Abstract
Green tea polyphenols may protect cells from UV damage through antioxidant activities and by stimulating the removal of damaged or cross-linked DNA. Recently, DNA repair pathways have been predicted as possible targets of epigallocatechin gallate (EGCG)-initiated signaling. However, whether and how green tea polyphenols can promote nucleotide excision repair and homologous recombination in diverse organisms requires further investigation. In this report, we used the budding yeast, Saccharomyces cerevisiae, as a model to investigate the effects of green tea extract on DNA repair pathways. We first showed that green tea extract increased the survival rate and decreased the frequency of mutations in yeast exposed to UVB-irradiation. Furthermore, green tea extract increased the expression of homologous recombination genes, RFA1, RAD51 and RAD52, and nucleotide excision repair genes, RAD4 and RAD14. Importantly, we further used a specific strand invasion assay to show that green tea extract promotes homologous recombination at double-strand breaks. Thus, green tea extract acts to preserve genome stability by activating DNA repair pathways in yeast. Because homologous recombination repair is highly conserved in yeast and humans, this study demonstrates yeast may be a useful platform for future research to investigate the underlying mechanisms of the bioactive compounds in DNA repair.
Similar content being viewed by others
Introduction
Over the past decades, many studies have demonstrated that green tea and its bioactive components are beneficial to human health. Most of the beneficial effects of green tea have been attributed to the robust anti-inflammatory and anti-tumorigenic effects of constituent polyphenolic catechins, such as (–)-epigallocatechin-3-gallate (EGCG), (–)-epigallocatechin (EGC), (–)-epicatechin-3-gallate (ECG) and (–)-epicatechin (EC)1,2,3,4. While these physiological effects are well documented, the mechanisms by which catechins may protect from tumorigenesis are not fully described.
Genomic instability leads to mutations and promotes cancer formation in complex organisms5. Moreover, maintenance of genome stability is critical for normal cell growth, and prevents damaged or mutated DNA from being inherited by the next generation in all organisms. In order to maintain genome stability, highly conserved DNA repair systems were evolved in ancient ancestors to minimize the cytotoxic and mutagenic effects of DNA damage. One of the most common DNA damaging agents, which can lead to mutagenesis and skin cancer in humans, is UV-irradiation from sunlight6. UV-irradiation causes multiple types of DNA damage, including oxidative damage7, cross-linking of bases8 and double-strand breaks (DSBs), which are most detrimental to genome integrity9. Accumulation of UV-induced DNA lesions undoubtedly increases the probability of cancer formation. Indeed, an early study has demonstrated that the incidence of UV-induced skin cancer in mice can be reduced by enhancing DNA repair by applying exogenous T4 endonuclease V, which can initiate the removal of UV-induced cyclobutane pyrimidine dimers (CPDs) from DNA10.
A number of phenolic compounds, including vanillin, cinnamaldehyde, coumarin and tannic acid, prevent mutagenesis in E. coli, by influencing DNA replication and/or repair after DNA damage11,12,13. Several studies have shown that green tea extract (GTE) contains polyphenols that may act as antioxidants to prevent DNA damage14,15,16. A recent study revealed that regular intake of green tea can lower lymphocytic DNA damage and increase the activity of 8-oxoguanine glycosylase (OGG1), a DNA glycosylase enzyme involved in base excision repair (BER) in human lymphocyte extracts17. Moreover, green tea polyphenols were also shown to enhance CPD removal in skin cells by nucleotide excision repair (NER) and decrease apoptosis in mice after UV exposure18,19,20. Interestingly, EGCG has been shown to prevent cell cycle progression in cancer cells21,22 and decrease UVB-induced oxidative stress in human skin cells23. Intriguingly, homologous recombination (HR) has been recently predicted as a possible target of EGCG in a bioinformatics study on breast cancer24. However, it is unknown whether green tea polyphenols can promote NER and HR in diverse model organisms that may be useful for further mechanistic studies.
