Abstract
Biocathodes where living microorganisms catalyse reduction of CO2 can potentially be used to produce valuable chemicals. Microorganisms harbouring hydrogenases may play a key role for biocathode performance since H2 generated on the electrode surface can act as an electron donor for CO2 reduction. In this study, the possibility of catalysing cathodic reactions by hydrogenotrophic methanogens, acetogens, sulfate-reducers, denitrifiers, and acetotrophic methanogens was investigated. The cultures were enriched from an activated sludge inoculum and performed the expected metabolic functions. All enrichments formed distinct microbial communities depending on their electron donor and electron acceptor. When the cultures were added to an electrochemical cell, linear sweep voltammograms showed a shift in current generation close to the hydrogen evolution potential (−1 V versus SHE) with higher cathodic current produced at a more positive potential. All enrichment cultures except the denitrifiers were also used to inoculate biocathodes of microbial electrolysis cells operated with H+ and bicarbonate as electron acceptors and this resulted in current densities between 0.1–1 A/m2. The microbial community composition of biocathodes inoculated with different enrichment cultures were as different from each other as they were different from their suspended culture inoculum. It was noteworthy that Methanobacterium sp. appeared on all the biocathodes suggesting that it is a key microorganism catalysing biocathode reactions.
Similar content being viewed by others
Introduction
Limitation of conventional energy sources and their impact on our climate, ecosystems and health has motivated a development of technologies that harvest renewable energy and allow environmentally friendly production of chemicals. The microbial electrolysis cell (MEC) is a modern, sustainable and promising approach to decrease electrode overpotentials, reduce the need for expensive metals catalysts, and allow direct generation of energy carriers at the cathode. MECs combine electrochemical systems with the catalytic ability of microorganisms for production of hydrogen1,2,3,4 and other valuable chemicals such as acetate5,6, methane7,8,9, caproate10, and alcohols11.
In MECs, microorganisms catalyse reactions on the anode, the cathode, or both electrodes simultaneously. For example, microorganisms could oxidize organic compounds and deliver electrons to the anode and other microorganisms present on the cathode could catalyse the reduction of hydrogen ions to hydrogen gas12. An external input voltage is applied to drive the reactions since the overall redox reaction is thermodynamically unfavourable. Microorganisms have different mechanisms for transferring electrons to or from electrodes. Some microorganisms have electrochemically active redox proteins on their outer membrane which can transfer electrons directly to the electrode. For example, Kim, et al.13 showed that Shewanella purefaciens could oxidize lactate and transfer electron to solid electrode in absence of a mediator. Reguera, et al.14 showed that Geobacter sulfurreducens produced conductive pili, nanowires, which could be used for transferring electrons from the cell surface to the surface of Fe(III) oxides. Rabaey, et al.15 showed that Pseudomonas aeruginosa produced soluble redox mediators, electron shuttles, which could be used by themselves or by other bacteria to enhance electron transfer between the cells and solid electrodes.
On a cathode, hydrogen can be produced abiotically3, biotically1, or enzymatically16. Microorganisms that produce hydrogen contain hydrogenases that catalyse the reversible reaction of 2 H+ + 2e− ↔ H2. It was previously shown that purified hydrogenases can enhance hydrogen production on a carbon electrode17,18,19. However, the enzymes are very unstable and usually lose their catalytic activity over time. Therefore, using whole cells can help to improve the stability of the system and enhance the hydrogen production reactions. Rozendal, et al.1 was the first to investigate the possibility of producing hydrogen using the ability of microorganisms for taking up electrons from the cathode in a MEC. They showed that bioanodes enriched on acetate and hydrogen could catalyse hydrogen production when the polarity of the electrodes was reversed.
Several microorganisms in pure cultures have been shown to catalyse cathode reactions in MECs. For example, Geobacter sulfurreducens was shown to catalyse hydrogen production at a potential between −0.8 and −1.0 V versus Ag/AgCl20. Desulfovibrio spp. have also been shown to catalyse H2 production21 and Methanobacterium spp. appear capable of catalysing reduction of CO2 to CH4 with a cathode as electron donor22. In enrichment cultures, many microbial taxa have been found on biocathodes and it is unclear which are involved in catalysing the electrochemical reactions. For example, on H2-producing cathodes, Croese, et al.23 described a community consisting of 46% Proteobacteria, 25% Firmicutes, and 17% Bacteroidetes. Desulfovibrio spp. appeared to play a key role in this community. Batlle-Vilanova, et al.24 enriched biocathodes dominated by Hoeflea sp. and Aquiflexum sp. Methanobacterium spp. have been observed on CH4-producing biocathodes8,25, and Acetobacterium spp. and Acetoanaerobium spp. have been observed on acetate-producing biocathodes25,26,27,28. These microorganisms are known hydrogenotrophs, which suggest that H2 is acting as an intermediate in electrode-attached biofilms producing CH4 and acetate. It is unclear, however, if these or other microorganisms in the diverse cathode communities, or even free enzymes29, catalyse H2 production on the cathode.
Biocathodes are usually started-up using a mixed culture inoculum, e.g. from wastewater and anaerobic digester sludge. However, the enrichment process is slow and can take several months25. One strategy to facilitate start-up is to pre-enrich a suitable microbial community and use it as inoculum. Previous research suggest hydrogenotrophic microorganisms are important on biocathodes and some studies have used pre-enriched H2-oxidizing bioanodes1,30 and hydrogenotrophic methanogens31 with promising results. However, detailed information about the microbial community composition in the inoculum and studies on which microorganisms in the inoculum are retained on the biocathode are lacking. Many different groups of hydrogenotrophic microorganisms exist and there is a lack of knowledge about how widespread the ability to catalyse biocathode reactions is.
The first goal of this study was to screen for the ability of four hydrogenotrophic cultures, enriched using different electron acceptors, to catalyse cathode reactions. One acetate-oxidizing methanogenic culture was also tested. The other goal was to investigate current generation and change in microbial community composition when some of the enrichment cultures were used as inoculums for biocathodes in MECs.
Results
Enrichment cultures
Growth
The change in optical density in the different enrichment cultures is shown in Figs 1–4. The cells grew quickly during the first 20 days of the experiment, except for one of the acetotrophic methanogenic enrichments (MgenA2), which had the slowest growth among the enrichments (Fig. 4). The sulfate-reducers (SR 1-2) had the fastest growth rate among the enrichments and started after 5 days (Fig. 3A). After approximately 70 days, the optical density reached to more stable values for all the enrichments suggesting that the cultures had reached a stationary phase where growth equalled decay (Figs 1–4). One of the hydrogenotrophic methanogenic enrichments (MgenH1) had an increased biomass concentration around day 90 due to removal of 140 mL for inoculation of a MEC (Fig. 1A).
