Abstract
Resulting from the nuclear fuel cycle, large amounts of depleted uranium (DU) tails are piling up, waiting for possible use or final disposal. To date, the recovery of the residual 235U isotope contained in DU has been conducted only marginally by physical processes. Relative isotope abundances are often mediated by biological processes, and the biologically driven U isotopic fractionation has been previously identified in reducing bacteria. Our results indicate that the cells of two microalgal strains (freshwater Chlamydomonas sp. (ChlGS) and marine Tetraselmis mediterranea (TmmRU)) took up DU from the exposure solutions, inducing U isotopic fractionation with a preference for the fissile 235U isotope over 238U. The n(235U)/n(238U) isotopic fractionation magnitudes (δ235) were 23.6 ± 12.5‰ and 370.4 ± 103.9‰, respectively. These results open up new perspectives on the re-enrichment of DU tailings, offering a potential biological alternative to obtain reprocessed natural-equivalent uranium. Additionally, the findings present implications for identifying biological signatures in the geologic records.
Similar content being viewed by others
Introduction
Motivated by the world’s continuously growing demand for energy and climate change becoming increasingly apparent, more efficient use of energy resources and diversification from fossil fuels (which are the principal cause of greenhouse gas emissions) has become a pressing need. Nuclear fission is an important alternative energy source to fossil fuels, as the energy conversion per gram of fuel is much higher and the carbon footprint is much lower. Moreover, compared to renewable alternative energies, nuclear power produces more energy, often at lower costs1. Nuclear electricity generation capacity in 2016 was 2.476 million GWh, approximately 11.5% of the total energy demand worldwide2. Therefore, nuclear power plays a key role in the gradual replacement of fossil fuels towards sustainable resources as part of the energy mix3: “We need nuclear power as a bridge toward a post-fossil-fuel future” (Professor Steven Chu, a Nobel Prize-winning physicist). Nuclear power generation is projected to increase and prevail to meet the future energy needs as a necessary stepping-stone towards a “clean” energy.
The basic nuclear fuel is initially uranium (U), whose consumption has been increasing rapidly, prompted by the nuclear energy demand. However, U resources on land are not unlimited4. The world´s present measured resources (5.7 Mt U) estimations are enough to last for the next 90 years5. Therefore, new viable sources of uranium are being sought. For example, extraction from seawater is gaining attention as an almost inexhaustible U fuel source4,6. Re-enrichment of depleted uranium (DU) is another secondary U fuel source.
Natural U consists of two main isotopes of interest: the fissile 235U (0.71 atomic %) and the fertile 238U (99.27 atomic %), along with the decay product 234U (0.0055 atomic %). In most cases, natural U needs different levels of enrichment depending on applications requirements (nuclear fuel range between a 3 and 5 atomic % of 235U7). The foundations of the nuclear fuel cycle are to obtain critical mass ‒ that leads to a sustained nuclear chain reaction ‒ of a fissionable nuclide in the nuclear fuel, such as the fissile 235U. One of the main hot spots in the nuclear fuel cycle is the tens of thousands of tons of DU tails (U resulting from the enrichment process, typically with ~0.0025-0.003 of 235U) sitting around with medium and low radioactive activity that must be confined with security in an already battered planet. For every ton of natural U enriched, approximately 130 kg of enriched fuel (~3.5 atomic % of 235U) is produced, and the balance is DU tails. The estimated world U requirements for 2017 are 65,014 Ut (76,671 t U3O8)8, with their subsequent major proportion resulting in DU. Annually, more than 50,000 tons of this byproduct swell the already substantial stockpiles only in U.S., Europe and Russia9. Even though DU has several uses, both civilian (i.e., radiation shielding of medical equipment or ballast in aircrafts) and military (particularly in ammunition), the estimated world’s stock is approximately 1.6 million tons of DU.
After processing, the by-product DU tails present the unique feature that they may be reprocessed and recycled to provide fresh nuclear fuel and reduce the volume of low-level wastes10. Several incentives of re-enrichment ‒ mainly in Russia11 and more recently by the U.S. Department of Energy ‒ have been put forward to recover the residual 235U contained in the DU and produce uranium with 235U natural contents (0.71 atomic %). However, current re-enrichment technology is only economically viable in centrifuge enrichment plants with spare capacity and low operation costs, and it involves high energy consumption and the associated CO2 emissions12. New technological developments pursuing a significant reduction of the environmental impact and greater U recycling-reprocessing would be desirable objectives considering the continuous increase of energy demand and the pressures upon energy sustainability13.