Yeast has been extensively used as an experimental organism for modern biology, especially due to its amenability to classical and molecular genetic methods. This organism is often used to associate genes with certain functions in eukaryotic cells, as it is genetically tractable and many biological mechanisms are highly conserved in humans25. For example, yeast models have been invaluable in dissecting the mechanisms by which phytochemicals in food (e.g. polyphenols from apples and tangeretin from orange peels) provide well-known benefits to human health26,27. In this study, we utilized the budding yeast Saccharomyces cerevisiae to investigate whether GTE could promote genome stability by regulating NER and HR, as most DNA repair pathways are highly conserved between yeast and humans28.
Results
Polyphenol content in green tea extract
We first determined the content of various compounds in GTE by HPLC. The concentrations of major catechins, including EGCG, EGC, ECG and EC, as well as gallic acid and caffeine in GTE (μg/100 ml) are shown in Table 1. The total catechin content in our GTE (~121 mg/100 ml) was comparable to those reported in a survey of nine green teas (74–216 mg/100 ml)14. Interestingly, our GTE had relatively low EGCG content, but much a higher level of EC. These differences with previous literature may be due to variations in production conditions and technologies14.
GTE treatment enhances the survival rate of UVB-irradiated cells
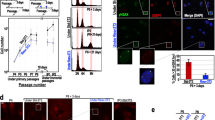
We examined how GTE and its major polyphenolic components, EGCG, EGC and caffeine, affect cell survival after UVB-irradiation. GTE- and EGCG-treated groups showed significant increases (P < 0.05 and P = 0.05, respectively) in cell survival rates, as compared to untreated and other groups at a dose of 200 J/m2 UVB (Fig. 1A). In contrast, the survival rates of cells treated with EGC, caffeine, or a combination of EGCG, EGC and caffeine did not differ from the control (Fig. 1A). To test whether GTE could enhance survival of UVB-irradiated cells at higher doses, cells were exposed to a UVB dose of 400 J/m2, and the survival rates were determined. Indeed, we observed a significantly higher (P < 0.05) survival rate in the GTE-treated group compared to control (Fig. 1B). Thus, cells treated with GTE were more tolerant to UV-damage as compared to untreated cells or those treated with certain pure compounds.
GTE-treatment enhances the survival of UVB-irradiated cells. Cells were irradiated with UVB at doses of (A) 200 J/m2 and (B) 400 J/m2. Relative percentage of surviving cells was calculated in comparison to UVB-irradiated controls. Data are presented as mean ± S.D. determined from at least two independent cultures measured in triplicate. *P < 0.05 compared to control. aP = 0.05 compared to control.
GTE promotes expression of NER genes
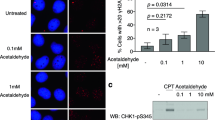
Green tea polyphenols have been shown to enhance the removal of highly mutagenic UV-induced adducts from DNA17,18,19, a process that is predominantly mediated by NER. The NER pathway is divided into two sub-pathways: global genomic NER (GG-NER), which repairs transcriptionally inactive or silent areas of the genome, and transcription-coupled NER (TC-NER), which repairs DNA lesions in the coding regions of active genes29. To understand which sub-pathway is influenced by GTE, we investigated the expression of genes involved in both NER sub-pathways. We first examined expression of RAD4 and RAD14, which encode key components of the GG-NER pathway30. RAD4 (the yeast homolog of human xeroderma pigmentosum C: XPC) encodes a DNA damage-binding protein that plays key roles in the early steps of GG-NER30. The product of RAD14 (a homolog of human xeroderma pigmentosum A: XPA) recognizes and binds to damaged DNA in both GG-NER and TC-NER31. GTE significantly enhanced (P < 0.05) expression of RAD4 from 20 min to 2 h, and RAD14 from 40 min to 2 h post-irradiation, as compared to the untreated group (Fig. 2A,B). However, GTE did not significantly affect expression of RAD26 (the yeast homolog of human Cockayne Syndrome B: CSB), which encodes an essential component of the TC-NER pathway (Fig. 2C). These results suggest that GTE may specifically upregulate the GG-NER repair pathway by activating genes encoding DNA-damage-sensing Rad14 and DNA-binding Rad4.