Hydrogenotrophic methanogens (MgenH)
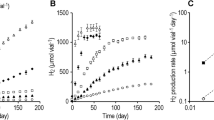
Hydrogen consumption rate and methane production rate are shown in Fig. 1B. Hydrogen was consumed with a rate of 0.6 mmol e−/d immediately after inoculation in both enrichments. The consumption rate increased slightly after 10 days for the next four weeks. Then, the rate varied between 0.8–4 mmol e−/d until the end of the experiment. The consumption rate was limited by the volume of H2 added during feeding since, in general, most of the H2 was consumed between two feeding occasions. In the beginning, feeding took place twice a week and in the end, once a week. MgenH1 had higher consumption rate at the end of the experiment because it contained a larger gas volume when 140 ml of liquid had been removed for inoculating an MEC. Both MgenH enrichments started to produce methane at a rate of approximately 0.8 mmol e−/d after 20 days. Then, the methane production rate varied between 0.2–2 mmol e−/d for the rest of the experiment. Very low amounts of acetate were detected in both enrichment cultures during the run.
Acetogens (Agen)
Figure 2 shows optical density, H2 consumption and volatile fatty acid (VFA) production rates, and concentrations of acetate and butyrate (the two major products) in the enrichments. VFA production rate was at its highest peak after 20 days (>2 mmol e−/d). Then, the production rate decreased for the next 40 days. Comparing the hydrogen consumption and VFA production shows that when hydrogen was consumed, VFA was produced. In the first 100 days of the run, acetate accumulated up to a concentration of about 70 mM-C. Then, the acetate concentration remained around 60–80 mM-C for the rest of run. Butyrate started to be produced at day 120 reaching a concentration of 20 mM-C in both enrichments at the end of the experiment.
Sulfate-reducers (SR)
Optical density, SO4−2 and H2 consumption rates in SR1 and SR2 are shown in Fig. 3A,B. In the SR enrichment cultures, sulfate was consumed at a rate of 1–1.5 mmol e−/d in the first 4 days after inoculation while the H2 consumption rate was between 1.5–2 mmol e−/d. Thereafter, sulfate consumption rate increased gradually over time, suggesting that the microorganisms capable of sulfate reduction increased in abundance in the enrichments. The same trend was observed for H2 consumption. On every occasion after addition of fresh medium to the enrichments, the sulfate consumption rate reached its highest rate. Then, SO4−2 consumption rate decreased gradually over the time as sulfate was consumed. The reason that the SO4−2 consumption rate is higher than the H2 consumption rate could be that the electron equivalent consumption was calculated considering only reduction of SO4−2 to H2S. However, in reality less reduced reaction products such as S0 may have been formed.
Nitrate-reducers (NR)
Figure 3C,D shows optical density, H2 and NO3− consumption rate in the nitrate-reducing enrichments. The NR enrichments started to consume nitrate at very low rate (<0.5 mmol e−/d) directly after inoculation. Then, NO3− and H2 consumption rates increased slightly up to 3.8–6.2 mmol e−/d and 1.8–2.2 mmol e−/d respectively at day 20 which showed that denitrifying microorganisms grew in the enrichments. The nitrate consumption rate was calculated based on the assumption that all nitrate was converted to nitrogen gas; however, nitrate and nitrous oxide could be also have been produced. Therefore, the amount of electron equivalents that were consumed is higher than the consumption rate of H2. Consumption of NO3− in both NR enrichments increased significantly whenever fresh nutrient medium was supplied. Then, the rate was decreased over the time until all the NO3− was consumed.
Acetotrophic methanogens (MgenA)
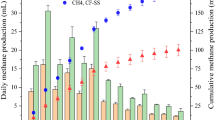
Optical density, acetate consumption rate, and methane production rate in the MgenA1 and MgenA2 enrichments are shown in Fig. 4. Significant increase in acetate consumption rate (1.2 mmol e−/d) after 5 days in MgenA1 shows that the microorganisms started to grow from the beginning of the experiment when acetate was the only electron donor available. However, in MgenA2, acetate consumption did not occur during the first 20 days (<0.3 mmol e−/d). This is also confirmed by the optical measurements, which showed a slow growth in MgenA2 during the first 20 days. In both enrichments, acetate consumption rate was at its peak when the new fresh medium, containing 20 mM acetate, was added. The rate gradually decreased over the time when the acetate was consumed. Methane started to be produced in MgenA1 enrichment after 15 days (1.2 mmol e−/d) which was faster compared to MgenA2 that started after 55 days (0.2 mmol e−/d). At the end of the experiment, the methane production rate in both MgenA cultures was significantly higher than the other methanogenic enrichments (MgenH 1–2). At the end of the experiment, methane production rate in MgenA2 was higher compared to MgenA1 because of 140 mL of the liquid had been removed from MgenA1 to inoculate an MEC.
Linear Sweep Voltammetry (LSV) and exchange current
Bioelectrochemical activity of the enrichments was evaluated using LSV. The exchange current density (i0) was calculated based on the Tafel equation for overpotentials greater than 0.4 V versus open circuit potential (OCP). Exchange current density for the different enrichments is showed in Figure S1. It increased after about one month in all enrichments except for NR and MgenA1 which increased after 47 and 96 days, respectively. In most of the subsequent tests, i0 in the presence of the microorganisms had higher value compared to the control. The improvement in exchange current density in the presence of microorganisms showed the ability of microorganisms to catalyse reduction reactions on a cathode. Linear sweep voltammograms of each enrichment at each time point results are shown in Figure S2.
