Abstract
Single-cell measurements of biochemical processes have advanced our understanding of cellular physiology in individual microbes and microbial populations. Due to methodological limitations, little is known about single-cell phosphorus (P) uptake and its importance for microbial growth within mixed field populations. Here, we developed a nanometer-scale secondary ion mass spectrometry (nanoSIMS)-based approach to quantify single-cell P uptake in combination with cellular CO2 and N2 fixation. Applying this approach during a harmful algal bloom (HAB), we found that the toxin-producer Nodularia almost exclusively used phosphate for growth at very low phosphate concentrations in the Baltic Sea. In contrast, the non-toxic Aphanizomenon acquired only 15% of its cellular P-demand from phosphate and ~85% from organic P. When phosphate concentrations were raised, Nodularia thrived indicating that this toxin-producer directly benefits from phosphate inputs. The phosphate availability in the Baltic Sea is projected to rise and therefore might foster more frequent and intense Nodularia blooms with a concomitant rise in the overall toxicity of HABs in the Baltic Sea. With a projected increase in HABs worldwide, the capability to use organic P may be a critical factor that not only determines the microbial community structure, but the overall harmfulness and associated costs of algal blooms.
Similar content being viewed by others
Introduction
Phosphorus (P) is an essential element for the growth of all living organisms, and the availability of P, in particular dissolved inorganic P (DIP, i.e. phosphate) limits productivity in a variety of ecosystems. The availability of DIP can play a vital role in the formation of harmful algal blooms in freshwater and brackish ecosystems1,2 and limit marine primary productivity3 as well as dinitrogen (N2) fixation4, the largest external input of nitrogen (N) to the ocean5. Microorganisms have developed a suite of features to cope with DIP scarcity, including adjustments of the cellular P requirements such as the replacement of phospholipids with sulfur-based lipids6, the intracellular storage of P for use at times of P starvation7, and the direct use of organic P sources8,9. In addition, organic P sources could be exploited indirectly through associated microorganisms (or the microbiome) that can hydrolyze DIP and make it available to their host10,11. The response of different microorganisms to DIP scarcity is likely distinct due to differences in the metabolic capabilities, and as a result has the potential to change microbial community structures. It is therefore crucial to understand the strategies used to circumvent DIP scarcity and to determine the importance of different P sources for microbial growth.
To assess the role of phosphorus for microbial growth, previous studies have performed radiolabeled (32P or 33P) phosphate and ATP uptake experiments, which provide important insights into the extent and mechanisms of DIP and DOP uptake at the single-cell level (e.g.12,13). Although microautoradiography combined with fluorescent in situ hybridization (MAR-FISH) links the identity of individual cells with DIP or DOP uptake, quantitative, cell-specific activities are commonly not reported (e.g.14,15,16,17) with some exceptions18. The combination of P radiotracers with cell flow-sorting has yielded quantitative insights into cellular P uptake rates for specific size classes/populations and phytoplankton taxa (e.g.12,19,20). While most studies have assessed DIP uptake, some have also used 33P-ATP as an organic P analogue. However, ATP can also be used as a carbon or nitrogen source21 and represents only a small fraction of the total organic P pool22,23. The DOP pool is diverse, and its concentration often exceeds that of DIP, suggesting that it is an important source of P in DIP-depleted environments24. The cell-specific alkaline phosphatase assay (e.g.25,26,27) has been used widely to study the use of organic P sources but is not a direct measure of uptake and cannot be easily linked to CO2 and N2 fixation rates in individual cells. Due to the complexity and the current methodological limitations, the quantitative importance of inorganic and organic P for growth of individual organisms in mixed environmental communities is largely unknown.