GTE treatment enhances expression of GG-NER genes following UVB irradiation. Expression levels of (A) RAD4, (B) RAD14 and (C) RAD26 are shown. Samples were collected before and after UVB-irradiation (200 J/m2) at the indicated time-points. White bars represent control groups and gray bars represent GTE groups. Data are presented as means ± S.D. *P < 0.05 compared to control at the same time-point, according to Student’s t-test.
GTE promotes HR repair
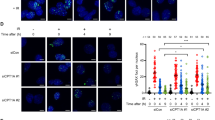
In addition to NER, HR is also involved in the repair of UV-induced DNA damage in yeast and human cells32,33, and its inactivation compromises yeast survival following UV-irradiation33. Thus, we further explored whether genes involved in the early stage of HR, including RFA1, RAD51 and RAD52, were also activated by GTE. We found that RFA1 expression was increased 20 min post-irradiation in GTE-treated cells, and expression was also elevated 3 h post-exposure (Fig. 3A). RAD51, which encodes a recombinase, also showed significant induction (P < 0.05) in cells treated with GTE at 3 h post-irradiation (Fig. 3B). Moreover, RAD52 expression levels were significantly elevated (P < 0.05) in GTE-treated cells at all times examined following exposure to UVB (Fig. 3C). These results suggest that the HR pathway is upregulated by GTE treatment in response to UV damage.
GTE treatment increases expression of HR genes following UVB irradiation. Expression levels of (A) RFA1, (B) RAD51 and (C) RAD52 are shown. Samples were collected before and after UVB-irradiation (200 J/m2) at the indicated time-points. Data are presented as means ± S.D. *P < 0.05 compared to control at the same time-point, according to Student’s t-test.
To assess whether GTE is able to enhance HR activity, we used a strand invasion/repair (SEI) assay (Fig. 4A). In this assay, a specific DSB can be induced at the MAT locus through the induction of an exogenous site-specific HO endonuclease. To determine whether GTE altered the efficiency of DSB repair by HR, we measured the initial strand-invasion phase using the SEI assay. This PCR-based assay allows us to measure the intermediates of the strand invasion/repair synthesis reaction (Fig. 4A). Our results show that at 60 min after DSB induction, SEI was dramatically enhanced in GTE-treated cells as compared to control cells (Fig. 4B). SEI peaked at 210 min in GTE-treated cells and declined thereafter, probably indicating the completion of HR repair. In control cells, SEI continued to increase slowly throughout the experimental duration. These results suggest that GTE enhances the rate of initial strand invasion and repair synthesis phase during HR, likely by inducing the expression of HR repair genes.
GTE promotes the HR activity. (A) Schematic of the MAT locus, HML, and HMR used for the single-end invasion (SEI) assay. The MAT locus, HML, and HMR share identical W, X and Z1 sequences. Galactose-induction of pGAL-HO endonuclease initiates the formation of a double-strand break at the HO cutting site. Repair of the double-strand break may occur via strand invasion by the HMRa region, which can be detected using qPCR with primers for SEI-F and SEI-R. (B) GTE enhanced HR repair at DSB sites, as measured by the SEI assay. White bars represent the control group and gray bars represent the GTE-treated group. Data are presented as means ± S.D.
UVB-induced CAN1 gene mutations suppressed by GTE
To investigate whether GTE-mediated induction of DNA repair genes results in suppression of UVB-induced gene mutations, we monitored the mutation rate of the CAN1 gene. We found that irradiation with 200 J/m2 UVB increased the frequency of Canr cells by about 40-fold in wild-type cells (Table 2). While treatment with GTE did not affect the spontaneous mutation rates, it markedly reduced the mutation frequency in irradiated cells, as compared to the irradiated control. GTE-treated cells exhibited a significantly lower CAN1 gene mutation rate upon irradiation (28-fold over non-irradiated), as compared to cells treated with pure compounds (36-fold for EGCG; 35-fold for caffeine; Table 2). These results suggest that total GTE is more potent for protecting and maintaining genome stability after UV exposure than any single functional compound that we tested. Finally, GTE-treatment did not reduce the gene mutation rate in HR-defective rad52∆ mutants (Table 2), an observation which is consistent with the notion that the protective effect of GTE is primarily through modulation of HR gene expression. Importantly, however, these results demonstrate that UVB-induced CAN1 gene mutations may be suppressed by GTE.
