Abstract
Transient hepatic steatosis upon liver resection supposes functional relationships between lipid metabolism and liver regeneration. Repin1 has been suggested as candidate gene for obesity and dyslipidemia by regulating key genes of lipid metabolism and lipid storage. Herein, we characterized the regenerative potential of mice with a hepatic deletion of Repin1 (LRep1−/−) after partial hepatectomy (PH) in order to determine the functional significance of Repin1 in liver regeneration. Lipid dynamics and the regenerative response were analyzed at various time points after PH. Hepatic Repin1 deficiency causes a significantly decreased transient hepatic lipid accumulation. Defects in lipid uptake, as analyzed by decreased expression of the fatty acid transporter Cd36 and Fatp5, may contribute to attenuated and shifted lipid accumulation, accompanied by altered extent and chronological sequence of liver cell proliferation in LRep1−/− mice. In vitro steatosis experiments with primary hepatocytes also revealed attenuated lipid accumulation and occurrence of smaller lipid droplets in Repin1-deficient cells, while no direct effect on proliferation in HepG2 cells was observed. Based on these results, we propose that hepatocellular Repin1 might be of functional significance for early accumulation of lipids in hepatocytes after PH, facilitating efficient progression of liver regeneration.
Similar content being viewed by others
Introduction
Initially identified as a candidate gene in the quantitative trait locus for facets of the metabolic syndrome in subcongenic rat strains1, further investigations showed that expression of the Replication initiator 1 (Repin1) in the liver and adipose tissue is significantly associated with obesity and dyslipidemia2,3,4,5. Repin1 is ubiquitously expressed with highest amounts in liver and intraabdominal adipose tissue1. Ruschke et al.2 showed that Repin1 in adipocytes regulates the expression of genes involved in adipogenesis, lipid droplet formation and fusion as well as glucose and fatty acid (FA) transport. Moreover, beneficial physiological consequences of a liver-restricted Repin1 deficiency (LRep1−/−), as lower body weight, reduced hepatic steatosis, increased energy expenditure and physical activity as well as improved insulin sensitivity, underline the significant role of Repin1 in glucose homeostasis and lipid metabolism6.
Altered regulation of metabolic genes and pathways involved in insulin signaling, lipid and glucose metabolism is functionally important for the initiation of regeneration upon hepatic insufficiency. Thereby, transient accumulation of lipids in the regenerating liver is a well-known phenomenon and appears to be essential for adequate liver regeneration after hepatic resection7,8,9,10. It might serve as energy source for subsequent metabolic events associated with the regenerative process and reconstruction of cell membranes11. Rapid increase in hepatic triglycerides (TG) results from enhanced lipolysis in peripheral adipose tissue and influx of non-esterified FAs into the liver12.
The significance of hepatic fat for liver regeneration is supported by multiple studies, showing that reduced hepatic adipogenesis13,14, disruption/inhibition of ß-oxidation15,16,17 or defects in efficient lipid uptake, transport and formation18,19 are substantially associated with inhibition of the regenerative response of the organ. However, some reports showed the opposite11,20 and thus, it is still controversially discussed whether fat or rather glucose is the main energy substrate after hepatectomy15,21,22. It is supposed that fat is the main energy substrate for liver regeneration only very early after resection, if glucose is not available. Furthermore, it is needed primarily by hepatocytes of acinar zone 1 which prefer fat as energy source21,22. Nevertheless, the complex mechanisms that regulate initiation and resolution of transient hepatic steatosis upon resection and the functional significance of these events for liver regeneration are not fully clarified.
On the contrary, there is considerable evidence that fatty livers are suboptimal for surgical resection or transplantation, predominantly because of their increased susceptibility to ischemia-reperfusion injury and their disrupted regenerative response23,24,25. However, depending on the experimental model used, discrepant data regarding the regenerative capacity of steatotic livers exist26,27,28,29,30.
Based on previous findings in LRep1−/− mice regarding altered hepatic lipid storage, we hypothesized that Repin1 plays a functional role in hepatic regeneration. Therefore, we investigated the regenerative potential of LRep1−/− after partial hepatectomy (PH).
Results
Hepatic Repin1 expression upon liver resection
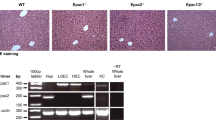
Quantitative RT-PCR analysis revealed almost no mRNA expression of Repin1 in LRep1−/− compared to wildtype mice, in which Repin1 level transiently decreased immediately after PH (Fig. 1A). Non-parenchymal cells were responsible for the remaining low Repin1 expression in LRep1−/− mice.
(A) Quantitative RT-PCR analysis of Repin1 mRNA expression in liver tissue of LRep1+/+ and LRep1−/− mice at different time points after PH (n = 6–7 per genotype and time point). Values are given as means ± SEM. Significance of differences between the groups of an individual time point was tested using Mann-Whitney Rank Sum test followed by Bonferroni correction (*p < 0.0083 vs. LRep1+/+ of the individual time point). Analysis of plasma activities of (B) alanine aminotransferase (ALT) and (C) glutamate dehydrogenase (GLDH) as well as (D) plasma albumin concentration in LRep1+/+ and LRep1−/− mice at different time points after PH (n = 5–8 per genotype and time point). Analysis of (E) TGs and (F) free FAs in plasma of LRep1+/+ and LRep1−/− mice at different time points after PH (n = 6–8 per genotype and time point). Significance of differences between the groups of an individual time point was tested using two way ANOVA, *p < 0.05 vs. LRep1+/+ 0 h).
Liver injury and liver function
Resection-associated liver injury, as given by a transient rise of liver enzymes upon PH, was significantly diminished in LRep1−/− animals with ALT values being about 1/3 lower than in LRep1+/+ plasma 24 h after PH (Fig. 1B). GLDH levels were also increasing upon 70% resection with a peak observed at 24 h, but with no significant differences between both genotypes (Fig. 1C). In contrast to that, albumin synthesis was maintained over the whole observation period with a minor decline in LRep1−/− mice at 72 h after PH (Fig. 1D).
Systemic and hepatic lipid profile
Hepatic Repin1 deficiency caused dyslipidemia in plasma at baseline (Fig. 1E and F) and altered hepatic lipid content after PH (Fig. 2). Whereas LRep1−/− mice showed higher plasma TG levels at baseline (Fig. 1E), liver resection initially induced a decrease of plasma TGs in both genotypes with lowest levels being present at 24 h. Afterwards systemic TGs were increasing again. A similar trend after PH with no significant differences between both mouse strains was observed for plasma FFA, in which a drop down of FFA has been seen at 24 h (Fig. 1F).
