönczy, P. Towards a molecular architecture of centriole assembly. Nat Rev Mol Cell Biol 13, 425–435, https://doi.org/10.1038/nrm3373 (2012).
Jana, S. C., Marteil, G. & Bettencourt-Dias, M. Mapping molecules to structure: unveiling secrets of centriole and cilia assembly with near-atomic resolution. Curr Opin Cell Biol 26, 96–106, https://doi.org/10.1016/j.ceb.2013.12.001 (2014).
Firat-Karalar, E. N. & Stearns, T. The centriole duplication cycle. Philos Trans R Soc Lond B Biol Sci 369, https://doi.org/10.1098/rstb.2013.0460 (2014).
Sugioka, K. et al. Centriolar SAS-7 acts upstream of SPD-2 to regulate centriole assembly and pericentriolar material formation. Elife 6, https://doi.org/10.7554/eLife.20353 (2017).
Nakazawa, Y., Hiraki, M., Kamiya, R. & Hirono, M. SAS-6 is a cartwheel protein that establishes the 9-fold symmetry of the centriole. Curr Biol 17, 2169–2174, https://doi.org/10.1016/j.cub.2007.11.046 (2007).
Kitagawa, D. et al. Structural basis of the 9-fold symmetry of centrioles. Cell 144, 364–375, https://doi.org/10.1016/j.cell.2011.01.008 (2011).
van Breugel, M., Wilcken, R., McLaughlin, S. H., Rutherford, T. J. & Johnson, C. M. Structure of the SAS-6 cartwheel hub from Leishmania major. Elife 3, e01812 (2014).
van Breugel, M. et al. Structures of SAS-6 suggest its organization in centrioles. Science 331, 1196–1199, https://doi.org/10.1126/science.1199325 (2011).
Guichard, P. et al. Cell-free reconstitution reveals centriole cartwheel assembly mechanisms. Nat Commun 8, 14813, https://doi.org/10.1038/ncomms14813 (2017).
Guichard, P. et al. Cartwheel architecture of Trichonympha basal body. Science 337, 553, https://doi.org/10.1126/science.1222789 (2012).
Cottee, M. A. et al. The homo-oligomerisation of both Sas-6 and Ana2 is required for efficient centriole assembly in flies. Elife 4, e07236, https://doi.org/10.7554/eLife.07236 (2015).
Hilbert, M. et al. SAS-6 engineering reveals interdependence between cartwheel and microtubules in determining centriole architecture. Nat Cell Biol 18, 393–403, https://doi.org/10.1038/ncb3329 (2016).
Hilbert, M. et al. Caenorhabditis elegans centriolar protein SAS-6 forms a spiral that is consistent with imparting a ninefold symmetry. Proc Natl Acad Sci USA 110, 11373–11378, https://doi.org/10.1073/pnas.1302721110 (2013).
Qiao, R., Cabral, G., Lettman, M. M., Dammermann, A. & Dong, G. SAS-6 coiled-coil structure and interaction with SAS-5 suggest a regulatory mechanism in C. elegans centriole assembly. EMBO J 31, 4334–4347, https://doi.org/10.1038/emboj.2012.280 (2012).
Keller, D. et al. Mechanisms of HsSAS-6 assembly promoting centriole formation in human cells. J Cell Biol 204, 697–712, https://doi.org/10.1083/jcb.201307049 (2014).
Klein, H. C., Guichard, P., Hamel, V., Gonczy, P. & Schwarz, U. S. Computational support for a scaffolding mechanism of centriole assembly. Sci Rep 6, 27075, https://doi.org/10.1038/srep27075 (2016).
Leidel, S., Delattre, M., Cerutti, L., Baumer, K. & Gönczy, P. SAS-6 defines a protein family required for centrosome duplication in C. elegans and in human cells. Nat Cell Biol 7, 115–125, https://doi.org/10.1038/ncb1220 (2005).
Dzhindzhev, N. S. et al. Two-step phosphorylation of Ana2 by Plk4 is required for the sequential loading of Ana2 and Sas6 to initiate procentriole formation. Open Biol 7, https://doi.org/10.1098/rsob.170247 (2017).
Dzhindzhev, N. S. et al. Plk4 phosphorylates Ana2 to trigger Sas6 recruitment and procentriole formation. Curr Biol 24, 2526–2532, https://doi.org/10.1016/j.cub.2014.08.061 (2014).