Operation of MECs
Current production in MECs
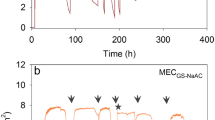
Four different MECs, inoculated with microorganisms from the MgenH1, Agen1, SR1, and MgenA1 enrichments, were operated over 8 weeks in order to investigate the catalytic ability of selected enrichments on a cathode over time. Figure 5 shows the current that was generated. The MgenHMEC generated a current density of about 0.2 A/m2 directly after inoculation followed by an increase up to 0.4 A/m2 during the next 6 days. However, the current dropped to 0.1 A/m2 for the next 5 days, followed by an increase to approximately 0.6 A/m2 after 29 days from the start-up. After 49 days, the potential was switched from −0.65 V to −0.8 V versus SHE and the current increased up to 0.8 A/m2. In AgenMEC, the current was generated directly after inoculation with a rate of about 0.2 A/m2 for the first 6 days. For the next 44 days, the current increased up to around 0.8 A/m2 before changing the potential from −0.65 V to −0.8 V versus SHE. After lowering the potential, the current increased up to around 1 A/m2. In MgenAMEC, current was generated at 0.1 A/m2 directly after inoculation followed by an increase up to 0.3 A/m2 after 4 days. Then, the current decreased to around 0.02 A/m2 for next 8 days. However, 12 days after inoculation, the current started to increase, and it reached to 0.8 A/m2. The current was varied between 0.8 to 0.9 A/m2 until the potential was switched from −0.65 V to −0.8 V versus SHE. After switching the potential, current generation increased to 1 A/m2 and varied between 0.8 to 1 A/m2 till the end of the experiment. The SRMEC behaved differently to the other MECs. The current reached up to 1 A/m2 already in the first 7 days after the inoculation. However, it dropped to 0.2 A/m2 before an increase again to a more stable value at around 0.7 ± 0.05 A/m2 for the rest of the experiment. Decreasing the potential from −0.65 V to −0.8 V versus SHE did not have a noticeable effect on current generation in SRMEC as opposed to the other MECs in this study.
Low concentrations of butyrate and lactate representing 4–5% of the charge passed in the MEC were observed in the SRMEC but not in the other MECs. Concentrations of other VFAs (formate, acetate, and propionate) were negligible. Hydrogen and methane were likely produced but could not be quantified, possibly because of small leakages through electrode- and membrane connections.
Cyclic Voltammetry (CV)
Figure 6 shows CV tests that were carried out during the operation of the MECs in order to evaluate the bioelectrochemical activity of the biocathodes. In all MECs, biological activity on the biocathode was observed directly after inoculation at day 1. In MgenHMEC, the hydrogen evolution peak near −1 V versus SHE gradually shifted to higher current at more positive potentials with longer operation time of the reactors. In AgenMEC, the current increased slightly at the potential of −1 V versus SHE after 18 days. However, at the end of the MEC operation, the current peak at −1 V versus SHE was much lower compared to previous CV tests even though the MEC was operated at −0.8 V versus SHE the last 12 days of the experiment. In SRMEC, a clear reduction peak was observed at −0.55 V versus SHE after inoculation. The next CV tests carried out on day 26, day 54, and day 66, showed reduction peaks at −0.42 V, −0.22 V, and −0.36 V versus SHE, respectively. The current that was generated at −1 V versus SHE at day 26 and day 54, increased noticeably and shifted more towards −0.9 V versus SHE compared to the beginning of the experiment. However, at day 66 the current that was generated at −1 V versus SHE was decreased markedly. The CV tests for MgenAMEC did not show any change over the operation period even though a current of about 1 A/m2 was generated in the MEC at the end of the experiment. This indicates that the microbial community was electrochemically active on the cathode, however, catalysis of cathodic reactions did not improve over time.
Microbial community analysis
A summary heatmap showing the relative abundance of the most abundant taxa in the suspended enrichment cultures and on the cathodes in the MECs is shown in Fig. 7. The values for the suspended cultures are calculated by merging the results from all samples taken during the enrichment. More detailed heatmaps for each enrichment culture are shown in Figure S3 (supplementary material). The inoculum contained a very diverse community. Geothrix, Zoogloea, and three SVs unclassified at the genus level but belonging to Burkholderiales, WCBH1-90, and OM90 were dominating. Neither of these taxa remained detectable in the enrichments, instead other taxa increased in abundance.
The MgenH enrichments were dominated by Methanobacterium sp. (76%), which is known as an archaeon using H2 and CO2 to produce methane in anaerobic digesters32. The genera Thauera, Pseudomonas, and Sporomusa were also occasionally found at relative abundances over 2%. Sporomusa spp. are known as hydrogenotrophic acetogens and are found in different anaerobic environments33,34. Thauera spp. are often found in aerobic and denitrifying environments35 and can use several substrates. It may have consumed and scavenged low levels of oxygen, which may have leaked into the cultures. Pseudomonas may also have served as oxygen scavengers36.
In the Agen enrichments, the most abundant sequences were mainly affiliated to Sporomusa (47%). Bilophila, Pseudomonas, and Desulfovibrio were also abundant. Bilophila is anaerobic bacteria37. Desulfovibrio spp. reduce sulfate with hydrogen or organic acids as electron donor38 and an anode as electron acceptor39.
In the SR enrichments, Desulfovibrio was the dominating genus (45%). Desulfomicrobium sp. (13%) and SV27 of the Selenomonadales order were also abundant, as well as Methanobacterium. Desulfomicrobium spp. are sulfate-reducing anaerobic bacterium, which can utilize hydrogen and several organic electron donors40. The Selenomonadales order contain homoacetogens such as Sporomusa sp.
In the MgenA enrichments, Methanosaeta, which is capable of producing methane from acetate41, was the dominating methanogen, however only present at 8.6% of relative abundance. Thauera, and Pseudomonas were also abundant and may have served as microaerobic scavengers of acetate and oxygen. Two unclassified sequences (SV10, SV29) belonging to the W5 class of the Cloacimonetes phylum were also abundant.
The NR enrichments were dominated by Paracoccus (58%) and Rehaibacterium (22%), which suggests they were capable of hydrogenotrophic denitrification. Paracoccus sp. has previously been observed on denitrifying biocathodes42. Stappia, Azoarcus, and SV23 of the Sphingobacteriales order were also abundant. Stappia and Azoarcus spp. are known to denitrify with a range of electron donors43,44.
The microbial communities that developed on the cathodes in the MECs were different from the suspended enrichment cultures. Methanobacterium sp. was an important taxon on all cathodes. The relative abundance ranged from 5% in the MgenAMEC to 32% in the MgenHMEC. Methanobacterium spp. have been observed on biocathodes in MECs in several previous studies25,45. Desulfovibrio sp. were present on all biocathodes although it decreased in abundance in the MEC inoculated with SR. It was previously shown that Desulfovibrio sp. are capable of catalysing H2-production on a cathode with a columbic efficiency close to 100%21 and sulfate-reducing bacteria appear to play a key role in microbial electrosynthesis46. Paracoccus, Thauera, and Stappia were also present at high relative abundance on several of the cathodes. These taxa likely served as scavengers of oxygen leaking into the MECs. The unclassified SV23, belonging to the Sphingobacteriales order and SV14 of the Rhizobiales order increased in relative abundance on all cathodes. Sphingobacteriales spp. have previously been found on biocathodes and may be involved in catalysing hydrogen generation26. Rhizobiales spp. were observed in methane-producing microbial electrosynthesis reactors where they may have functioned as methane oxidizers47. Two SVs of the Coriobacteriales order were particularly abundant (54% in total) on the cathode in the MEC inoculated with the SR culture. Bacteria in this order have previously been shown to play a role in the catalysis of anaerobic reduction reactions on biocathodes48. For the MEC inoculated with Agen culture, it is worth noting that Sporomusa sp., which were highly abundant in the suspended enrichment culture (47%) and are known to catalyse microbial electrosynthesis of acetate6, made up only 0.5% of the reads on the cathode.