(A) Quantification of TGs in liver tissue of LRep1+/+ and LRep1−/− mice at different time points after PH using a standard kit (Values are given as means ± SEM; n = 6–8 per genotype and time point, Mann-Whitney Rank Sum test followed by Bonferroni correction, ***p < 0.001 vs. LRep1+/+ 24 h). (B) Representative Oil Red O stained frozen liver sections of LRep1+/+ and LRep1−/− mice 24 h after PH. Scale bars represent 100 µm. (C) Histomorphometric analysis of semi-thin toluidine blue stained liver sections to quantify fat deposition as percentage of blue stained area compared with the total section area. Values are given as means ± SEM (n = 3–6 per genotype and time point; Mann-Whitney Rank Sum test followed by Bonferroni correction). (D) Representative semi-thin toluidine blue stained liver sections of LRep1+/+ and LRep1−/− mice 24 h after PH. Scale bars represent 100 µm. (E) Quantitative analysis of the number, area and size of lipid droplets in hepatocytes of LRep1+/+ and LRep1−/− mice 24 h after PH by means of transmission electron microscopy and representative images (lipid droplets are marked with white asterisks). Scale bars represent 5 µm. Values are given as means ± SEM (n = 5 per genotype; t- test, ***p < 0.001 vs. LRep1+/+). (F) Quantitative analysis of TG groups as well as total amount of liver TGs (insert) at baseline (0 h), 24 and 48 h after PH in LRep1+/+ and LRep1−/− mice using LC-MS. The first number is equivalent to the carbon numbers and the second number represents the numbers of double bonds. Values are given as means ± SEM (n = 6 per genotype and time point; total TGs: two way ANOVA, ***p < 0.001 vs. LRep1+/+; TG groups: Mann-Whitney Rank Sum test followed by Bonferroni correction).
Liver TGs and lipids were analyzed by different methods. Biochemical quantification of hepatic TG demonstrated a PH-induced steep rise in TG of LRep1+/+ livers with a peak value 24 h after PH (Fig. 2A). In contrast, livers of LRep1−/− revealed significantly reduced TG at 24 h after PH (Fig. 2A). Additionally, Oil red O stained frozen liver sections (Fig. 2B) showed also a much lesser extent of fat positive area in LRep1−/− tissue. Moreover, fat deposition was quantified by histomorphometric analysis of semi-thin toluidine blue stained liver sections (Fig. 2C). This analysis also confirmed the dramatic increase of hepatic lipids 24 h after resection in wildtype mice, whereas lipid accumulation was much less pronounced in LRep1−/− mice. This means that 24 h after PH livers of LRep1+/+ mice consisted of ~20% lipid droplets compared to ~10% in LRep1−/− tissue, whereas livers of sham mice (0 h) simply contained <1% lipids (Fig. 2C). Representative toluidine blue stained liver sections (Fig. 2D) display this distinct difference in fat content between both genotypes at 24 h. Further electron microscopic analysis of the lipid droplets (Fig. 2E) revealed a significantly reduced number and total area of lipid droplets in LRep1−/− compared to LRep1+/+ hepatocytes 24 h after PH, with no difference in droplet size. Representative electron microscopic images displaying the higher number of lipid droplets in wildtype livers compared to LRep1−/− tissue (Fig. 2E, white asterisks).
For the time points with the most striking differences in lipid content between both genotypes, we analyzed the pattern of TG subgroups using LC-MS (Fig. 2F). Generally, the most prominent TG groups during early liver regeneration were TG 52:3 and 52:4. Compared to wildtype LRep1−/− mice exhibited markedly lower levels of these TG groups at 24 h after resection, but showed a delayed increase 48 h after resection, when in wildtype livers the levels of these TG groups have already declined. Measurement of total TG levels with this method confirmed the above mentioned results of significantly lower levels in LRep1−/− compared to LRep1+/+ livers at 24 h after resection. In LRep1−/− mice a delayed increase of TG at 48 h (Fig. 2F, inserted graph) was detected.
Hepatic glycogen content
As the liver plays an important role in glucose homeostasis and storage, we evaluated the kinetics of glycogen content upon resection by different methods. Analysis of glycogen in liver tissue homogenates revealed a rapid resection-induced loss of glycogen storage in both mouse strains, with almost no existing glycogen at 6 h after PH (Fig. 3A and B). Thereafter, glycogen content replenished slowly and was fully restored 7 d after PH. This could also be verified by PAS staining of LRep1−/− and LRep1+/+ livers (Fig. 3A), impressively showing the initial loss and continuous increase of glycogen after PH. Hepatic Repin1 deficiency was accompanied by a significantly reduced glycogen content at 3 d after PH (Fig. 3B). Representative electron microscopic images of LRep1+/+ also display high glycogen content (white crystalline structures) at basal (0 h) and almost no glycogen deposition 24 h after PH (Fig. 3C).
(A) Representative images of PAS-stained liver sections of in LRep1+/+ and LRep1−/− mice at different time points after PH. Scale bars represent 100 µm. (B) Quantitative analysis of liver glycogen content in LRep1+/+ and LRep1−/− mice at different time points after PH. Values are given as means ± SEM (n = 6–8 per genotype and time point). Significance was tested by two-way ANOVA followed by the appropriate post hoc comparison (Holm-Sidak method), **p < 0.01 vs. LRep1+/+ 3 d. (C) Transmission electron microscopic images of LRep1+/+ under physiological conditions (0 h) and 24 h after PH showing loss of glycogen (0 h, white deposition) upon PH. Scale bars represent 5 µm.
Regenerative capacity
Quantitative analysis of BrdU-stained hepatocytes (Fig. 4A) and non-parenchymal cells (Fig. 4B) as well as counting of mitotic figures (Fig. 4C) were performed as indices of hepatocellular regenerative response upon PH. Livers of LRep1-/- displayed attenuated cell proliferation, as indicated by significantly reduced BrdU incorporation in hepatocytes 48 h after PH compared to LRep1+/+ mice (Fig. 4A,B). Representative immunohistochemical images of hepatic tissue in LRep1+/+ and LRep1−/− livers at 48 h (Fig. 4A) and 3 d (Fig. 4B) after PH demonstrate the high number of BrdU-positive cells as characteristic sign of both parenchymal and non-parenchymal cell proliferation in the regenerating LRep1+/+ liver, whereas livers of LRep1−/− showed less BrdU-positive cells.
Quantitative analysis of proliferation by means of (A) BrdU-positive hepatocytes, (B) BrdU-positive non-parenchymal cells and (C) mitotic figures (each given as cells/HPF) in livers of LRep1+/+ and LRep1−/− mice at different time points after PH. Values are given as means ± SEM (n = 6–8 per genotype and time point; *p < 0.0083 vs. LRep1+/+ of the individual time point, Mann-Whitney Rank Sum test followed by Bonferroni correction). Representative immunohistochemical images (lower panel) of hepatic tissue in LRep1+/+ and LRep1−/− 48 h (A) as well as 3 days (B) after resection, displaying the marked rise of BrdU-positive cells, particularly in LRep1+/+, as characteristic sign of cell proliferation in the regenerating liver. Scale bars represent 100 µm. (D) Percent regenerated liver weight relative to preoperative liver weight in livers of LRep1+/+ and LRep1−/− mice at different time points after PH. The weight of the regenerated liver was used to calculate the growth of residual liver lobes as ratio of regenerated liver/preoperative liver weight × 100 (%). Pre-operative liver weight was assumed to be 4.7% of body weight for LRep1−/− and 5.0% of body weight for LRep1+/+.
To characterize proliferation in more detail, hepatocytes with mitotic figures were counted at the indicated time points (Fig. 4C). Interestingly, 48 h after resection LRep1−/− showed a markedly higher number of hepatocytes being in mitosis compared to LRep1+/+.
There was a constant increase of liver weight upon PH in both genotypes with return to almost pre-operative values (Fig. 4D), with LRep1−/− genotype being associated with a slightly lower liver weight (88.7% ± 2.9%) at day 7 after PH compared to wildtype (95.4% ± 3.3%).