Discussion
In the present study, we found that GTE enhances expression of DNA repair genes in response to UVB exposure. Interestingly, GTE was specifically found to enhance HR activity in order to protect against UV-induced gene mutations. Such effects on gene expression are likely to contribute to the enhanced survival observed in GTE-treated cells.
DNA repair plays an essential role in protecting the genome from endogenous and exogenous damage. When DNA is damaged, it must be rapidly and efficiently repaired. In humans, the XPC protein forms a damage recognition complex with HR23B to detect UV-induced DNA lesions34,35. XPA subsequently interacts with replication protein A (RPA) to bind and remove damaged DNA. Thus, cells that carry defective XPC and XPA are extremely sensitive to UV, and have very low NER activity36. Patients with mutations in these genes are predisposed to skin cancer and other systemic conditions. We have demonstrated that GTE activates expression of the RAD4 and RAD14 genes within minutes of UV exposure in wild-type yeast. Given that Rad4 and Rad14 are responsible for recognizing UV-damaged nucleotides in the indiscriminate genome-wide NER repair pathway, activation of these two genes may help to maintain the overall stability of the genome. Thus, our findings identify GTE as a possible supplement to enhance GG-NER to combat UV-initiated damage.
HR repair at the MAT locus in yeast is highly dependent on Rad52 and Rad51 proteins, and has been used extensively to assess HR activity. The RFA1, RAD51 and RAD52 genes are required for UV-induced HR repair in both yeast and human cells37,38. We demonstrated that GTE stimulates expression of RFA1, RAD51 and RAD52, and this increased expression may translate into enhanced HR-mediated repair of DSBs (Fig. 4B). UV-irradiation may result in a single-strand break, and pairing of the exposed single-stranded DNA with homologous DNA allows HR repair to occur39. Consistent with this idea, we observed that a key HR gene, RAD51, was induced after (60 min after UVB treatment, Fig. 4B) the time at which we observed activation of NER genes (20 min after UVB treatment, Fig. 2A,B); this upregulated expression of RAD51 then persisted for more than 3 h. Importantly, we observed that HR repair required nearly 4 h to complete according to our SEI assay (Fig. 4). Thus, GTE may promote HR repair when DSBs are encountered. Together, our results suggest that GTE treatment can sequentially enhance NER and HR DNA repair to allow cells to recover from UV-induced DNA damage.
Previous reports have argued that mutagenesis is induced shortly after irradiation, due to faulty repair or lack of repair before or during DNA replication40. Our data suggest that GTE positively regulates the expression of repair genes for at least 3 h (Figs 3 and 4), and long-term treatment with GTE suppresses UVB-induced mutagenesis (Table 2). It is well documented that delayed mutations can arise many cell generations after UV damage, thereby increasing the gene mutation rate in the genome40,41,42,43,44. Thus, our findings indicate that continuous application of GTE to the cells decreases the incidence of delayed mutations, contributing to improved survival and genome stability in the cells.
A previous study in yeast showed that cooperative action of all apple components has more anti-aging power than individual components27. Similarly, while EGCG has been suggested to be the major bio-effective factor in GTE, we observed that GTE was more effective than EGCG at promoting cell survival and reducing gene mutation rate. This improved effect may be due to the existence of other components in GTE, such as chlorophylls and pheophytin, which also function as antioxidants, anti-genotoxic and tumor-suppressing agents44,45,46,47. Thus, our results suggest that maintenance of genome stability after UV-damage by GTE may not be solely an effect of EGCG; instead, a combination of bioactive compounds in GTE may function together to suppress UVB-induced genome instability in cells.
In conclusion, we show that GTE activates specific DNA repair and promotes genome stability in yeast. As such mechanisms are well-conserved between yeast and human, this study demonstrates that the yeast model may be a useful platform for future research on the underlying mechanisms of bioactive compounds in DNA repair.