Here, we developed a new nanoSIMS-based, single-cell approach to directly quantify DIP uptake rates. These are measured simultaneously with C and N assimilation in the same cell and can therefore yield the contribution of DIP to the total P demand and growth of microorganisms. Meanwhile, measured single-cell elemental ratios allow for a mass balance to determine the importance of P storage and organic P utilization. Using this approach, we determined the quantitative importance of different P sources for three different harmful algal bloom (HAB) organisms in the Baltic Sea. The Baltic Sea suffers from seasonally recurring cyanobacterial HABs. These extensive blooms are usually dominated by one of three genera: (1) Nodularia spumigena, a producer of the toxin Nodularin28, (2) Dolichospermum sp., which in some cases can produce the toxin Microcystin29, and (3) Aphanizomenon sp., considered to be non-toxic28. But Aphanizomenon’s capacity to form dense blooms can contribute to, for example, oxygen depletion upon degradation and economic losses from reduced tourism (for review see30). In recent years, these noxious HABs have covered an area more than one-half of the entire Baltic Sea (~200,000 km2; ref.31). The HABs develop under conditions of seasonal shallow thermal stratification and are believed to be stimulated by excess DIP after the spring phytoplankton bloom has exhausted the dissolved inorganic nitrogen (DIN) pool1. Although the excess DIP is drawn down to nanomolar concentrations, the HABs continue to increase in size and intensity for several weeks32. It has been suggested that organic P33 as well as internal P storage7 are key factors in maintaining the HAB growth. While culture studies have suggested that Nodularia is superior to Aphanizomenon at using organic P8,9, the quantitative importance of these P sources in the environment is currently not known.
Results and Discussion
Single-cell approach
We developed a single-cell approach to measure the cellular DIP uptake rate using nanometer-scale secondary ion mass spectrometry (nanoSIMS). The measurement relies on the incorporation of radio-labeled (33P) phosphate into cellular biomass and the subsequent beta decay of the short-lived radioactive 33P (half-life ~25 d) to the stable 33S. The 33S/32S ratio is then used to calculate the DIP uptake rate (see methods). The challenge of this approach is to achieve high specific activities of 33P to attain enough 33P mass so as to significantly exceed the natural abundance of 33S (~0.75 atomic percent (at%)) in the cellular biomass (Supplementary Fig. S1; see methods). In addition to measuring a stable isotope, an advantage of measuring S isotopes instead of P isotopes is that sulfur is better ionized (~10-fold)34 by the nanoSIMS cesium (Cs+) primary ion beam resulting in increased ion counts and therefore improved statistics for determining the cellular DIP uptake rates. The measured cellular DIP uptake rate can be directly compared to cellular C and N assimilation rates of individual cells in mixed field populations when incubated with multiple isotopes in the same incubation. To test the developed method, we incubated a laboratory culture of Nodularia spumigena (KAC64) and field-collected colonies of Aphanizomenon sp. with 13CO2 and DI33P under DIP concentrations naturally occurring at the beginning of summer in the Baltic Sea (~150–300 nM DIP). The subsequent nanoSIMS analysis revealed the significant enrichment of both 13C and 33S isotopes in individual cells of the filamentous cyanobacteria indicating the cellular uptake and incorporation of CO2 and DIP into biomass (Fig. 1) with not all cells being enriched in 33S. This heterogeneity was observed for both the Nodularia culture and the field-collected Aphanizomenon and for both CO2 fixation and DIP uptake (Fig. 2a,b). Similar heterogeneity was previously reported for clonal35,36,37 and field-collected populations38,39,40 and was attributed to phenotypic heterogeneity.
Single-cell imaging of cyanobacterial filaments. 13CO2 fixation and DI33P uptake as indicated by the 13C/12C and 33S/32S ratios measured via nanoSIMS in filaments of a Nodularia culture and filaments of Aphanizomenon collected in June from the Baltic Sea, both under sufficiently high DIP concentrations. White outlines indicate the cyanobacterial cells. NA = natural abundance. Scale bars are 3 µm for all images. An example of unenriched cells is given in Supplementary Fig. S2.
Single-cell isotopic enrichments and uptake rates of cyanobacteria. Cellular 13C/12C and 33S/32S enrichments by (a) a Nodularia culture and (b) Aphanizomenon collected in June from the Baltic Sea after incubations with 13CO2 and DI33P. The dashed lines indicate the detection limit of the isotopic enrichment, and cellular isotope ratios at or below the detection limit (DL) are shown in the faded colors. The CO2 fixation and DIP uptake rates for (c) the Nodularia culture and (d) Aphanizomenon were calculated from the isotopic enrichments (see methods) for all cells, with rates below the DL shown in faded colors. DL 13C/12C: 0.0117, DL 33S/32S: 0.0084. NA: natural abundance (0.01 for 13C/12C and 0.0078 for 33S/32S).