Hepatic lipogenesis and lipolysis
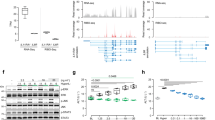
In order to evaluate the impact of hepatic de novo lipogenesis in the process of regeneration, especially upon Repin1 deficiency, we analyzed mRNA expression of the hepatic fatty acid synthase (Fasn) as a regulatory gene essential for efficient lipogenesis (Fig. 5A). In wildtype mice Fasn mRNA expression transiently decreased after PH and thereafter constantly increased with values at day 3 and 7 being much higher than pre-PH (Fig. 5A). A similar kinetic could be observed in LRep1−/− livers. Because of higher basal expression, Repin1 deficiency resulted in a steeper decline of Fasn mRNA immediately after resection compared to wildtype mice. The extent of increase in Fasn expression at the later observation time points was less prominent in LRep1−/− than in wildtype mice (Fig. 5A). Additionally, we observed almost no resection-induced alterations in regulation of hepatic β-oxidation in both genotypes, as shown by almost constant Pparα mRNA expression, except for the time point 48 h (Fig. 5B). At that time point, Pparα mRNA expression was reduced in LRep1−/− compared to wildtype mice.
Quantitative RT-PCR analysis of (A) Fasn, (B) Pparα, (C) Cd36, (D) Fatp5, (E) Fatp2 and (F) Fabp1 mRNA expression in liver tissue of LRep1+/+ and LRep1−/− mice at different time points after PH (n = 6–8 per genotype and time point). Values are given as means ± SEM. Significance of differences between the groups of an individual time point was tested by Mann-Whitney Rank Sum test followed by Bonferroni correction.
Hepatic FA transport
Next, we examined the expression of different hepatic lipid transporters. Here, Cd36 mRNA expression tended to be lower in LRep1−/− early after PH, particularly at 48 h (Fig. 5C). Also mRNA expression of the liver-specific membrane associated FA transport protein 5 (Fatp5) was found to be markedly diminished in LRep1−/− at 6 h (Fig. 5D), though the difference did not reach statistical significance. However, mRNA expression of other membrane associated transport proteins like Fatp2 (Fig. 5E), intracellular FA binding proteins like Fabp1 (Fig. 5F) or the lipid droplet fusion protein Snap23 (data not shown) were not significantly altered between both experimental groups.
In vitro analyses
To analyze whether Repin1 deficiency directly influences hepatocellular proliferation, we performed a BrdU proliferation assay with HepG2 cells using different concentrations of siRNA for Repin1 (siRepin1). In contrast to the nonsense siRNA for the Luciferase gene (siLuci), higher Repin1 siRNA concentrations resulted in Repin1 deficiency (Fig. 6A). However, this had no inhibiting effect on the proliferative capacity of HepG2 cells (Fig. 6B).
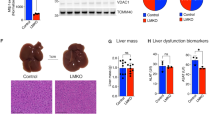
(A) Repin 1 mRNA expression of HepG2 cells 72 h after treatment with 0, 10, 20, 40, 80 and 160 nM siRNA specific for Luciferase (siLuci) and Repin1 (siRepin1) (n = 3–4 per group and concentration) and (B) proliferation of these cells using a BrdU proliferation assay (n = 9 per group and concentration). Values are given as means ± SEM (n = 4 per genotype and time point; *p < 0.0083 vs. LRep1+/+ of the individual time point, Mann-Whitney Rank Sum test followed by Bonferroni correction. (C) FACS analysis of Cd36 of primary isolated wildtype hepatocytes (Wt), which additionally were transfected with siLuci or siRepin1, and of hepatocytes of LRep1−/− mice under normal (left) and steatotic (right) culture conditions for a period of 72 h. Data are presented as box plots indicating the median, the interquartile range in form of a box and the minimum and maximum as whiskers. #p < 0.05 vs. Wt siLuci of the individual culture condition, Mann-Whitney Rank Sum test. (D) Repin1 mRNA expression of primary hepatocytes of the different genotypes and treatment groups as described above. *p < 0.01 vs. Wt; #p < 0.05 vs. Wt siLuci of the individual culture condition, Mann-Whitney Rank Sum test. (E) Oil Red O staining of primary isolated wildtype hepatocytes (Wt), which additionally were transfected with siLuci or siRepin1, and of hepatocytes of LRep1−/− mice under steatotic culture conditions for a period of 24 and 72 h. Scale bars represent 50 µm. (F) Scanning and electron microscopic images of primary isolated Wt hepatocytes (Wt), which additionally were transfected with a siRNA for Luciferase (siLuci) and Repin1 (siRepin1) as well as of LRep1−/− hepatocytes after 72 h of FA exposure. Compared to controls lipid droplets (white asterisks) appear smaller in Repin1-deficient hepatocytes. Scale bars represent 10 µm.
As CD36 is a membrane protein, we additionally performed FACS analysis of CD36 on the cell surface of primary isolated hepatocytes of LRep1−/− and wildtype mice (LRep1+/+) under normal (conventional) and, to simulate transient steatosis as in the early phase of liver regeneration, under steatotic culture conditions (Fig. 6C). Wildtype hepatocytes were also treated with 20 nM siRepin1/siLuci to evaluate the impact of a spontaneously induced Repin1 deficiency on CD36 cell surface expression. Compared to the corresponding wildtype control (Wt or Wt siLuci) Repin1 mRNA expression was significantly reduced in primary hepatocytes of LRep1−/− mice as well as in siRepin1 treated primary wildtype cells (Fig. 6D). Generally, in vitro steatosis was accompanied by a slightly increased Repin1 expression in wildtype hepatocytes (Fig. 6D). Although transfection efficiency was lower in steatotic cells compared to normal cultured hepatocytes, siRNA treatment resulted in a significantly decreased Repin1 expression (Fig. 6D). FACS analysis of normal cultured cells revealed a significantly reduced CD36 positivity only of cells with siRNA-induced Repin1 deficiency (Fig. 6C). In vitro steatosis induced cell surface CD36 expression in each analyzed group of cells. A reduced CD36 expression was only observed upon spontaneously (siRNA-) induced Repin1 deficiency (Fig. 6C). Irrespective of the culture condition, CD36 expression values of LRep1−/− hepatocytes displayed a high variance. Of most interest, Oil Red O staining showed reduced lipid accumulation in both Repin1 deficient hepatocyte cultures upon FFA exposition for 24 h (Fig. 6E). However, with prolonged FFA exposure time of 72 h lipid accumulation increased with just minor differences between controls and Repin1-deficient groups (Fig. 6E).
We additionally performed SEM (Fig. 6F) and TEM (not shown) analysis of differentially cultured and treated primary hepatocytes after FA exposure for 72 h to characterize hepatocellular lipid uptake/load and intracellular lipid distribution in more detail. In general, in contrast to normal culture (not shown) in vitro steatosis induced formation of numerous lipid droplets of different size in wildtype cells (Fig. 6F, Wt and Wt siLuci). Noticeably, deficiency of Repin1 resulted in much smaller intracellular lipid droplets (Fig. 6F, Wt siRepin1 and LRep1−/−). TEM analysis revealed that lipid droplets are solely located intracellular (not shown).
Discussion
Recent studies investigated distinct roles of the zinc finger protein Repin1 in adipocytes2,4,31. In human adipose tissue, Repin1 mRNA expression significantly correlates with body fat content and adipocyte size2,5. Relationships between Repin1 expression in adipose tissue and metabolic parameters also suggest a functional role for Repin1 in the liver. As alterations in systemic and hepatic metabolism are important modulators of the physiological regenerative response to hepatic insufficiency, we therefore elucidated the potential role of Repin1 in liver regeneration after partial hepatectomy using LRep1−/− mice6.
