Abstract
LncRNA play important roles in regulation of host immune and inflammation responses in defending bacterial infection. Clostridium perfringens (C. perfringens) type C is one of primary bacteria leading to piglet diarrhea and other intestinal inflammatory diseases. For the differences of host immune capacity, individuals usually show resistance and susceptibility to bacterial infection. However, whether and how lncRNAs involved in modulating host immune resistance have not been reported. We have investigated the expression patterns of ileum lncRNAs of 7-day-old piglets infected by C. perfringens type C through RNA sequencing. A total of 16 lncRNAs and 126 mRNAs were significantly differentially expressed in resistance (IR) and susceptibility (IS) groups. Many lncRNAs and mRNAs were identified to regulate resistance and susceptibility of piglets through immune related pathways. Five lncRNAs may have potential function on regulating the expressions of cytokines, these lncRNAs and cytokines work together to co-regulated piglet immune response to C. perfringens, affecting host resistance and susceptibility. These results provide valuable information for understanding the functions of lncRNA and mRNA in affecting piglet diarrhea resistance of defensing to C. perfringens type C, these lncRNAs and mRNAs may be used as the important biomarkers for decreasing C. perfringens spread and diseases in human and piglets.
Similar content being viewed by others
Introduction
Diarrhea is one of the important reasons leading piglet death, causing the huge economic losses in worldwide pig industries, especially the newborn and suckling piglets1. Recently, Clostridium perfringens (C. perfringens) type C is considered as an increasingly pathogenic bacteria of pig source and one of the important causes for high morbidity and mortality in neonatal piglet diarrhea2,3,4, and have become the substantial problem to hinder the health development of livestock industry. C. perfringens type C includes two types of toxins, α and β, which have been proposed to act as spreading factors that destroy the physical properties of tissue matrices and intercellular spaces, thereby aiding in the spread of bacteria within host5.
C. perfringens type C is also a common part of microbiota in pig intestinal tract, sows can transfer this bacterium to piglets by breast milk and feces. Clinical diseases caused by C. perfringens type C can be divided into acute and chronic course. The acute course mainly causes sudden death of 7-day-old neonatal piglets with characteristics of depression and bloody diarrhea. While chronic disease can persist more than one week, which is characterized by intermittent or persistent diarrhea with or without blood and dehydration. Generally, it affects the growth and development of sick piglets, even stiff pigs and death3. The contaminated pig is considered as the dangerous infector of C. perfringens type C, becoming an important cause of foodborne illness and zoonotic disease6. Human can be usually acquired through the consumption of C. perfringens-contaminated pork products or the direct contact with infected piglets7. Decreasing the prevalence and severity of C. perfringens in pig herds may effectively reduce spread and control transmission of bacteria from pig products to humans and to the environment, especially in large pork-producing and consuming countries.
In the process of bacterial infections, the severity of infection is impacted by pathogenicity of microbiota and its interaction with host immunity defense system8. The hypoimmune piglets are weaker and more susceptible to C. perfringens type C infection than hyperimmune ones for the lower immunity defense system. Currently, few frontier views are available to treat or reverse C. perfringens type C infection disease, the main reason is the poor understanding on the regulatory mechanism of host defense against C. perfringens type C infection. Therefore, exploring the potential mechanism of different resistances in piglets infected by C. perfringens type C will help to screen highly resistant piglets to decrease death rate, which may provide the new slight in methods of preventing and treating these infectious diseases.
LncRNAs are a type of transcripts with a length of more than 200 nucleotides and with no obvious potential to encode functional protein. Specific lncRNAs have been clearly known to participate in some important biological processes, such as development9,10, posttranscriptional regulation11, and immune diseases12,13, which have been drawn the increasing attention. Recently, identifying functional mRNAs of hosts regulated C. perfringens infection are clearly under the way, such as broiler chicken14,15. As the important posttranscriptional pathogenesis, these transcripts and their associated orchestrated networks are implicated in mediating complex pathological mechanisms of C. perfringens infection. To date, little studies have examined the dysregulated lncRNAs and their target genes in regulating resistance and susceptibility of piglets exposed to C. perfringens type C.
This study aims to identify and compare ileum lncRNA and mRNA expression profiles of resistant and sensitive piglets infected by C. perfringens type C infection using Ribo-Zero RNA-seq. An amount of differentially expressed lncRNAs and mRNAs were identified between resistant and sensitive piglets, which might play important roles in regulating piglet resistance to C. perfringens type C infection, though the potential roles of lncRNAs should be further validated. These results propose as a reliable model for exploring host resistance mechanisms in defensing C. perfringens type C infection, which may provide valuable foundation for further breeding diarrhea-resistance piglet strain.
Materials and Methods
Bacterial culture
The C. perfringens type C strain (CVCC 2032) was obtained from the China Veterinary Culture Collection Center and used in this study. The bacterium was cultured at 37 °C in the bouillon culture-medium (HopeBio, Qingdao, China) for 16 h with shaking before used for infection. The colony-forming units (CFUs) of C. perfringens type C was determined by plate colony counting method, and finally an expected concentration of 1 × 109 CFU/mL C. perfringens type C medium was used to inoculate piglets.
Animal experiment
All procedures described here were approved by the experimental license from Gansu Research Center of Swine Production Engineering and Technology, Gansu Agricultural University, in agreement with the relevant guidelines and regulations imposed by the Administration of Affairs Concerning Experimental Animals. Animals were humanely sacrificed as necessary to ameliorate suffering.
The candidate piglets were the descendants of seronegative Yorkshire sows × Landrace boar from health nucleus herd (confirmed by history and seronegative sows) in Dingxi city, Gansu province of China. Fecal samples of all piglets were collected and detected negative for Escherichia coli, Salmonella and C. perfringens tested by commercial enzyme-linked immuno sorbent assay (ELISA) kits (Jiancheng Bioengineering Institute, Nanjing, China) at the times of selection, transportation and inoculation. Finally, a total of 30 suckling piglets at 7 days old were screened, 5 of 30 piglets were randomly selected as control group (IC), the remaining 25 piglets were challenged by oral gavage of 1 mL 1 × 109 CFU/mL C. perfringens type C medium for five consecutive days. Every piglet was housed in one pen separately to avoid cross infection, and they were raised in appropriate condition of climate-controlled and fully isolation, receiving water and diets ad libitum. During the period of infection, piglets were monitored for clinical signs and fecal consistency 3–4 times daily. Fecal consistency was scored based on the visual observation of symptoms traits: 0 = normal, solid feces, 1 = slight diarrhea, soft and loose feces, 2 = moderate diarrhea, semi-liquid feces, 3 = severe diarrhea, liquid and unformed feces16. Grouping criteria were performed as following: recording fecal consistency score of every defecation of each piglet, then summing and ranking total scores of each piglet, at last, combining with the clinical signs, the top five piglets with the highest and lowest fecal scores were designated as susceptibility (IS) and resistance (IR) groups, respectively.