U is a ubiquitous element, present at significant amounts in the Earth’s crust14. U is not biologically linked with any type of life, yet various mechanisms through which U is biotically processed are common in the environment, for instance: biosorption, bioaccumulation, biomineralization, and biotransformation15,16,17,18,19. Consequently, microbial communities can also have dramatic effects on U mobilization/immobilization20,21. However, the study of in vivo and biologically mediated U isotope fractionation constitute research areas still to be explored. Traditionally, natural 235U and 238U variability, i.e., differential isotopic behaviours, has gone unaddressed and is assumed invariant owed to the small relative differences in mass of the isotopes22,23. Driven by the advent of technological advances in analytical measurements, the growing field of isotopic fractionation revealed considerable variations of U isotope ratios in natural settings (e.g., ores, granites, corals, seawater22,23,24). Therefore, significant U isotopic fractionation might take place at the Earth´s surface22 and represent a powerful tool in environmental, geological, marine, life and energy sciences. Biological U isotopic fractionation in nature has been linked to bacteria adept at inducing U(VI) biotic reduction7,25,26,27, as it was recently found that the redox reaction is responsible for the isotopic fractionation and is not related to the U uptake inside the cells26. Biologically U(VI) reduction studies resulted in the accumulation of 238U in the reduced product, except Rademancher et al.27 found 235U enrichment in the resultant U(IV). In another framework, human neuron-like cells in vitro achieved an isotopic fractionation of natural U with preferential intracellular uptake of 235U isotope28.
Hence, natural physicochemical processes leading to isotopic fractionation, both mass dependent and independent29,30, might take place during U uptake by the cells. This raises the question of whether different microalgal species give rise to U isotopic fractionation. Recently, evidence of U fractionation has been obtained during Chlamydomonas cells’ U uptake in a U acid mine drainage medium, suggesting a 235U enrichment31,32. Here, we have studied the U isotopic fractionation during DU uptake in two marine and freshwater Chlorophyta strains. As enrichment of the fissile 235U is expected in the cellular pellet, DU was used to address the potential in U reprocessing. Changes in the 235U/238U ratios of the extracellular and cellular U were investigated for 24 days in an extremophile Chlamydomonas (ChlGS strain) isolated from a U mining pond and a marine Tetraselmis (TmmRU strain). These results represent a potential tool for U recycling and reprocessing and may entail implications in the study of U isotopes in natural samples.
Results
We performed two independent experiments, each with one of the strains of interest, the Chlamydomonas (ChlGS) and Tetraselmis (TmmRU) Chlorophyta strains, in different media supplemented with DU. ChlGS is an extremophile isolated from an acid U mine tailings pond, tolerant up to 25 mg U L−1 and other metals, and artificially selected for U uptake33. The isotopic ratio n(235U)/n(238U) with a value of 0.007375 ± 0.000013 found in the mine water (see Supplementary Fig. S1) was far above the consensus natural abundance 235U/238U [0.007198-0.007202]34, suggesting a possible enrichment process in the U mine pond. Conversely, TmmRU is a seawater strain, only previously exposed to naturally occurring trace U35 and subsequently selected for U tolerance. ChlGP cell replicates were exposed to 4 mg L−1 DU ~ 0.0050 atomic 235U freshwater stock solution and TmmRU to 2 mg L−1 DU ~ 0.0022 atomic 235U marine stock solution. The analytical procedure and sample resin purification validation were accomplished by the analyses of procedure control solutions (certified IRMM-053 material) between samples during the measure sessions to correct the bias induced during the inductively coupled plasma mass spectrometry (ICP-MS) measurements. The average n(235U)/n(238U) values found in the control solution was 0.007112 ± 0.000024 (σ, n = 34) (Fig. S2), these values were used to calculate the discriminatory factor of each sample (for definition, see SI Material and Methods).
Within each bioassay, twelve independent trials were conducted, and the cellular pellet incorporated from 6.1 to 78.5 µg U for ChlGS and from 0.57 to 3.22 µg U for TmmRU (Table 1). ChlGS cells exhibited an increased U uptake performance until day 24, virtually the entire U amount in dissolution. Conversely, TmmRU cells incorporated 1.5–5% of the U present in the exposure solution. DU stock solutions were isotopically characterized before the exposition, and the average n(235U)/n(238U) ratio values found for the stock marine and freshwater solutions were 0.005058 ± 0.000083 (σ, n = 3) and of 0.002172 ± 0.000023 (σ, n = 3), respectively. The n(235U)/n(238U) ratios were determined in the cellular pellets and the extracellular solutions for each independent trial at different times: 3, 12 and 24 days (Figs 1, 2 and Table 1). The IRMM-053 material bracketed the samples for mass bias correction. No significant difference in the n(235U)/n(238U) ratio between days was found in either bioassay (P > 0.5).
Freshwater ChlGS strain bioassay n(235U)/n(238U) (%) ratios and SD values over time. Cellular pellets (red circles) and supernatants (blue triangles) isotopic ratios were analysed by independent assays in groups of four at 3, 12, and 24 days. Solid lines correspond to the mean, and dashed lines correspond to the reproducibility (σ, n = 12), of the n(235U)/n(238U) values in the independent experiments for the cellular pellet (red lines) and supernatant (blue lines). Significant differences in the n(235U)/n(238U) values of supernatants and pellets appeared in the data according to a t-test (t = 6.17, df = 21.49, ***P < 0.001) (software package R version)69.
