Abstract
Terrestrial ecosystems in the maritime Antarctic experienced rapid warming during the latter half of the 20th century. While warming ceased at the turn of the millennium, significant increases in air temperature are expected later this century, with predicted positive effects on soil fungal diversity, plant growth and ecosystem productivity. Here, by sequencing 16S ribosomal RNA genes in 40 soils sampled from along a 1,650 km climatic gradient through the maritime Antarctic, we determine whether rising air temperatures might similarly influence the diversity of soil bacteria. Of 22 environmental factors, mean annual surface air temperature was the strongest and most consistent predictor of soil bacterial diversity. Significant, but weaker, associations between bacterial diversity and soil moisture content, C:N ratio, and Ca, Mg, PO43− and dissolved organic C concentrations were also detected. These findings indicate that further rises in air temperature in the maritime Antarctic may enhance terrestrial ecosystem productivity through positive effects on soil bacterial diversity.
Similar content being viewed by others
Introduction
During the latter half of the 20th century, air temperatures in the maritime Antarctic rose more rapidly than in any other region of the Southern Hemisphere (c. 0.2 °C increases in mean near surface annual temperature per decade)1 and then stabilised before the turn of the millennium2. Warming in the region led to the collapse of ice shelves, the retreat of glaciers, and increases in the frequencies of precipitation1,3, the growth of bryophytes4, the occurrence of invasive species5 and the ranges of native plants6,7. Much less is known, however, about how warming and its associated environmental changes might influence the diversity of soil micro-organisms (i.e., bacteria, archaea and microeukarya) across the region. This knowledge gap is significant, as changes to microbial diversity may affect terrestrial ecosystem functioning due to the important roles of many taxa as autotrophs, saprotrophs, pathogens and symbionts. In addition, climate models, assuming only moderate anthropogenic greenhouse gas emissions, predict that in the latter half of this century the maritime Antarctic will warm at similar rates to those observed between the 1950s and late 1990s8.
Soil fungal diversity in the region is positively associated with air temperature, with warming-induced changes in fungal community composition being predicted to enhance ecosystem productivity through effects on nutrient cycling9. At present, however, little is known of how further climate warming will influence the diversity of bacteria in maritime Antarctic soils. Current knowledge about the regional-scale factors influencing soil bacterial diversity in maritime Antarctica derives predominantly from a survey conducted between 2003 and 200510,11,12,13,14,15. Sanger sequencing, microarray analyses and denaturing gradient gel electrophoresis of bacterial 16S rRNA genes in soils from this survey indicate that alpha diversity decreases, and community composition changes, with increasing latitude10,11,15. However, as soil was sampled from five to eight locations between the Falkland Islands and the Ellsworth Mountains in the continental Antarctic, with temperature being recorded at three sites10,11,12,13,14,15, the environmental drivers of this spatial pattern in bacterial diversity remain unclear.
Here, we test the hypothesis that the diversity of maritime Antarctic soil bacteria is significantly associated with reductions in temperature, liquid water and nutrient availability at higher latitudes10. We studied bacterial diversity in 40 soils sampled during the 2007–2008 austral spring and summer from along a 1,650 km climatic gradient encompassing almost the entire maritime Antarctic (Fig. 1, Supplementary Table S1). The soils that were sampled were devoid of vegetation, and are hence typical of maritime Antarctic terrestrial ecosystems. For each sample, 22 environmental factors were measured, including soil pH, electrical conductivity, moisture content, ion and element concentrations and mean annual surface air temperature (MASAT), which was derived from the Regional Atmospheric Climate Model16. The diversity of soil bacterial communities was characterised using high-throughput phylogenetic marker gene sequencing. Significant associations between environmental factors and the diversity and composition of bacterial communities in the region were then identified.
Locations of sampling sites along the climatic gradient. Site names, latitudes, longitudes, mean annual surface air temperatures (MASAT), altitudes and soil pH values are shown in Supplementary Table S1. MASAT for 2007 are shown as a colour gradient. Upper, middle and lower insets show MASAT, soil C:N ratio and Mg concentration as functions of latitude, respectively. The image was generated using ArcGIS v. 10.363.
