Abstract
To relate microbial diversity with various host traits of interest (e.g., phenotypes, clinical interventions, environmental factors) is a critical step for generic assessments about the disparity in human microbiota among different populations. The performance of the current item-by-item α-diversity-based association tests is sensitive to the choice of α-diversity metric and unpredictable due to the unknown nature of the true association. The approach of cherry-picking a test for the smallest p-value or the largest effect size among multiple item-by-item analyses is not even statistically valid due to the inherent multiplicity issue. Investigators have recently introduced microbial community-level association tests while blustering statistical power increase of their proposed methods. However, they are purely a test for significance which does not provide any estimation facilities on the effect direction and size of a microbial community; hence, they are not in practical use. Here, I introduce a novel microbial diversity association test, namely, adaptive microbiome α-diversity-based association analysis (aMiAD). aMiAD simultaneously tests the significance and estimates the effect score of the microbial diversity on a host trait, while robustly maintaining high statistical power and accurate estimation with no issues in validity.
Similar content being viewed by others
Introduction
The human microbiome studies have been accelerated by the recent advances in high-throughput sequencing technologies1,2,3 which enabled an unbiased characterization of all microbes from different organs (e.g., gut, mouth, skin, vagina, etc.) of the human body. One of the most fundamental steps in microbiome studies is to survey the disparity in microbial diversity among different populations (e.g., case vs. control, treatment vs. placebo, or smoking vs. non-smoking). For instance, reduced microbial diversity has been found to be associated with various host phenotypes, such as obesity4, fatty liver disease4, type II diabetes5, inflammatory bowel diseases6 and additional disorders7,8. Clinical interventions (e.g., antibiotic use) and environmental factors (e.g., diet, smoking, delivery mode) have also been found to shift up or down the microbial diversity9,10. For such microbial diversity association analyses, the most commonly used approach is to relate α-diversity (within-sample microbial diversity) with a host trait of interest based on traditional statistical methods (e.g., fitting a linear regression model for the association between α-diversity and a continuous trait (e.g., body mass index (BMI)) or a logistic regression model for the association between α-diversity and a binary trait (e.g., disease/treatment status) with or without covariate adjustments). Such α-diversity-based association analysis offers systematic statistical inference facilities including the effect estimates of microbial diversity on a host trait (e.g., regression coefficient estimates) as well as hypothesis testing tools (e.g., p-values). As a result, we can comprehensively assess which population has higher or lower microbial diversity with the extent of the disparity as well as whether it is statistically significant or not.
However, many of the recent microbial community-level association tests continued to ignore some of the fundamental elements of statistical inference. For example, MiRKAT11, MiSPU12 and OMiAT13 produce only p-values without any effect estimation facilities (i.e., purely a test for significance). Although they boast about statistical power increase, it is difficult to lead to any novel clinical interventions or public health promotion programs based solely on p-values. To explain, suppose that we found a significant difference in a microbial community (e.g., bacterial kingdom) between diseased and healthy populations using MiRKAT, MiSPU or OMiAT. However, here, the only available conclusion is that the two populations are simply different in microbial community composition with no further understanding about how the difference exists. Instead, α-diversity-based association analysis provides effect estimation on the disparity in direction and size of the microbial diversity among different populations (e.g., the diseased population is considerably lower in microbial diversity) which are essential to better understand microbial communities (e.g., lower microbial diversity may indicate higher morbidity) and make plans (e.g., plans to recover microbial diversity to normality). In ecology, α-diversity has also been widely used as a guideline for community ecologists and conservation biologists to make plans to preserve natural ecosystems or restore perturbed communities14,15,16.
Notably, a variety of α-diversity metrics can be considered in the analysis. Different α-diversity metrics reflect different views on the true diversity and they perform differently. For example, Richness (also known as Observed), Shannon17 and Simpson18 indices are non-phylogenetic metrics (i.e., based solely on abundance information) which weight relatively rare, mid-abundant and abundant species, respectively. Accordingly, they are suitable when associated species are rare, mid-abundant and abundant species, respectively. In contrast, phylogenetic diversity (PD)19, phylogenetic entropy (PE)20 and phylogenetic quadratic entropy (PQE)21,22 are phylogenetic metrics (i.e., based on both abundance and phylogenetic information) which weight relatively rare, mid-abundant and abundant species, respectively. The phylogenetic metrics are suitable when associated species have disparity in both abundance and phylogeny, where PD, PE and PQE are suitable when associated species are rare, mid-abundant and abundant species, respectively. In reality, associated species can be rare or abundant, or they can have disparity in phylogeny rather than abundance or vice versa. However, it is highly difficult to predict which situation among such various possible association patterns is the one for our study and to choose a single optimal α-diversity metric to use. This is because of the unknown nature of the true association. The approach of cherry-picking a test which has the smallest p-value or the largest effect size after running multiple item-by-item α-diversity-based association analyses is not statistically valid (e.g., do not correctly control type I error) because the multiplicity (i.e., multiple testing) issue is not properly accounted for23. Therefore, a valid statistical method which robustly suits various unknown association patterns is needed.
In this paper, I introduce a novel adaptive microbial diversity association test, namely, adaptive microbiome α-diversity-based association analysis (aMiAD), which robustly maintains high statistical power and accurate microbial diversity effect score estimation throughout various association patterns while satisfying the requisite validity issue. aMiAD employs the minimum p-value from multiple candidate item-by-item α-diversity-based association analyses as its test statistic and estimate its own p-value and microbial diversity effect score based on a residual-based permutation method. The use of minimum p-value statistic is to adaptively approach the highest power and the most accurate microbial diversity effect score estimation among multiple candidate analyses, while the residual-based permutation method based on the minimum p-value statistic is to robustly satisfy the validity issue (e.g., correctly controlling type I error) with no distributional assumption to be satisfied. Three non-phylogenetic metrics, Richness, Shannon, Simpson indices and three phylogenetic metrics, PD, PE and PQE are selected as the candidate α-diversity metrics for aMiAD because of their distinguished features which properly modulate abundance and phylogenetic information.