The similarity of the microbial communities in different samples were visualized using principal coordinate analysis (Fig. 8), which showed that samples from cultures enriched on the same substrates grouped together but were clearly separated from samples enriched on other substrates. The samples collected after 63 days of MEC operation are different from the suspended enrichment and from each other. An exception is the SR enrichment, were the liquid sample from the MEC is very similar to the suspended enrichment culture while only the cathode sample has diverged. The different cathode samples do not cluster together but depend on the inoculum.
Principle coordinate analysis based on a matrix of pairwise dissimilarities between samples calculated the beta component of Hill numbers of order 171.
Discussion
Enrichments
In general, the five enrichment cultures functioned as intended. The hydrogenotrophic cultures consumed hydrogen and produced methane (MgenH) and acetate (Agen), and reduced sulfate (SR) and nitrate (NR) while the acetate-oxidizing culture (MgenA) produced methane. The enrichments also formed distinct microbial communities depending on their electron donor and electron acceptor (Fig. 8). In the Agen and SR enrichments, addition of 2-bromoethanesulfonate was generally an effective strategy for inhibiting methanogens. However, in the end of the experimental run Methanobacterium appeared and methane was produced in low rates (0.02 mmol/d) which suggesting 2-bromoethanesulfonate was degraded over time. Degradation has previously been observed to occur, particularly under aerobic conditions49.
The Agen enrichments started to produce butyrate around day 120 when acetate had accumulated to its maximum concentration (Fig. 2C). No butyrate was detected in the other enrichments, which suggests that the accumulation of acetate and inhibition of methanogens played key roles for butyrate production. A modelling study suggested that acetate reduction to butyrate with hydrogen as electron donor is unlikely due to thermodynamic constraints50. The reduction of acetate to ethanol after which ethanol serves as electron donor for chain elongation of acetate to butyrate could be a possible pathway51. However, no ethanol was observed in the enrichments. Raes, et al.52 suggested that the slow kinetics of acetate reduction to ethanol in comparison to the ethanol oxidation rate during acetate chain elongation could explain why ethanol is not observed. However, they also suggested that butyrate might be produced via acetyl-CoA reduction with hydrogen or a cathode serving as electron donor. Sporomusa sp., which dominated the Agen enrichments, can produce both acetate and ethanol when fed with H2 and CO253. Thus, acetate and ethanol produced by the Sporomusa sp. may have been used to produce butyrate by other members of the community.
Cathode catalysis
All the enrichment cultures seemed to improve catalysis of the hydrogen evolution reaction on the cathode, although the results for the LSV tests were variable. All cultures also contained taxa that have previously been observed on microbial electrodes. Methanobacterium was highly abundant in the MgenH enrichment. Cheng, et al.8 showed that both a mixed culture dominated by Methanobacterium sp. and a pure culture of Methanobacterium palustre could catalyse methane production on a biocathode. Several studies have also shown that Methanobacterium spp. are selected for on biocathodes25,45,54. In the Agen enrichment, a Sporomusa sp. was highly abundant. Nevin, et al.55 showed that several Sporomusa spp. could use a cathode to produce organic acids. The SR enrichment was dominated by Desulfovibrio spp., which are known to catalyse H2 generation on biocathodes21,46. The MgenA enrichments, contained Pseudomonas spp, which are known to mediate electron transfer to an anode by production of soluble redox mediators15. MgenA also contained Methanosaeta, which has been observed on biocathodes56,57. The NR enrichments were dominated by Paracoccus spp., which have been observed on denitrifying biocathodes42. Thus, all the enrichments contained microbial taxa that have previously been reported to be electrochemically active. A previous study also reported that hydrogenases released by cells during culturing can adsorb to electrodes and catalyse hydrogen generation29. In this study, hydrogenotrophic microorganisms were enriched. These microorganisms must have contained hydrogenses, which could have been released during cell lysis.
MEC operation
The MECs generated current densities between 0.1–1 A/m2 at a potential of −0.65 V versus SHE directly after inoculation. This can be compared to other strategies used to start-up biocathodes. One strategy that has been investigated in several studies is the pre-enrichment of bioanodes followed by a change in potential so the electrodes operate as biocathodes. This was used successfully by Rozendal, et al.1, who enriched a hydrogen-oxidizing bioanode, then lowered the potential to −0.7 V versus SHE and obtained a biocathode generating a current density of about 1.1 A/m2. Enriching bioanodes on acetate and switching to cathode operation has been a less successful strategy. Saheb-Alam, et al.25 obtained a current density of 0.016 ± 0.007 A/m2 and it took over 170 days of operation as a biocathode before a current density of 0.6–3.6 A/m2 could be obtained. In comparison, direct start-up of the biocathode from sewage and anaerobic sludge only took 83 days25. Pisciotta, et al.58 enriched bioanodes on acetate in a sediment microbial fuel cell. After conversion to biocathodes the electrode generated 0.02 A/m2 at −0.7 V versus SHE and 0.002 A/m2 at −0.439 V versus SHE. Higher current densities have been obtained with biocathodes started up using other types of enrichment cultures. Villano, et al.31 used hydrogenotrophic methanogens and obtained 0.75 A/m2 at −0.75 V versus SHE. Both the strategy and the current density was similar to the MgenH enrichment tested in our study. However, the microbial community composition was not investigated by Villano, et al.31.
Clear reduction peaks in the CV curves day 1 and onward confirms that the cultures used to inoculate the MEC contained redox active components. Among the MECs, the SRMEC had a large number of redox peaks which are comparable to the ones observed by Aulenta, et al.21 for biocathodes inoculated by Desulfovibrio sp. However, on the biocathode in SRMEC, Desulfovibrio sp. was identified at a very low relative abundance. Instead, in the liquid phase, Desulfovibrio sp. was the second most abundant taxa. It has been previously shown that Desulfovibrio sp. contains enzymes that can directly transfer electron from a cathode59,60. It is possible that free enzymes in the liquid could be the reason of observing significant redox curves in the SRMEC. However, the Coriobacteriales spp., which were highly enriched on the cathode could also have catalysed cathodic reactions. Members of this order are known to be anaerobic and carry out fermentation61 and have also been found on biocathodes48.