Results
Associations between latitude and environmental factors
We observed a significant increase (r2 = 65%, P < 0.001) in MASAT between southeast Alexander Island at 72°S (MASAT −11 °C) and Signy Island at 60 °S (MASAT −4 °C), with a 0.58 °C increase in air temperature for each degree decrease in latitude (Fig. 1; Supplementary Table S2). In addition, we found that the ratio of soil organic C to N decreased at lower latitudes (r2 = 41%, P < 0.001; Fig. 1; Supplementary Table S2), which was principally owing to the highest C:N ratios being recorded in four soils from the most southerly location along the gradient (Fig. 1). An association was also found between latitude and soil Mg concentration, with higher concentrations of the element in soils from lower latitudes (r2 = 27%, P < 0.001; Fig. 1; Supplementary Table S2). None of the other environmental factors were significantly associated with latitude (Supplementary Table S2).
Factors influencing soil bacterial alpha diversity
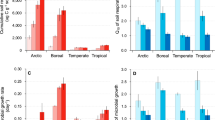
Stepwise multiple regression modelling indicated that MASAT was the strongest and most consistent predictor of the numbers of observed and predicted (Chao1) bacterial taxa, as well as the phylogenetic diversity of each community (Table 1; Fig. 2). For every degree Celsius increase in MASAT, stepwise multiple regression models indicated an additional 32.8 observed taxa, 136.8 predicted taxa and a 1.3 unit increase in Faith’s Phylogenetic Diversity values (Table 1). All three measures of soil bacterial alpha diversity were also positively, but less closely, associated with soil moisture concentration (Table 1). Lastly, the number of observed OTUs was positively associated with Mg concentration, and the two other measures of alpha diversity were negatively associated with concentrations of water extractable PO43−, or dissolved organic C and Ca (Table 1). Soil pH value, which ranged from pH 5.1–7.9 (Supplementary Table S1), was not found to influence bacterial alpha diversity.
Factors influencing soil bacterial beta diversity
Stepwise PERMANOVA analyses indicated that MASAT was the strongest predictor of differences in bacterial community composition between soils (Table 2). Significant associations were also detected between changes in soil bacterial community composition and the concentrations of Mg, moisture, dissolved organic C and soil C:N ratio (Table 2). Soil pH had no influence on beta diversity.
Univariate regression analyses showed that the relative abundances of members of the Acidobacteria (OTU 6), Chitinophagaceae (OTU 19, Bacteroidetes), Gemmatimonadetes (OTUs 34 and 36) and Comamonadaceae (OTU 44, Betaproteobacteria) were positively associated with MASAT (Fig. 3a–e), and that members of the Chloracidobacterium (OTUs 1 and 2, Acidobacteria) and Dechloromonas (OTU 46, Betaproteobacteria) were negatively associated with MASAT (Fig. 3f–h). The relative abundances of four bacterial taxa were found to be related to soil C:N ratio, with a member of the Chitinophagaceae (OTU 14) being positively associated with C:N ratio (Fig. 4a), and a Comamonadaceae (OTU 43), a Flavobacterium (OTU 26, Bacteroidetes) and a Zymomonas (OTU 39, Alphaproteobacteria) population being negatively associated with C:N ratio (Fig. 4b–d). Similarly, another member of the Chitinophagaceae (OTU 18) and a representative of the Xanthomonadaceae (OTU 48, Gammaproteobacteria) were found to be positively associated with dissolved organic C concentration (Fig. 4e,f). Members of the Acidobacteria (OTU 5) and the Flavobacteriaceae (OTU 25, Bacteroidetes) were relatively more abundant in soils with high Mg concentrations (Fig. 4g,h).
Soil bacterial community composition
In addition to members of the Acidobacteria, Bacteroidetes, Gemmatimonadetes and Proteobacteria, representatives of the Actinobacteria and Cyanobacteria were abundant in maritime Antarctic soils (Fig. 5). Those of the Candidate phylum AD3, Chloroflexi and Firmicutes were also abundant in soils at several locations but were more sporadically distributed (Fig. 5). The bacterial taxa detected were closely related to those observed in other surveys of soils from Antarctica or other cold and arid environments (Supplementary Figs S1–S5).
Heatmap illustrating the abundances of the dominant (>5%) bacterial taxa recorded in each soil as well as those that were significantly associated with the predictor variables shown in Table 2.