The ileum tissues of fifteen piglets from IR, IS and IC groups were collected and flushed cleanly with PBS buffer (pH 7.4), and then quickly frozen in liquid nitrogen and stored at −80 °C until RNA extraction. Concurrently, the body weight, heart, liver, spleen, lung and kidney of piglets in IR, IS and IC groups were recorded. Blood samples of each piglets were collected from precaval vein every day, and stored at −80 °C.
Total RNA isolation, library construction and lncRNA sequencing
Total RNA samples were isolated from ileum tissues using TRIzol™ reagent (Invitrogen, USA) and quantified by Nanodrop equipment. Purity and integrity of RNA extracts were assessed using the NanoPhotometer® spectrophotometer (IMPLEN, CA, USA) and RNA Nano6000 Assay Kit of the Bionalyzer 2100 system (Agilent Technologies, CA, USA), which were then used for library preparation.
Approximately 3 μg rRNA-depleted RNA (Ribo-Zero RNA) was acquired from total RNA by Epicentre Ribo-zero™ rRNA Removal Kit (Epicentre, USA) and cleaned up by ethanol precipitation to prepare sequencing library. Subsequently, strand-specific RNA sequencing libraries were generated from Ribo-Zero RNA by NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (NEB, Ipswich, MA, UK) to capture all transcripts with and without poly A. The library fragments of preferentially 150–200 bp in length were purified with AMPure XP system (Beckman Coulter, Beverly, USA). At last, library qualities were assessed on the Agilent Bioanalyzer 2100 system.
After clustering using the TruSeq PE Cluster Kit v3-cBot-HS (Illumina®), RNA libraries were sequencing on Illumina Hiseq4000 platform (Illumina, San Diego, CA, USA) to generated 150 bp paired-end (PE150) reads at the Novogene Bioinformatics Institute (Beijing, China).
Identification of different expressed lncRNA and mRNA
After quality control, the paired-end clean reads were mapped to the pig reference genome sequence (Sus scrofa 10.2) by TopHat217 and were assembled by Scripture18 and Cufflinks19 in a reference-based approach.
The coding potentials of transcripts were predicted by four tools named Coding-Non-Coding-Index (CNCI)20, Coding Potential Calculator (CPC)21, Pfam-scan v1.3 (E-value < 0.001)22, and phylogenetic codon substitution frequency (phyloCSF) v2012102823 to distinguish mRNA from lncRNA. Transcripts, which were predicted by any one of these four tools, were filtered out, those without coding potential were defined as candidate lncRNA.
The FPKMs (fragments per kilo-base of exon per million fragments mapped) of lncRNA and mRNA were calculated by Cuffdiff19. Gene FPKMs were computed by summing FPKMs of transcripts in each group. Differential expression levels were determined using a model based on the negative binomial distribution model. Transcripts with a corrected P-value < 0.05 were assigned as significantly differentially expressed.
Target gene prediction
Cis and trans analyses were used to predict the target genes of differentially expressed lncRNAs. The target genes of lncRNA in cis role were predicted by lncRNAs regulation on expression of their neighboring protein-coding genes, which were close to 10 K upstream and downstream regions of lncRNA24. The target genes of lncRNA in trans role were identified by expression levels of lncRNA and mRNA, the expressed correlation between lncRNAs and coding genes was calculated based on Pearson’s correlation coefficient, the Pearson’s correlation coefficient (|r| > 0.95) were selected.
In structure, lncRNAs can form special secondary structures to regulate expression of mRNAs. Thus, the secondary structures of lncRNAs were predicted based on the free energy using RNAFold web server online software (http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi).
Function enrichment prediction
Analyses of Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathway (https://www.kegg.jp/kegg/kegg1.html) were applied to investigate the potential roles of differentially expressed lncRNA target genes and mRNAs. The P value < 0.05 were considered as significantly enriched.
Real-time quantitative PCR (qPCR) validation
Total RNA samples of 15 ileum tissues used for RNA-seq were processed to synthesize cDNA using reverse transcriptase Kit (TaKaRa, Dalian, China). A total of 3 lncRNAs (LNC_001066, LNC_001186 and ENSSSCT00000032859) and 5 mRNAs (TNFRSF11A, TLR8, IRAK3, LCP2 and CYP1A1) were used to perform used to qPCR detection for validating the accuracy of RNA sequencing.
In addition, blood samples of 15 piglets from IR, IS and IC groups were collected at the 1, 2, 3, 4 and 5 days after infection, respectively. After lysing red blood cells from blood, total RNA samples were isolated from leukocytes using TRIzol reagent, and then used to reverse transcribe and synthesize cDNA using reverse transcriptase Kit (TaKaRa, Dalian, China). Finally, four cytokine genes (interleukin IL-1β, interferon IFN-α, tumor necrosis factor TNF-α, and nuclear factor NF-κB) were selected to quantify relative-expression levels to explore changes of cytokines.
The specific amplification primers of these genes, cytokines and housekeeping GAPDH gene were designed using NCBI website BLAST online software (Supplementary Table S1), the qPCR detection was qualified using 2−∆∆Ct value methods25. The qPCR reaction was performed in 20 µL system involved 9.5 µL 2 × SYBR Green Realtime PCR Master Mix (TaKaRa, Dalian, China), 1 µL forward and reverse primers, 1 µL cDNA and 7.5 µL RNase free ddH2O using LightCycler 480II Real-Time PCR System. The cycling conditions included an initial activation denaturation (95 °C for 3 min), and followed by 30 cycles (95 °C for 15 s (denaturation), 60° ± 1 °C for 15 s (annealing), 72 °C for 20 s (extension)). Each biological replicate was comprised of three technical replicates.