Cell-specific rates were calculated for both cyanobacteria using the labeling percentage of 13C and 33P in the CO2 and DIP pool, the natural abundance of 13C and 33S, the incubation time and the cell size (Fig. 2c,d). The distribution of cellular rates is similar between the two cyanobacteria species with relatively high DIP uptake rates as compared to the canonical Redfield ratio suggesting that DIP concentrations were sufficient for both culture and field-collected colonies. As a proof of principle, these experiments demonstrate that cellular DIP uptake can be quantified using DI33P incorporation and subsequent nanoSIMS analysis of the cellular 33S/32S ratio simultaneously with carbon and/or nitrogen assimilation.
Importance of P sources during harmful algal bloom
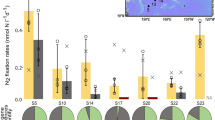
We used the new method to determine the quantitative importance of DIP for the growth of three abundant HAB organisms at very low DIP concentrations in the summer. Our sampling took place during a cyanobacterial bloom that developed in early August 2015 when DIN was nearly depleted in the surface waters and the DIP concentration was ~250 nM (Fig. 3). The bloom covered large expanses of the Baltic Sea (Fig. 3a). At the Swedish National Monitoring Station B1 (Fig. 3a) the cyanobacterial bloom consisted mainly of Aphanizomenon sp., consistent with recurring summer blooms throughout the central Baltic32,41. Dolichospermum sp. and Nodularia spumigena, both (potential) toxin-producers, constituted about one to two orders of magnitude less biomass than Aphanizomenon sp., respectively (Fig. 3b). The development of the HAB caused a drawdown of DIP concentrations (Fig. 3b) from ~250 nM to 20–30 nM. This drawdown induced a strong decrease of the colony-based DIP uptake rates in the diazotrophic cyanobacteria from ~37 pmol colony−1 day−1 at ~150 nM DIP to ~7 pmol colony−1 d−1 at 34 nM DIP within 48 hours (Fig. 3c). Phosphate additions that raised the DIP concentration to >1 µM restored the uptake rates to those observed at a concentration of ~150 nM DIP. Logarithmic curve fitting (best fit) was used to determine the apparent half-saturation constant (KM value) of ~60 nM DIP at very low DIP concentrations (Fig. 3c). This KM value should be viewed cautiously, because these colonies also entrap organisms such as diatoms, flagellates and heterotrophic bacteria. DIP concentrations at the beginning of the bloom (~250 nM DIP) exceeded this apparent KM value for phosphate uptake indicating that the onset of the bloom was not limited by DIP. In contrast, cyanobacteria became P-stressed after the onset and throughout the bloom, when concentrations had dropped to ~30 nM DIP (08 Aug 2015). Despite these very low DIP concentrations, the HAB increased in size and intensity (Fig. 3b).
Harmful algal bloom (HAB) and DIP uptake rates in the Baltic Sea in summer 2015. (a) Composite satellite image (05–12 August) of cyanobacterial bloom (MODIS-Aqua; remote sensing reflectance at 555 nm), white dot: monitoring station B1. (b) DIP and total organic phosphorus (TOP) concentrations at station B1 (own data and open access data under the Creative Commons Attribution 4.0 at www.smhi.se), extent of the surface (black) and subsurface (white) bloom (data provided by72) and the cyanobacterial community structure during the sampling campaign; (c) colony-based DIP uptake rates during the bloom (circles = liquid scintillation counting, triangles = digital autoradiography). The dashed line shows the best fit which was used for the KM determination.
Single-cell nanoSIMS analyses of Nodularia, Aphanizomenon and Dolichospermum showed that all three cyanobacteria were actively fixing CO2 and N2 while also taking up DIP during the HAB (Figs 4 and 5). The measured CO2 and N2 fixation as well as the DIP uptake rates are likely conservative due to the use of fixatives during the sample processing42,43. Intriguingly, a fraction of the assimilated DIP appeared to be accumulating in granule-type areas of the cells despite DIP scarcity (Fig. 4). These granule-type areas were also hot-spots of freshly assimilated CO2 and N2 (Fig. 4). The exact nature of these granules is currently not known. Cyanophycin, a common storage product in cyanobacteria under P limitation44, itself does not contain phosphorus. However, freshly assimilated P is possibly stored as polyphosphates (polyP) in the same location as cyanophycin. PolyP formation and storage has traditionally been viewed as a ‘luxury’, where uptake occurs at DIP-replete conditions for use at times of P starvation45. However, more recent evidence suggests that polyP might play an important role for P cycling in phytoplankton during DIP scarcity13,46,47. Although the exact mechanisms are not known yet, it appears that phytoplankton conserve polyP during P starvation relative to other cellular P pools47. Nonetheless, the quantity of P stored in the observed granules is small (max. ~5%) here compared to the overall cellular DIP uptake.