Moyer, T. C., Clutario, K. M., Lambrus, B. G., Daggubati, V. & Holland, A. J. Binding of STIL to Plk4 activates kinase activity to promote centriole assembly. J Cell Biol 209, 863–878, https://doi.org/10.1083/jcb.201502088 (2015).
Ohta, M. et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat Commun 5, 5267, https://doi.org/10.1038/ncomms6267 (2014).
Rogala, K. B. et al. The Caenorhabditis elegans protein SAS-5 forms large oligomeric assemblies critical for centriole formation. Elife 4, e07410, https://doi.org/10.7554/eLife.07410 (2015).
Shimanovskaya, E., Qiao, R., Lesigang, J. & Dong, G. The SAS-5 N-terminal domain is a tetramer, with implications for centriole assembly in C. elegans. Worm 2, e25214, https://doi.org/10.4161/worm.25214 (2013).
David, A. et al. Molecular basis of the STIL coiled coil oligomerization explains its requirement for de-novo formation of centrosomes in mammalian cells. Sci Rep 6, 24296, https://doi.org/10.1038/srep24296 (2016).
Slevin, L. K., Romes, E. M., Dandulakis, M. G. & Slep, K. C. The mechanism of dynein light chain LC8-mediated oligomerization of the Ana2 centriole duplication factor. J Biol Chem 289, 20727–20739, https://doi.org/10.1074/jbc.M114.576041 (2014).
Bianchi, S. et al. Interaction between the Caenorhabditis elegans centriolar protein SAS-5 and microtubules facilitates organelle assembly. Mol Biol Cell 29, 722–735, https://doi.org/10.1091/mbc.E17-06-0412 (2018).
Megraw, T. PP2A targets SAS-5 in centriole assembly. Dev Cell 20, 416–417, https://doi.org/10.1016/j.devcel.2011.03.021 (2011).
Kitagawa, D. et al. PP2A phosphatase acts upon SAS-5 to ensure centriole formation in C. elegans embryos. Dev Cell 20, 550–562, https://doi.org/10.1016/j.devcel.2011.02.005 (2011).
Song, M. H., Liu, Y., Anderson, D. E., Jahng, W. J. & O’Connell, K. F. Protein phosphatase 2A-SUR-6/B55 regulates centriole duplication in C. elegans by controlling the levels of centriole assembly factors. Dev Cell 20, 563–571, https://doi.org/10.1016/j.devcel.2011.03.007 (2011).
Kitagawa, D., Busso, C., Flückiger, I. & Gönczy, P. Phosphorylation of SAS-6 by ZYG-1 is critical for centriole formation in C. elegans embryos. Dev Cell 17, 900–907, https://doi.org/10.1016/j.devcel.2009.11.002 (2009).
Lettman, M. M. et al. Direct Binding of SAS-6 to ZYG-1 Recruits SAS-6 to the Mother Centriole for Cartwheel Assembly. Dev Cell 25, 284–298, https://doi.org/10.1016/j.devcel.2013.03.011 (2013).
Adl, S. M. et al. The revised classification of eukaryotes. J Eukaryot Microbiol 59, 429–493, https://doi.org/10.1111/j.1550-7408.2012.00644.x (2012).
Wishart, D. S., Sykes, B. D. & Richards, F. M. The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31, 1647–1651 (1992).
Kay, L. E., Torchia, D. A. & Bax, A. Backbone dynamics of proteins as studied by 15N inverse detected heteronuclear NMR spectroscopy: Application to staphylococcal nuclease. Biochemistry 28, 8972–8979 (1989).
Berlow, R. B., Dyson, H. J. & Wright, P. E. Functional advantages of dynamic protein disorder. FEBS Lett 589, 2433–2440, https://doi.org/10.1016/j.febslet.2015.06.003 (2015).
Fuxreiter, M. Fuzziness in Protein Interactions-A Historical Perspective. J Mol Biol 430, 2278–2287, https://doi.org/10.1016/j.jmb.2018.02.015 (2018).
Arbesu, M., Iruela, G., Fuentes, H., Teixeira, J. M. C. & Pons, M. Intramolecular Fuzzy Interactions Involving Intrinsically Disordered Domains. Front Mol Biosci 5, 39, https://doi.org/10.3389/fmolb.2018.00039 (2018).