Statistical analysis
The experimental data was displayed as the mean ± standard error of mean (SEM). One-way ANOVA was performed to calculate statistical significance followed by Duncan to independently compare each C. perfringens type C treatment group to the control group. All statistical analyses were conducted by using correlation test (Student’s t-test) using SPSS 18.0 (SPSS Inc., Chicago, IL, USA), and P < 0.05 or P < 0.01 represent significance level.
Results
Difference of body and organ index among IR, IS and IC groups
We had made the statistics analysis about the differences of body and organ weights of piglets among IR, IS and IC groups. As shown in Fig. 1, body and organ weights of infection group (IR and IS) were significantly lower than those of control group (P < 0.01), meanwhile, piglets in the IR group had significantly higher levels of body weight, heart, liver and spleen than piglets in the IS group (P < 0.01), however, the weights of liver, lung and kidney were not statistically significant difference between IR and IS groups (P > 0.05).
Analyses of differentially expressed lncRNAs
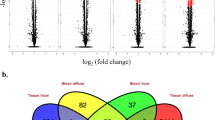
To identify C. perfringens-responsive lncRNAs, the normalized expression of lncRNAs was compared between the IR and IS treatment groups. After C. perfringens type C infection, a total of 359 lncRNAs and 2588 mRNAs were significantly expressed in the IR vs IC group (Supplementary Table S2, Fig. 2A), as well as 419 lncRNAs and 3283 mRNAs in the IS vs IC group (Supplementary Table S3, Fig. 2A).
Analysis of differentially expressed lncRNAs and mRNA of piglets among the IR, IS and IC groups. (A) Venn diagram analysis of differently expressed lncRNA and mRNA; (B) Volcano plot analysis of differently expressed lncRNA and mRNA. Note: (1) Abscissa represents log2 (fold-change), and ordinate represents −log10 (P value); (2) Red dots denote the significant differentially expressed up-regulation and down-regulation transcripts, respectively; (3) Blue dots denote no differentially expressed transcripts.
We conducted a Venn diagram for lncRNA and mRNA to understand the differential expression degree. As shown in Fig. 2A, a total of 16 lncRNAs and 126 mRNAs were found to differently express in the IR vs IS group, in which, 6 lncRNAs and 41 mRNAs were up-regulated, 10 lncRNAs and 85 mRNAs were down-regulated (Fig. 2B,C). There were 4 lncRNAs and 28 mRNAs were common differentially expressed among the IR, IS and IC groups after C. perfringens type C infection. Compared with IC group, 10 lncRNAs and 100 mRNAs were significantly differentially expressed in the IR vs IS group, which were used as the main resource for next analyses, and the details of these 10 lncRNAs were summarized and presented in Table 1.
Prediction of lncRNA and mRNA functions
To better understand the functions of lncRNAs, we firstly predicted the potential target genes of 10 lncRNAs, and meanwhile screened the differentially expressed target genes in IR vs IS groups compared to IC group, the screened results were presented in the Supplementary Table S4. While, there were no target genes predicted by three lncRNAs, LNC_000139, LNC_001415 and ALDBSSCT0000004597, which may be the reason of incomplete information for the updating pig genomic annotation.
In general, lncRNA can act as signal molecules to regulate downstream gene transcription, as decoy molecules to play a role in blocking molecules, as guide molecules to combine with proteins, or as scaffold molecules to accurately control signal transduction and molecular dynamics in multiple biological processes. LncRNA exert regulatory function by three kinds of modes, i.e. local single chain structure, local secondary structural motifs, and target molecular interaction of particularly tertiary structural motifs26,27. Recently, it is difficult to acquire tertiary structure of lncRNA, secondary structure prediction of lncRNA target molecules are helpful to study their functional mechanisms in some degree, lncRNA with different secondary structures can exert different functions. Therefore, the structural properties of these 10 lncRNAs were predicted by statistical analyses, which may help to verify the subsequent function and to increase our understanding to their regulation modes. Finally, 9 differentially expressed lncRNAs were successfully predicted secondary structures (Fig. 3), except for LNC_001066 with excessive length. As demonstrated, the secondary structures of these lncRNAs mainly include stem loop, hairpin loop, multibranch loop et al., these loops are formed by with many unpaired bases, which can match with small molecules to regulate functions of RNAs and compounds by base pairing, thus each lncRNA may regulate multiple target genes.
The secondary structures prediction of 9 screened differentially expressed lncRNAs. Note: A–O represents lncRNAs: LNC_000139, LNC_001415, LNC_000231, LNC_001186, ENSSSCT00000018610, ENSSSCT00000032859, ALDBSSCT0000004597, ALDBSSCT0000009442 and ALDBSSCT0000007865, LNC_001066 is not predicted for the length more than 10000 nt.
Furthermore, the GO and KEGG enrichment analyses were performed to predict the potential functions of lncRNAs. Except for 3 lncRNAs without predicted target genes, the remaining 7 lncRNAs were performed to predict target genes, which were presented in Table 2. These target genes were found to significantly enrich in 86 and 204 GO terms through cis and trans functions, respectively, in IR vs IS treatment group compared to IC group. In which, the main immune associated GO functions included MHC class I protein complex and transcription factor activity in cellular component, immune response, macrophage tolerance induction, antigen processing and presentation, and regulation of I-κB kinase/NF-κB signaling in biological process (Fig. 4A) (Supplementary Table S5), the top 20 enriched pathways of lncRNA target genes mainly included some immune and inflammatory related pathways, such as cell adhesion molecules, inflammatory bowel disease (IBD), T cell receptor signaling pathway, natural killer cell mediated cytotoxicity (Fig. 4B) (Supplementary Table S6).
Functional enrichment analyses of lncRNA target genes identified in IR vs IS group. (A) GO function annotation of the screened lncRNAs target genes. The x-axis indicates the detail terms and the y-axis indicates gene numbers. (B) KEGG signaling pathways of the screened lncRNAs target genes. The x-axis indicates the gene ratio and the y-axis indicates the name of the KEGG pathway. The size of the dot indicates the number of target genes, and the color of the dot indicates different P value (Fisher’s Exact Test).