Discussion
Our analyses, based on 40 soils sampled from along a 1,650 km climatic gradient, indicate that air temperature is the strongest and most consistent factor explaining latitudinal changes in maritime Antarctic soil bacterial diversity. In accordance with other studies in the region showing positive associations between air temperature and soil fungal diversity9, decreased bryophyte and lichen diversity in more southerly habitats17,18 and increased plant growth rates in response to warming in the latter half of the 20th century4,6, the data reported here indicate that the warming which is predicted to occur towards the end of the current century under moderate greenhouse gas emission scenarios8 is likely to lead to increased diversity of soil bacterial communities across the region. These predicted changes to soil bacterial diversity are likely to be effected by bird19 and human vectors5, as well as by wind currents, which are thought to be capable of transporting bryophyte spores, pollen and microbes from South America to the maritime Antarctic, and southwards along the Antarctic Peninsula20,21,22.
Although bacterial diversity is probably more closely associated with soil temperature than air temperature, the inaccessible nature of the majority of the sites studied here – some of which had not previously been visited – precluded the long-term measurement of soil temperatures and their inclusion in regression models as predictors for bacterial diversity. However, previous research indicates that, despite air and soil temperatures becoming decoupled during winter in maritime Antarctica owing to the insulating effects of snow and ice cover23, air temperature is the best predictor for soil temperature in the region24 (Supplementary Fig. S6). Rising air and soil temperatures might also affect the frequency of freeze-thaw cycles at soil surfaces, which short-term laboratory studies have found to have marginal effects on maritime Antarctic bacterial community composition12. However, rises in temperature will increase the availability of liquid water, a key driver of biological diversity in Antarctic terrestrial ecosystems25. Indeed, in this study, soil moisture was positively associated with the number of bacterial taxa in soil and was a significant predictor of changes to the relative frequencies of taxa between soils. In these frigid and usually arid habitats, growth is typically limited to periods between late spring and late summer, when soil temperatures rise above 0 °C and liquid water becomes available23,26, with the increased number of bacterial taxa in warmer and wetter soils likely reflecting the enhanced potential for metabolic activity for a wider range of taxa during late austral spring and summer9,18. Although rising air temperatures across the region might lead to accelerated soil thaw and increased meltwater and summertime precipitation27, amplifying the effects of climate warming on soil bacterial diversity, we cannot discount the possibility that changing atmospheric circulation patterns, such as increased frequencies of easterly winds bringing cold, dry air from continental Antarctica28, will force the drying of maritime Antarctic soils, limiting the effects of rising air temperatures on soil bacterial diversity.
Increased soil bacterial diversity associated with rising air temperatures can be expected to enhance terrestrial ecosystem productivity by accelerating the mineralisation rates of C and N and other elements in soil that limit the growth of other organisms, in particular higher plants and bryophytes12,13,29. The analyses here suggest that future rises in air temperature in the region will result in increases in the abundances of specific bacterial taxa, notably members of the Acidobacteria – a frequent phylum in more northerly maritime Antarctic soils10 – and the Gemmatimonadetes, Chitinophagaceae and Comamonadaceae. Increases in the abundances of these taxa might be expected to enhance the decomposition of chitin and other soil organic compounds14,30 and, given that a relative of OTU 34, Gemmatimonas sp. AP64, is capable of photosynthesis31, perhaps also to increase the fixation of atmospheric C into soil. However, of the three taxa that were found here to decrease in abundance in warmer soils, two (OTUs 1 and 2, both Chloracidobacterium) are close relatives of the photoheterotroph Candidatus Chloracidobacterium thermophilum32, suggesting decreased C fixation into warmer maritime Antarctic soils. Reductions in these taxa may hence potentially counteract any changes arising from increases in the abundances of members of the Gemmatimonadetes, representing a shift in the structure of the microbial photoautotroph community.