Materials and Methods
GTE preparation
Green tea powder (purchased from a local tea company in Taiwan) (1.35 g) was mixed with warm (60 °C) distilled water (100 ml). Extraction was carried out for 20 min at 60 °C under constant stirring. The mixture was then cooled in an ice bath before being centrifuged at 4000 rpm for 5 min. The resulting supernatant was filter sterilized through a 0.22 μm MilliporeTM filter, yielding the GTE. Caffeine, catechin and gallic acid content were determined by HPLC using a Luna® C18 reverse-phase analytical column (4.6 mm i.d. ×250 mm, 5 μm particle size; Phenomenex Inc. Torrance, CA)48.
Yeast strains and growth conditions
The wild type Saccharomyces cerevisiae strain, BY4741 (MATa his3Δ leu2Δ met15Δ ura3Δ) (Euroscarf, Denmark), was used in this study. Cells were cultured in yeast extract-peptone-dextrose (YPD) media with or without EGCG (200 μM), epigallocatechin (EGC, 690 μM), caffeine (0.7 mM) or GTE, which was added at 3.5 ml GTE in 5 ml YPD culture broth. Cells were cultured for 16 h at 30 °C with shaking. Saturated cultures were used to inoculate fresh media, and the new cultures were incubated for a further 3 h at 30 °C to reach exponential phase (2 − 3 × 107 cells/ml) prior to initiating experiments.
Cell survival assay
Exponential-phase YPD cultures were diluted to an appropriate concentration, before being seeded onto YPD plates. Plates with the same cell densities, as determined by OD 600 nm, were treated with or without UVB irradiation using a UVB light source (200 or 400 J/m2; 365 nm peak; UVP, USA). Cells were then immediately incubated at 30 °C for 3 days in the dark. Relative survival was calculated as the ratio of colonies arising in irradiated versus control plates.
Gene expression
Total RNA was extracted by the acid-phenol method49. Briefly, cells were collected and resuspended in 500 μl TES buffer (10 mM Tris-HCl pH 7.5, 10 mM EDTA, 0.5% SDS). The cell suspension was mixed with 400 μl of warm acid phenol, and incubated at 65 °C for 20 min with vortexing at 5 min intervals. Following centrifugation, the supernatant was extracted twice with acid phenol and once with chloroform. Total RNA was reverse transcribed into cDNA using random primers and the Superscript II® kit (Invitrogen). Real-time PCR was performed using the ABI StepOne PlusTM system. The sequences of primers used to amplify each gene are listed in Table 3. Data were normalized to ACT1 and are presented relative to unirradiated controls. All gene expression experiments were carried out in triplicate, and two independent studies were performed.
Single-end invasion (SEI) assay
The SEI assay was performed as previously described50. Wild-type cells (MATα his3Δ leu2Δ met15Δ ura3Δ) were transformed with galactose-regulated HO nuclease expression plasmids (pGAL-HO; Trp1). Transformed cells were used to inoculate in S.C.-Trp media containing 2% raffinose, with or without GTE. Cultures were incubated at 30 °C for 24 h, to a final OD 600 of 0.6–0.8. HO endonuclease was induced by adding galactose to 2%. After 60 min, HO nuclease expression was repressed by addition of glucose to 2%. The repair intermediates were detected by real-time PCR using the extracted DNA from the cultures as templates. The fold-increase of SEI in the HO endonuclease-induced cells was calculated relative to the non-induced control. SEI assays were carried out in triplicate, and two independent studies were performed. The SEI primer sequence is shown in Table 3.
Gene mutation assay
Gene mutation rates were determined as previously described51. Briefly, five to ten independent colonies were randomly selected and grown in YPD media. Cells were then plated onto either YPD to evaluate plating efficiency or synthetic complete arginine-dropout plates containing 60 mg/L canavanine. Canavanine-resistant mutants (Canr) were counted and the median mutation rates were measured52,53. The average fold-increase in gene mutation rate was calculated relative to wild-type cells without UV treatment as a control.
Statistical analysis
Data are presented as the mean ± S.D. Comparisons were performed using Student’s t-test. Gene mutation experiments were evaluated using the Mann-Whitney method. Statistical significance was set as P < 0.05.