The rest of the paper is organized as follows. The methodological details for aMiAD can be found in the following Methods section. Then, extensive simulations and real data applications are addressed in the Results section. I finally discuss possible extensions for the use of aMiAD in the Discussion section.
Methods
I first organize related notations and models. Then, I address details on the six candidate α-diversity metrics, Richness, Shannon17, Simpson18, PD19, PE20 and PQE21,22. Finally, I delineate the test statistic and microbial diversity effect score of aMiAD and the residual permutation-based computational algorithm. While the application of aMiAD can be much broader (e.g., extendable to generalized linear models), I describe aMiAD to relate microbial diversity with a continuous (e.g., BMI) or a binary (e.g., disease/treatment status) trait.
Here, I notify that the α-diversity referred in this paper considers different types of operational taxonomic units (OTUs) in the bacterial kingdom per biological sample (e.g., human, mouse), indicating within-sample diversity of OTUs in the bacterial kingdom. However, in practice, any subunits (e.g., species or other lower-level microbial taxa) in a different microbial assemblage (e.g., kingdom of archaea, fungi, protists or viruses, phylum of firmicutes or bacteroidetes) can be considered.
Models and notations
Suppose that there are n samples, p OTUs in a microbial community (e.g., bacterial kingdom) and q covariates (e.g., age, gender). Let Yi denote a continuous (e.g., BMI) or a binary (e.g., disease/treatment status) trait, Zij denote OTUs, and Xik denote covariates for i = 1, …, n, j = 1, …, p and k = 1, …, q. To relate OTUs in a community with a host trait while adjusting for covariate effects, I consider a multiple linear regression model equation (1) for a continuous trait and a multiple logistic regression model equation (2) for a binary trait.
where β0 is a regression coefficient for the intercept, αk’s are regression coefficients for the effect of q covariates (e.g., age, gender), h (Zi) is a function which characterizes the relationship between OTUs and a host trait, and ∈i is an error term which is independently and identically distributed with a mean zero and a variance of σ2. Here, we are particularly interested in testing the null hypothesis, H0: h (Zi) = 0; that is, no association between OTUs and a host trait.
Notably, we can flexibly specify h (Zi) to reflect different patterns of the relationship. For example, the linear relationship between OTUs and a host trait can be surveyed by setting h (Zi) = \(\sum _{{\rm{j}}=1}^{{\rm{p}}}\,{{\rm{\beta }}}_{{\rm{j}}}{{\rm{Z}}}_{{\rm{ij}}}\), while diverse non-linear relationships can be surveyed by the use of non-linear transformations of OTUs (e.g., polynomials or splines)24,25. Furthermore, any positive semi-definite kernel function can be used for h (Zi), where MiRKAT11 has especially been credited with establishing a kernel machine regression framework for distance-based community-level association analysis. Among diverse alternatives, I formulate h (Zi) as a function of α-diversity metric equation (3) for the ultimate goal of inferring the effect of microbial diversity on a host trait.
where γ is an index for a chosen α-diversity metric (e.g., Richness, Shannon, Simpson, PD, PE, PQE), β(γ) is a regression coefficient for the α-diversity metric and D(γ) i’s are the values of the α-diversity metric for i = 1, …, n.
α-diversity indices
α-diversity is an intuitive and natural index which summarizes the extent of microbial diversity in a community. A variety of α-diversity metrics have been proposed, and they are classified into non-phylogenetic and phylogenetic metrics. The non-phylogenetic metrics are constructed based solely on microbial abundance information, while the phylogenetic metrics further utilize phylogenetic tree information. I here survey three non-phylogenetic metrics, Richness, Shannon17 and Simpson18 indices, and three phylogenetic metrics, PD19, PE20 and PQE21,22.
To begin with non-phylogenetic metrics, Richness, Shannon and Simpson indices are weighted variants based on the generalized diversity framework, known as the effective number of types (or Hill number), which quantifies how many effective types of interest exist in a community26,27,28. Here, the effective number of types (Dw) equation (4) is defined as the inverse of the mean weighted proportional abundance26,27.
where p is the total number of OTU types present in a community, rj is the relative abundance (i.e., proportion) of the j-th OTU for j = 1, …, p and w (\(\in {\mathbb{R}}\)) is the weight for the proportions (also known as the order of the diversity) which needs to be pre-specified.
Notably, with different pre-specifications for the order of the diversity (w) equation (4), different α-diversity metrics can be derived. In particular, when w = 0, D0 equals to p (i.e., the total number of OTU types present in a community) which is known as Richness (DRichness) equation (5).
where p is the total number of OTU types present in a community. When w = 1, D1 cannot be defined; hence, the mathematical limit of \({{\rm{l}}{\rm{i}}{\rm{m}}}_{{\rm{w}}\to 1}{{\rm{D}}}_{{\rm{w}}}=\exp (-\sum _{{\rm{j}}=1}^{{\rm{p}}}{{\rm{r}}}_{{\rm{j}}}\,{\rm{l}}{\rm{n}}\,{{\rm{r}}}_{{\rm{j}}})\)26,27 which is the weighted geometric mean proportional abundance is alternatively employed. Then, Shannon index (DShannon) equation (6) is derived by taking the logarithm to \({{\rm{l}}{\rm{i}}{\rm{m}}}_{{\rm{w}}\to 1}{{\rm{D}}}_{{\rm{w}}}\)17.
where p is the total number of OTU types present in a community and rj is the proportion of the j-th OTU for j = 1, …, p. When w = 2, D2 equals to \({(\sum _{{\rm{j}}=1}^{{\rm{p}}}{{{\rm{r}}}_{{\rm{j}}}}^{2})}^{-1}\), which is the weighted arithmetic mean proportional abundance known as Inverse Simpson index26,27. Then, Simpson index (DSimpson) equation (7) is derived by taking the minus of the inverse of D2, −D2−1 18.
where p is the total number of OTU types present in a community and rj is the proportion of the j-th OTU for j = 1, …, p.