Marine TmmRU strain bioassay n(235U)/n(238U) ratios and SD values over time. Red circles and blue triangles represent the n(235U)/n(238U) ratios for the cellular pellets and supernatants, respectively, of independent assays analysed at 3, 12 and 24 days. Solid red and blue lines depict the mean n(235U)/n(238U) values obtained for the cellular pellet and supernatant, respectively, in 12 independent experiments. Dashed lines depict the reproducibility (σ, n = 12). Cellular pellets and supernatant values were significantly different according to the Wilcoxon test (W = 0.82, ***P < 0.001) (software package R version)69.
The δ235 (U) values, equation (S3), of each independent experiment were calculated from the n(235U)/n(238U) ratio between the cellular pellet and the supernatant (Table 1). For both microalgal species tested during this investigation, we observed a significant shift in U isotopic composition towards lighter values of δ235. Magnitudes of U removal from the dissolution by cells uptake were of δ235(U) pellet-supernatant ≈ 22.2‒25.5‰ (ChlGS) and 305.2‒445.6‰ (TmmRU). Meanwhile, both studies found microalgal pellets to be considerably enriched in the isotopically light 235U isotope, whilst the supernatants showed a depletion in 235U. This implies an isotopic fractionation towards a lighter composition in the cell pellet in comparison to the supernatant. Additionally, these differences represent a down-blending of the isotope 235U in the cellular adjoin media of the isotope 235U, which necessarily was consumed in the process (Table 1).
Discussion
Our results demonstrate that the two Chlorophyta microalgal strains studied fractionate DU. Each strain displayed a different enrichment factor but both reflect a strong fractionation of the light 235U isotope in the cellular pellet. The marine strain TmmRU, despite presenting higher cell volume and organic weight36,37, presented a lower U uptake rate. However, the enrichment factor was significantly higher than in the freshwater ChlGS, despite being exposed to a lower 235U content DU (~0.0022 235U). The similar U isotopic composition found in the cellular pellets of the different replicates of each strain, irrespective of the culture time and the total U incorporated by the cells, raises the issues of preferential U uptake paths. The pathways leading to U uptake by the cells are not well documented18, and the joint complexity derivate of the simultaneously of several processes28,38 ‒ redox reactions, ligand exchange, diffusion, adsorption ‒ make interpretations of U fractionation origin extremely complex. Furthermore, U speciation and redox state may influence the fractionation process. Whatever the route, as previously described by Baselga-Cervera et al.39, U is bound to the outer wall and transported across the cell wall and membrane, becoming distributed inside the cell. These data suggest that U carried by the cellular pellet accomplishes isotopic partitioning, resulting in 235U being concentrated by the cells and the surrounding environment being enriched in 238U.
Consistent with our results, the isotopic ratio n(235U)/n(238U) value found in the U mine water suggest a possible enrichment process. Only microbial life has been detected in this pond39,40, and therefore biomass does not pass to higher trophic levels. Thus, 235U, as one of the lighter isotopes of U, could preferentially enter cells. When cells break, U enriched in 235U might be liberated to the water. The remaining U enriched in 238U bound to the cell wall might sediment in the bed of the pond. Additionally, bacteria that might be present in this pond and can contribute to this result. Reductive bacteria can induce U fractionation; the reaction products (U(IV)) are enriched in 238U, rendering the residual dissolved U enriched in 235U25,41. Combined effects of different microorganisms may have led to this result.
Microbial isotopic behaviour in elements with higher mass numbers, such as U, is typically poorly studied compared to light elements because of the need for more sophisticated and precise analyses. Recent evidence in the field has demonstrated U isotopic fractionation mediated by bacteria and neuron-like human cells. Biotic reduction studies with metal-reducing bacterial isolates show an enrichment in the heavier 238U isotope into the solid U(IV) byproduct26,42,43 that is not dependent on microbial sorption. Conversely, the isotopic behaviour displayed by neuron-like human cells showed a preferential 235U incorporation28. Paredes et al.28 suggested two possible isotopic fractionation processes based on the enrichment direction and U bioaccumulation: a mass-dependent “zero-point energy” mechanism29 and the mass-independent “nuclear field shift”. In the case of our study, as the two processes are not mutually exclusive and operate in the same fractionation direction, we cannot determine the precise contribution of each mechanism proposed. High-precision determination of other U isotopes, such as 234U ‒ commonly concentrated during 235U enrichment ‒ could provide insights into the fractionation mechanism at the cellular level and the proposed biological preference for lighter isotopes, but it is complex due to the abundances limitation [natural 234U abundance is comprehended between 0.000050–0.000059]. Reported isotope fractionation ratios for 235U/238U during U reduction by bacteria have ranged from −0.31 to −0.99‰26,42,43, and 0.38 ± 0.13‰ in neuron-like line cells29. We found that the cellular pellet was enriched in 235U relative to the supernatant with DU by 23.6 ± 12.5‰ and 370.4 ± 103.9‰ for the ChlGS and TmmRU strains, respectively. This fractionation behaviour is consistent in its direction with that observed in neuron-like cells, but the fractionation factors are significantly higher. One potential application of the observed microalgae-induced U isotopic variation is for DU waste re-enrichment along the U fuel cycle for nuclear fission energy. The current DU stockpiles worldwide would render one-third of natural-equivalent U after several cycles of re-enrichment, reducing DU tail and U-mine production. Currently, only centrifuge separation and gaseous diffusion have operated at commercial scale, even though several enrichment processes have been demonstrated historically and in the laboratory44,45,46,47. Both enrichment processes present important drawbacks: water reactivity by-products rendering hazardous compounds highly corrosive, as well as significant amounts of energy consumption, considerable costs and generate DU as a low-level waste product48. Reprocessing U tails with current techniques, despite its potential, is not economically and energetically feasible and does not ease the problem of final tail disposition49. Particularly interesting would be to develop a viable biological process to enrich DU, recovering natural-equivalent U from tailings waiting for final disposal without high energy demand and the associated carbon footprint. Hypothetically, multiple stages of microalgal bioaccumulation of DU, cell harvesting and U resuspension into the next higher step would enrich the DU to the desired amount, up to the 235U natural content or even above. The predicted advantages of this biological process are the reduced cost and low energy requirements, turning DU in a resource that even the U-tail problem decreases only marginally. Further experimentation may resolve the expected value range for U fractionation mediated by microalgal bioaccumulation and the efficiency of the enrichment process. In addition, other microalgal species might display a different isotopic behaviour, considering the significant differences exhibited by the two Chlorophyta species.