Single-cell imaging of cyanobacterial filaments collected during the cyanobacterial bloom in 2015. CO2 (13C/12C) and N2 (15N/14N) fixation and DIP (33S/32S) uptake measured through nanoSIMS for Nodularia, Dolichospermum and Aphanizomenon. The cellular P distribution is shown as the 31P/12C ratio. White outlines indicate the cyanobacterial cells. NA = natural abundance. Scale bars are 5 µm for all nanoSIMS images. Dolichospermum images have been size-adjusted to conserve the relative size proportions between the different cyanobacteria. For style reasons, these images have been framed with black bars.
Single-cell uptake rates of cyanobacterial filaments. Single-cell CO2 and N2 fixation versus DIP uptake by Nodularia (a,b), Dolichospermum (c,d) and Aphanizomenon (e,f) under very low DIP concentrations. Solid lines indicate canonical Redfield ratios. Cellular rates at or below detection limit are indicated by faded colors (a,b,e and f) or by grey (c,d).
Within each of the three populations of diazotrophic cyanobacteria, we observed considerable heterogeneity in C, N and P-uptake between cells, even within filaments (Supplementary Fig. S3), as has previously been observed for environmental populations38,39,40. In fact, the observed heterogeneity within a filament was often as large as the variability between filaments, and we statistically show that enough cells have been measured so that the results obtained here are representative for the different cyanobacteria (Supplementary Figs S4–S6). Most notable, however, was the difference in DIP uptake between the organisms. In Nodularia, CO2 and N2 fixation activity had a clear correlation to DIP uptake with ratios slightly higher than the Redfield ratio (Fig. 5a,b) while Dolichospermum showed a fairly weaker relationship of DIP uptake to CO2 and N2 fixation rates (Fig. 5c,d). Surprisingly, CO2 and N2 fixation had no relationship to DIP uptake in Aphanizomenon (Fig. 5e,f) indicating that P sources other than DIP were more important to its growth.
Baltic Sea filamentous cyanobacteria seem capable of adjusting their cellular elemental ratios to alleviate P stress7,32. A changing stoichiometry (C:P and/or N:P) could be the result of either using previously stored P or decreasing cellular P demand. To investigate whether P storage or a decreased cellular P demand could explain the difference between the organisms, we used energy-dispersive X-ray spectroscopy (EDS) to measure the elemental ratios of individual cells in colonies of all three cyanobacteria (Fig. 6).
Single-cell elemental imaging of cyanobacterial filaments. Scanning electron microscopic (SEM) images of Nodularia, Dolichospermum and Aphanizomenon filaments with elemental imaging for carbon, nitrogen and phosphorus obtained via energy-dispersive X-ray spectroscopy (EDS). Scale bar size is indicated in the SEM image and is consistent within each of the cyanobacteria.
The elemental ratios of all three cyanobacteria were already above Redfield at the beginning of our incubations indicating that P storage was not an important contributor to alleviating P stress, and therefore did not contribute to the increase in size and intensity of the cyanobacterial HAB in August 2015. However, we observed a shift in the elemental ratios within 24 hours. The C:P and N:P ratios of all three cyanobacteria increased by up to 45% and 200%, respectively (Supplementary Table S1) suggesting that the cellular P content decreased relative to C and N. While this changing stoichiometry likely contributed to alleviating or adjusting to P stress7,32, it cannot explain the differences between the cyanobacteria as the changes in the elemental ratios are similar for all three of them (Supplementary Table S1).