Increasing air temperatures in maritime Antarctica are thought not only to enhance plant growth and soil fungal diversity4,6,9, but also to accelerate mineralisation rates and soil nutrient cycling33. Maritime Antarctic soils in which there are greater availabilities of inorganic nutrients might be expected to support more active34 and perhaps more diverse bacterial communities. This expectation was only weakly supported by the multiple regression analyses here, which indicated a positive effect of soil Mg concentration on the observed number of bacterial taxa, with univariate regressions also indicating increased abundances of members of the Acidobacteria and Flavobacteriaceae in soils containing higher concentrations of the element. In contrast, the multiple regressions indicated that the concentrations of dissolved organic C, water extractable PO43− and Ca were all negatively associated with the predicted numbers of soil bacterial species and phylogenetic diversity, suggesting that the positive effects of warming on maritime Antarctic soil bacterial diversity may be counteracted by increased concentrations of soil nutrients associated with more productive soil and plant communities33. These findings corroborate observations from the Arctic, showing reductions in bacterial diversity and evenness in nutrient-amended soils associated with increased relative abundances of copiotrophic Alphaproteobacteria and Betaproteobacteria35,36. The substantial decline reported here in phylogenetic diversity associated with increasing dissolved organic C concentrations in soil is likely to have arisen, at least in part, from increases in the abundances of specific taxa such as members of the Bacteroidetes and Gammaproteobacteria, close relatives of which exhibit copiotrophic behaviour29. Univariate regressions also indicated that soils with low C:N ratios, which form in warmer environments owing to faster organic matter decomposition37, and may be indicative of nutrient-rich habitats, were preferred by Zymomonas, Flavobacterium and a member of the Comamonadaceae, representatives of phyla frequent in soils with high rates of C turnover29.
In agreement with previous studies in the maritime Antarctic that have found no influence of soil pH on soil bacterial alpha diversity or community composition10,15, or in which pH has been identified as an only marginally significant predictor (P > 0.06) for the frequencies of C and N cycling genes14, soil pH had no influence on bacterial alpha or beta diversity in the present study. This is in stark contrast to continental scale studies across North America and South America, which report highly significant first order polynomial relationships between soil pH and bacterial diversity, and no effects of mean annual temperature on diversity, in 88–98 soils sampled from Argentina through to northern Alaska38,39. This disparity is probably due to the absence from the analyses here of strongly acidic soils, in which there are highly significant linear increases in bacterial diversity between pH 3.5–5.0 in the Americas38,39. In contrast, no changes in bacterial diversity are recorded in North American and South American soils over the pH range of the soils studied here (pH 5.1–7.9)38,39. Nevertheless, why temperature influences soil bacterial diversity in maritime Antarctica and not in the Americas remains to be resolved. One possible explanation is that the much harsher environmental conditions encountered in maritime Antarctic soils, such as desiccation, low temperatures (<0 °C for approximately eight months each year and annual minima of between −15 °C and −40 °C) and wide temperature fluctuations (annual ranges 35–65 °C)23 exert stronger selection pressures on bacterial taxa than in the soils sampled from across South America and North America, of which only six are exposed to annual mean temperatures of <0 °C38,39. This view is supported by the finding here that the taxa of bacteria present in maritime Antarctic soils – which are phylogenetically similar to those previously recorded in the soils of the region10,40,41,42 – have affinities with taxa encountered in other cold and arid environments, such as ice from the McMurdo Dry Valleys in Continental Antarctica43, a Lake Vostok ice core44 and hyper-arid soils from the Chilean Andes45.
Conclusions
The data reported here indicate that temperature is the predominant factor determining the diversity of bacterial communities in maritime Antarctic soils. Future rises in air temperature, predicted to occur later this century under moderate greenhouse gas emission scenarios8, are thus likely to lead to increased numbers of bacterial species in the soils of the region, enhancing terrestrial ecosystem productivity. However, increased concentrations of soil nutrients such as PO43−, Ca and dissolved organic C, which are negatively associated with bacterial diversity, may counteract the positive effects of rising air temperatures on the numbers of bacterial species in soil.
Methods
Soil sampling
Soils without plant cover were sampled from between Signy Island (60°S) in the South Orkney Islands and Alexander Island (72°S) in the southern maritime Antarctic in the 2007–2008 austral spring and summer. The uppermost c. 50 mm of soil was collected in DNA/RNAase treated plastic tubes from each of five locations at each site and was bulked. Soils were immediately snap frozen by immersion in a mixture of dry ice and ethanol (c. −80 °C) and were maintained at this temperature until they were processed.