The importance of Repin1 for FA uptake in adipocytes has been implied previously2,4,6,31 which is in line with our data, as hepatic deficiency of Repin1 causes reduced lipid content in hepatocytes during the early phase of liver regeneration. This was accompanied by a delayed cell proliferation. As fat accumulates concomitantly with cellular proliferation32 and Repin1 deficiency itself showed no direct effect on liver cell proliferation, it can be assumed that the reduced and shifted fat accumulation in LRep1−/− mice in turn impacts extent and course of hepatocellular proliferation, but finally without impairing liver weight recovery.
Generally, discrepant data concerning the effects of (chronic) steatosis on liver regeneration exist. However, several groups suggest that steatosis does not impair the regenerative response or even an induction of mild steatosis may be beneficial for surgical outcome of hepatectomies30, while others showed the opposite24,33,34,35. Much controversy continues on the predominant energy substrate (glucose or fat) for liver regeneration which complies on the availability of glucose21 but might primarily relate to the hepatic energy status15,21. Thus, with increasing resected hepatic mass and a decreasing energy charge, the liver predominantly utilizes FA as energy source15 and therefore transiently accumulates fat after PH7,36. Many studies dealing with a decreased hepatic fat accumulation imply a significant role for acute hepatic steatosis in liver regeneration11,13,14,18,37,38. Moreover, for several decades it has been known that infusion of lipids and carnitine following PH stimulated the initiation of liver regeneration16,39,40. Consistent with previous studies19,41 we also hypothesize that a lower temporary fat accumulation, as seen upon Repin1 deficiency, is linked to a delayed DNA synthesis in liver cells. Using different methods, we verified a significant decrease in lipids/TGs at 24 h after resection in LRep1−/− mice, while TG accumulation peaks in wildtype mice. However, LRep1−/− mice showed a compensatory shifted increase of TG at 48 h after PH. Phenotypic characterization of LRep1−/− mice6 suggests that reduced TG accumulation in the liver is due to decreased expression of genes involved in lipid uptake and formation. At the molecular level, we demonstrated that during liver regeneration absence of Repin1 resulted in a temporarily reduced mRNA expression of the FA transporter Cd36 and Fatp5, while expression of other hepatic lipid transfer or binding genes were similar between both genotypes. FATP and CD36 have been suggested to be a functional unit42. Impaired and delayed FA uptake, resulting in lower hepatic TGs, was also shown in Cd36-, Fatp2- and Fatp5-deficient mice43,44,45,46. However, as no difference in circulating lipids (TG, FFA) was observed between both genotypes, it can also be suggested that FA transport was compensated by other mechanisms or was still sufficient to protect against dyslipidemia in LRep1−/− mice. To analyze lipid uptake and formation as well as cell surface CD36 protein expression, we additionally performed in vitro experiments with primary hepatocytes. In contrast to primary LRep1−/− hepatocytes, siRNA-mediated downregulation of Repin1 in vitro alone caused a significant reduction of cell surface CD36 expression in normal, but also steatotic hepatocytes. Interestingly, we also noticed distinct defects in efficient lipid accumulation and formation upon Repin1 deficiency by using fat staining and electron microscopy. However, differences disappeared after long-term exposure to FAs, indicating different underlying mechanisms. Hepatic FA uptake is complex and suggested to be a dual-kinetic process, consisting of a rapid, carrier-mediated, and a delayed, passive diffusional phase37,42. Thus, it can be supposed that Repin1 deficiency results in impairment of the facilitated, protein-mediated, phase of hepatic FA uptake (e.g. for acute conditions, like liver resection) but with preservation of the diffusional phase (e.g. upon chronic FA exposure).
Of interest, we noted that simultaneously with the onset of transient hepatic steatosis upon PH, the resection-induced liver injury is most severe. In turn, a diminished liver injury was obvious in LRep1−/−, probably due to a lower lipid load.
Due to the fact that the main source of TGs in hepatocytes during regeneration does not arise from de novo lipogenesis47, but rather from FA uptake from plasma and thus, lipolysis of systemic fat, a rise in circulating and hepatic FAs can be observed11,38. It is also well known, that loss of systemic adipose storages occurs in proportion to the extent of hepatic resection/insufficiency38. Although it is suggested that de novo lipogenesis provides an initial supply of FAs at the very early phase (0–6 h)19, expression profile of Fasn in this study indicates that de novo lipogenesis is not involved in the early phase of transient fat accumulation, neither in wildtype nor in LRep1−/− mice. However, Repin1 deficiency rather resulted in an impaired rise of Fasn expression in the later regenerative phase, when hepatic lipogenesis becomes more essential, suggesting a promoting role for Repin1 in lipogenesis. Interestingly, this would indicate a causal connection between the observed transient down-regulation of Repin1 and suppressed lipogenesis following PH.
By analysis of Pparα, a transcriptional regulator of FA oxidation48, prominent alterations in β-oxidation of FAs could not be observed during the course of liver regeneration.
Cell cycle of hepatocytes and fat accumulation are subjected to a well-defined dynamic. PH induces four waves of proliferation which are linked to three waves of fat accumulation32. In contrast to the DNA synthesis, which was significantly diminished in LRep1−/− hepatocytes at 48 h, mitosis index at 48 h was increased upon Repin1 deficiency. Remarkably, hepatocyte mitosis occurs consistently during a time of day determined by the circadian clock49. Also performing surgery between 10:00 and 12:00 am, Zou et al.32 showed that mitosis always peaks at 6:00 am, with a first mitosis peak observed at 44 h after resection. Therefore, we postulate that in this study at 48 h the mitosis peak of LRep1+/+ hepatocytes as well as a probably decreased mitosis index of LRep1−/− cells are already past and just a delayed and shifted compensatory increase in the number of mitotic LRep1−/− hepatocytes is present. In addition, the division of hepatocytes is so rapid, taking no more than 30 minutes21 that the mitotic index is a suboptimal indicator for the evaluation of the regenerative condition. Furthermore, as the first wave of hepatocyte DNA synthesis is present at 36–48 h after PH10, it also can be assumed that the differences between both genotypes are even more noticeable for earlier DNA synthesis peaks between 36 h and 44 h after PH.
Interestingly, hypoglycemia frequently occurs after partial liver resection36. This metabolic response is suggested to initiate signals that promote liver regeneration as dextrose supplementation suppresses PH-induced proliferation39,50, mainly due to inhibition of the release of free FAs from systemic adipose stores15. As also shown by Lai et al.21, glycogen disappeared in the early post-hepatectomy regenerative phase and reappeared constantly until day 7. Although Repin1 deficiency resulted in reduced basal glucose uptake in adipocytes due to lower Glut1 expression2, LRep1−/− livers restored glycogen nearly at the same degree as LRep1+/+ and showed a lower glycogen content only at day 3 after PH. To account for changes in lipid availability compensatory pathways for generation of intracellular energy sources have to be upregulated. As in LRep1−/− mice the regenerative response is just timely delayed, it can be suggested that LRep1−/− hepatocytes also use glucose as energy substrate.