Importantly, by the formula equation (4), we can infer that as the value of w increases, relatively abundant OTUs are weighted, but it is vice versa as the value of w decreases27. Therefore, Richness, Shannon and Simpson indices weight relatively rare, mid-abundant and abundant OTUs, respectively; hence, they are also suitable when associated OTUs are rare, mid-abundant and abundant, respectively.
In contrast, the phylogenetic metric, PD19, utilizes phylogenetic tree information while considering only the incidence (i.e., presence/absence) information of OTUs. Specifically, PD (DPD) is defined as the sum of the lengths of the branches for the OTUs present in a community equation (8).
where p is the total number of OTU types present in a community and 1j is the length of all the branches that belong to the j-th OTU for j = 1, …, p. Therefore, PD is suitable when associated OTUs have high disparity in phylogeny rather than in abundance. Given that prevalent OTUs are likely to be present in all samples, PD is also suitable especially for rare OTUs which have high disparity in the classification of presence/absence.
PE20 equation (9) and PQE21,22 equation (10) are phylogenetic generalizations of the Shannon and Simpson indices, which incorporate all differing microbial abundance information (i.e., beyond the incidence (presence/absence) information for PD) while weighting relatively mid-abundant and abundant OTUs.
where p is the total number of OTU types present in a community, 1j is the length of all the branches that belong to the j-th OTU and rj is the proportion of the j-th OTU for j = 1, …, p. Therefore, PE and PQE are suitable when associated OTUs have high disparity in phylogeny, where they are relatively mid-abundant and abundant, respectively.
The above α-diversity metrics are the most fundamental and widely used, and they were sufficient in my simulations and real data analyses. Yet, the potential extension to other α-diversity metrics is addressed later in Discussion.
aMiAD
aMiAD is constructed based on the score test29 of the linear equation (1) or logistic equation (2) regression model, which surveys the association between each of the α-diversity metrics and a host trait while adjusting for covariates. Here, the unstandardized score statistic (U(γ)) is formulated with equation (11).
where γ is an index for a chosen α-diversity metric (e.g., Richness, Shannon, Simpson, PD, PE, PQE) and \({\hat{{\rm{\mu }}}}_{{\rm{i}},0}\) is the fitted value under the null hypothesis, which is estimated as \({\widehat{{\rm{\beta }}^{\prime} }}_{0}+{\sum }_{{\rm{k}}=1}^{{\rm{q}}}{{\rm{X}}}_{{\rm{i}}{\rm{k}}}{\widehat{{\rm{\alpha }}^{\prime} }}_{{\rm{k}}}\) for the linear regression model equation (1) or \({{\rm{logit}}}^{-1}({\widehat{{\rm{\beta }}^{\prime} }}_{0}+{\sum }_{{\rm{k}}=1}^{{\rm{q}}}{{\rm{X}}}_{{\rm{ik}}}{\widehat{{\rm{\alpha }}^{\prime} }}_{{\rm{k}}})\) for the logistic regression model equation (2), where \({\widehat{{\rm{\beta }}^{\prime} }}_{0}\) and \({\widehat{{\rm{\alpha }}^{\prime} }}_{{\rm{k}}}\) are maximum likelihood estimates (MLEs) under the null hypothesis. This unstandardized score statistic (U(γ)) is sufficient to estimate the p-value (P(γ)) based on my residual permutation-based method (see Computational algorithm) because its mean and standard error are evaluated under the null hypothesis equivalently for both the observed and null (i.e., permuted) statistic values resulting in no change in their relative comparison25. Yet, the mean and standard error under the null hypothesis are also estimated to derive the standardized score statistic (\({{\rm{U}}}_{({\rm{\gamma }})}^{\ast }\)). The standardized score statistic (\({{\rm{U}}}_{({\rm{\gamma }})}^{\ast }\)) is asymptotically related to the regression coefficient (β(γ)) equation (3) and tells effect direction and size of a chosen α-diversity metric29,30. I denote \({{\rm{U}}}_{({\rm{\gamma }})}^{\ast }\) as MiDivES(γ) and use it as the effect score of a chosen α-diversity metric.
Here, the score test equation (11) with its resulting p-value (P(γ)) and effect score (MiDivES(γ)) handles α-diversity metrics one-by-one. Yet, as described above, the performance differs according to the choice of α-diversity metric and the true underlying association pattern. Because of the unknown nature of the true association pattern, we cannot predict which α-diversity index is the optimal choice to our study in advance. Therefore, in order to robustly suit various association patterns, I propose a data-driven adaptive test, aMiAD. The test statistic of aMiAD (TaMiAD) is the minimum p-value from multiple item-by-item α-diversity-based association analyses equation (12).
where γ is an index for a metric in a set of multiple candidate α-diversity metrics (Γ), where Γ = {Richness, Shannon, Simpson, PD, PE, PQE}, and P(γ) is the estimated p-value for the use of each α-diversity metric (γ ∈ Γ). Here again, TaMiAD equation (12) is the test statistic of aMiAD, and this minimum p-value (i.e., \({{\rm{T}}}_{{\rm{a}}{\rm{M}}{\rm{i}}{\rm{A}}{\rm{D}}}={min}_{\gamma \epsilon {\rm{\Gamma }}}{{\rm{P}}}_{(\gamma )}\) equation (12)) itself is not the p-value I report for aMiAD. The approach of cherry-picking the minimum p-value among multiple candidate analyses (i.e., \({{\rm{T}}}_{{\rm{a}}{\rm{M}}{\rm{i}}{\rm{A}}{\rm{D}}}={min}_{\gamma \epsilon {\rm{\Gamma }}}{{\rm{P}}}_{(\gamma )}\) equation (12)) and reporting it (i.e., \({{\rm{T}}}_{{\rm{a}}{\rm{M}}{\rm{i}}{\rm{A}}{\rm{D}}}={min}_{\gamma \epsilon {\rm{\Gamma }}}{{\rm{P}}}_{(\gamma )}\) equation (12)) as it is cannot correctly control type I error rates because of the inherent multiplicity (i.e., multiple testing) issue23. I use a residual permutation-based method (see Computational algorithm) based on the minimum p-value statistic equation (12) to estimate the p-value for aMiAD (denoted as PaMiAD).