Soil physicochemical characteristics
Analyses of the concentrations of soil moisture, elements and water-extractable ions were performed on 4 mm-sieved soils, as described previously9.
Air temperature data
Mean annual surface air temperature (MASAT) data for each site, gridded at a horizontal resolution of 55 × 55 km for the year 2007, were derived from the Regional Atmospheric Climate Model16. Although long-term soil temperature measurements for the majority of the sites studied here are unavailable, an analysis of five years of air and soil temperatures at Mars Oasis, the southernmost site sampled, shows that there is a close correlation between daily mean air temperature measured at 1 m above ground level and soil surface temperatures at 0–5 cm depth (Supplementary Fig. S6).
DNA extraction, PCR amplification and 454 pyrosequencing
Total DNA was extracted from soils as previously described9. PCRs were performed on 2 µl DNA extracts, in 1× High Fidelity PCR Buffer (Invitrogen), with 100 nM of each dNTP (Invitrogen), 2 mM MgSO4 (Invitrogen), 1 unit of Platinum® Taq High Fidelity (Invitrogen), and 400 nM of each universal bacterial primer, made up to 30 µl total volume with molecular biology grade water. The forward primer was 27F (5′ AGAGTTTGATCCTGGCTCAG), 5′-labelled with the 454 FLX sequencing primer adapter B sequence. The reverse primer was 338R (5′ TGCTGCCTCCCGTAGGAGT), 5′-labelled with a sample specific barcode sequence46 and the 454 FLX sequencing primer adapter A sequence. Thermocycling conditions were as follows: 94 °C for 2 min; five cycles touchdown at 94 °C for 30 s, 60–56 °C for 30 s, 68 °C for 45 s, and then 30 cycles of 94 °C for 30 s, 55 °C for 30 s, 68 °C for 30 s and a final extension step at 68 °C for 8 min. Amplicons were purified and normalised to 25 ng DNA per sample using a SequalPrepTM Normalization Plate Kit according to the manufacturer’s instructions (ThermoFisher Scientific). Amplicons were then pooled for 454 GS-FLX Titanium pyrosequencing (Roche), which was performed at the NERC Biomolecular Analysis Facility (University of Liverpool, UK).
Processing of sequence data
Sequences were quality filtered and dereplicated using the QIIME script split_libraries.py with the homopolymer filter deactivated47 and then checked for chimeras against the October 2013 release of the GreenGenes database using UCHIME48 v. 3.0.617. Homopolymer errors were corrected using Acacia49 v. 1.48. Using QIIME, sequences were then clustered at 97% similarity using UCLUST50 and cluster representatives were randomly selected. GreenGenes taxonomy51 (October 2013 release) was then assigned to the cluster representatives using BLAST, and tables with the abundances of each Operational Taxonomic Unit (OTU) and its taxonomic assignment in each sample were generated. In addition, full length sequences that were identified as the nearest BLAST matches for each OTU were aligned and a midpoint rooted phylogenetic tree was generated.
A plot of the observed number of taxa relative to the number of sequences per sample showed that the sequencing did not account for all of the taxa present in the soils (Supplementary Fig. S7). Rarefaction52 was hence used to calculate the expected number of taxa for an equal number of sequences per sample. The numbers of reads were rarefied to the nearest multiple of 50 sequences below the minimum number of sequences per sample (5,734) by re-sampling the OTU table. All comparisons of diversity were subsequently based on rarefied datasets comprising 5,700 sequences per sample. Inspection of the 95% confidence intervals for each rarefied sample (Supplementary Fig. S7) showed that there was considerably more between-than within-sample variation, indicating that the mean alpha diversity values used in subsequent analyses adequately represented the within-sample diversity to facilitate robust comparisons between samples. The mean number of observed OTUs, predicted OTUs (Chao1)53 and Faith’s Phylogenetic Diversity Index values54 were calculated using QIIME.