Data Availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Jankun, J., Selman, S. H., Swiercz, R. & SkrzypczakJankun, E. Why drinking green tea could prevent cancer. Nature 387, 561–561 (1997).
Balentine, D. A., Wiseman, S. A. & Bouwens, L. C. M. The chemistry of tea flavonoids. Crit Rev Food Sci 37, 693–704 (1997).
Sharma, P. et al. Tea polyphenols for the prevention of UVB-induced skin cancer. Photodermatol Photo 34, 50–59 (2018).
Yang, C. S., Chung, J. Y., Yang, G. Y., Chhabra, S. K. & Lee, M. J. Tea and tea polyphenols in cancer prevention. J Nutr 130, 472s–478s (2000).
Aguilera, A. & Gomez-Gonzalez, B. Genome instability: a mechanistic view of its causes and consequences. Nat Rev Genet 9, 204–217 (2008).
Brash, D. E. et al. A Role for Sunlight in Skin-Cancer - Uv-Induced P53 Mutations in Squamous-Cell Carcinoma. P Natl Acad Sci USA 88, 10124–10128 (1991).
Beehler, B. C., Przybyszewski, J., Box, H. B. & Kuleszmartin, M. F. Formation of 8-Hydroxydeoxyguanosine within DNA of Mouse Keratinocytes Exposed in Culture to Uvb and H2o2. Carcinogenesis 13, 2003–2007 (1992).
Fornace, A. J., Kohn, K. W. & Kann, H. E. DNA Single-Strand Breaks during Repair of Uv Damage in Human Fibroblasts and Abnormalities of Repair in Xeroderma Pigmentosum. P Natl Acad Sci USA 73, 39–43 (1976).
Limoli, C. L., Giedzinski, E., Bonner, W. M. & Cleaver, J. E. UV-induced replication arrest in the xeroderma pigmentosum variant leads to DNA double-strand breaks, gamma-H2AX formation, and Mre11 relocalization. P Natl Acad Sci USA 99, 233–238 (2002).
Yarosh, D. et al. Pyrimidine Dimer Removal Enhanced by DNA-Repair Liposomes Reduces the Incidence of Uv Skin-Cancer in Mice. Cancer Res 52, 4227–4231 (1992).
Ohta, T. Modification of Genotoxicity by Naturally-Occurring Flavorings and Their Derivatives. Crit Rev Toxicol 23, 127–146 (1993).
Ohta, T., Watanabe, M., Shirasu, Y. & Inoue, T. Post-Replication Repair and Recombination in Uvra Umuc Strains of Escherichia-Coli Are Enhanced by Vanillin, an Antimutagenic Compound. Mutat Res 201, 107–112 (1988).
Takahashi, K., Sekiguchi, M. & Kawazoe, Y. Effects of vanillin and o-vanillin on induction of DNA-repair networks: modulation of mutagenesis in Escherichia coli. Mutat Res 230, 127–134 (1990).
Henning, S. M. et al. Catechin content of 18 teas and a green tea extract supplement correlates with the antioxidant capacity. Nutr Cancer 45, 226–235 (2003).
Yen, G. C. & Chen, H. Y. Antioxidant Activity of Various Tea Extracts in Relation to Their Antimutagenicity. J Agr Food Chem 43, 27–32 (1995).
Hamiltonmiller, J. M. T. Antimicrobial Properties of Tea (Camellia-Sinensis L). Antimicrob Agents Ch 39, 2375–2377 (1995).
Ho, C. K., Choi, S. W., Siu, P. M. & Benzie, I. F. F. Effects of single dose and regular intake of green tea (Camellia sinensis) on DNA damage, DNA repair, and heme oxygenase-1 expression in a randomized controlled human supplementation study. Mol Nutr Food Res 58, 1379–1383 (2014).
Meeran, S. M., Akhtar, S. & Katiyar, S. K. Inhibition of UVB-Induced Skin Tumor Development by Drinking Green Tea Polyphenols Is Mediated Through DNA Repair and Subsequent Inhibition of Inflammation. J Invest Dermatol 129, 1258–1270 (2009).