Our findings might also have implications for the identification of biotic signatures using isotope tools to study ancient microbial life and understand global uranium flux. Significant uncertainties remain regarding isotopic signatures owing to the ambiguity in the interpretation of the signs. Development of analytical techniques has opened the possibility of studying small U isotope natural variation. Uranium is the heaviest element for which natural variations have been reported50. U isotopic fractionation in nature occurs, with opposite directions, in both anoxic/euxinic and oxic environments and can be associated with chemical transformations such as adsorption, speciation or redox chemistry22. Microorganisms probably have a part in the U cycling and deposit formation in nature51,52,53. The large isotope fractionation that occurs during microalgal U uptake suggests the role of microalgae in the conservative behaviour of U and a tight range of isotope composition in modern oceans. For geological implications, recent studies of biotic redox transformation of U with metal-reducing bacteria induced U isotopic fractionation enriching for 238U in the reaction products25,26, contradicting previous studies27 and consistent with environmental studies and U-reduced depositional samples22,41,43,53,54. U isotopic fractionation mediated by bacterial enzymatic reduction raises as a tangible biosignature in the rock record for specific metabolic groups and onto the timing emergence of specific metabolisms. In oxic sedimentary environments, sample fractionation occurs towards lighter isotope composition, and intriguingly, banded-iron formation samples present the lightest U composition studied22. The banded-iron formations’ lighter values could indicate microbial phototroph co-precipitation by adsorption of iron and U, supporting the previously suggested microbial implication55. Thus, the preferential accumulation of the fissile 235U isotope in sediments might be a proxy for the activity and presence of microalgae. U fractionation biological fingerprints in ancient sedimentary rocks would provide insights into ancient microbial activity and establish temporal constraints. Additionally, our findings might provide insights into the Oklo phenomenon on the assumption of a microbial contribution in the initiation of the natural nuclear reactors40,56,57. Several factors probably contribute to these unique chain reactions. Two thousand million years ago – the same time proposed for Oklo’s event58,59,60,61 – 235U made up approximately 3% of the natural U, a condition that makes possible the starting of a fission reaction. However, a U-rich mineral deposit needs to be formed to obtain a critical mass. Most likely, the presence of increasing oxygen in the Earth’s atmosphere enabled the U flux and subsequent concentration in U ore bodies62. The direction and magnitude of the observed microalgal U isotopic fractionation during bioaccumulation could support the biological hypothesis of the origin of the natural reactors.
Methods
Microalgal species and culture conditions
We used two Chlorophyta, a freshwater environmental extremophile, Chlamydomonas saelices nov. sp. (strain ChlSG) isolated from a uranium mining pond in Saelices el Chico, Salamanca, Spain (previously described in depth by García-Balboa et al.39 and Baselga-Cervera31), and the marine Tetraselmis mediterranea (Lucksch) R.E. Norris, Hori & Chihara (strain TmmRU) isolated in the east-coastal waters of Sardinia (Italy). Both strains were selected for U tolerance and placed in the Spanish Algae Bank (Spanish acronym BEA) with the access numbers BEA/D04-12 and BEA/IDA/0062 for ChlSG and TmmRU, respectively63,64. Strains were propagated asexually in cell culture flasks (Greiner; Bio-One Inc., Longwood, NJ, USA) with BG-11 medium (Sigma-Aldrich Chemie, Taufkirchen, Germany)) for ChlGS and Guillard’s F/2 medium (Sigma-Aldrich Chemie, Taufkirchen, Germany) for TmmRU (media were prepared according to the manufacturers’ directions). Strains were sub-cultured by serial transfers to fresh medium once every 20 days to ensure mid-log-exponential growth, and the axenicity was regularly tested. Culture conditions were continuous illumination at 80 µmol m−2 s−1 over the wavelength range from 400 to 700 nm and a temperature of 22 °C.