The estimated microbial diversity effect score of aMiAD, namely, adaptive microbial diversity effect score (aMiDivES) equation (13), is the standardized score statistic value based on the α-diversity metric which results in the minimum p-value among multiple candidate analyses, which is then further standardized by its mean and standard error under the null hypothesis.
where γm is an index of the metric which results in the minimum p-value in a set of multiple candidate α-diversity metrics (Γ), where Γ = {Richness, Shannon, Simpson, PD, PE, PQE}, MiDivES(γm) is an estimated microbial diversity effect score for the α-diversity metric which results in the minimum p-value, E(MiDivES(γm), 0) and SE(MiDivES(γm), 0)) are the mean and standard error of MiDivES(γm) under the null hypothesis. Here again, aMiDivES is the E(MiDivES(γm) which is further standardized by its mean (E(MiDivES(γm), 0)) and standard error (SE(MiDivES(γm), 0)) under the null hypothesis equation (13), and the genuine microbial diversity effect score of the test reaching the minimum p-value (i.e., MiDivES(γm)) is not the microbial diversity effect score I report for aMiAD. I use a residual permutation-based method (see Computational algorithm) to estimate the mean (E(MiDivES(γm), 0)) and standard error (SE(MiDivES(γm), 0)).
Computational algorithm
The computational algorithm to estimate the p-value (PaMiAD) and the effect score (aMiDivES) of aMiAD is based on a residual-based permutation method which randomly shuffles the residuals estimated from the null model, which reflects the null situation of no association. It is constructed based on the score statistic equation (11) and its derivatives equations (12) and (13) which do not require MLE; hence, we can avoid heavy computation and no convergence error in the iterative algorithm for MLE. It is non-parametric; hence, the outcomes are robustly valid with no underlying distributional assumption to be satisfied. The approach based on the minimum p-value statistic and a residual-based permutation method has also been widely used in prior studies11,12,13,25,31, where the validity issue was robustly satisfied. Detailed procedures can be found in (Supplementary S1 Text).
Ethics approval and consent to participate
Not applicable. This study involves only secondary analyses. All utilized microbiome datasets are publicly and freely available which do not require any ethics approval and consent to participate.
Results
Simulations
I conducted simulation experiments under a wide range of scenarios in order to evaluate and compare item-by-item α-diversity-based association tests and aMiAD in terms of hypothesis testing (i.e., type I error and power) and effect score estimation (i.e., central tendency, dispersion and accuracy). I also evaluate the approach of cherry-picking a test which has the smallest p-value (denote it as Minimum P) or the largest effect size (i.e., the largest deviation from zero effect) (denote it as Largest ES) among multiple item-by-item α-diversity-based association analyses in terms of the validity issues of properly controlled type I error and the central tendency and dispersion of microbial diversity effect scores under the null hypothesis. I also evaluate other existing adaptive community-level association tests (i.e., Optimal MiRKAT (OMiRKAT)11, adaptive MiSPU (aMiSPU)12 and OMiAT13) in terms of hypothesis testing only (i.e., type I error and power) as they do not provide any effect estimation facilities. I applied default settings for the implementation of their software package (aMiAD ver. 1.0, MiRKAT ver. 1.0.1, MiSPU ver. 1.0, and OMiAT ver. 5.3), as suggested.
Simulation design
I simulated microbiome data according to prior studies11,13,25 which reflect real OTUs’ proportions and dispersion on the basis of the Dirichlet-multinomial distribution32. In particular, I used real gut microbiome data33 from 35 fecal samples (collected from non-obese diabetic (NOD) mice at 6 weeks of age in the control group with no antibiotic treatment) for 353 OTUs (after removing OTUs with proportional mean abundance ≤10−4) to estimate the proportions and dispersion parameter. Then, simulation data were iteratively generated from the Dirichlet-multinomial distribution with the pre-specified values of the estimated proportions and dispersion parameter and the total reads per sample of 1,000 for small (n = 50) and large (n = 100) sample sizes, respectively11,13,25. Then, binary outcomes were generated based on the logistic regression model equation (14)11,13.