Mnich, C. D. et al. Green tea extract reduces induction of p53 and apoptosis in UVB-irradiated human skin independent of transcriptional controls. Exp Dermatol 18, 69–77 (2009).
Schwarz, A. et al. Green tea phenol extracts reduce UVB-induced DNA damage in human cells via interleukin-12. Photochem Photobiol 84, 350–355 (2008).
Huang, C. H. et al. EGCG inhibits protein synthesis, lipogenesis, and cell cycle progression through activation of AMPK in p53 positive and negative human hepatoma cells. Mol Nutr Food Res 53, 1156–1165 (2009).
Khafif, A., Schantz, S. P., Al-Rawi, M., Edelstein, D. & Sacks, P. G. Green tea regulates cell cycle progression in oral leukoplakia. Head Neck-J Sci Spec 20, 528–534 (1998).
Katiyar, S. K., Afaq, F., Perez, A. & Mukhtar, H. Green tea polyphenol (-)-epigallocatechin-3-gallate treatment of human skin inhibits ultraviolet radiation-induced oxidative stress. Carcinogenesis 22, 287–294 (2001).
Song, X., Zhang, M., Chen, L. & Lin, Q. Bioinformatic Prediction of Possible Targets and Mechanisms of Action of the Green Tea Compound Epigallocatechin-3-Gallate Against Breast Cancer. Front Mol Biosci 4, 43 (2017).
Botstein, D. & Fink, G. R. Yeast: an experimental organism for modern biology. Science 240, 1439–1443 (1988).
Xiang, L. et al. Anti-aging effects of phloridzin, an apple polyphenol, on yeast via the SOD and Sir2 genes. Biosci Biotechnol Biochem 75, 854–858 (2011).
Palermo, V. et al. Apple Can Act as Anti-Aging on Yeast Cells. Oxid Med Cell Longev (2012).
Hughes, T. R. Yeast and drug discovery. Funct Integr Genomics 2, 199–211 (2002).
Mellon, I., Spivak, G. & Hanawalt, P. C. Selective Removal of Transcription-Blocking DNA Damage from the Transcribed Strand of the Mammalian Dhfr Gene. Cell 51, 241–249 (1987).
Guzder, S. N., Sung, P., Prakash, L. & Prakash, S. Yeast DNA-Repair Gene Rad14 Encodes a Zinc Metalloprotein with Affinity for Ultraviolet-Damaged DNA. P Natl Acad Sci USA 90, 5433–5437 (1993).
Guzder, S. N., Sommers, C. H., Prakash, L. & Prakash, S. Complex formation with damage recognition protein Rad14 is essential for Saccharomyces cerevisiae Rad1-Rad10 nuclease to perform its function in nucleotide excision repair in vivo. Mol Cell Biol 26, 1135–1141 (2006).
Aboussekhra, A. & Al-Sharif, I. S. Homologous recombination is involved in transcription-coupled repair of UV damage in Saccharomyces cerevisiae. Embo J 24, 1999–2010 (2005).
Toussaint, M., Wellinger, R. J. & Conconi, A. Differential participation of homologous recombination and nucleotide excision repair in yeast survival to ultraviolet light radiation. Mutat Res-Gen Tox En 698, 52–59 (2010).
Li, L., Lu, X. Y., Peterson, C. & Legerski, R. XPC interacts with both HHR23B and HHR23A in vivo. Mutat Res-DNA Repair 383, 197–203 (1997).
Sugasawa, K. et al. A multistep damage recognition mechanism for global genomic nucleotide excision repair. Gene Dev 15, 507–521 (2001).
Masson, C. et al. Global genome repair is required to activate KIN17, a UVC-responsive gene involved in DNA replication. P Natl Acad Sci USA 100, 616–621 (2003).
Lettier, G. et al. The role of DNA double-strand breaks in spontaneous homologous recombination in S. cerevisiae. Plos Genet 2, 1773–1786 (2006).
Lambert, S. & Lopez, B. S. Inactivation of the RAD51 recombination pathway stimulates UV-induced mutagenesis in mammalian cells. Oncogene 21, 4065–4069 (2002).