Prior to experiments, both microalgal cultures of ChlGS and TmmRU were analysed to ensure the absence of U before exposure to DU. No U was detected in either of the cultures.
Exposure experiments to DU
Here, we studied the U fractionation behaviour of two microalgal species artificially selected for U recovery63,64: the freshwater extremophile Chlamydomonas (strain ChlGS) isolated from a well-studied U acid mine pond31,39 and the marine Tetraselmis (strain TmmRU) isolated from the Mediterranean sea only sparsely exposed to naturally occurring trace 235U. Two different sets of experiments were performed with two different strains ChlGS and TmmRU. In both bioassays, twelve replicates were established in cell culture flasks with 20 mL of depleted uranium medium with approximately 2 × 10 6 cells and 0.5 × 10 6 cells per mL for ChlGS and TmmRU, respectively. In the case of ChlGS, the depleted uranium medium consisted in a preparation of 4 mg L‒1 of U with an isotopic relation n(235U)/n(238U) of 0.005058 ± 0.000083 in bi-distilled water sterilized and enriched with BG-11. In the TmmRU bioassay, the depleted uranium medium consisted of a preparation of 2 mg L‒1 of U with an isotopic ratio n(235U)/n(238U) of 0.002172 ± 0.000023 in bi-distilled water sterilized and enriched with F/2. Our experiments were performed in aqueous systems, oxidizing conditions and below Σ pH 7–8, conditions were the mobile U(VI) is predominantly found as uranyl ions (UO2+2) or hydroxyl complexes18,65,66. pH was selected at the onset of the microalgal fractionation experiments to have predominantly soluble U species, so ligands of the cell walls would attract U cations though chemical sorption67. The U concentration for TmmRU was obtained from the dose-response curve and represented the higher dose before the IC50 values for growth inhibition. In ChlGS strain, the U concentration used was selected based upon previous U uptake experiments33. At different times along the cells’ growth curve (3, 12 and 24 days), four independent cultures were centrifuged at 4000 rpm for 15 minutes, and the two phases obtained (supernatant and pellet) of each replicate were frozen. Throughout the experiments, due to the media alkalization induced by the cells’ photosynthetic activity, some U precipitation could take place.
The experimental media were supplied by the Energy, Environmental and Technological Research Center (Spanish acronym CIEMAT). All the samples were sent to CIEMAT for isotopic analysis and total U analyses. The isotopic relation results were obtained by averaging the data obtained in three analyses.
Uranium fractionation characterization
Algal pellets (mg) and filtered supernatants (5 mL) were treated by microwave acid digestion with 5 mL of 65% HNO3 and 30% H2O2 (4:1 v/v). For pellet samples, before microwave treatment, the sample-reagent mixture was sonicated. After digestion, the sample solution was evaporated almost to dryness, and the residue was dissolved to 5 mL with 3 M HNO3. One millilitre of this solution was brought to a final volume of 10 mL with MilliQ water to quantify the U by quadrupole-based ICP-MS (Q-ICP-MS) using the external calibration and internal standard method. The remaining volume was used for isotopic analysis after purifying U with UTEVA resin according to revised protocols reported by Carter et al.68 and Weyer et al.22, summarized as follows. The sample solution was loaded onto a previously washed and pre-conditioned column. The column was rinsed with 2 × 5 mL of 3 M HNO3, 6 mL of 9 M HCl and 5 × 3 mL of 5 M HCl. Uranium was eluted in 5 × 3 mL of 0.02 M HCl. The collected uranium fraction was evaporated and the residue re-dissolved in 2 mL of concentrated HNO3, and the uranium was again evaporated. This last residue was dissolved in HNO3 2%. Matrix separation and U concentration were checked in the fractions collected by Q-ICP-MS (iCAP Q, Thermo Scientific). Procedural blanks of the entire sample treatment procedure, including digestion and purification, were performed in every sample batch.
Due to the high uranium content, purified samples were diluted with HNO3 2% before isotope ratio measurements to a concentration that matched that of the isotopic standard IRMM-053.
Isotope ratios were determined in a double-focusing sector field ICP-MS (Element 2, Thermo Scientific) equipped with a single collector and using a desolvating system (Cetac Aridus II) for sample introduction. This unit stabilized the ion beam and provided a 5-fold enhancement in sensitivity (10 × 106 cps per ng.g-1). Measurements were carried out under optimized conditions of the overall system to obtain the maximum accuracy and precision in isotopic measurements (for detail information of the procedure validation, see SI Material and Methods). For data acquisition, standard bracketing was used placing every sample between two isotopic standards (IRMM-053) and washing with HNO3 2% before every sample and standard (see experimental settings in Supplementary Table S1).
Potential isotope fractionation due to chromatographic extraction was evaluated by measuring the 235U/238U ratio in the isotopic CRM as well as in the DU before and after passing through the column using the same elution protocol as in the samples.
Data Availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information Files).