In addition, we further analyzed the enriched GO terms and KEGG pathways of 100 differentially expressed mRNAs. Results showed that a total of 19 significantly enriched GO terms (Corrected P Value < 0.05) and 549 GO terms were detected in IR vs IS treatment group (Supplementary Table S7). For example, the significantly enriched GO terms were mainly found in biological process of mucosal immune response, innate immune response in mucosa, immune response, defense response to gram-positive bacterium, and in cellular component of extracellular region. Meanwhile, 12 significantly enriched KEGG pathways main including toll-like receptor signaling pathway, antigen processing and presentation, glycosphingolipid biosynthesis, protein digestion and absorption, as well as some immune-related pathways of chemokine signaling pathway, NF-κB signaling pathway, cytokine-cytokine receptor interaction, and MAPK signaling pathway (P < 0.05) (Supplementary Table S8). Importantly, some immune-related genes were also found significantly differentially expressed and enriched in these bacterial infection associated pathways, such as TLR8, LBP and SPP1 in toll-like receptor signaling pathway, HSP70 and CD8A in antigen processing and presentation, CXCL9, CXCL10, CCR5 and CCL17 in chemokine signaling pathway.
To validate the accuracy lncRNA sequencing data, a total of 3 lncRNAs and 5 mRNAs were selected to perform the qPCR detection. As shown in Fig. 5, the results of qPCR detection were perfectly matched to sequencing data, suggesting that the sequencing data was accurate.
Expression of cytokine genes
The relative expressions of IL-1β, IFN-α, TNF-α, and NF-κB over the course of the infection were detected by qPCR method (Fig. 6). Compared to IC group, the expression levels of IL-1β and NF-κB were downregulated both in the IR and IS groups at the 1dpi, while IFN-α and TNF-α were upregulated in the IR and IS groups. in which, the expression of IFN-α was significant difference between the IR group and IC group (P < 0.05). At the 3dpi, expression levels of TNF-α and NF-κB in the IS group and IL-1β in the IR group were significantly increased in comparison with the IC group (P < 0.05). At the 5dpi, in the IR group, the expression levels of IFN-α and TNF-αwere significantly upregulated than those in the IS group, while expressions of IL-1β and NF-κB in the IR group were significantly downregulated than those in the IS group. The results showed that C. perfringens type C infection significantly affect the expressions of these cytokine genes in piglets among IR, IS and IC groups, the high expressed cytokines may hint various degrees of inflammation response in piglets, which may be associated with the piglet immune resistant to C. perfringens type C infection.
Discussion
C. perfringens has been recognized as one of the widespread potential bacterial pathogens, which can result in many infectious diseases, seriously influencing human and animal heaths. C. perfringens type C can cause piglet hemorrhagic diarrhea, even in human. Generally, the occurrences of these diseases are mainly caused by the close contact with C. perfringens type C-infected piglets or -polluted environments. The pathogenic ability of C. perfringens type C is various in infecting different piglet individuals and causing inflammatory diseases, in turn, piglets also have multifarious immune abilities to resist C. perfringens type C infection, these differences largely depend on host-pathogen interactions and host immune tolerance to bacteria28. Recently, lncRNA are considered as an important regulatory factor in modulating host inflammatory and immune responses against bacterial and viral infection27,29,30. Therefore, identifying C. perfringens type C resistant-associated lncRNAs and mRNAs and these potential functions in defensing bacterial infection were essential to further explore regulatory mechanism of host and prevention infectious diseases caused by C. perfringens type C.
C. perfringens type C infection cause some inflammatory and immune diseases, while the intensity and persistence of these diseases were associated with the resistance differences of piglet individuals to pathogen infection. Indeed, C. perfringens type C infection have affected the growing development of inoculated piglets, the inoculated piglets exhibited the growth delay and deficiency, the weights of body, heart, liver and spleen of piglets in the IR group were higher than those in the IS group (P < 0.01), suggesting that potential resistant piglets showed fewer adverse effects on growth and development than sensitive piglets. In addition, C. perfringens type C infection had changed the expression levels of several inflammatory cytokine genes. The dysregulated expressions of cytokines play a positive role in contributing to host immune and inflammatory responses in the intestine damage and host defense the invasion of various microorganisms, which may be associated with many autoimmunity diseases31,32,33,34,35. The IFN-α, TNF-α31, IL-1β32 and NF-κB33 are crucial mediators of inflammation, researches had reported that TNF-α and IL-1 could induce biological activities of IL-8 in various types of inflammation34. In our study, the overexpressed IFN-α, TNF-α and downregulated of IL-1β and NF-κB in the IR group were significantly different with these in the IS group after C. perfringens type C infection. The results showed that the lower inflammatory responses were activated in the resistant piglets, in other words, resistant piglets may have the better ability in reducing inflammatory responses induced by C. perfringens type C infection and may be more beneficial for improving their prognosis.
A total of 10 lncRNAs and 100 mRNAs were found significantly differentially expressed between IR vs IS group. We found that after C. perfringens type C infection, the differentially expressed lncRNAs LNC_001066, LNC_000231, LNC_001186 and ENSSSCT00000032859 were found to regulate some immune-related target genes interleukin 1 receptor associated kinase 3 (IRAK3), TLR8, LCP2, TNFRSF11A, CYP1A1 and SLA-1, meanwhile, these target genes were also significantly differentially expressed in the IR vs IS group, the differentially expressed lncRNA target genes were mainly enriched in some key signal transduction and immune-related signaling pathways, such as toll-like receptor signaling pathway, NF-κB signaling pathway and cytokine-cytokine receptor interaction, which were considered to be associated with bacterial infection, especially C. perfringens36,37. These results may hint the potential effects of lncRNAs and target genes on signal transduction and cytokine. To further explore the potential functions of lncRNAs, we predicted the possible regulatory relationships by constructing a network diagram, which based on the following criteria: firstly, the immune related target genes of differentially expressed lncRNAs in cis and trans roles were predicted through target gene prediction, then we search the downstream inflammation related cytokine genes of immune target genes through KEGG signaling pathway database and some related articles. Based on the methods, a potential relationship among the 7 lncRNAs, target genes and inflammation related cytokines was evaluated and presented in the Supplementary Figure. As shown, the differentially expressed lncRNAs had potential regulatory relationship with their target genes. Furthermore, these target genes directly or indirectly affected the expressions of inflammation related cytokine genes IL-1β, IFN-α, TNF-α, NF-κB, CCL5, and so on, for example, the upregulated LNC_001066 could improve the expression of target genes TLR8, IRAK3 and LCP2, overexpression of TLR8, IRAK3 and LCP2 genes directly or indirectly affected the expressions of TRAF6, NF-κB and MAPK, respectively, which may trigger the secretions of cytokines IFN-α, IL-8 and TNF-α through NF-κB and MAPK signaling pathways. LNC_000231 could directly upregulated the expression of TNFRSF11A, TRAF3 gene could be activated by the upregulated TNFRSF11A, and further affect the expression level of NF-κB, all of them participated in regulating host inflammatory and immune responses to C. perfringens type C infection. This analysis was performed to reveal the potential relationship between the differentially expressed lncRNAs, target genes and cytokine changes, which might provide some new perspectives for understanding the transcriptional regulation of piglet immune response to bacterial infection.