Our results demonstrate that acute changes in lipid metabolism have specific effects on the course of liver regeneration with Repin1 being important for regulation of lipid load. Although hepatocytes of LRep1−/− maintain the ability to incorporate lipids after PH, however, the number of lipid droplets and finally the TG content is substantially lower than in wildtype. Consequently, insufficient amounts of energy might cause timely delayed proliferation of LRep1−/− hepatocytes. However, this had no impact on liver weight recovery which might also caused by compensatory hypertrophy8. As also discussed by others, a decreased expression of FA transporters seems to offer a partial explanation for a lower lipid load, but it has not clearly been determined if this effect is either due to decreased FA uptake or altered TG synthesis51. Besides, a more rapid turnover of TGs in the absence of Repin1 could also account for reduced TG accumulation in these mice. Vice versa, it can be assumed that reduced hepatic lipid content after resection may hamstring hepatocytes from regulating genes involved in metabolic pathways that are required for sufficient recovery. Because liver regeneration is a complex and multi-factorial process, more probably a combination of hepatic lipid load and genetic and metabolic changes determines recovery from surgery.
Due to its contribution to increased lipid accumulation and adipocyte hypertrophy, augmented Repin1 expression may play a fundamental role in adipose tissue dysfunction and its related metabolic diseases52. Whereas therapeutic strategies to reduce Repin1 expression are of interest in human obesity, the availability of lipids during regeneration after liver surgery is of high importance. Thus, Repin1 modulation may be a novel strategy to interfere in this process, particularly in case of limited liver regeneration of pre-existing liver injury.
Methods
Mice
Male mice with a hepatocyte-restricted, Cre-loxP-mediated Repin1 deletion (LRep1AlbCre; LRep1−/−) and wildtype littermates (LRep1+/+) (background C57BL/6 N) at an age of 10–16 weeks and with a body weight of 25–30 g were kept at a 12 h day and night cycle on water and standard laboratory chow ad libitum. Experiments were approved by the local government Landesamt für Landwirtschaft, Lebensmittelsicherheit und Fischerei Mecklenburg-Vorpommern (LALLF M-V/TSD/7221.3-1.1-099/12) and conducted in accordance with the German legislation on protection of animals and EU-directive 2010/63/EU.
Surgical procedure and experimental groups
Mice were anesthetized by breathing isoflurane (1.5 vol%) and subjected to a 68% PH as described previously53,54. Sham- operated animals without hepatic resection served as group 0 h (n = 6 per genotype). The animals were allowed to recover from anesthesia and surgery under a red warming lamp and were held in single cages until the subsequent experiments followed at postoperative hours 6, 24 and 48 as well as days 3 and 7 (n = 7–8 per genotype and time point). Animals were sacrificed under ketamine/xylazine anesthesia (90/7 mg/kg bw ip) at the indicated time points (7–8 animals per time point) and blood as well as liver tissue samples were taken. The remnant livers were harvested, weighed and processed for subsequent analyses. The weight of regenerated liver was used to calculate the growth of residual liver lobes according to weight of regenerated liver/preoperative liver weight × 100 (%). As determined by sham-operated animals, preoperative liver weight was assumed as 4.7% of body weight for LRep1−/− and 5.0% of body weight for LRep1+/+. Additionally, to analyze the regenerative response, 5-bromo-2-deoxyuridine (BrdU; 50 mg/kg bw ip) was injected 1 h prior to harvest of liver tissue. BrdU incorporation into DNA was analyzed by immunohistochemistry.
Hematological measurements and plasma enzyme levels
Blood samples were collected at final time points by retrobulbar sinus puncture. Blood cells were assessed with an automated cell counter (Sysmex KX-21, Sysmex). Activities of alanine aminotransferase (ALT), aspartate aminotransferase (AST) and glutamate dehydrogenase (GLDH) in ethylenediaminetetraacetic acid (EDTA) plasma were measured spectrophotometrically as indicators of hepatocellular disintegration and necrosis using cobas c 111 analyzer (Roche Diagnostics; Rotkreuz, Switzerland) according to the manufacturer’s instructions.
Assays
EDTA plasma further served for the analysis of albumin as a parameter of liver function using a commercially available enzyme-linked immunosorbent assay kit in accordance with the manufacturer’s instructions (Assaypro, MO, USA). Measurements of plasma triglycerides (TG) and free fatty acids (FFA), serving as indicators of systemic dyslipidemia, were performed using the TG (Cayman Chemical Company, MI, USA; 10010303) and FFA (abcam, ab65641) assay kit methods according to the manufacturer’s instructions. Glycogen levels in liver samples were measured using a glycogen assay kit (Abnova; KA0861) according to the manufacturer’s instructions with 10 mg of homogenized frozen liver tissue. For measurement of hepatic TG, Lipids were extracted from mouse liver biopsies using the commercially available LabAssay Triglyceride (WAKO Pure Chemicals, Kyoto, Japan).
Histopathology/Cell staining
Lipid accumulation in 8 µm frozen liver sections and cultured primary hepatocytes was visualized by Oil Red O staining. Briefly, liver sections were rinsed with 60% 2-propanol and stained for 15 min with freshly prepared and filtered Oil Red O (Sigma) solution (0,2% (w/v) in 60% 2-propanol diluted from a 0,5% (w/v) Oil Red O stock solution in 100% 2-propanol). After rinsing in 60% 2-propanol and distilled water, slides were counterstained with hematoxylin. Primary hepatocytes grown on cover slips were fixed in 4% phosphate buffered formalin (Grimm med. Logistik, Torgelow, Germany) for 5 min. After washing with PBS and water, the cells were then stained for 7 min using the same Oil Red O solution as described above. After rinsing extensively in water the cells were counterstained with hematoxylin.
Liver tissue was fixed in 4% phosphate buffered formalin for two to three days, embedded in paraffin, and cut into 5 µm thick sections. Periodic acid-Schiff (PAS) reaction was used to visualize the presence and distribution of carbohydrates, particularly glycogen, in liver cells. Therefore, slides were treated with 0.1% periodic acid solution for 5 min, washed and subsequently incubated with Schiff’s Reagent for 15 min at RT. After vigorous washing nuclei were counterstained with hematoxylin. The intensity of purple color in PAS-stained liver sections corresponds to the amount of glycogen in hepatocytes. Digital images were taken with a Color View II FW camera (Color 10 View, Munich, Germany).
Immunohistochemistry
For analysis of DNA-incorporated BrdU in liver cells, 5 µm paraffin sections collected on poly-L-lysine-coated glass slides were incubated with monoclonal mouse anti-BrdU antibody (1:50; M0744, Dako) overnight at 4 °C followed by incubation with HRP-conjugated goat anti-mouse immunoglobin (LSAB kit plus; Dako). Sites of peroxidase-binding were detected by DAB (Dako). Sections were counterstained with hematoxylin. BrdU-positive hepatocellular and non-parenchymal nuclei (discriminated by morphology and intrahepatic location) were counted in a blinded manner within 30 consecutive high power fields (HPF) (x40 objective, numerical aperture 0.65) and are given as cells/HPF. BrdU-stained liver sections were also used to count mitotic figures (visible chromosome aggregation) in a blinded manner within 30 consecutive high power fields (HPF) (x40 objective, numerical aperture 0.65) and are given as mitotic figures/HPF.
Conventional RT-PCR analysis
RNA isolation and conventional Real-time PCR gene expression analysis was performed as described previously55 using SYBR Green I (Roche, Mannheim, Germany) detection of amplified dsDNA strands applying the LightCycler System 1.5 (Roche) (Table 1). All data were calculated by using the comparative ddCt method and expression values were normalized to the expression levels of the GAPDH housekeeping gene. Target gene expression was compared to wildtype C57BL/6 N liver tissue pool.