References
Karakosta, C., Pappas, C., Marinakis, V. & Psarras, J. Renewable energy and nuclear power towards sustainable development: Characteristics and prospects. Renew. Sustain. Energy Rev. 22, 187–197 (2013).
Gitzel, T. & Bowden, P. World Nuclear Association. Chernobyl Accident 1986 (2001).
Brook, B. W. et al. Why nuclear energy is sustainable and has to be part of the energy mix. Sustain. Mater. Technol. 1–2, 8–16 (2014).
Lu, Y. Uranium extraction: Coordination chemistry in the ocean. Nat. Chem. 6, 175–177 (2014).
World Nuclear Association. Uranium Supplies: Supply of Uranium - World Nuclear Association. Available at, http://www.world-nuclear.org/information-library/nuclear-fuel-cycle/uranium-resources/supply-of-uranium.aspx. (Accessed: 20th November 2017) (2016).
Liu, C. et al. A half-wave rectified alternating current electrochemical method for uranium extraction from seawater. Nat. Energy 2, 17007 (2017).
Czerwinski, K. & Polz, M. Uranium enrichment using microorganisms. US Patent 7, 452–703 (2008).
World Nuclear Association. World Nuclear Power Reactors 2007-08 and Uranium Requirements. World Nucl. Assoc. 1–7 (2009).
Kok, K. D. Nuclear engineering handbook. (CRC Press, 2009).
Netter, P. Reprocessing of spent oxide fuel from nuclear power reactors. in Nuclear Fuel Science and Engineerin 459–504 (2002).
Diehl, P. Re-enrichment of West European Depleted Uranium Tails in Russia. WISE Uranium Proj. (2004).
NEA, O. & IAEA. Management of Depleted Uranium: a joint report. (2001).
Dang, D. H., Novotnik, B., Wang, W., Georg, R. B. & Evans, R. D. Uranium Isotope Fractionation during Adsorption, (Co)precipitation, and Biotic Reduction. Environ. Sci. Technol. 50, 12695–12704 (2016).
Markich, S. J. Uranium Speciation and Bioavailability in Aquatic Systems: An Overview. Sci. World J. 2, 707–729 (2002).
Fortin, C., Dutel, L. & Garnier-Laplace, J. Uranium Complexation and uptake by a green alga in raltion to chemical speciation; the importance of the free uranyl ion. Environ. Toxicol. Chem. 23, 974 (2004).
Li, X., Hu, H., Yu, J. & Zhao, W. Selection of Suitable Microalgal Species for Sorption of Uranium in Radioactive Wastewater Treatment. Huan Jing Ke Xue 37, 1858–63 (2016).
Manikandan, N., Prasath, C. S. S. & Prakash, S. Biosorption of uranium and thorium by Marine micro algae. 40, 121–124 (2011).
Newsome, L., Morris, K. & Lloyd, J. R. The biogeochemistry and bioremediation of uranium and other priority radionuclides. Chem. Geol. 363, 164–184 (2014).
Kalinowski, B. E. et al. Microbial leaching of uranium and other trace elements from shale mine tailings at Ranstad. Geoderma 122, 177–194 (2004).
Suzuki, Y., Kelly, S. D., Kemner, K. M. & Banfield, J. F. Radionuclide contamination: Nanometre-size products of uranium bioreduction. Nature 419, 134–134 (2002).
Petrie, L., North, N. N., Dollhopf, S. L., Balkwill, D. L. & Kostka, J. E. Enumeration and characterization of iron(III)-reducing microbial communities from acidic subsurface sediments contaminated with uranium(VI). Appl. Environ. Microbiol. 69, 7467–79 (2003).
Weyer, S. et al. Natural fractionation of 238U/235U. Geochim. Cosmochim. Acta 72, 345–359 (2008).
Stirling, C. H., Andersen, M. B., Potter, E. K. & Halliday, A. N. Low-temperature isotopic fractionation of uranium. Earth Planet. Sci. Lett. 264, 208–225 (2007).
Keegan, E. et al. The provenance of Australian uranium ore concentrates by elemental and isotopic analysis. Appl. Geochemistry 23, 765–777 (2008).
Basu, A., Sanford, R. A., Johnson, T. M., Lundstrom, C. C. & Löffler, F. E. Uranium isotopic fractionation factors during U(VI) reduction by bacterial isolates. Geochim. Cosmochim. Acta 136, 100–113 (2014).
Stylo, M. et al. Uranium isotopes fingerprint biotic reduction. Proc. Natl. Acad. Sci. USA 112, 5619–24 (2015).
Rademacher, L. K. et al. Experimentally determined uranium isotope fractionation during reduction of hexavalent U by bacteria and zero valent iron. Environ. Sci. Technol. 40, 6943–6948 (2006).
Paredes, E. et al. Evidence of isotopic fractionation of natural uranium in cultured human cells. Proc. Natl. Acad. Sci. USA 113, 14007–14012 (2016).
Wiederhold, J. G. Metal Stable Isotope Signatures as Tracers in Environmental Geochemistry. Environ. Sci. Technol. 49, 2606–2624 (2015).