where X1i and X2i are two covariates (e.g., age and gender) simulated from the normal distribution with mean 50 and standard deviation (SD) 5 and the Bernoulli distribution with success probability 0.5, respectively, β is a scalar value (\(\in {\mathbb{R}}\)) which determines the effect direction and size of the associated OTUs in a set Λ, where Zij is an OTU count and wi is a weight for the phylogenetic disparity defined as the sum of the branch lengths for present OTUs divided by the sum of the branch lengths for absent OTUs, and ‘scale’ is the standardization function to have mean 0 and SD 111,13,25. To estimate empirical type I error rate and the mean (as a measure of central tendency) and variance (as a measure of dispersion) of microbial diversity effect scores under the null hypothesis, I set β = 0. To estimate statistical power and the accuracy of effect scores, I set β from the uniform distribution between −3 and 3 (i.e., Unif(−3, 3)). Here, the R2 value between β values randomly generated from Unif(−3, 3) and microbial diversity effect scores estimated from each method was used as a measure of estimation accuracy. The set of associated OTUs in the community (Λ) was selected with four different scenarios: (1) Λ = {OTUs in bottom 20% in abundance}, (2) Λ = {A random 20% of OTUs}, (3) Λ = {OTUs in top 20% in abundance}, (4) Λ = {OTUs in a cluster among 7 clusters partitioned by partitioning-around-medoids (PAM) algorithm}, respectively. The first three scenarios mimic the situations when rare, mid-abundant and abundant OTUs, respectively, are associated. For the fourth scenario, I used PAM algorithm34 to partition all OTUs in the community into 7 clusters based on their cophenetic distances. Here, the number of clusters, 7, was selected by maximizing the average silhouette width from 5 to 10 candidate numbers of clusters35,36. I randomized the choice of an associated cluster among the 7 clusters to avoid arbitrary choice13,25, whereas the outcomes for each of the 7 clusters can be found in Supporting Information (Fig. S1). The fourth scenario mimics the situation when phylogenetically close OTUs are associated.
Simulation results
Type I error
I estimate that the empirical type I error rates are well-controlled at the significance level of 0.05 for aMiAD, as well as all item-by-item α-diversity-based association tests and adaptive community-level association tests (OMiRKAT, aMiSPU and OMiAT), for both small (n = 50) and large (n = 100) sample sizes (Table 1). However, the cherry-picking approaches (i.e., Minimum P and Largest ES) show overly inflated empirical type I error rates for both small (n = 50) and large (n = 100) sample sizes (Table 1), indicating the violation of the requisite validity issue in hypothesis testing.
Central tendency and dispersion of effect scores under the null hypothesis
I estimate that the means of microbial diversity effect scores under the null hypothesis are around zero, indicating no bias in the estimation, for all surveyed tests and for both small (n = 50) and large (n = 100) sample sizes (Table 2). I also estimate that the variances of microbial diversity effect scores under the null hypothesis are around one for aMiAD, as well as all the item-by-item α-diversity-based association tests, for both small (n = 50) and large (n = 100) sample sizes (Table 2). However, the cherry-picking approaches (i.e., Minimum P and Largest ES) show overly inflated variance estimates for both small (n = 50) and large (n = 100) sample sizes (Table 2), indicating over-estimation of effect size.
Power and estimation accuracy
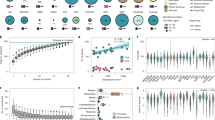
To begin with comparing the performance of α-diversity-based association tests, Richness estimates the greatest power and R2 values when rare OTUs are associated for both small (n = 50) (Figs 1A,C and (S1)) and large (n = 100) (Figs 1B,D and (S1)) sample sizes, while the Shannon index estimates the greatest power and R2 values when mid-abundant OTUs are associated for both small (n = 50) (Figs 1A,C and (S2)) and large (n = 100) (Figs 1B,D and (S2)) and the Simpson index estimates the greatest power and R2 values when abundant OTUs are associated for both small (n = 50) (Figs 1A,C and (S3)) and large (n = 100) (Figs 1B,D and (S3)), which are explained by their abundance weighting schemes. When phylogenetically close OTUs are associated (i.e., OTUs in a random cluster among the 7 clusters partitioned by the PAM algorithm are associated), the phylogenetic metrics (i.e., PD, PE and PQE) estimates greater power and R2 values than the non-phylogenetic metrics (i.e., Richness, Shannon and Simpson) for both small (n = 50) (Figs 1A,C and (S4)) and large (n = 100) (Figs 1B,D and (S4)) sample sizes, where PE estimates the greatest power and R2 values. This is because the phylogenetic metrics further incorporate phylogenetic information, while the non-phylogenetic metrics are based only on abundance information. To be more detailed, the performance also varies by which cluster among the 7 clusters partitioned by PAM algorithm is selected (see Supporting Information (Fig. S1)). That is, the Shannon index estimates the greatest power and R2 values when OTUs in the first cluster are associated (Fig. S1A–D(C1)), PE estimates the greatest power and R2 values when OTUs in the second, third, fifth and sixth clusters are associated (Fig. S1A–D(C2, C3, C5, C6)), and PQE estimates the greatest power and R2 values when OTUs in the fourth cluster are associated (Fig. S1A–D(C4, C7)).
Estimated powers and R2 values. (A) Estimated powers for the α-diversity-based association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 50). (B) Estimated powers for the α-diversity-based association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 100). (C) Estimated powers for the adaptive association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 50). (D) Estimated powers for the adaptive association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 100). (E) Estimated R2 values for the α-diversity-based association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 50). (F) Estimated R2 values for the α-diversity-based association tests for the scenarios, where rare (S1), mid-abundant (S2), abundant (S3) and phylogenetically close (S4) OTUs are associated (n = 100). **S1. Λ = {OTUs in bottom 20% in abundance}; S2. Λ = {A random 20% of OTUs}; S3. Λ = {OTUs in top 20% in abundance}; S4. Λ = {OTUs in a random cluster (average) among the 7 clusters partitioned by PAM}.
Although it may not be feasible to reflect all possible true association patterns in the natural world to our simulations, the most meaningful observation here is that aMiAD adaptively approaches the greatest power and R2 values among different item-by-item analyses throughout all surveyed scenarios (Figs 1A–D and S1A–D), while the performance for each α-diversity metric considerably fluctuates (Figs 1A–D and S1A–D). In reality, the true association scenario is mostly unknown, while a variety of scenarios are also likely to exist. Thus, aMiAD is attractive due to its high adaptivity and robustness to better cope with the unknown nature.