Liver lipidomics
TGs were analyzed by LC-MS on a 6550 iFunnel QTOF mass spectrometer (Agilent Technologies, Waldbronn, Germany) equipped with a Dual Agilent Jet Stream (AJS) electrospray source. Electrospray parameters were as follows: gas temperature, 200 °C; drying gas flow, 11 l/min; nebulizer pressure, 35 psig; sheath gas temperature, 350 °C; sheath gas flow, 12 l/min; capillary voltage, 4000 V; nozzle voltage, 500 V; and fragmentor voltage, 350 V. Mass spectrometric analysis was done in positive ion mode with a scan range of m/z 100–1100 and a scan rate of 4 spectra/s. Data was acquired with intensity thresholds of 10 counts/0.001% and stored in profile mode. TG standards were obtained from Larodan Fine Chemicals AB (Malmö, Sweden). Frozen liver tissue samples (about 30 mg) were homogenized in an ice cold mixture of 400 µl of methanol and 130 µl of water in a FastPrep® 24 homogenizer (MP Biomedicals, Santa Ana, USA) for 20 s at speed 6.0 using lysing matrix D. An aliquot of the homogenate was spiked with internal standard and extracted with chloroform as described previously56. The chloroform extract was evaporated to dryness and reconstituted in 2-propanol. Chromatographic separation of the TGs was carried out at 70 °C on a Poroshell 120 RP18 column (2.1 × 100 mm, 2.7 μm particle size, Agilent) coupled to a 1290 Infinity UHPLC System (Agilent). Gradient runs of mobile phase A (20 mM ammonium acetate in acetonitrile:water 95:5 (v/v)) and mobile phase B (20 mM ammonium acetate in 2-propanol) were programmed as follows: 0–2 min, 5–15% B; 2–4 min, 15% B; 4–8 min, 15–50% B; 8–11 min, 50–60% B; 11–12 min, 60% B, 12–13 min, 60–90% B ;13–21 min, 90% B. The column was reconditioned to initial conditions (5% B) for 5 min. TGs were analyzed as ammonium adducts [M+NH4]+ using triheptadecanoin as internal standard. Calibration samples were prepared in aqueous BSA in the concentration range from 0.0625 to 75 nmol/10 µl and were worked up as described above and analyzed together with the unknown samples. Calibration curves based on internal standard calibration were obtained by weighted (1/x) quadratic regression for the peak-area ratio of the analyte to the internal standard against the amount of the analyte. The concentration of the analytes in unknown samples was obtained from the regression curve. Assay accuracy and precision were determined by analyzing quality controls that were prepared like the calibration samples.
Electron microscopy
For electron microscopy, hepatocytes were cultured on 13 mm circular supports cut from Melinex plastic film sheets (no. L4103, Plano, Wetzlar, Germany) with a manual punching tool. The Melinex supports were sterilized by immersion in ethanol and air-dried under a cell culture hood after washes with sterile water. For fixation, cultured hepatocytes or small blocks of liver tissue were immersed in a fixative containing 2% glutaraldehyde and 1% paraformaldehyde in 0.1 M sodium phosphate buffer pH 7.3. Prior to resin embedding for transmission electron microscopy (TEM) samples were washed three times with 0.1 M sodium phosphate buffer and were postfixed with a solution of 1% aqueous osmiumtetroxide for 1 hour. Following washes in distilled water, the cells or tissue blocks were dehydrated in a graded series of acetone completed with two pure acetone steps. Next the specimens were infiltrated with epoxy resin (Epon 812, Serva, Heidelberg, Germany) starting with 1:1 mixture of acetone and resin overnight, continued with pure resin for 4 hours and transfer to rubber molds on the following day. After curing of the resin at 60 °C for 2 days, semithin sections (0.5 µm) and thin sections (50–70 nm) were cut on a ultramicrotome (Ultracut E, Reichert&Jung, Wien, Austria) using diamond knifes (Diatome, Biel, Switzerland). Semithin sections were stained with toluidine blue to visualize the tissue structure and for further use in morphometric measurements (see below). Thin sections for ultrastructural inspection were cut from these areas, transferred to 300 mesh copper grids and were stained with lead citrate and uranyl acetate. The grids were examined in a Zeiss EM902 electron microscope (Carl Zeiss, Oberkochen, Germany) operated at 80 kV. Digital images were acquired with a side-mounted 1 × 2k FT-CCD Camera (Proscan, Scheuring, Germany) using iTEM camera control and imaging software (Olympus-SIS, Münster, Germany) with calibrated morphometric measurement tools to determine e.g. lipid droplet size and numbers.
In addition, for histomorphometric analysis of semi-thin toluidine blue stained liver sections, images of 10–15 random low power fields (10x magnification, Olympus BX 51, Hamburg, Germany) were acquired with a Color View II FW camera (Color View, Munich, Germany) and evaluated using an image analysis system (Adobe Photoshop). Fat deposition was quantified as percentage of blue stained area compared with the total section area.
For scanning electron microscopy (SEM), hepatocytes cultured on glass coverslips (no. L40971, Plano, Wetzlar, Germany) or on Melinex supports were fixed as detailed above, washed with sodium phosphate buffer and dehydrated with a graded series of ethanol or acetone. The cells were critical point dried using an Emitech K850 critical point dryer (Emitech Ltd. Ashford, UK) with CO2 as an intermedium. Specimens were mounted on SEM stubs with adhesive carbon tape (Plano, Wetzlar, Germany) and were coated with evaporated carbon in a Leica EM SCD 500 coater (Leica Microsystems, Vienna, Austria). The cellular structure was viewed with a field-emission SEM, Zeiss Merlin VP compact (Carl Zeiss Microscopy, Jena Germany) operated at 10 kV and digital images with a size of 1024 × 768 pixels were recorded.
Isolation and culture of primary hepatocytes and HepG2 cell line
Hepatocytes were isolated from male LRep1−/− and LRep1+/+ mice at an age of 6 weeks as described previously55. Cells were plated on collagen A (200 µg/ml in PBS; Biochrom, Berlin, Germany) coated cell culture dishes with Williams E medium (PAN Biotech, Aidenbach, Germany) containing L-glutamine, 10% (v/v) FCS (for the first 4 hours, subsequently FCS free) and 100 nM dexamethasone (Sigma Aldrich) and incubated in a 5% CO2 humidified atmosphere at 37 °C.
The human hepatocellular carcinoma cell line HepG2 was cultured in Dulbecco’s modified Eagle’s medium (DMEM, high glucose) supplemented with 10% (v/v) FCS at 37 °C in 5% CO2.
In vitro transfection
For siRNA transfection cells were seeded in 6-well petri dishes at a density of 1 × 105 (HepG2) or 2.5 × 105 (primary hepatocytes) cells per well. To transfect HepG2 cells the media was changed 24 h after seeding. Repin1 siRNAs or Luciferase siRNAs were transfected using AtuFECT01 (Silence Therapeutics, Berlin, Germany) at final siRNA concentrations of 0, 10, 20, 40, 80 and 160 nM. Primary LRep1+/+ hepatocytes were transfected simultaneously with seeding with 20 nM siRNA. After an incubation period of 4 h at 37 °C the medium was changed again and the cells were cultivated until further analyses were performed.