Statistical analyses
Relationships between latitude, mean annual surface air temperature (MASAT), altitude and the soil physicochemical characteristics were identified using Pearson’s correlations. The influence of MASAT and the soil physicochemical factors on the alpha diversity metrics (i.e., the numbers of observed and predicted [Chao1] OTUs, and Faith’s Phylogenetic Diversity Index values) were assessed using stepwise multiple regression analyses. The influence of MASAT and the soil physicochemical parameters on the composition of bacterial communities (beta diversity) was assessed using Permutational Multivariate Analysis of Variance55 (PERMANOVA) as implemented in the Vegan R package56. Parsimonious PERMANOVA models were built by forward selection of significant predictors and the OTU relative abundances were Hellinger transformed prior to analysis. All analyses were implemented using R version 3.2.357.
Phylogeographical analyses
Bacterial 16S rRNA gene amplicon sequences from the 2003–2005 survey of maritime Antarctic soils10 (Genbank ACC: EF219488 – EF221599) were downloaded and aligned with the GreenGenes database (October 2013 release) and the most dominant OTUs from the present study using PyNAST58. All sequences were then inserted into the full GreenGenes phylogeny (October 2013 release) by maximum parsimony using Arb59. Sequences neighbouring the OTU sequences from this study were selected and then aligned in MEGA660 using MUSCLE61. Phylogenetic trees were inferred in MEGA6 using the maximum likelihood method based on the Jukes-Cantor model62.
Data Availability
The 16S rRNA gene amplicon sequences associated with this study have been deposited in the NCBI SRA under accession: PRJNA213362.
References
Adams, B. et al. The instumental peroid. In: Turner, J., et al. (eds) Antarcticclimate change and the environment. pp. 183–298. Scientific Committee on Antarctic Research, Scott Polar Research Institute (2009).
Turner, J. et al. Absence of 21st century warming on Antarctic Peninsula consistent with natural variability. Nature 535, 411–415 (2016).
Mulvaney, R. et al. Recent Antarctic Peninsula warming relative to Holocene climate and ice-shelf history. Nature 489, 141–144 (2012).
Royles, J. et al. Plants and soil microbes respond to recent warming on the Antarctic Peninsula. Curr. Biol. 23, 1702–1706 (2013).
Frenot, Y. et al. Biological invasions in the Antarctic: extent, impacts and implications. Biol. Rev. Camb. Philos. Soc. 80, 45–72 (2005).
Fowbert, J. A. & Smith, R. L. L. Rapid population increases in native vascular plants in the Argentine Islands, Antarctic Peninsula. Arct. Alp. Res. 26, 290–296 (1994).
Convey, P., Hopkins, D. W., Roberts, S. J. & Tyler, A. N. Global southern limit of flowering plants and moss peat accumulation. Polar Res. 30, 8929 (2011).
Bracegirdle, T. J., Connolley, W. M. & Turner, J. Antarctic climate change over the twenty first century. J. Geophys. Res. 113, D03103 (2008).
Newsham, K. K. et al. Relationship between soil fungal diversity and temperature in the maritime Antarctic. Nat. Clim. Chang. 6, 182–186 (2016).
Yergeau, E., Newsham, K. K., Pearce, D. A. & Kowalchuk, G. A. Patterns of bacterial diversity across a range of Antarctic terrestrial habitats. Environ. Microbiol. 9, 2670–2682 (2007).
Yergeau, E. et al. Environmental microarray analyses of Antarctic soil microbial communities. ISME J. 3, 340–351 (2009).
Yergeau, E. & Kowalchuk, G. A. Responses of Antarctic soil microbial communities and associated functions to temperature and freeze-thaw cycle frequency. Environ. Microbiol. 10, 2223–2235 (2008).
Yergeau, E. et al. Shifts in soil microorganisms in response to warming are consistent across a range of Antarctic environments. ISME J. 6, 692–702 (2012).
Yergeau, E., Kang, S., He, Z., Zhou, J. & Kowalchuk, G. A. Functional microarray analysis of nitrogen and carbon cycling genes across an Antarctic latitudinal transect. ISME J. 1, 163–179 (2007).
Yergeau, E. et al. Size and structure of bacterial, fungal and nematode communities along an Antarctic environmental gradient. FEMS Microbiol. Ecol. 59, 436–451 (2007).
Van Lipzig, N. P. M., Van Meijgaard, E. & Oerlemans, J. Evaluation of a regional atmospheric model using measurements of surface heat exchange processes from a site in Antarctica. Mon. Weather Rev. 127, 1994–2011 (1999).