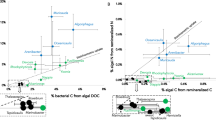
Apart from DIP, organic P compounds can serve as P sources for cyanobacterial growth. Since the organic P pool is very diverse, it is nearly impossible to measure the direct uptake of total organic P. Nevertheless, if the total P demand for cellular growth is compared with the measured DIP uptake, the contribution of total organic P can be calculated. We calculated the mass balance of total P demand for cellular growth using the C-based growth rates (from 13CO2 assimilation) and the changing elemental stoichiometries over the incubation time (see Methods). This mass balance predicts the CO2:DIP uptake ratio that would be necessary to sustain all growth by DIP (Fig. 7a). The predicted CO2:DIP uptake ratio is then compared to the experimental CO2:DIP uptake ratio obtained from the nanoSIMS analysis to reveal the importance of DIP and organic P. If the ratios are similar, (nearly) all growth can be sustained by the experimentally measured DIP uptake. However, if the experimental ratio is higher than the predicted ratio (i.e. less DIP uptake relative to carbon was measured), not all growth can be sustained by DIP. Since the changing elemental composition has already been taken into account at this point (including potential storage), the difference between the predicted and the experimental P uptake must be satisfied by organic P sources.
Cellular C:P ratios, CO2:DIP uptake ratios and C-based growth rates. (a) C:P biomass ratios in August at the start (t0) and end (t24) of the incubation (changes between these two time points were significantly different for all three cyanobacteria, Kruskal Wallis test p < 0.05) and the predicted (pre.) and experimentally (exp.) measured CO2:DIP-uptake ratios; lines = medians, boxes = 25th and 75th percentiles, bars = standard deviation. Outliers (indicated by numbers in parentheses below number of measured cells (n)) are not plotted here for easier visualization but were all included in the analysis. (b) Growth rates based on 13CO2 incorporation at in situ and added DIP (+1 µM) concentrations; lines = medians, boxes = 25th and 75th percentiles, bars = standard deviation, black circles = outliers). All changes in growth rates between the two different DIP concentrations were significant for all three cyanobacteria, Kruskal Wallis test, p < 0.05. Numbers in parentheses below X-axis indicate the number of measured cells. (a,b) Nodularia (red, Nod), Dolichospermum (white, Dol) and Aphanizomenon (blue, Aph). Cellular elemental ratios at t0 and t24 can also be found in Supplementary Table S1.
The predicted CO2:DIP uptake ratio was very similar to the experimental, isotope-based ratio of CO2 fixation to DIP uptake for Nodularia (Fig. 7a) indicating that this cyanobacterium relied almost entirely on DIP uptake for its growth, even at very low DIP concentrations. In contrast, a massive mismatch was observed between the predicted and the experimental ratio for Aphanizomenon (Fig. 7a) indicating substantial utilization of organic P: approximately 85% of its cellular P was obtained from organic P while only 15% was acquired from DIP uptake. Dolichospermum showed an intermediate strategy (Fig. 7) with about 40% of phosphorus originating from organic P and 60% from DIP. This segregation of phosphorus usage in field populations is surprising as cultured isolates of all three cyanobacteria are (genetically) capable of using organic P48,49,50,51. Further, this result from the field is at odds with several previous laboratory-based, culture studies suggesting that Nodularia outcompetes Aphanizomenon for organic P sources8,9. One mechanism to increase an organism’s competitiveness is to make use of the surrounding microbiome (or phycosphere). Interactions between a ‘host’ and its microbiome include but are not limited to mutualism and competition52. Associated microorganisms of the filamentous, non-heterocystous cyanobacterium Trichodesmium, for example, possess an array of tools for the hydrolysis of DIP from organic P through alkaline phosphatases which could then potentially be taken up by Trichodesmium itself11. The cyanobacterium Aphanizomenon ovalisporum uses the production of the toxin cylindrospermopsin to induce extracellular alkaline phosphatase activity in its surrroundings53. The thereby cleaved DIP could then be used by A. ovalisporum to satisfy its P demand. Alkaline phosphatases cleave the DIP (i.e. phosphate or Pi) from phosphoester bonds, and can exist as extracellular, membrane-bound, periplasmic, or cytoplasmic enzymes, although the majority appears to be extracellular and cytoplasmic in marine bacteria54. In our experiments, the DIP assimilated by Nodularia (as measured by the 33P incorporation) can originate from the initial 33P-labeled DIP pool (Supplementary Fig. S7; pathway (1) as well as from non-labeled DIP that has been released from organic P by extracellular hydrolysis by Nodularia or hydrolysis by surrounding microorganisms. Only the direct uptake from the initial 33P-labeled DIP pool will be captured by our 33P-DIP method. Since the measured 33P-labeled DIP uptake by Nodularia almost entirely accounts for their cellular P demand, it is very unlikely that they used a substantial amount of organic P (Supplementary Fig. S7; pathway 2–4). The indirect use of DOP by Aphanizomenon (pathways 3 and 4 in Supplementary Fig. S7) cannot be completely excluded here but is unlikely given the sparse epibiotic population of Aphanizomenon filaments38,55.