In general, the microbial communities that developed on the cathodes in the MECs were different from the MgenH, Agen, SR, and MgenA enrichments used as inoculum. However, they were also different to each other, which shows that the inoculum had a strong effect on how the biocathodes developed during the 63 days of operation. One similarity was the increase in Methanobacterium sp. on all the biocathodes except MgenHMEC, where it was already present in high relative abundance in the inoculum. This further confirms previous observations that this taxon is highly selected for on biocathodes reducing CO2/HCO3− 8,25,54.
Materials and Methods
Inoculum, nutrient medium, and enrichment setup
Duplicate glass bottles (325 mL total volume each) were used for enriching five different cultures. Activated sludge from a municipal wastewater treatment plant (1 mL) was added to the bottles as inoculum. The bottles were filled up to 250 mL with a nutrient medium as described by Marshall, et al.26. The goal was to enrich hydrogenotrophic cultures performing methanogenesis, acetogenesis, sulfate reduction, nitrate reduction, as well as an acetate-oxidizing methanogenic culture. To accomplish this, the nutrient medium was amended with different electron acceptors and in some cases 2-bromoethanesulfonate to inhibit methanogens, as described in Table S1. The bottles were sealed with rubber caps and the head space (70 mL) was sparged with Ar/CO2 gas (85%/15%) to remove oxygen. Then, the head space of the hydrogenotrophic bottles were filled with pure hydrogen gas at an overpressure of 160–180 kPa. During sampling, the amount of liquid that was extracted from the enrichment cultures was replaced by fresh medium. Every 3 weeks, fresh medium (20 mL) containing 20 mM NaSO4, NaNO3, and sodium acetate was added to the sulfate-reducing, denitrifiers and acetate-oxidizing enrichments. Sterile syringes (12 mL) and needles were used for collecting samples, sparging of gas, and addition of fresh medium for each enrichment.
Analytical methods
The gas phase was analysed by gas chromatography (micro-GC, Agilent) one/two times a week. Samples from the liquid were collected one/two times a week and were analysed by high performance liquid chromatography (HPLC) equipped with a UV detector (Shimadzu) and an Aminex HPX-87H column (BioRad), with 5 mM H2SO4 eluent pumping at 0.5 mL/min. Cell growth was measured using optical density (OD) measured at 600 nm wavelength which correlates directly with the cell concentration62,63.
Electrochemical screening
Linear sweep voltammetry (LSV) was conducted occasionally with 4 mL mixed liquor collected from the enrichment cultures. The test was carried out in a double-chamber electrochemical cell (each chamber being 2 mL) containing a cation exchange membrane (CMI-7000, Membranes International Inc.), graphite foil electrodes, and a Ag/AgCl reference electrode. LSV was swept from open circuit potential (OCP) to −1.2 V versus SHE at a scan rate of 2 mV/s. Exchange current density was calculated using the Tafel equation \(({\rm{\eta }}={\rm{A}}\times \,\mathrm{ln}(\frac{i}{{i}_{0}}))\) based on the region of the voltammogram representing an overpotential greater than 0.4 V versus OCP. The symbols in the equation have the following meanings: \({\rm{\eta }}\) is the overpotential, A is the Tafel slope, \(i\) is the current density, and \({i}\) 0 is the exchange current density.
MEC setup
A glass double-chamber MEC, with a total volume of 340 mL in each chamber, was used to test the ability of some of the enrichment cultures to colonize a cathode. The cathode chamber was filled up to 280 mL with a mixed liquid from the enrichment bottle (140 mL) and the nutrient medium buffer (140 mL) described above. Bicarbonate or hydrogen ions were the only electron acceptor available in the cathode chamber. A graphite foil electrode (Alpha Aesar, 43083-1 mm thick, 39.2 cm2) was installed as working- and counter electrode in each chamber. An Ag/AgCl reference electrode with an offset of 0.197 V versus the standard hydrogen electrode (SHE) was installed in the working-chamber. The two chambers were separated by a cation exchange membrane (CMI-7000, Membranes International Inc.). The working electrode potential was controlled using Wenking M lab potentiostat and the current was recorded by MlabSci470c sequencer multichannel potentiostat software (version 4.7.0). Since the electrode potential has a strong effect on the magnitude of the cathodic current31,64, the MEC were operated at both −0.65 V and −0.8 V versus SHE. The bioelectrochemical activity of the cathode was investigated using cyclic voltammetry (CV). CV was done with scan limits of 0.7 V and −1.0 V versus SHE at a scan rate of 1 mV/s.
Microbial community analysis
Samples (10 mL) from the enrichment cultures were collected on a sterile membrane filter (Sartorius Stedim Biotech, 47 mm ϕ, 0.2 μm) for microbial community analysis. Moreover, biocathodes were harvested from MECs for microbial community analysis at the end of the experiment. Samples were stored at −20 °C prior to DNA extraction. DNA was extracted using the Fast DNA kit for soil (MP Biomedical). The 16S rRNA genes were amplified in duplicates using the forward primer 515’F (GTGBCAGCMGCCGCGGTAA) and the reverse primer 806 R (GGACTACHVGGGTWTCTAAT) to amplify V4 region sequences of the bacterial and archaeal 16S rRNA genes65 with dual index labelling according to Kozich, et al.66. Duplicate PCR reactions were carried out in 20 μL volume using 1 μL of target DNA, 17 μL of AccuPrime Pfx SuperMix (Life Technologies), and 1 μL of the forward and reverse primers, respectively. The PCR was conducted using a Bio-Rad T100 Thermal Cycler with a program consisting of activation (95 °C, 5 min); 30 cycles of denaturation (95 °C, 20 sec), annealing (50 °C, 20 sec) and elongation (68 °C, 60 sec), and final elongation (68 °C, 10 min). The products were purified (MagJET NGS Cleanup and Size Selection Kit, ThermoFischer Scientific), normalized per concentration, as measured by a Qubit Fluorometer (ThermoFischer Scientific), and pooled prior to sequencing on an Illumina MiSeq using the Miseq Reagent Kit V3.
The sequence reads were processed in USearch (v. 10) using the Unoise algorithm67 to generate sequences variants (SVs), which are analogous to operational taxonomic units68. Taxonomic classification was done using the Sintax algorithm69 with the Midas database70 (v. 123). Raw data files were deposited to the National Center of Biotechnology Information’s (NCBI) Sequence Read Archive (SRA) database available online with study accession of PRJNA482287. Analysis of the sequencing results was carried out using qDiv (github.com/omvatten/qDiv).