To compare aMiAD with the three adaptive community-level association tests (OMiRKAT, aMiSPU and OMiAT) (Figs 1E,F and S1E,F), OMiAT estimates the greatest power values for most of the scenarios except that aMiAD estimates the greatest power values for small sample size (n = 50) when abundant OTUs (Figs 1E and (S3)) and OTUs in the second cluster among the 7 clusters partitioned by the PAM algorithm are associated (Fig. S1E(C2)), aMiSPU estimates the greatest power values when OTUs in the fourth cluster are associated for both small (n = 50) (Fig. S1E(C4)) and large (n = 100) (Fig. S1F(C4)) sample sizes and OMiRKAT estimates the greatest power values when OTUs in the seventh cluster are associated for both small (n = 50) (Fig. S1E(C7)) and large (n = 100) (Fig. S1F(C7)) sample sizes. To summarize, we may conclude that OMiAT is most robustly powerful. However, once again, OMiAT, as well as OMiRKAT and aMiSPU, does not provide any effect estimation facilities; hence, its interpretability and usability are limited.
Real data applications
The disparity in microbial diversity between control and antibiotic treatment groups
Cox et al. (2013) performed microbiota-profiling studies to survey if the gut microbiota affected during maturity by antibiotic treatment leads to continued metabolic consequences37. To demonstrate the use of aMiAD, I analyzed a part of the original data, which surveys the effect of antibiotic treatment with low-dose penicillin (LDP) on microbial diversity of the gut microbiota. In particular, I compared microbial diversity of the bacterial kingdom between two groups of mice, 8 control and 7 antibiotic treatment mice. To summarize the sampling and profiling procedures while details are found in the original literature37, the 8 control mice are 8 germ-free mice to whom cecal microbiota from mice with no treatment were transferred and the 7 antibiotic treatment mice are 7 germ-free mice to whom cecal microbiota from LDP-treated mice were transferred. Fecal samples from the 8 control and 7 antibiotic treatment mice were collected after 23 days of the transfer, and the V4 region of the bacterial 16S rRNA gene was targeted in the amplicon sequencing with barcoded fusion primers38. Then, the QIIME pipeline2 was used to quantify OTUs and construct their phylogenetic tree. The OTUs were rarefied using the software package, phyloseq39 due to the varying total reads per sample40. 59 OTUs were included in the analysis after removing OTUs which are not present in any sample after random subsampling of the rarefaction39. Here, only a few OTUs (i.e., 59 OTUs), which may not represent the entire ecosystem, were analyzed because of some data quality issues (e.g., small sample size, low sequencing depth and the antibiotic treatment effect which can substantially reduce microbial abundance/diversity).
We can first visually observe in the box-plots (Fig. 2A) that all the α-diversity metrics are lower for the antibiotic treatment group than the control group, while PD and then Richness show the greatest disparity. Correspondingly, we can observe negative estimated effect scores for all α-diversity metrics, indicating microbial diversity is lower for the antibiotic treatment group than the control group, where the disparity is especially significant for PD (p-value: <0.001) and Richness (p-value: <0.001) indices (Fig. 2B). aMiAD estimates that microbial diversity is significantly different between the two groups (p-value: 0.001), where the microbial diversity is lower for the antibiotic treatment group than the control group (aMiDivES: −2.028 < 0) (Fig. 2B).
The comparison between the control (Con) and antibiotic treatment (Ant) groups. (A) The box-plots for each α-diversity metric (i.e., Richness, Shannon, Simpson, PD, PE, PQE). (B) The outcomes for the item-by-item (i.e., Richness, Shannon, Simpson, PD, PE, PQE) and adaptive (i.e., aMiAD) α-diversity-based association tests.
The disparity in microbial diversity between non-diseased and diseased groups
Environmental exposures (e.g., antibiotic use) during maturation have been associated with immunological and metabolic development through the mechanisms involved in the interaction between microbiota and host41. Type 1 diabetes (T1D) is one of the most common autoimmune diseases, which is caused by pancreatic β-cell destruction. T1D often appears in the pediatric age, and its incidence rate is globally increasing42. Livanos et al., (2016) performed microbiota-profiling studies to survey if the gut microbiota mediates the effect of antibiotic treatment on T1D onset33. To demonstrate the use of aMiAD, I analyzed a part of the original data, which surveys if the microbial diversity of gut microbiota altered by antibiotic treatment is differential by T1D status. To summarize the sampling and profiling procedures33, 19 NOD mice were exposed to the antibiotic (specifically, therapeutic-dose pulsed antibiotic) treatment, then, their fecal samples were collected after 6 weeks of the exposure. The V4 region of the bacterial 16S rRNA gene was targeted in the amplicon sequencing with barcoded fusion primers38 and the QIIME pipeline2 was used to quantify OTUs and construct their phylogenetic tree. The OTUs were rarefied using the software package, phyloseq39 due to the varying total reads per sample40. 390 OTUs were included in the analysis after removing OTUs which are not present in any sample after random subsampling of the rarefaction39.
We can first visually observe in the box-plots (Fig. 3A) that the phylogenetic metrics (PD, PE and PQE) show a greater disparity than the non-phylogenetic metrics (Richness, Shannon and Simpson), where PQE and then PE show the greatest disparity. Here, we can also observe that the microbial diversity is lower for the T1D group than the non-diseased group for all α-diversity metrics but the Shannon index (Fig. 3A). Correspondingly, PQE (p-value: 0.012) and PE (p-value: 0.015) estimate significant p-values with negative effect direction (Fig. 3B). The Shannon index is the only metric which estimates positive effect direction (Fig. 3B). This indicates that item-by-item analyses are substantially sensitive to (e.g., the decision on significance and/or effect direction can even be reversed by) the choice of α-diversity metric. aMiAD estimates that microbial diversity is significantly different between the two groups (p-value: 0.048), where the microbial diversity is lower for the T1D group than the non-diseased group (aMiDivES: −1.619 < 0) (Fig. 3B).