Peat, H. J., Clarke, A. & Convey, P. Diversity and biogeography of the Antarctic flora. J. Biogeogr. 34, 132–146 (2006).
Green, T. G. A., Sancho, L. G., Pintado, A. & Schroeter, B. Functional and spatial pressures on terrestrial vegetation in Antarctica forced by global warming. Polar Biol. 34, 1643–1656 (2011).
Lewis, L. R. et al. First evidence of bryophyte diaspores in the plumage of transequatorial migrant birds. PeerJ 2, e424 (2014).
Marshall, W. A. Biological particles over Antarctica. Nature 383, 680–680 (1996).
Wilson, K., Sprent, J. I. & Hopkins, D. W. Nitrification in Antarctic soils. Nature 385, 404–404 (1997).
Biersma, E. M. et al. Low genetic variation between South American and Antarctic populations of the bank-forming moss Chorisodontium aciphyllum (Dicranaceae). Polar Biol. 41, 599–610 (2018).
Convey, P., Coulson, S. J., Worland, M. R. & Sjöblom, A. The importance of understanding annual and shorter-term temperature patterns and variation in the surface levels of polar soils for terrestrial biota. Polar Biol. 41, 1587–1605 (2018).
Salamene, S. et al. Correlation between atmospheric physical factors and soil temperature of Keller Peninsula, King George Island, Antarctica. In Proceedings 19 th World Congress of Soil Sciences, Brisbane, Australia: Soil Solutions for a Changing World (ed. Gilkes, R.) 13–14 (Crawley, International Union of SoilSciences, 2010).
Kennedy, A. D. Water as a limiting factor in the antarctic terrestrial environment: a biogeographical synthesis. Arct. Alp. Res. 25, 308 (1993).
Dennis, P. G. et al. Warming constrains bacterial community responses to nutrient inputs in a southern, but not northern, maritime Antarctic soil. Soil Biol. Biochem. 57, 248–255 (2013).
Turner, J., Lachlan-Cope, T., Colwell, S. & Marshall, G. J. A positive trend in western Antarctic Peninsula precipitation over the last 50 years reflecting regional and Antarctic-wide atmospheric circulation changes. Ann. Glaciol. 41, 85–91 (2005).
Robinson, S. A. et al. Rapid change in East Antarctic terrestrial vegetation in response to regional drying. Nat. Clim. Chang. 8, 879–884 (2018).
Fierer, N., Bradford, M. A. & Jackson, R. B. Toward an ecological classification of soil bacteria. Ecology 88, 1354–64 (2007).
Bodenhausen, N. et al. Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS One 8, e56329 (2013).
Zeng, Y. et al. Functional type 2 photosynthetic reaction centers found in the rare bacterial phylum. Gemmatimonadetes. Proc. Natl. Acad. Sci. USA 111, 7795–7800 (2014).
Garcia Costas, A. M. et al. Complete genome of Candidatus Chloracidobacterium thermophilum, a chlorophyll-based photoheterotroph belonging to the phylum. Acidobacteria. Environ. Microbiol. 14, 177–190 (2012).
Hill, P. W. et al. Vascular plant success in a warming Antarctic may be due to efficient nitrogen acquisition. Nat. Clim. Chang. 1, 50–53 (2011).
Harris, J. M. & Tibbles, B. J. Factors affecting bacterial productivity in soils on isolated inland nunataks in continental Antarctica. Microb. Ecol. 33, 106–123 (1997).
Campbell, B. J., Polson, S. W., Hanson, T. E., Mack, M. C. & Schuur, E. A. G. The effect of nutrient deposition on bacterial communities in Arctic tundra soil. Environ. Microbiol. 12, 1842–1854 (2010).
Koyama, A., Wallenstein, M. D., Simpson, R. T. & Moore, J. C. Soil bacterial community composition altered by increased nutrient availability in Arctic tundra soils. Front. Microbiol. 5, 516 (2014).
Callesen, I., Raulund-Rasmussen, K., Westman, C. J. & Tau-Strand, L. Nitrogen pools and C:N ratios in well-drained Nordic forest soils related to climate and soil texture. Boreal Environ. Res. 12, 681–692 (2007).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 103, 626–631 (2006).