References
Rozendal, R. A., Jeremiasse, A. W., Hamelers, H. V. M. & Buisman, C. J. N. Hydrogen production with a microbial biocathode. Environ Sci Technol 42, 629–634, https://doi.org/10.1021/Es071720+ (2008).
Rozendal, R. A., Hamelers, H. V. M., Euverink, G. J. W., Metz, S. J. & Buisman, C. J. N. Principle and perspectives of hydrogen production through biocatalyzed electrolysis. Int J Hydrogen Energ 31, 1632–1640, https://doi.org/10.1016/j.ijhydene.2005.12.006 (2006).
Liu, H., Grot, S. & Logan, B. E. Electrochemically assisted microbial production of hydrogen from acetate. Environ Sci Technol 39, 4317–4320, https://doi.org/10.1021/Es050244p (2005).
Escapa, A., Mateos, R., Martinez, E. J. & Blanes, J. Microbial electrolysis cells: An emerging technology for wastewater treatment and energy recovery. From laboratory to pilot plant and beyond. Renew Sust Energ Rev 55, 942–956, https://doi.org/10.1016/j.rser.2015.11.029 (2016).
Marshall, C. W., Ross, D. E., Fichot, E. B., Norman, R. S. & May, H. D. Long-term Operation of Microbial Electrosynthesis Systems Improves Acetate Production by Autotrophic Microbiomes. Environ Sci Technol 47, 6023–6029, https://doi.org/10.1021/Es400341b (2013).
Nevin, K. P., Woodard, T. L., Franks, A. E., Summers, Z. M. & Lovley, D. R. Microbial Electrosynthesis: Feeding Microbes Electricity To Convert Carbon Dioxide and Water to Multicarbon Extracellular Organic Compounds. Mbio 1, e00103–00110, https://doi.org/10.1128/mBio.00103-10 (2010).
Villano, M., Monaco, G., Aulenta, F. & Majone, M. Electrochemically assisted methane production in a biofilm reactor. J Power Sources 196, 9467–9472, https://doi.org/10.1016/j.jpowsour.2011.07.016 (2011).
Cheng, S. A., Xing, D. F., Call, D. F. & Logan, B. E. Direct Biological Conversion of Electrical Current into Methane by Electromethanogenesis. Environ Sci Technol 43, 3953–3958, https://doi.org/10.1021/es803531g (2009).
Clauwaert, P. & Verstraete, W. Methanogenesis in membraneless microbial electrolysis cells. Appl Microbiol Biotechnol 82, 829–836, https://doi.org/10.1007/s00253-008-1796-4 (2009).
Van Eerten-Jansen, M. C. A. A. et al. Bioelectrochemical Production of Caproate and Caprylate from Acetate by Mixed Cultures. Acs Sustain Chem Eng 1, 513–518, https://doi.org/10.1021/sc300168z (2013).
Sharma, M. et al. Bioelectrocatalyzed reduction of acetic and butyric acids via direct electron transfer using a mixed culture of sulfate-reducers drives electrosynthesis of alcohols and acetone. Chem Commun 49, 6495–6497, https://doi.org/10.1039/c3cc42570c (2013).
Jeremiasse, A. W., Hamelers, E. V. M. & Buisman, C. J. N. Microbial electrolysis cell with a microbial biocathode. Bioelectrochemistry 78, 39–43, https://doi.org/10.1016/j.bioelechem.2009.05.005 (2010).
Kim, B. H. et al. Electrochemical activity of an Fe(III)-reducing bacterium, Shewanella putrefaciens IR-1, in the presence of alternative electron acceptors. Biotechnol Tech 13, 475–478, https://doi.org/10.1023/A:1008993029309 (1999).
Reguera, G. et al. Extracellular electron transfer via microbial nanowires. Nature 435, 1098–1101, https://doi.org/10.1038/nature03661 (2005).
Rabaey, K., Boon, N., Hofte, M. & Verstraete, W. Microbial phenazine production enhances electron transfer in biofuel cells. Environ Sci Technol 39, 3401–3408, https://doi.org/10.1021/es048563o (2005).
Morozov, S. V. et al. Bioelectrocatalytic hydrogen production by hydrogenase electrodes. Int J Hydrogen Energ 27, 1501–1505, https://doi.org/10.1016/S0360-3199(02)00091-5 (2002).
Vignais, P. M., Billoud, B. & Meyer, J. Classification and phylogeny of hydrogenases. Fems Microbiol Rev 25, 455–501, https://doi.org/10.1111/j.1574-6976.2001.tb00587.x (2001).
Lojou, E. & Bianco, P. Electrocatalytic reactions at hydrogenase-modified electrodes and their applications to biosensors: From the isolated enzymes to the whole cells. Electroanal 16, 1093–1100, https://doi.org/10.1002/elan.200403002 (2004).
Vincent, K. A., Parkin, A. & Armstrong, F. A. Investigating and exploiting the electrocatalytic properties of hydrogenases. Chem Rev 107, 4366–4413, https://doi.org/10.1021/cr050191u (2007).
Geelhoed, J. S. & Stams, A. J. M. Electricity-Assisted Biological Hydrogen Production from Acetate by Geobacter sulfurreducens. Environ Sci Technol 45, 815–820, https://doi.org/10.1021/es102842p (2011).
Aulenta, F., Catapano, L., Snip, L., Villano, M. & Majone, M. Linking Bacterial Metabolism to Graphite Cathodes: Electrochemical Insights into the H2-Producing Capability of Desulfovibrio sp. ChemSusChem 5, 1080–1085, https://doi.org/10.1002/cssc.201100720 (2012).
Beese-Vasbender, P. F., Grote, J. P., Garrelfs, J., Stratmann, M. & Mayrhofer, K. J. J. Selective microbial electrosynthesis of methane by a pure culture of a marine lithoautotrophic archaeon. Bioelectrochemistry 102, 50–55, https://doi.org/10.1016/j.bioelechem.2014.11.004 (2015).
Croese, E., Pereira, M. A., Euverink, G.-J. W., Stams, A. J. M. & Geelhoed, J. S. Analysis of the microbial community of the biocathode of a hydrogen-producing microbial electrolysis cell. Appl Microbiol Biotechnol 92, 1083–1093, https://doi.org/10.1007/s00253-011-3583-x (2011).