The comparison between the non-diseased (Non) and diseased (T1D) groups. (A) The box-plots for each α-diversity metric (i.e., Richness, Shannon, Simpson, PD, PE, PQE). (B) The outcomes for the item-by-item (i.e., Richness, Shannon, Simpson, PD, PE, PQE) and adaptive (i.e., aMiAD) α-diversity-based association tests.
Discussion
The recent microbial community-level association tests might be more powerful, where we, especially, observed in Simulations that OMiAT is most robustly powerful (Figs 1E,F and S1E,F). However, they do not provide any effect estimation facilities; hence, any further information about the disparity in microbial community composition is not accessible. Instead, aMiAD additionally estimates microbial diversity effect score, which can further enhance the interpretability. Here, I briefly discuss that other ANOVA-based methods (e.g., mvabund43) cannot directly adjust potential confounding effects (e.g., age, gender), while the regression-based methods (e.g., MiRKAT, MiSPU, OMiAT, aMiAD) can easily adjust them.
I chose the six α-diversity metrics, Richness, Shannon17, Simpson18, PD19, PE20 and PQE21,22, as the candidate α-diversity metrics for aMiAD because of their distinguished features44. However, we are not restricted to these metrics, and other α-diversity metrics might be considered. For example, Chao145 and ACE46, can be used to further modulate the extent of the rarity of association OTUs. Chao1 and ACE utilize abundance information as “≥2 or <2 reads” and “≥10 or <10 reads”, respectively, while Richness utilizes it as presence (i.e., ≥1 reads) or absence (i.e., 0 read). Thus, we may expect that Chao1 might be suitable when the extent of the rarity is relatively lower than the one for Richness, but relatively higher than the one for ACE. The Inverse Simpson index can also be considered by replacing the original Simpson index. Yet, I heuristically determined to use the original Simpson index as the Inverse Simpson index did not show any better performance. Notably, novel statistical estimates for α-diversity have still been proposed while further addressing the issues of missing species, sampling noise, experimental noise and so forth47,48,49,50,51,52. Any α-diversity metrics can be easily employed in my software package, aMiAD, through user options.
In this paper, I introduced aMiAD which adaptively approaches to the highest power and the most accurate microbial diversity effect score estimation among multiple item-by-item α-diversity-based association analyses. aMiAD also robustly satisfies the requisite validity issues in hypothesis testing and effect score estimation. Although I proposed aMiAD to relate microbial diversity with a continuous (e.g., BMI) or binary (e.g., disease/treatment status) trait of interest, it would be extendable to different types of trait (e.g., survival, multinomial trait)25,53,54,55. Moreover, an extension to the linear mixed effect model56/generalized linear mixed effect model57 is needed for correlated (e.g., family-based or longitudinal) study designs.
Data Availability
The utilized microbiome data are publicly available at the European Bioinformatics Institute (EBI) database (https://www.ebi.ac.uk, accession code: ERP016357)33 and the Sequence Read Archive (SRA) repository (https://www.ncbi.nlm.nih.gov/sra, accession code: SRP042293)37. The software package, aMiAD, is freely available at https://github.com/hk1785/aMiAD.
References
Hamady, M. & Knight, R. Microbial community profiling for human microbiome projects: Tools, techniques. Genome Res. 19(7), 1141–52 (2009).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–6 (2010).
Thomas, T., Gilbert, J. & Meyer, F. Metagenomics - a guide from sampling to data analysis. Microb. Inform. Exp. 2, 3 (2012).
Arslan, N. Obesity, fatty liver disease and intestinal microbiota. World J. Gastroenterol. 20(44), 16452–63 (2014).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012).
Knights, D., Lassen, K. G. & Xavier, R. J. Advances in inflammatory bowel disease pathogenesis: linking host genetics and the microbiome. Gut 62, 1505–10 (2013).
Bajaj, J. S. et al. Salivary microbiota reflects changes in gut microbiota in cirrhosis with hepatic encephalopathy. Hepatology 62, 1260–71 (2015).
Liu, M. et al. Oxalobacter formigenes-associated host features and microbial community structures examined using the American Gut Project. Microbiome 5, 108 (2017).
Charlson, E. S. et al. Disordered microbial communities in the upper respiratory tract of cigarette smokers. PLOS One 5, 12 (2010).
Bokulich, N. A. et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl. Med. 8, 343–82 (2016).
Zhao, N. et al. Testing in microbiome-profiling studies with MiRKAT, the microbiome regression-based kernel association test. Am. J. Hum. Genet. 96, 797–807 (2015).
Wu, C., Chen, J., Kim, J. & Pan, W. An adaptive association test for microbiome data. Genome Med. 8, 56 (2016).
Koh, H., Blaser, M. J. & Li, H. A powerful microbiome-based association test and a microbial taxa discovery framework for comprehensive association mapping. Microbiome 5, 45 (2017).
Connell, J. H. Diversity of tropical rainforests and coral reefs. Science 199, 1304–10 (1978).
Brook, B. W., Sodhi, N. S. & Ng, P. K. L. Catastrophic extinctions follow deforestation in Singapore. Nature 424, 420–6 (2003).
Gotelli, N. J. et al. Patterns and causes of species richness: a general simulation model for macroecology. Ecol. Lett. 12(9), 873–86 (2009).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27(379–423), 623–56 (1948).
Simpson, E. H. Measurement of diversity. Nature 163, 688 (1949).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Allen, B., Kon, M. & Bar-Yam, Y. A new phylogenetic diversity measure generalizing the Shannon index and its application to phyllostomid bats. Am. Nat. 174(2), 236–43 (2009).
Rao, C. R. Diversity and dissimilarity coefficients: a unified approach. Theor. Popul. Biol. 21(1), 24–43 (1982).