Durant, S. T. et al. UV radiation induces delayed hyperrecombination associated with hypermutation in human cells. Mol Cell Biol 26, 6047–6055 (2006).
Dahle, J. & Kvam, E. Induction of delayed mutations and chromosomal instability in fibroblasts after UVA-, UVB-, and X-radiation. Cancer Res 63, 1464–1469 (2003).
Myung, K. & Kolodner, R. D. Induction of genome instability by DNA damage in Saccharomyces cerevisiae. DNA Repair 2, 243–258 (2003).
Smith, S., Gupta, A., Kolodner, R. D. & Myung, K. Suppression of gross chromosomal rearrangements by the multiple functions of the Mre11-Rad50-Xrs2 complex in Saccharomyces cerevisiae. DNA Repair 4, 606–617 (2005).
Suzuki, K., Ojima, M., Kodama, S. & Watanabe, M. Radiation-induced DNA damage and delayed induced genomic instability. Oncogene 22, 6988–6993 (2003).
Higashi-Okai, K. & Okai, Y. Potent suppressive activity of chlorophyll a and b from green tea (Camellia sinensis) against tumor promotion in mouse skin. J UOEH 20, 181–188 (1998).
Higashi-Okai, K., Yamazaki, M., Nagamori, H. & Okai, Y. Identification and antioxidant activity of several pigments from the residual green tea (Camellia sinensis) after hot water extraction. J UOEH 23, 335–344 (2001).
Okai, Y. & Higashi-Okai, K. Potent anti-inflammatory activity of pheophytin a derived from edible green alga, Enteromorpha prolifera (Sujiao-nori). Int J Immunopharmacol 19, 355–358 (1997).
Suzuki, Y. & Shioi, Y. Identification of chlorophylls and carotenoids in major teas by high-performance liquid chromatography with photodiode array detection. J Agric Food Chem 51, 5307–5314 (2003).
Yang, D. J., Hwang, L. S. & Lin, J. T. Effects of different steeping methods and storage on caffeine, catechins and gallic acid in bag tea infusions. J Chromatogr A 1156, 312–320 (2007).
Chomczynski, P. & Sacchi, N. Single-Step Method of Rna Isolation by Acid Guanidinium Thiocyanate Phenol Chloroform Extraction. Anal Biochem 162, 156–159 (1987).
Tsukuda, T. et al. INO80-dependent chromatin remodeling regulates early and late stages of mitotic homologous recombination. DNA Repair (Amst) 8, 360–369 (2009).
Huang, D. & Koshland, D. Chromosome integrity in Saccharomyces cerevisiae: the interplay of DNA replication initiation factors, elongation factors, and origins. Genes Dev 17, 1741–1754 (2003).
Lea, D. E. & Coulson, C. A. The Distribution of the Numbers of Mutants in Bacterial Populations. J Genet 49, 264–285 (1949).
Hall, B. M., Ma, C. X., Liang, P. & Singh, K. K. Fluctuation AnaLysis CalculatOR: a web tool for the determination of mutation rate using Luria-Delbruck fluctuation analysis. Bioinformatics 25, 1564–1565 (2009).
Acknowledgements
This research partially supported by Ministry of Science and Technology, Taiwan, under Grant NSC 98-2313-B-002-058-MY2 and by internal support from the College of Bioresources and Agriculture, Natioanl Taiwan University
Author information
Authors and Affiliations
Contributions
Y.C.L. conceived and designed the research study and wrote and revised the manuscript. S.Y.C. performed the SEI assay and the data analysis. H.Y.C. carried out the cell survival mutation assay. T.H.C. analysed the compositions of GTE. Y.J.L. assisted the experiments. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chong, S.Y., Chiang, HY., Chen, TH. et al. Green tea extract promotes DNA repair in a yeast model. Sci Rep 9, 3842 (2019). https://doi.org/10.1038/s41598-019-39082-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-39082-9
This article is cited by
-
Mitigative effect of green tea extract against mercury(II) chloride toxicity in Allium cepa L. model
Environmental Science and Pollution Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.