Batlle-Vilanova, P. et al. Assessment of biotic and abiotic graphite cathodes for hydrogen production in microbial electrolysis cells. Int J Hydrogen Energ 39, 1297–1305, https://doi.org/10.1016/j.ijhydene.2013.11.017 (2014).
Saheb-Alam, S. et al. Effect of Start-Up Strategies and Electrode Materials on Carbon Dioxide Reduction on Biocathodes. Appl Environ Microb 84, e02242–17, https://doi.org/10.1128/AEM.02242-17 (2018).
Marshall, C. W., Ross, D. E., Fichot, E. B., Norman, R. S. & May, H. D. Electrosynthesis of Commodity Chemicals by an Autotrophic Microbial Community. Appl Environ Microb 78, 8412–8420, https://doi.org/10.1128/Aem.02401-12 (2012).
LaBelle, E. V., Marshall, C. W., Gilbert, J. A. & May, H. D. Influence of Acidic pH on Hydrogen and Acetate Production by an Electrosynthetic Microbiome. Plos One 9, e109935, https://doi.org/10.1371/journal.pone.0109935 (2014).
Jourdin, L., Lu, Y., Flexer, V., Keller, J. & Freguia, S. Biologically Induced Hydrogen Production Drives High Rate/High Efficiency Microbial Electrosynthesis of Acetate from Carbon Dioxide. Chemelectrochem 3, 581–591, https://doi.org/10.1002/celc.201500530 (2016).
Deutzmann, J. S., Sahin, M. & Spormann, A. M. Extracellular Enzymes Facilitate Electron Uptake in Biocorrosion and Bioelectrosynthesis. Mbio 6, e00496-1510.1128/mBio.00496-15 (2015).
Liang, D. W. et al. Effects of bicarbonate and cathode potential on hydrogen production in a biocathode electrolysis cell. Front Env Sci Eng 8, 624–630, https://doi.org/10.1007/s11783-013-0584-2 (2014).
Villano, M. et al. Bioelectrochemical reduction of CO2 to CH4 via direct and indirect extracellular electron transfer by a hydrogenophilic methanogenic culture. Bioresource Technol 101, 3085–3090, https://doi.org/10.1016/j.biortech.2009.12.077 (2010).
Demirel, B. & Scherer, P. The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Reviews in Environmental Science and Bio/Technology 7, 173–190, https://doi.org/10.1007/s11157-008-9131-1 (2008).
Balk, M. et al. (Per)chlorate reduction by an acetogenic bacterium, Sporomusa sp., isolated from an underground gas storage. Appl Microbiol Biotechnol 88, 595–603, https://doi.org/10.1007/s00253-010-2788-8 (2010).
Breznak, J. A., Switzer, J. M. & Seitz, H.-J. Sporomusa termitida sp. nov., an H2/CO2-utilizing acetogen isolated from termites. Arch. Microbiol. 150, 282–288, https://doi.org/10.1007/bf00407793 (1988).
Thomsen, T. R., Kong, Y. & Nielsen, P. H. Ecophysiology of abundant denitrifying bacteria in activated sludge. Fems Microbiol Ecol 60, 370–382, https://doi.org/10.1111/j.1574-6941.2007.00309.x (2007).
Xiao, Z., Awata, T., Zhang, D. & Katayama, A. Denitrification by Pseudomonas stutzeri coupled with CO2 reduction by Sporomusa ovata with hydrogen as an electron donor assisted by solid-phase humin. J Biosci Bioeng 122, 307–313, https://doi.org/10.1016/j.jbiosc.2016.02.002 (2016).
Baron, E. J. et al. Bilophila wadsworthia, gen. nov. and sp. nov., a Unique Gram-negative Anaerobic Rod Recovered from Appendicitis Specimens and Human Faeces. Microbiology 135, 3405–3411, https://doi.org/10.1099/00221287-135-12-3405 (1989).
Heidelberg, J. F. et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat Biotechnol 22, 554, https://doi.org/10.1038/nbt959 (2004).
Saheb-Alam, S., Persson, F., Wilen, B. M., Hermansson, M. & Modin, O. Effects of storage on mixed-culture biological electrodes. Sci Rep 5, 18433, https://doi.org/10.1038/srep18433 (2015).
Hippe, H., Vainshtein, M., Gogotova, G. I. & Stackebrandt, E. Reclassification of Desulfobacterium macestii as Desulfomicrobium macestii comb. nov. Int J Syst Evol Microbiol 53, 1127–1130, https://doi.org/10.1099/ijs.0.02574-0 (2003).
Smith, K. S. & Ingram-Smith, C. Methanosaeta, the forgotten methanogen? Trends Microbiol 15, 150–155, https://doi.org/10.1016/j.tim.2007.02.002 (2007).
Virdis, B. et al. Biofilm stratification during simultaneous nitrification and denitrification (SND) at a biocathode. Bioresource Technol 102, 334–341, https://doi.org/10.1016/j.biortech.2010.06.155 (2011).
Weber, C. F. & King, G. M. Physiological, ecological, and phylogenetic characterization of Stappia, a marine CO-oxidizing bacterial genus. Appl Environ Microb 73, 1266–1276, https://doi.org/10.1128/AEM.01724-06 (2006).
Lee, D.-J., Wong, B.-T. & Adav, S. S. Azoarcus taiwanensis sp. nov., a denitrifying species isolated from a hot spring. Appl Microbiol Biotechnol 98, 1301–1307, https://doi.org/10.1007/s00253-013-4976-9 (2014).
Mateos, R., Sotres, A., Alonso, R. M., Escapa, A. & Morán, A. Impact of the start-up process on the microbial communities in biocathodes for electrosynthesis. Bioelectrochemistry 121, 27–37, https://doi.org/10.1016/j.bioelechem.2018.01.002 (2018).
Xiang, Y., Liu, G., Zhang, R., Lu, Y. & Luo, H. Acetate production and electron utilization facilitated by sulfate-reducing bacteria in a microbial electrosynthesis system. Bioresource Technol 241, 821–829, https://doi.org/10.1016/j.biortech.2017.06.017 (2017).
Babanovaa, S. et al. The effect of membrane type on the performance of microbial electrosynthesis cells for methane production. J Electrochem Soc 164, H3015–H3023 (2017).
Kobayashi, H., Fu, Q., Maeda, H. & Sato, K. Draft genome sequence of a novel Coriobacteriaceae sp. strain, EMTCatB1, reconstructed from the metagenome of a thermophilic electromethanogenic biocathode. Genome Announcements 5, e00022–00017, https://doi.org/10.1128/genomeA.00022-17 (2017).