Abe, M., Suzuki, T., Fujii, Y., Hada, M. & Hirao, K. An ab initio molecular orbital study of the nuclear volume effects in uranium isotope fractionations. J. Chem. Phys. 129, 164309 (2008).
Baselga-Cervera, B. Estudios de evolución rápida en microorganismos bajo condiciones selectivas y de cambio global: algunos modelos y aplicaciones en biología [dissertation’s thesis]. (Universidad Complutense de Madrid, 2017).
Costas, E., Baselga, B. & Tarin, F. A new process for the fractionation of uranium. Nucl. España 360, 74–77 (2015).
Baselga-Cervera, B., Romero-López, J., García-Balboa, C., Costas, E. & López-Rodas, V. Improvement of the Uranium Sequestration Ability of a Chlamydomonas sp. (ChlSP Strain) Isolated From Extreme Uranium Mine Tailings Through Selection for Potential Bioremediation Application. Front. Microbiol. 9, 523 (2018).
Condon, D. J., Mclean, N., Noble, S. R. & Bowring, S. A. Isotopic composition (238U/235U) of some commonly used uranium reference materials. Geochim. Cosmochim. Acta 74, 7127–7143 (2010).
Davies, R. V., Kennedy, J., McIlroy, R. W., Spence, R. & Hill, K. M. Extraction of uranium from sea water. Nature 203, 1110–1115 (1964).
Creswell, L. Phytoplankton Culture for Aquaculture Feed. SRAC Publ. 1–16, https://doi.org/10.1039/b914312b (2010).
Lavens, P. & Sorgeloos, P. Manual on the Production and Use of Live Food for Aquaculture. (Food and Agriculture Organization of the United Nations, 1996).
Kafantaris, F.-C. A. & Borrok, D. M. Zinc isotope fractionation during surface adsorption and intracellular incorporation by bacteria. Chem. Geol. 366, 42–51 (2014).
Baselga-Cervera, B., López-Rodas, V., García-Balboa, C. & Costas, E. Microalgae: the first nuclear engineers? An. Real Acad. Farm. 79, 634–645 (2013).
García-Balboa, C. et al. Rapid adaptation of microalgae to bodies of water with extreme pollution from uranium mining: An explanation of how mesophilic organisms can rapidly colonise extremely toxic environments. Aquat. Toxicol. 144–145, 116–123 (2013).
Bopp, C. J., Lundstrom, C. C., Johnson, T. M. & Glessner, J. J. G. Variations in 238U/235U in uranium ore deposits: Isotopic signatures of the U reduction process? Geology 37, 611–614 (2009).
Basu, A. Isotopic Fractionation of Chromium and Uranium During Abiotic and. (University of Illinois, 2013).
Stirling, C. H., Andersen, M. B., Warthmann, R. & Halliday, A. N. Isotope fractionation of 238U and 235U during biologically-mediated uranium reduction. Geochim. Cosmochim. Acta 163, 200–218 (2015).
Seko, M., Miyake, T., Inada, K. & Takeda, K. Uranium Isotope Enrichment by Chemical Method. Nucl. Technol. 50, 178–186 (1980).
Ling, L. & Zhang, W. Enrichment and Encapsulation of Uranium with Iron Nanoparticle. J. Am. Chem. Soc. 137, 2788–2791 (2015).
Morel, J., Etcheverry, M. & Riazuelo, G. Uranium enrichment measurement by X- and γ-ray spectrometry with the “URADOS” process. Appl. Radiat. Isot. 49, 1251–1257 (1998).
Maomi, S., Hunihiko, T. & Tetsuya, M. The chromatographic uranium enrichment process by Asahi chemical. AIChE Symp. Ser.; (United States) 78, 221 (1982).
World Nuclear Association. Uranium Enrichment | Enrichment of uranium - World Nuclear Association. World Nuclear Association Available at, http://www.world-nuclear.org/information-library/nuclear-fuel-cycle/conversion-enrichment-and-fabrication/uranium-enrichment.aspx#.UWrver-IRAs. (Accessed: 21st November 2017) (2016).
Diehl, P. Re-enrichment of depleted uranium tails in Gaseous Diffusion Plants. (2007).
Hoefs, J. Stable isotope geochemistry. (Springer, 2015).
Anderson, D. M. & Keafer, B. A. An endogenous annual clock in the toxic marine dinoflagellate Gonyaulax tamarensis. Nature 325, 616–617 (1987).
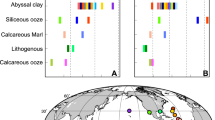
Zheng, Y., Anderson, R. F., Geen, A., Van & Fleisher, M. Q. Preservation of particulate non-lithogenic uranium in marine sediments. Geochemica Cosmochim. Acta 66, 3085–3092 (2002).
Brennecka, G. A., Borg, L. E., Hutcheon, I. D., Sharp, M. A. & Anbar, A. D. Natural variations in uranium isotope ratios of uranium ore concentrates: Understanding the 238U/235U fractionation mechanism. Earth Planet. Sci. Lett. 291, 228–233 (2010).