After C. perfringens type C infection, the expressions of LNC_001066 in the IR group were significantly upregulated compared with IS group, while the expressions of LNC_000231 was only found expressed in the IS group, the expressions of IRAK3 and TLR8 in the IR group were significantly higher than those in the IS group. To further explore the regulatory roles of lncRNAs, we predicted the potential relationship between lncRNA, target genes and cytokines. As shown in the Table 2, LNC_001066 and LNC_000231 co-regulated the expressions of immune-related target genes, IRAK3 and TLR8, to participate in the progress of host immune response in defensing bacterial infection through apoptosis and toll-like receptor signaling pathway, respectively. TLR8 is one of toll-like receptors (TLRs) family, which is the first line of host defense against invading pathogen invasion. TLR8 can recognize distinct pathogen-associated molecular patterns in intracellular and play a critical role in host innate immune responses after bacterial infection38. TLR8 can activate a series of cascade reaction, including inducing the expressions of IRAK and TRAF639,40, and improve the productions of IFN-α, NF-κB through NF-κB and MAPK signaling pathways41. As the core immune regulator, the activations of NF-κB and MAPK pathways trigger the expressions of proinflammatory cytokines and chemokines, including the secretion of TNF-α, IFN-α, IL-1β and IL-838,42. The suppressed TLRs could reduce systemic inflammatory response caused by bacterial infection. It was predicted that TLR8 had potential relationship with the expressions of cytokines IFN-α and IL-1β through mediating expressions of TRAF6. C. perfringens infection suppressed the expression of TLR8 in inoculated piglets, which was relatively higher expressed in IR group than that in IS group, interestingly, IFN-α was correspondingly higher expressed in IR group than in IS group, as well as expression of IL-1β at 1–3 dpi.
Beside this, LNC_000231 could target expression of TNFRSF11A gene, as known as receptor activator of NF-κB (RANK). TNFRSF11A can affect intracellular signal transduction process through several TRAFs and may lead to the activation of various signaling pathways, including the NF-κB, MAPK and PI3K/AKT cascades43. The overexpression of TNFRSF11A can interacted with TNF receptor-associated factors (TRAF) family members, such as TRAF3 and TRAF6 to activate the NF-κB-mediated apoptosis44. In our study, the expressions of TNFRSF11A were significantly upregulated after piglets inoculated with C. perfringens type C, and the expression in the IS group was higher than that in the IR group, suggesting that high expression of TNFRSF11A might trigger apoptosis process of intestinal cell45 and had an adverse effect in host defense the invasion of C. perfringens type C.
Lymphocyte cytosolic protein 2 (LCP2) is one of the immune genes targeted by LNC_001066 and involve in NK cell-mediated cytotoxicity pathway. LCP2 plays important roles in promoting normal T-cell development and activation through MAPK signaling pathway46. Study found that the up-regulated LCP2 could promote angiogenesis during precancerous lesion formation and activated the production of excessive amounts of proinflammatory cytokines, such as IFN-γ and TNF-α47,48, the loss of LCP2 revealed a variable degree of abnormal intestinal vasculature in human and exhibited the hemorrhagic symptoms of subcutaneous and intraperitoneal tissues in mice49.The expression of LCP2 in the IS group was lower than in the IR group, correspondingly, the expressions of IFN-α and TNF-α were also lower in the IS group, this results might cause some adverse symptom and resulted in the occurrence of damage lesions in C. perfringens type C infected piglets, including cell cycle regulation disorders and decreased local immunity50.
ENSSSCT00000032859 targeted the expression of SLA-1 by antigen processing and presentation pathway, which was overexpressed in the IR group, and was not detected in the IS group. Study had reported that SLA-1 could bind with toxic polypeptide and induced a series of immune responses by recognizing CD8+ cell, E. coli F18-resistant piglets exhibited high expression levels of SLA-1, which was related to several immune functions and could help to defense E. coli F18-related porcine gastrointestinal tract diseases51. The results also suggested that the high expression of SLA-1 in resistant piglets may have the stronger capacity to defend C. perfringens type C infection.
The cytochrome P450 family 1 subfamily A member 1 (CYP1A1), directly targeted by LNC_001186, have some special relevance to metabolic activation and detoxication52, CYP1A1 may stimulate the productions of inflammatory cytokines in immune and inflammatory responses, including TNF-α, IL-1β, IL-6, CXCL5 and so on53. The activated CYP1A1 is associated with tissue toxicity and carcinogenesis, such as gastrointestinal tract54. Study found that intestine CYP1A1 contributed to a metabolic “shield” protecting host from ingesting carcinogens by interplaying with TLRs ligands55. Therefore, the higher expressed CYP1A1 in the IS group might have some association with the expression of IL-1β, the higher expressed IL-1β reflected that the sensitive piglets might suffer more intense immune responses during C. perfringens type C infection.