Rago, L., Guerrero, J., Baeza, J. A. & Guisasola, A. 2-Bromoethanesulfonate degradation in bioelectrochemical systems. Bioelectrochemistry 105, 44–49, https://doi.org/10.1016/j.bioelechem.2015.05.001 (2015).
González-Cabaleiro, R., Lema, J. M., Rodríguez, J. & Kleerebezem, R. Linking thermodynamics and kinetics to assess pathway reversibility in anaerobic bioprocesses. Energ Environ Sci 6, 3780–3789, https://doi.org/10.1039/c3ee42754d (2013).
Angenent, L. T. et al. Chain Elongation with Reactor Microbiomes: Open-Culture Biotechnology To Produce Biochemicals. Environ Sci Technol 50, 2796–2810, https://doi.org/10.1021/acs.est.5b04847 (2016).
Raes, S. M. T., Jourdin, L., Buisman, C. J. N. & Strik, D. P. B. T. B. Continuous long-term bioelectrochemical chain elongation to butyrate. Chemelectrochem 4, 386–395, https://doi.org/10.1002/celc.201600587 (2016).
Ammam, F., Tremblay, P.-L., Lizak, D. M. & Zhang, T. Effect of tungstate on acetate and ethanol production by the electrosynthetic bacterium Sporomusa ovata. Biotechnol Biofuels 9, 163, https://doi.org/10.1186/s13068-016-0576-0 (2016).
van Eerten-Jansen, M. C. et al. Microbial community analysis of a methane-producing biocathode in a bioelectrochemical system. Archaea 2013, 481784, https://doi.org/10.1155/2013/481784 (2013).
Nevin, K. P. et al. Electrosynthesis of Organic Compounds from Carbon Dioxide Is Catalyzed by a Diversity of Acetogenic Microorganisms. Appl Environ Microb 77, 2882–2886, https://doi.org/10.1128/aem.02642-10 (2011).
Yang, H.-Y. et al. Temperature dependence of bioelectrochemical CO2 conversion and methane production with a mixed-culture biocathode. Bioelectrochemistry 119, 180–188, https://doi.org/10.1016/j.bioelechem.2017.10.002 (2018).
De Vrieze, J. et al. Biomass retention on electrodes rather than electrical current enhances stability in anaerobic digestion. Water Res 54, 211–221, https://doi.org/10.1016/j.watres.2014.01.044 (2014).
Pisciotta, J. M., Zaybak, Z., Call, D. F., Nam, J. Y. & Logan, B. E. Enrichment of Microbial Electrolysis Cell Biocathodes from Sediment Microbial Fuel Cell Bioanodes. Appl Environ Microb 78, 5212–5219, https://doi.org/10.1128/Aem.00480-12 (2012).
Moreno, C., Franco, R., Moura, I., Le Gall, J. & Moura, J. J. G. Voltammetric studies of the catalytic electron‐transfer process between the Desulfovibrio gigas hydrogenase and small proteins isolated from the same genus. Eur J Biochem 217, 981–989, https://doi.org/10.1111/j.1432-1033.1993.tb18329.x (1993).
Nivière, V., Hatchikian, E. C., Bianco, P. & Haladjian, J. Kinetic studies of electron transfer between hydrogenase and cytochrome c3 from Desulfovibrio gigas. Electrochemical properties of cytochrome c3. Biochimica et Biophysica Acta (BBA) - Bioenergetics 935, 34–40, https://doi.org/10.1016/0005-2728(88)90105-3 (1988).
Goodfellow, M. et al. Bergey’s Manual of Systematic Bacteriology. Second Edition. The Actinobacteria. Part A and B., Vol. 5 (Springer, 2012).
Biesta-Peters, E. G., Reij, M. W., Joosten, H., Gorris, L. G. M. & Zwietering, M. H. Comparison of Two Optical-Density-Based Methods and a Plate Count Method for Estimation of Growth Parameters of Bacillus cereus. Appl Environ Microb 76, 1399–1405, https://doi.org/10.1128/Aem.02336-09 (2010).
Shao, J., Xiang, J., Axner, O. & Ying, C. Wavelength-modulated tunable diode-laser absorption spectrometry for real-time monitoring of microbial growth. Appl. Opt. 55, 2339–2345, https://doi.org/10.1364/AO.55.002339 (2016).
Liu, H., Song, T., Fei, K., Wang, H. & Xie, J. Microbial electrosynthesis of organic chemicals from CO2 by Clostridium scatologenes ATCC 25775T. Bioresour Bioprocess 5, 7, https://doi.org/10.1186/s40643-018-0195-7 (2018).
Hugerth, L. W. et al. DegePrime, a Program for Degenerate Primer Design for Broad-Taxonomic-Range PCR in Microbial Ecology Studies. Appl Environ Microb 80, 5116–5123, https://doi.org/10.1128/Aem.01403-14 (2014).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Appl Environ Microb 79, 5112–5120, https://doi.org/10.1128/Aem.01043-13 (2013).
Edgar, R. C. UNOISE2: Improved error-correction for Illumina 16S and ITS amplicon reads. bioRxiv, https://doi.org/10.1101/081257 (2016).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. Isme Journal 11, 2639–2643, https://doi.org/10.1038/ismej.2017.119 (2017).
Edgar, R. C. SINTAX, a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv, https://doi.org/10.1101/074161 (2016).
McIlroy, S. J. et al. MiDAS: the field guide to the microbes of activated sludge. Database-Oxford, doi:ARTN bav06210.1093/database/bav062 (2015).
Jost, L. Partitioning diversity into independent alpha and beta components. Ecology 88, 2427–2439, https://doi.org/10.1890/06-1736.1 (2007).
Acknowledgements
The project was funded by the Swedish Research Council (grant VR 2012-5167). The Genomics core facility at the University of Gothenburg is acknowledged for support and use of their equipment.
Author information
Authors and Affiliations
Contributions
S.S.-A. and O.M. wrote the main manuscript text, S.S.-A. carried out most of the experimental work, all authors analysed the data and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saheb-Alam, S., Persson, F., Wilén, BM. et al. A variety of hydrogenotrophic enrichment cultures catalyse cathodic reactions. Sci Rep 9, 2356 (2019). https://doi.org/10.1038/s41598-018-38006-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-38006-3
This article is cited by
-
Biogas potential of organosolv pretreated wheat straw as mono and co-substrate: substrate synergy and microbial dynamics
Scientific Reports (2024)
-
Biogas Upgrading by Hydrogenotrophic Methanogens: An Overview
Waste and Biomass Valorization (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.