Lauber, C. L., Hamady, M., Knight, R. & Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 75, 5111–5120 (2009).
Newsham, K. K., Pearce, D. A. & Bridge, P. D. Minimal influence of water and nutrient content on the bacterial community composition of a maritime Antarctic soil. Microbiol. Res. 165, 523–530 (2010).
Chong, C. W. et al. High levels of spatial heterogeneity in the biodiversity of soil prokaryotes on Signy Island, Antarctica. Soil Biol. Biochem. 42, 601–610 (2010).
Chong, C. W., Convey, P., Pearce, D. A. & Tan, I. K. P. Assessment of soil bacterial communities on Alexander Island (in the maritime and continental Antarctic transitional zone). Polar Biol. 35, 387–399 (2012).
Bidle, K. D., Lee, S., Marchant, D. R. & Falkowski, P. G. Fossil genes and microbes in the oldest ice on earth. Proc. Natl. Acad. Sci. USA 104, 13455–13460 (2007).
Raymond, J. A., Christner, B. C. & Schuster, S. C. A bacterial ice-binding protein from the Vostok ice core. Extremophiles 12, 713–717 (2008).
Costello, E. K., Halloy, S. R. P., Reed, S. C., Sowell, P. & Schmidt, S. K. Fumarole-supported islands of biodiversity within a hyperarid, high-elevation landscape on Socompa Volcano, Puna de Atacama, Andes. Appl. Environ. Microbiol. 75, 735–747 (2009).
Hamady, M., Walker, J. J., Harris, J. K., Gold, N. J. & Knight, R. Error-correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat. Methods 5, 235–237 (2008).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Bragg, L., Stone, G., Imelfort, M., Hugenholtz, P. & Tyson, G. W. Fast, accurate error-correction of amplicon pyrosequences using Acacia. Nat. Methods 9, 425–426 (2012).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Gotelli, N. J. & Colwell, R. K. Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecol. Lett. 4, 379–391 (2001).
Chao, A. Estimating the population size for capture-recapture data with unequal catchability. Biometrics 43, 783 (1987).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Oksanen, J. et al. Vegan. Community ecology package. R package version 2.4–2 (2017).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing (2015).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–7 (2010).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Jukes, T. H. & Cantor, C. R. Evolution of protein molecules. In: Munro, H. N. Mammalian protein metabolism. pp. 21–132. Academic Press (1969).
Anon. ArcGIS version 10. 3, http://desktop.arcgis.com/en/arcmap/10.3, Environmental Systems Research Institute (ESRI) (2011).
Acknowledgements
This work was funded by a NERC Antarctic Funding Initiative grant (NE/D00893X/1; AFI 7/05). P.G. Dennis acknowledges support from a University of Queensland Early Career Researcher Award. Logistical support was provided by the British Antarctic Survey and the Royal Navy (HMS Endurance). P.T. Fretwell, V.A. Laudicina, V. Ord, P. Coates, M. Dunn, P. Torode, M. Jobson, A. Clark, J. Wake, R. Hall, G. Marshall, M. Biszczuk and K. Bazeley provided data and technical support. All are gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
D.W.H., K.K.N., S.P.R. and A.G.O. secured funding. P.G.D., K.K.N. and D.W.H. collected soil samples. P.G.D. performed the analyses. K.K.N. and P.G.D. wrote the manuscript, which was approved by the other authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dennis, P.G., Newsham, K.K., Rushton, S.P. et al. Soil bacterial diversity is positively associated with air temperature in the maritime Antarctic. Sci Rep 9, 2686 (2019). https://doi.org/10.1038/s41598-019-39521-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-39521-7
This article is cited by
-
Biogeographic survey of soil bacterial communities across Antarctica
Microbiome (2024)
-
Microbial metabolic responses and CO2 emissions differentiated by soil water content variation in subarctic tundra soils
Journal of Microbiology (2022)
-
Influence of prokaryotic microorganisms on initial soil formation along a glacier forefield on King George Island, maritime Antarctica
Scientific Reports (2021)
-
Bacterial Community Composition and Diversity Respond to Nutrient Amendment but Not Warming in a Maritime Antarctic Soil
Microbial Ecology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.