BrdU-proliferation assay
HepG2 proliferation was measured by a BrdU Cell Proliferation ELISA (Roche Applied Sciences, Mannheim, Germany). For this purpose, HepG2 were trypsinized 48 h after transfection and seeded on a 96-well-plate at a density of 4 × 103 cells per well. After attachment BrdU labeling solution was added and the cells were grown for additional 24 h. Fixation, staining and measurement of cell proliferation was performed according to manufacturer’s instructions.
In vitro steatosis
For induction of steatosis primary LRep1−/− and LRep1+/+ hepatocytes as well as primary transfected LRep1+/+ hepatocytes were exposed to a mixture of long-chain FFA (oleic acid and sodium palmitate at a ratio of 2:1). A stock solution of 50 mM of the FFA-mixture prepared in Williams E medium containing 1% BSA was diluted in medium to obtain a final concentration of 0.5 mM. The FFA mixture was added to hepatocytes ~24 h after seeding and transfection for a period of 72 h. The medium with FFA mixture was renewed daily.
FACS analysis
Hepatocellular cell surface CD36 expression levels were investigated by flow cytometry analyses, conducted with a FACSCalibur cytometer (BD Biosciences) running CellQuest (BD Biosciences) acquisition and analysis software. For analysis of CD36 expression primary hepatocytes were harvested after culture for 96 h, washed and incubated at 4 °C for 30 min with 20 µl anti-CD36-FITC antibody (Santa Cruz, sc-13572 FITC) or mouse IgA-FITC (sc-3900) as isotype control in FACS buffer containing PBS with 0.5% BSA. Cells were washed again and resuspended in FACS buffer for measurement. The number of CD36 positive cells (%) in each experimental group was normalized to the number of CD36 positive cells (%) of normal cultured LRep1+/+ hepatocytes.
Statistical analysis
Results are presented as mean ± standard error of the mean (SEM) or as box plots indicating the median, the interquartile range in form of a box, and the minimum and maximum as whiskers. All statistical analyses were performed using SigmaPlot 12.0 (Systat Software Inc., Erkrath, Germany). After testing for normality and equal variance across groups differences between the two groups and time points were assessed by two-way ANOVA followed by the appropriate post hoc comparison (Holm-Sidak method) including Bonferroni probabilities to compensate for multiple comparisons, p < 0.05 was used to define statistical significance. If the data were not normally distributed, pairwise comparison was performed using Mann-Whitney Rank Sum test including Bonferroni probabilities to compensate for multiple comparisons, and thus statistical significance was set at p < 0.0083. For reasons of clarity and comprehensiveness, only statistically significant differences and p-values p < 0.05 between the groups of an individual time point are indicated in the figures.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Klöting, N., Wilke, B. & Klöting, I. Triplet repeat in the Repin1 3′-untranslated region on rat chromosome 4 correlates with facets of the metabolic syndrome. Diabetes. Metab. Res. Rev. 23, 406–410 (2007).
Ruschke, K. et al. Repin1 maybe involved in the regulation of cell size and glucose transport in adipocytes. Biochem. Biophys. Res. Commun. 400, 246–251 (2010).
Heiker, J. T. & Klöting, N. In vitamins and Hormones 91, 97–105 (2013).
Hesselbarth, N. et al. Repin1 Deficiency in Adipose Tissue Improves Whole-body Insulin Sensitivity, and Lipid Metabolism. Int J Obes 41, 1815–1823 (2017).
Krüger, J. et al. Metabolic effects of genetic variation in the human REPIN1 gene. Int. J. Obes., https://doi.org/10.1038/s41366-018-0123-0 (2018).
Kern, M. et al. Liver-restricted Repin1 deficiency improves whole-body insulin sensitivity, alters lipid metabolism, and causes secondary changes in adipose tissue in mice. Diabetes 63, 3295–3309 (2014).
Farrell, G. C. Probing Prometheus: Fat fueling the fire? Hepatology 40, 1252–1255 (2004).
Kachaylo, E. et al. PTEN Down-Regulation Promotes β-Oxidation to Fuel Hypertrophic Liver Growth After Hepatectomy in Mice. Hepatology 66, 908–921 (2017).
Trotter, N. L. A Fine Structure Study of Lipid in Mouse Liver Regenerating After Partial Hepatectomy. J. Cell Biol. 21, 233–244 (1964).
Michalopoulos, G. K. Liver Regeneration. J. Cell Physiol. 213, 286–300 (2007).
Newberry, E. P. et al. Altered Hepatic Triglyceride Content After Partial Hepatectomy Without Impaired Liver Regeneration in Multiple Murine Genetic Models. Hepatology 48, 1097–1105 (2008).
Tijburg, L., Nyathi, C., Meijer, G. & Geelen, M. Biosynthesis and secretion of triacylglycerol in rat liver after partial hepatectomy. Biochem J 277, 723–728 (1991).
Shteyer, E., Liao, Y., Muglia, L. J., Hruz, P. W. & Rudnick, D. A. Disruption of hepatic adipogenesis is associated with impaired liver regeneration in mice. Hepatology 40, 1322–1332 (2004).
Walldorf, J. et al. Propranolol impairs liver regeneration after partial hepatectomy in C57Bl/6-mice by transient attenuation of hepatic lipid accumulation and increased apoptosis. Scand. J. Gastroenterol. 45, 468–476 (2010).
Nakatani, T. et al. Differences in predominant energy substrate in relation to the resected hepatic mass in the phase immediately after hepatectomy. J Lab Clin Med 97, 887–898 (1981).
Holeček, M. Nutritional modulation of liver regeneration by carbohydrates, lipids, and amino acids: a review. Nutrition 15, 784–788 (1999).
Anderson, S. P. et al. Delayed liver regeneration in peroxisome proliferator-activated receptor-α-null mice. Hepatology 36, 544–554 (2002).
Fernandez, M. A. et al. Caveolin-1 Is Essential for Liver Regeneration. Science (80-.). 96, 1628–1632 (2006).
Kohjima, M. et al. Delayed liver regeneration after partial hepatectomy in adipose differentiation related protein-null mice. J. Hepatol. 59, 1246–1254 (2013).
Rao, M. S., Peters, J. M., Gonzalez, F. J. & Reddy, J. K. Hepatic regeneration in peroxisome proliferator-activated receptor α-null mice after partial hepatectomy. Hepatol. Res. 22, 52–57 (2002).
Lai, H.-S., Chen, W.-J. & Chen, K.-M. Energy substrate for liver regeneration after partial hepatectomy in rats: effects of glucose vs fat. J. Parenter. Enter. Nutr. 16, 152–156 (1992).
Gebhardt, R. Metabolic zonation of the liver: Regulation and implications for liver function. Pharmacol. Ther. 53, 275–354 (1992).
Behrns, K. E. et al. Hepatic steatosis as a potential risk factor for major hepatic resection. J. Gastrointest. Surg. 2, 292–298 (1998).
Markus, S. & Pierre-Alain, C. Failure of regeneration of the steatotic rat liver: disruption at two different levels in the regeneration pathway. Hepatology 31, 35–42 (2000).
Abshagen, K., Mertens, F., Eipel, C. & Vollmar, B. Limited therapeutic efficacy of thrombopoietin on the regeneration of steatotic livers. Int. J. Clin. Exp. Pathol. 6 (2013).
Abanobi, S. E., Lombardi, B. & Shinozuka, H. Stimulation of DNA Synthesis and Cell Proliferation in the Liver of Rats Fed a Choline-devoid Diet and Their Suppression by Phenobarbital. Cancer Res. 42, 412–415 (1982).