Warwick, R. M. & Clarke, K. R. New ‘biodiversity’ measures reveal a decrease in taxonomic distinctness with increasing stress. Mar. Ecol. Prog. Ser. 129(1), 301–5 (1995).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57(1), 289–300 (1995).
Lin, X. et al. Kernel machine SNP-set analysis for censored survival outcomes in genome-wide association studies. Genet. Epidemiol. 35, 620–31 (2011).
Koh, H., Livanos, A. E., Blaser, M. J. & Li, H. A highly adaptive microbiome-based association test for survival traits. BMC Genom. 19, 210 (2018).
Hill, M. O. Diversity and evenness: a unifying notation and its consequences. Ecology 54, 427–32 (1973).
Tuomisto, H. A diversity of beta diversities: straightening up a concept gone awry. Part 1. Defining beta diversity as a function of alpha and gamma diversity. Ecography 33, 2–22 (2010).
Li, H. Microbiome, metagenomics, and high-dimensional compositional data analysis. Annu. Rev. Stat. Appl. 2, 73–94 (2015).
Rao, C. R. Large sample tests of statistical hypotheses concerning several parameters with applications to problems of estimation. Math. Proc. Camb. Philos. Soc. 44(1), 50–7 (1948).
Wang, K. & Huang, J. A score-statistic approach for the mapping of quantitative-trait loci with sibships of arbitrary size. Am. J. Hum. Genet. 70, 412–24 (2002).
Pan, W., Kim, J., Zhang, Y., Shen, X. & Wei, P. A powerful and adaptive association test for rare variants. Genetics 4, 1081–95 (2014).
Mosimann, J. E. On the compound multinomial distribution, the multivariate β-distribution, and correlations among proportions. Biometrika 49(1-2), 65–82 (1962).
Livanos, A. E. et al. Antibiotic-mediated gut microbiome perturbation accelerates development of type 1 diabetes in mice. Nat. Microbiol. 1, 6140 (2016).
Reynolds, A. P., Richard, G., De La Iglesia, B. & Rayward-Smith, V. J. Clustering rules: A comparison of partitioning and hierarchical clustering algorithms. J. Math. Model. Algorithms 5, 474–504 (2016).
Calinski, T. & Harabasz, J. A dendrite method for cluster analysis. Comm. Statist. Theory Methods 3, 1–27 (1974).
Hennig, C. & Liao, T. F. How to find an appropriate clustering for mixed-type variables with application to socio-economic stratification. Appl. Statist. 62(3), 309–69 (2013).
Cox, L. M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–21 (2013).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–4 (2012).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLOS One 8, 4 (2013).
Weiss, S. et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5, 27 (2017).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–93 (2012).
Diamond Project Group. Incidence and trends of childhood type 1 diabetes worldwide 1990–1999. Diabetic Medicine 23, 857–66 (2006).
Wang, Y., Naumann, U., Wright, S. T. & Warton, D. I. mvabund – an R package for model-based analysis of multivariate abundance data. Methods Ecol. Evol. 3, 471–74 (2012).
McCoy, C. O. & Matsen, F. A. IV Abundance-weighted phylogenetic diversity measures distinguish microbial states and are robust to sampling depth. PeerJ 1, e157 (2013).
Chao, A. Non-parametric estimation of the number of classes in a population. Scand. J. Stat. 11, 265–70 (1984).
Chao, A. & Lee, S. Estimating the number of classes via sample coverage. J. Am. Stat. Assoc. 87, 210–17 (1992).
Lemos, L. N., Fulthorpe, R. R., Triplett, E. W. & Roesch, L. F. Rethinking microbial diversity analysis in the high throughput sequencing era. J. Microbiol. Methods 86(1), 42–51 (2011).
Li, K., Bihan, M., Yooseph, S. & Methé, B. A. Analyses of the microbial diversity across the human microbiome. PLOS One 7, 6 (2012).
Bunge, J., Willis, A. & Walsh, F. Estimating the number of species in microbial diversity studies. Annu. Rev. Stat. App. 1, 427–45 (2014).
Birtel, J., Walser, J., Pichon, S., Bürgmann, H. & Mattews, B. Estimating bacterial diversity for ecological studies: methods, metrics, and assumptions. PLOS One 10, 4 (2015).
Willis, A. & Bunge, J. Estimating diversity via frequency ratios. Biometrics 71(4), 1042–49 (2015).
Kaplinsky, J. & Arnaout, R. Robust estimates of overall immune-repertoire diversity from high-throughput measurements on samples. Nat. Commun. 7, 11881, https://doi.org/10.1038/ncomms11881 (2016).
Plantinga, A. et al. MiRKAT-S: a community-level test of association between the microbiota and survival times. Microbiome 5, 17 (2017).
Zhan, X. et al. A small-sample multivariate kernel machine test for microbiome association studies. Genet. Epidemiol. 21, 210–20 (2017).
Zhan, X., Plantinga, A., Zhao, N. & Wu, M. C. A fast small-sample kernel independence test for microbiome community-level association analyses. Biometrics 73(4), 1453–63 (2017).
Laird, N. M. & Ware, J. H. Random-effects models for longitudinal data. Biometrics 38, 963–73 (1982).
Breslow, N. E. & Clayton, D. G. Approximate inference in generalized linear mixed models. J. Am. Stat. Assoc. 88, 9–25 (1993).
Acknowledgements
The author is grateful to Prof. Ni Zhao at Johns Hopkins University and Prof. Amy Willis at University of Washington and the anonymous reviewers for their insightful observations and comments.
Author information
Authors and Affiliations
Contributions
H.K. is the only author who contributes to every aspect of this work.
Corresponding author
Ethics declarations
Competing Interests
The author declares no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koh, H. An adaptive microbiome α-diversity-based association analysis method. Sci Rep 8, 18026 (2018). https://doi.org/10.1038/s41598-018-36355-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-36355-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.