In addition, TNIP1 (TNF-α induced protein 3 (TNFAIP3 or A20) interacting protein 1) could be targeted by ENSSSCT00000018610. Overexpression of TNIP1 gene interact with TNFAIP3 to restrict NF-κB activation, then further reduced autoimmunity response by TNF-independent signals56 and prevented intestine immune and inflammatory diseases57,58,59. Study reported that TNFAIP3 repressed cell apoptosis by TNF and IL-1β, and affected proinflammatory gene expressions by directly acting on TRAF6 signaling molecule60. EFR3A and SPOCD1 gene were regulated by ALDBSSCT0000007865 and ALDBSSCT0000009442, respectively. EFR3A plays a role in controlling G protein-coupled receptor (GPCR) activity by affecting receptor phosphorylation, the mutation of this gene is associated with the early step of colorectal tumorigenesis61, changed expression of EFR3A contributes to host immune response against endocytosis of infectious bursal disease virus (IBDV) for preventing and decreasing infections62. SPOCD1 protein product belongs to TFIIS family transcription factors, SPOCD1 plays important functions in mediating inhibition of cancer cell proliferation63 and inducting cell apoptosis in breast, lung, gastric and pancreatic cancer cell lines through activation of p5364 and TGF-β signaling pathways65,66, the biological process involved by these genes ultimately plays a crucial role in the regulation of host inflammatory response.
The above results strongly implied the relationship among expression of lncRNA, target genes and cytokine changes, suggesting dysregulated lncRNAs affected inflammatory and proinflammatory cytokine expression by triggering the functional immune-related genes, meanwhile, expression differences of these molecules ultimately reflected in the resistance and susceptibility of host during pathogenic bacteria invasion. The piglets with higher resistance may have more abilities to reduce and counteract the damage caused by bacterial infection, though the definite information of how these lncRNAs regulate host immune response by affecting inflammatory cytokine changes need to be further validated.
Conclusions
In this study, we have comprehensive compared the lncRNA expression patterns in intestinal inflammation response of piglets with different immune resistance in defensing C. perfringens type C infection. These significantly expressed lncRNA can trigger the target genes to influence cytokine expression, and further contribute to the different abilities of piglet resistance to bacterial infection or induce the injure of the inflammation diseases. This study will provide the pivotal resources for further exploring the resistance and susceptibility of hosts to C. perfringens infection.
References
Huang, X. Y. et al. Effect of Genetic Diversity in Swine Leukocyte Antigen-DRA Gene on Piglet Diarrhea. Genes. 7, 36, https://doi.org/10.3390/genes7070036 (2016).
Songer, J. G., Pfeffer, M., Truyen, U. & Gaastra, W. Clostridia as agents of zoonotic disease. Vet microbiol. 140, 399–404, https://doi.org/10.1016/j.vetmic.2009.07.003 (2010).
Songer, J. G. & Uzal, F. A. Clostridial enteric infections in pigs. J Vet Diagn Invest. 17, 528–536, https://doi.org/10.1177/104063870501700602 (2005).
Chan, G. et al. The epidemiology of Clostridium perfringens type A on Ontario swine farms, with special reference to cpb2-positive isolates. BMC Vet Res. 8, 156, https://doi.org/10.1186/1746-6148-8-156 (2012).
Matsushita, O. & Okabe, A. Clostridial hydrolytic enzymes degrading extracellular components. Toxicon. 39, 1769–1780, https://doi.org/10.1016/S0041-0101(01)00163-5 (2001).
Scharff, R. L. Economic burden from health losses due to foodborne illness in the United States. J Food Prot. 75, 123, https://doi.org/10.4315/0362-028X.JFP-11-058 (2012).
Popescu, F. et al. Susceptibility of primary human endothelial cells to C. perfringens beta-toxin suggesting similar pathogenesis in human and porcine necrotizing enteritis. Vet microbiol. 153, 173–177, https://doi.org/10.1016/j.vetmic.2011.02.017, Epub 2011 Feb 23 (2011).
Zanella, R. et al. Identification of loci associated with tolerance to Johne’s disease in Holstein cattle. Anim genet. 42, 28–38, https://doi.org/10.1111/j.1365-2052.2010.02076.x (2011).
Paralkar, V. R. et al. Lineage and species-specific long noncoding RNAs during erythro-megakaryocytic development. Blood. 123, 1927–1937, https://doi.org/10.1182/blood-2013-12-544494, Epub 2014 Feb 4 (2014).
Zhao, W. et al. Systematic identification and characterization of long intergenic non-coding RNAs in fetal porcine skeletal muscle development. Sci Rep. 5, 8957, https://doi.org/10.1038/srep08957 (2015).
Kretz, M. et al. Suppression of progenitor differentiation requires the long noncoding RNA ANCR. Genes Dev. 26, 338–343, https://doi.org/10.1101/gad.182121.111, Epub2012 Feb 2 (2012).
Zhou, Y. et al. Integrative Analysis Reveals Enhanced Regulatory Effects of Human Long Intergenic Non-Coding RNAs in Lung Adenocarcinoma. J Genet Genomics. 42, 423–436, https://doi.org/10.1016/j.jgg.2015.07.001, Epub 2015 Jul 10 (2015).
Cui, W. et al. Discovery and characterization of long intergenic non-coding RNAs (lincRNA) module biomarkers in prostate cancer: an integrative analysis of RNA-Seq data. BMC genomics. 16, 7, S3, https://doi.org/10.1186/1471-2164-16-S7-S3, Epub 2015 Jun 11 (2015).
Kim, D. K. et al. Transcriptional profiles of host-pathogen responses to necrotic enteritis and differential regulation of immune genes in two inbreed chicken lines showing disparate disease susceptibility. Plos one. 9, e114960, https://doi.org/10.1371/journal.pone.0114960, eCollection2014 (2014).
Sarson, A. J. et al. Gene expression profiling within the spleen of Clostridium perfringens-challenged broilers fed antibiotic-medicated and non-medicated diets. BMC genomics. 10, 260, https://doi.org/10.1186/1471-2164-10-260 (2009).
Yang, Q. L., Kong, J. J., Wang, D. W., Zhao, S. G. & Gun, S. B. Swine Leukocyte Antigen-DQA Gene Variation and Its Association with Piglet Diarrhea in Large White, Landrace and Duroc. Asian-Australas J Anim Sci. 26, 1065–1071, https://doi.org/10.5713/ajas.2013.13067 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36, https://doi.org/10.1186/gb-2013-14-4-r36 (2014).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol. 28, 503–510, https://doi.org/10.1038/nbt.1633, Epub 2010 May 2 (2010).
Trapnell, C. et al. Transcript assembly and abundance estimation from RNA-Seq reveals thousands of new transcripts and switching among isoforms. Nature Biotechnol. 28, 511–515, https://doi.org/10.1038/nbt.1621, Epub 2010 May 2 (2010).