Farrell, G. C., Robertson, G. R., Leclercq, I. & Horsmans, Y. Liver regeneration in obese mice with fatty livers: Does the impairment have relevance for other types of fatty liver disease? Hepatology 35, 731 (2002).
Picard, C. et al. Steatosis is not sufficient to cause an impaired regenerative response after partial hepatectomy in rats. J. Hepatol. 36, 645–652 (2002).
Leclercq, I. A. et al. Defective hepatic regeneration after partial hepatectomy in leptin-deficient mice is not rescued by exogenous leptin. Lab. Investig. 86, 1161–1171 (2006).
Sydor, S. et al. Steatosis does not impair liver regeneration after partial hepatectomy. Lab. Investig. 93, 20–30 (2013).
Kunath, A. et al. Repin1 deficiency improves insulin sensitivity and glucose metabolism in db/db mice by reducing adipose tissue mass and inflammation. Biochem. Biophys. Res. Commun. 478, 398–402 (2016).
Zou, Y. et al. Four Waves of Hepatocyte Proliferation Linked with Three Waves of Hepatic Fat Accumulation during Partial Hepatectomy-Induced Liver Regeneration. PLoS One 7, e30675 (2012).
Yang, S. Q., Lin, H. Z., Mandal, A. K., Huang, J. & Diehl, A. M. Disrupted signaling and inhibited regeneration in obese mice with fatty livers: Implications for nonalcoholic fatty liver disease pathophysiology. Hepatology 34, 694–706 (2001).
Veteläinen, R., van Vliet, A. K. & van Gulik, T. M. Severe Steatosis Increases Hepatocellular Injury and Impairs Liver Regeneration in a Rat Model of Partial Hepatectomy. Ann. Surg. 245, 44–50 (2007).
Hamano, M. et al. Lipid overloading during liver regeneration causes delayed hepatocyte DNA replication by increasing ER stress in mice with simple hepatic steatosis. J. Gastroenterol. 49, 305–316 (2014).
Rudnick, D. A. & Davidson, N. O. Functional Relationships between Lipid Metabolism and Liver Regeneration. Int. J. Hepatol. 2012, 549241 (2012).
Brasaemle, D. L. CELL BIOLOGY: Enhanced: A Metabolic Push to Proliferate. Science (80-.). 313, 1581–1582 (2006).
Gazit, V. et al. Liver Regeneration is Impaired in Lipodystrophic fld Mice. Hepatology 52, 2109–2117 (2010).
Holecek, M. & Simek, J. Effect of the infusion of glucose, itralipid and nutramin on the initiation of rat liver regeneration after partial hepatectomy. Physiol Bohemoslov 37, 467–473 (1988).
Bláha, V., Šimek, J. & Zadák, Z. Liver regeneration in partially hepatectomized rats infused with carnitine and lipids. Exp. Toxicol. Pathol. 44, 165–168 (1992).
Fernández-Rojo, M. A. et al. Caveolin-1 orchestrates the balance between glucose and lipid-dependent energy metabolism: Implications for liver regeneration. Hepatology 55, 1574–1584 (2012).
Ehehalt, R. et al. Translocation of long chain fatty acids across the plasma membrane – lipid rafts and fatty acid transport proteins. Mol. Cell. Biochem. 284, 135–140 (2006).
Febbraio, M. et al. A null mutation in murine CD36 reveals an important role in fatty acid and lipoprotein metabolism. J. Biol. Chem. 274, 19055–62 (1999).
Hajri, T., Han, X. X., Bonen, A. & Abumrad, N. A. Defective fatty acid uptake modulates insulin responsiveness and metabolic responses to diet in CD36-null mice. J. Clin. Invest. 109, 1381–1389 (2002).
Doege, H. et al. Targeted Deletion of FATP5 Reveals Multiple Functions in Liver Metabolism: Alterations in Hepatic Lipid Homeostasis. Gastroenterology 130, 1245–1258 (2006).
Falcon, A. et al. FATP2 is a hepatic fatty acid transporter and peroxisomal very long-chain acyl-CoA synthetase. Am. J. Physiol. - Endocrinol. Metab. 299, E384–E393 (2010).
Chakravarthy, M. V. et al. “New” hepatic fat activates PPARα to maintain glucose, lipid, and cholesterol homeostasis. Cell Metab. 1, 309–322 (2005).
Bechmann, L. P. et al. The interaction of hepatic lipid and glucose metabolism in liver diseases. J. Hepatol. 56, 952–964 (2012).
Matsuo, T. et al. Control Mechanism of the Circadian Clock for Timing of Cell Division in Vivo. Science (80-.). 302, 255–259 (2003).
Weymann, A. et al. p21 is Required for Dextrose-Mediated Inhibition of Mouse Liver Regeneration. Hepatology 50, 207–215 (2009).
Mashek, D. G. Hepatic Fatty Acid Trafficking: Multiple Forks in the Road. Adv. Nutr. 4, 697–710 (2013).
Blüher, M. Adipose Tissue Dysfunction in Obesity. Exp Clin Endocrinol Diabetes 117, 241–250 (2009).
Abshagen, K., Eipel, C., Menger, M. D. & Vollmar, B. Comprehensive Analysis of the Regenerating Mouse Liver: An In Vivo Fluorescence Microscopic and Immunohistological Study. J. Surg. Res. 134 (2006).
Greene, A. K. & Puder, M. Partial Hepatectomy in the Mouse: Technique and Perioperative Management. J. Investig. Surg. 16, 99–102 (2003).
Reetz, J. et al. Development of Adenoviral Delivery Systems to Target Hepatic Stellate Cells In Vivo. PLoS One 8 (2013).
Wu, H., Southam, A. D., Hines, A. & Viant, M. R. High-throughput tissue extraction protocol for NMR- and MS-based metabolomics. Anal. Biochem. 372, 204–212 (2008).
Acknowledgements
We thank Maren Nerowski, Eva Lorbeer, Dorothea Frenz, Berit Blendow, Ute Schulz, Laura Grüner and Monika Seiler for their excellent technical assistance. We thank Georg Damm for providing an in vitro steatosis protocol. This research was supported in part by the Robert Bosch Foundation, Stuttgart, by the Federal Ministry of Education and Research (BMBF, Germany) within the research network Systems Medicine of the Liver (LiSyM, grant number 031L0037) and FKZ:01E01501; DZD:82DZD00601 as well as by the Deutsche Forschungsgemeinschaft (SFB1052/2 B04). Marie Liebig was supported by the Deutsche Forschungsgemeinschaft (AB 453/2-1).
Author information
Authors and Affiliations
Contributions
N.K. and K.A. conceived the research ideas and supervised the project. B.D., K.A., M.L., A.W., B.G., U.S., U.H. and M.F. performed the experiments. B.D., M.F., U.H., B.V. and K.A. analyzed the data. K.A., N.K., B.V. and M.S. wrote and edited the manuscript. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abshagen, K., Degenhardt, B., Liebig, M. et al. Liver-specific Repin1 deficiency impairs transient hepatic steatosis in liver regeneration. Sci Rep 8, 16858 (2018). https://doi.org/10.1038/s41598-018-35325-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35325-3
Keywords
This article is cited by
-
Regenerative Hepatic Effect of Red Propolis Extract Administration After Partial Hepatectomy in Rats
Revista Brasileira de Farmacognosia (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.