Cutter, A. D. Divergence times in Caenorhabditis and Drosophila inferred from direct estimates of the neutral mutation rate. Mol Biol Evol 25, 778–786, https://doi.org/10.1093/molbev/msn024 (2008).
Marques, S. R. et al. An essential role of the basal body protein SAS-6 in Plasmodium male gamete development and malaria transmission. Cell Microbiol 17, 191–206, https://doi.org/10.1111/cmi.12355 (2015).
Mayer, C., Slater, L., Erat, M. C., Konrat, R. & Vakonakis, I. Structural analysis of the Plasmodium falciparum erythrocyte membrane protein 1 (PfEMP1) intracellular domain reveals a conserved interaction epitope. J Biol Chem 287, 7182–7189, https://doi.org/10.1074/jbc.M111.330779 (2012).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6, 277–293 (1995).
Garrett, D. S., Powers, R., Gronenborn, A. M. & Clore, G. M. A common sense approach to peak peaking in two-, three-, and four-dimensional spectra using automatic computer analysis of contour diagrams. J. Magn. Res. 95, 214–220 (1991).
Goddard, T. D. & Kneller, D. G. SPARKY3, University of California, San Francisco.
Wishart, D. S. & Sykes, B. D. The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J. Biomol. NMR 4, 171–180 (1994).
Vakonakis, I., Langenhan, T., Promel, S., Russ, A. & Campbell, I. D. Solution structure and sugar-binding mechanism of mouse latrophilin-1 RBL: a 7TM receptor-attached lectin-like domain. Structure 16, 944–953, https://doi.org/10.1016/j.str.2008.02.020 (2008).
Webb, B. & Sali, A. Comparative Protein Structure Modeling Using MODELLER. Curr Protoc Bioinformatics 47, 5 6 1–32, https://doi.org/10.1002/0471250953.bi0506s47 (2014).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958, https://doi.org/10.1002/prot.22711 (2010).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of Simple Potential Functions for Simulating Liquid Water. Journal of Chemical Physics 79, 926–935, https://doi.org/10.1063/1.445869 (1983).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular-Dynamics with Coupling to an External Bath. Journal of Chemical Physics 81, 3684–3690, https://doi.org/10.1063/1.448118 (1984).
GROMACS User Manual version 5.1, www.gromacs.org (2015).
Darden, T., York, D. & Pedersen, L. Particle Mesh Ewald - an N.Log(N) Method for Ewald Sums in Large Systems. Journal of Chemical Physics 98, 10089–10092, https://doi.org/10.1063/1.464397 (1993).
Essmann, U. et al. A Smooth Particle Mesh Ewald Method. Journal of Chemical Physics 103, 8577–8593, https://doi.org/10.1063/1.470117 (1995).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. Journal of Computational Chemistry 18, 1463–1472, https://doi.org/10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H (1997).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J Mol Graph 14(33–38), 27–38, https://doi.org/10.1016/0263-7855(96)00018-5 (1996).
Tribello, G. A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED 2: New feathers for an old bird. Comput Phys Commun 185, 604–613, https://doi.org/10.1016/j.cpc.2013.09.018 (2014).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10, 845–858, https://doi.org/10.1038/nprot.2015.053 (2015).
Acknowledgements
We are grateful to Nick Soffe for maintenance of the solution NMR facility, as well as to Leanne Slatter for assistance with protein production. We thank the Biotechnology and Biological Sciences Research Council UK (BB/J008265/1) and the Medical Research Council UK (MR/N009274/1) for their support. J.B. was funded by a PhD studentship from the Biotechnology and Biological Sciences Research Council UK to the Oxford Interdisciplionary Bioscience Doctoral Training Partnership. The Oxford Biochemistry NMR facility was supported by the Wellcome Trust (094872/Z/10/Z), the Wellcome Institutional Strategic Support Fund, the EPA Cephalosporin Trust and the John Fell OUP Research Fund.
Author information
Authors and Affiliations
Contributions
J.B. conceived, performed and analysed experiments. I.B. performed and analysed experiments. M.C.E. and I.V. conceived and analysed experiments. P.C.B. and M.M. directed the simulations. J.B. and I.V. wrote the manuscript and prepared figures.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Busch, J.M.C., Erat, M.C., Blank, I.D. et al. A dynamically interacting flexible loop assists oligomerisation of the Caenorhabditis elegans centriolar protein SAS-6. Sci Rep 9, 3526 (2019). https://doi.org/10.1038/s41598-019-40294-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-40294-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.