Sun, L. et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 41, e166, https://doi.org/10.1093/nar/gkt646, Epub 2013 Jul 27 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–349, https://doi.org/10.1093/nar/gkm391 (2007).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–D301, https://doi.org/10.1093/nar/gkr1065, Epub 2011 Nov 29 (2012).
Lin, M. F., Jungrei, I. & Kellis, M. PhyloCSF_ a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics. 27, i275–i282, https://doi.org/10.1093/bioinformatics/btr209 (2011).
Bruegge, J.Z., Einspanier, R. & Sharbati, S. A Long Journey Ahead: Long Non-coding RNAs in Bacterial Infections. Front Cell Infect Microbiol. 7, 95, https://doi.org/10.3389/fcimb.2017.00095 eCollection2017 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods. 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Molecular Cell. 43(6), 904–914, https://doi.org/10.1016/j.molcel.2011.08.018 (2011).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat Rev Genet. 10, 155–159, https://doi.org/10.1038/nrg2521 (2009).
Laine, A. L., Burdon, J. J., Nemri, A. & Thrall, P. H. Host ecotype generates evolutionary and epidemiological divergence across a pathogen metapopulation. Proc Biol Sci. 281, https://doi.org/10.1098/rspb.2014.0522 (2014).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science. 341, 789–792, https://doi.org/10.1126/science.1240925, Epub 2013 Aug 1 (2013).
Atianand, M. K. et al. A Long Noncoding RNA lincRNA-EPS Acts as a Transcriptional Brake to Restrain Inflammation. Cell. 165, 1672–1685, https://doi.org/10.1016/j.cell.2016.05.075 (2016).
Leppkes, M., Roulis, M., Neurath, M. F., Kollias, G., Becker, C. Pleiotropic functions of TNF-α in the regulation of the intestinal epithelial response to inflammation. Int Immunol. 26, 509–515, https://doi.org/10.1093/intimm/dxu051, Epub 2014 May 12 (2014).
Alnabhani, Z. et al. Pseudomonas fluorescens alters the intestinal barrier function by modulating IL-1β expression through hematopoietic NOD2 signaling. Inflamm Bowel Dis. 21, 543–555, https://doi.org/10.1097/MIB.0000000000000291 (2015).
Zhang, G. L. et al. Association of the NFKBIA gene polymorphisms with susceptibility to autoimmune and inflammatory diseases: a meta-analysis. Inflamm Res. 60, 11–18, https://doi.org/10.1007/s00011-010-0216-2, Epub 2010 May 21 (2011).
Miyamoto, K., Matsukawa, A., Ohkawara, S., Takagi, K. & Yoshinaga, M. IL8 is involved in homologous TNF-α, but not in IL1ß-induced neutrophil infiltration in rabbits. Inflamm Res. 46, 472–477, https://doi.org/10.1007/s000110050227 (1997).
Baccala, R., Hoebe, K., Kono, D. H., Beutler, B. & Theofilopoulos, A. N. TLR-dependent and TLR-independent pathways of type I interferon induction in systemic autoimmunity. Nat Med. 13, 543–551, https://doi.org/10.1038/nm1590 (2007).
Vallabhapurapu, S. & Karin, M. Regulation and Function of NF-κB Transcription Factors in the Immune System. Annu Rev Immunol. 27, 693–733, https://doi.org/10.1146/annurev.immunol.021908.132641 (2009).
Truong, A. D., Hong, Y. H. & Lillehoj, H. S. RNA-seq profiles of immune related genes in the spleen of necrotic enteritis-afflicted chicken lines. Asian-Australas J Anim Sci. 28(10), 1496–1511, https://doi.org/10.5713/ajas.15.0143 (2015).
Lu, Y. et al. Expression profiles of genes in Toll-like receptor-mediated signaling of broilers infected with Clostridium perfringens. Clin Vaccine Immunol. 16(11), 1639–1647, https://doi.org/10.1128/CVI.00254-09 (2009).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol. 11, 373–384, https://doi.org/10.1038/ni.1863, Epub 2010 Apr 20 (2010).
Heil, F. et al. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science. 303, 1526–1529, https://doi.org/10.1126/science.1093620 (2004).
Sloane, J. A., Blitz, D., Margolin, Z. & Vartanian, T. A clear and present danger: endogenous ligands of Toll-like receptors. Neuromolecular Med. 12, 149–163, https://doi.org/10.1007/s12017-009-8094-x, Epub 2009 Oct 14 (2010).
Cherfils-Vicini, J. et al. Triggering of TLR7 and TLR8 expressed by human lung cancer cells induces cell survival and chemoresistance. J Clin Invest. 120, 1285, https://doi.org/10.1172/JCI36551, Epub 2010 Mar8 (2010).
Darnay, B. G., Besse, A., Poblenz, A. T., Lamothe, B. & Jacoby, J. J. TRAFs in RANK signaling. Adv Exp Med Biol. 597, 152–159, https://doi.org/10.1007/978-0-387-70630-6_12 (2007).
Bharti, A. C., Takada, Y., Shishodia, S. & Aggarwal, B. B. Evidence that receptor activator of nuclear factor (NF)-kappaB ligand can suppress cell proliferation and induce apoptosis through activation of a NF-kappaB-independent and TRAF6-dependent mechanism. J Biol Chem. 279, 6065–6076, https://doi.org/10.1074/jbc.M308062200 (2004).
Knesebeck, A. V. D. et al. RANK (TNFRSF11A) Is Epigenetically Inactivated and Induces Apoptosis in Gliomas. Neoplasia. 14, 526–534. PMID: 22787434 (2012).
Cuadros, M. et al. Identification of a Proliferation Signature Related to Survival in Nodal Peripheral T-Cell Lymphomas. J Clin Oncol. 25, 3321–3329, https://doi.org/10.1200/JCO.2006.09.4474 (2007).
Siggs, O. M. et al. Quantitative reduction of the T cell receptor adapter protein SLP-76 unbalances immunity and immune regulation. J Immunol. 194, 2587–2595, https://doi.org/10.4049/jimmunol.1400326 (2015).
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354–357, https://doi.org/10.1093/nar/gkj102 (2005).