Bopp, C. J. et al. Uranium 238 U/235 U Isotope Ratios as Indicators of Reduction: Results from an in situ Biostimulation Experiment at Rifle, Colorado, USA. Environ. Sci. Technol. 44, 5927–5933 (2010).
Kappler, A., Pasquero, C., Konhauser, K. O. & Newman, D. K. Deposition of banded iron formations by anoxygenic phototrophic Fe(II)-oxidizing bacteria. Geology 33, 865 (2005).
Nagy, B. et al. Role of organic matter in the Proterozoic Oklo natural fission reactors, Gabon, Africa. Geology 21, 655–658 (1993).
Dutkiewicz, a, George, S. C., Mossman, D. J., Ridley, J. & Volk, H. Oil and its biomarkers associated with the Palaeoproterozoic Oklo natural fission reactors, Gabon. Chem. Geol. 244, 130–154 (2007).
Roth, E. The discovery and study of the nuclear reactor in Oklo. J. Radioanal. Chem. 37, 65–78 (1977).
Fujii, Y. et al. The nuclear interaction at Oklo 2 billion years ago. Nucl. Phys. B 573, 377–401 (2000).
Gauthier-Lafaye, F., Weber, F. & Ohmoto, H. Natural fission reactors of Oklo. Economic Geology 84, 2286–2295 (1989).
Flambaum, V. V. & Shuryak, E. V. Limits on cosmological variation of strong interaction and quark masses from big bang nucleosynthesis, cosmic, laboratory and Oklo data. Phys. Rev. D - Part. Fields, Gravit. Cosmol. 65, 11 (2002).
Gauthier-Lafaye, F., Holliger, P. & Blanc, P.-L. Natural fission reactors in the Franceville basin, Gabon: A review of the conditions and results of a “critical event” in a geologic system. Geochim. Cosmochim. Acta 60, 4831–4852 (1996).
Costas Costas, E., López Rodas, V., García Balboa, C., De Miguel Fernandez, L. & Baselga Cervera, B. Método de recuperación y enriquecimiento de uranio mediante bioacumulación en microalgas mejoradas genéticamente. WO patent (2012).
López Rodas, V., Costas Costas, E., García Balboa, C. & Romero López, J. Microorganismo de la especie Tetraselmis mediterránea (CEPA TmmRU) y su uso para la producción de uranio enriquecido. Spanish patent (2017).
Choppin, G., Liljenzin, J.-O., Rydberg, J. & Ekberg, C. Behavior of Radionuclides in the Environment. In Radiochemistry and Nuclear Chemistry 753–788 (2013).
Zavodska, L., Kosorinova, E., Scerbakova, L. & Lesny, J. Environmental chemistry of uranium. HVISSN 1418–7108 HU ISSN 1418-7108: HEJ Manuscript no: ENV-081221-A (2008).
Kalin, M., Wheeler, W. N. & Meinrath, G. The removal of uranium from mining waste water using algal/microbial biomass. J. Environ. Radioact. 78, 151–177 (2004).
Carter, H. E., Warwick, P., Cobb, J. & Longworth, G. Determination of uranium and thorium in geological materials using extraction chromatography. Analyst 124, 271–274 (1999).
R Development Core Team. R. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing 1, 409 (2011).
Acknowledgements
We thank Prof. Mª T. Miras-Portugal for her comments, to J. L. Valle from Westinghouse and F. Tarín from ENUSA (Empresa Nacional del Uranio S.A.) for proof-reading the manuscript and discussion. Thanks are given to L. de Miguel and Eva Salgado for their technical support. We are also grateful to CIEMAT institution (Centro de Investigaciones Energéticas, Medioambientales y Tecnológicas) and especially to Abel Yllera de Llano, Ana Isabel Barrado y Estefania Conde Vilda. We acknowledge the national founding of Spanish Secretaría de Estado, Investigación, Desarrollo e Innovación (grant CTM-2013-44366-R).
Author information
Authors and Affiliations
Contributions
B.B., C.G., V.L. and E.C. conceived the presented idea and the experiments. B.B., C.G., V.L. and E.C. carried out the experiments. M.F. contributed to reagents/materials/analysis tools and samples analyzed. B.B., C.G., V.L., M.F. and E.C. contributed to the interpretation of the results. B.B., M.F. and C.G. wrote the manuscript supervised by V.L. and E.C. B.B., C.G., M.F., V.L. and E.C. made important intellectual contributions.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baselga-Cervera, B., García-Balboa, C., López-Rodas, V. et al. Evidence of microalgal isotopic fractionation through enrichment of depleted uranium. Sci Rep 9, 1973 (2019). https://doi.org/10.1038/s41598-019-38740-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-38740-2
This article is cited by
-
An exploratory study on the possibilities of microalgal biotechnology to obtain the essential 6Li isotope as fusion fuel
Biotechnology for Biofuels and Bioproducts (2023)
-
Rapid Colonization of Uranium Mining-Impacted Waters, the Biodiversity of Successful Lineages of Phytoplankton Extremophiles
Microbial Ecology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.