Abtahian, F. et al. Regulation of Blood and Lymphatic Vascular Separation by Signaling Proteins SLP-76 and Syk. Science. 299, 247–251, https://doi.org/10.1126/science.1079477 (2003).
Chen, D., Yang, K., Zhang, G., Mei, J. & Xiang, L. Screen and analysis of key disease genes for precancerous lesions of oral buccal mucosa induced by DMBA in golden hamsters. Oncol Lett. 2, 265–271, https://doi.org/10.3892/ol.2010.228 (2011).
Ye, L. et al. Investigation of the relationship between SLA-1 and SLA-3 gene expression and susceptibility to Escherichia coli F18 in post-weaning pigs. Comp Immunol Microbiol Infect Dis. 35, 23–30, https://doi.org/10.1016/j.cimid.2011.09.006, Epub 2011 Oct 22 (2012).
Alexandrov, K., Rojas, M. & Satarug, S. The critical DNA damage by benzo(a)pyrene in lung tissues of smokers and approaches to preventing its formation. Toxicol Lett. 198, 63–68, https://doi.org/10.1016/j.toxlet.2010.04.009, Epub 2010 Apr 24 (2010).
Uno, S. et al. Oral benzo pyrene in Cyp1 knockout mouse lines: CYP1A1 important in detoxication, CYP1B1 metabolism required for immune damage independent of total-body burden and clearance rate. Mol Pharmacol. 69, 1103–1114, https://doi.org/10.1124/mol.105.021501 (2006).
Vogel, C. F. A. et al. Transgenic Overexpression of Aryl Hydrocarbon Receptor Repressor (AhRR) and AhR-Mediated Induction of CYP1A1, Cytokines, and Acute Toxicity. Environ Health Perspect. 124, 1071–1083, https://doi.org/10.1289/ehp.1510194, Epub 2016 Feb 5 (2016).
Do, K. N., Fink, L. N., Jensen, T. E., Gautier, L. & Parlesak, A. TLR2 Controls Intestinal Carcinogen Detoxication by CYP1A1. PloS one. 7, e32309, https://doi.org/10.1371/journal.pone.0032309, Epub 2012 Mar 19 (2012).
Cheng, L. Q., Zhang, D. M. & Chen, B. Tumor necrosis factor α-induced protein-3 protects zinc transporter 8 against proinflammatory cytokine-induced downregulation. Exp Ther Med. 12, 1509–1514, https://doi.org/10.3892/etm.2016.3457 (2016).
Murphy, S. F., Rhee, L., Nero, T. M. & Boone, D. L. 736 Intestinal Epithelial-Cell (IEC) Specific Expression of Tumor Necrosis Factor Alpha-Induced Protein 3 (TNFAIP3) Reveals Vital Role for IECs in Mediating Intestinal Inflammation. Gastroenterology. 144, S132, https://doi.org/10.1016/S0016-5085(13)60476-X (2013).
Vereecke, L. et al. PS2-048. A20/TNFAIP3 in intestinal homeostasis and inflammation. Cytokine. 56, 75, https://doi.org/10.1016/j.cyto.2011.07.208 (2011).
Vereecke, L., Beyaert, R. & Loo, G. V. The ubiquitin-editing enzyme A20 (TNFAIP3) is a central regulator of immunopathology. Trends Immunol. 30, 383–391, https://doi.org/10.1016/j.it.2009.05.007, Epub 2009 Jul 28 (2009).
Boone, D. L. et al. The ubiquitin-modifying enzyme A20 is required for termination of Toll-like receptor responses. Nat Immunol. 5, 1052–1060, https://doi.org/10.1038/ni1110 (2004).
Zhou, D. et al. Exome Capture Sequencing of Adenoma Reveals Genetic Alterations in Multiple Cellular Pathways at the Early Stage of Colorectal Tumorigenesis. Plos one. 8, e53310, https://doi.org/10.1371/journal.pone.0053310, Epub 2013 Jan 2 (2013).
Hui, R. K. & Leung, F. C. Differential Expression Profile of Chicken Embryo Fibroblast DF-1 Cells Infected with Cell-Adapted Infectious Bursal Disease Virus. PloS one. 10, e0111771, https://doi.org/10.1371/journal.pone.0111771 eCollection2015 (2015).
Liang, J., Zhang, H., Hu, J., Liu, Y. & Li, Z. SPOCD1 promotes cell proliferation and inhibits cell apoptosis in human osteosarcoma. Mol Med Rep. 17, 3218–3225, https://doi.org/10.3892/mmr.2017.8263, Epub2017 Dec 12 (2018).
Hubbard, K., Catalano, J., Puri, R. K. & Gnatt, A. Knockdown of TFIIS by RNA silencing inhibits cancer cell proliferation and induces apoptosis. BMC Cancer. 8, 133, https://doi.org/10.1186/1471-2407-8-133 (2008).
Cha, Y., Kim, D. K., Hyun, J., Kim, S. J. & Park, K. S. TCEA3 binds to TGF-beta receptor I and induces Smad-independent, JNK-dependent apoptosis in ovarian cancer cells. Cell Signal. 25, 1245–1251, https://doi.org/10.1016/j.cellsig.2013.01.016, Epub 2013 Jan 26 (2013).
Zhu, M. et al. Exome Array Analysis Identifies Variants in SPOCD1 and BTN3A2 That Affect Risk for Gastric Cancer. Gastroenterology. 152, 2011–2021, https://doi.org/10.1053/j.gastro.2017.02.017, Epub 2017 Feb 27 (2017).
Acknowledgements
The work was supported by the discipline construction fund project of Gansu Agricultural University (GSAU-XKJS-2018—041).
Author information
Authors and Affiliations
Contributions
X.H. conceived and designed the study, analyzed the data and manuscript writing. W.S. made the statistical analysis and revised of the manuscript. S.G. approved of the version to be published and provided funding support for the conduct of the research. Z.Y. did the laboratory work in the expression, H.S., Q.Y., P.W., and S.L. contributed to collected data of the research. L.L., S.Z., and participated in the analysis and interpretation of data. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, X.Y., Sun, W.Y., Yan, Z.Q. et al. Novel Insights reveal Anti-microbial Gene Regulation of Piglet Intestine Immune in response to Clostridium perfringens Infection. Sci Rep 9, 1963 (2019). https://doi.org/10.1038/s41598-018-37898-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37898-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.