In essence, our field study shows that internally stored P did not play a substantial role in extending the HAB through late summer as elemental ratios were already around Redfield at the start of the bloom. The plasticity in the cellular elemental stoichiometry likely contributed to the growth of all three cyanobacteria with elemental ratios exceeding Redfield already a few days after the onset of the bloom. Most remarkably, however, organic P is used unequally by the three cyanobacteria with Nodularia relying almost exclusively on DIP and Aphanizomenon retrieving the majority of its cellular P from organic P. When DIP was added to incubations, Aphanizomenon did not grow faster than at very low DIP concentrations (Fig. 7b) suggesting that Aphanizomenon’s growth is independent of the availability of DIP during late summer. In contrast, the growth rate of Nodularia was stimulated when DIP was added to our incubations from 0.35 to 0.59 d−1 (Mann-Whitney test, p < 0.01) (Fig. 7b). Interestingly, Dolichospermum showed only a slightly enhanced growth rate resembling its intermediate DIP usage (Fig. 7). Overall our results indicate that the different strategies used to overcome summertime DIP scarcity are mostly based on differences in organic P usage between the different cyanobacteria. While several factors control the spatial and temporal distribution of the different cyanobacteria in the Baltic Sea, we hypothesize that its efficiency to use organic P leads to the numerical dominance of Aphanizomenon in the central Baltic during the stratified summer period when the three cyanobacteria co-occur (Fig. 3b and56). In contrast, the growth of Nodularia appears to benefit from the availability of DIP. A modelling study found that the best fit to observations of Nodularia blooms in the Baltic Sea could only be explained if DIP input by turbulent mixing and upwelling events were considered in addition to excess DIP in the water at the start of summer57. Further, a mesocosm study in the Baltic Sea found that growth of Nodularia was enhanced relative to Aphanizomenon and Dolichospermum when DIP was added to the water58. Both studies are in agreement with our results and together suggest that occurrences and abundances of Nodularia could largely benefit from increased DIP availability.
Modelling projections of the environmental conditions in the Baltic Sea under changing climate and nutrient inputs in the 21st century reveal that most scenarios (including business-as-usual) will lead to increased nutrient availability in surface waters, in particular phosphorus59. Furthermore, a major fraction of the phosphorus supply to the Baltic Sea is from deeper waters that accumulate DIP released from anoxic/hypoxic sediments60. Increases in anoxia and hypoxia have thus kept P quota in the Baltic Sea at the same or increased levels even when inputs have been reduced60. Our results imply that when the overall DIP availability increases and/or DIP is introduced to surface waters during summer stratification (e.g., by turbulent mixing, upwelling or point sources), out of the three cyanobacteria, the toxin-producer Nodularia would benefit most and potentially outcompete other HAB organisms. An increased availability or external input of DIP could thus trigger or increase blooms of Nodularia and shift non-toxic blooms toward more toxic ones by changing the community structure. The extent, intensity and frequency of HABs worldwide are predicted to increase under global climate change61 and eutrophication via anthropogenic activity62. The capability of HAB organisms to use organic P may be a critical factor that not only determines the microbial community structure, but the overall harmfulness and associated costs of algal blooms.
Our new approach allows for the simultaneous measurement of cellular DIP uptake and C and N assimilation rates in individual cells. The combination of these rates with the cellular elemental stoichiometry enabled us to quantify the importance of DIP, P storage and organic P for the growth of harmful cyanobacteria in a mixed environmental community. This approach could be applied in other low-phosphate ecosystems to determine the importance of different P sources for microbial growth and the potential role of organic P in the success of bloom-forming organisms.
Methods
Sampling and experimental set-up
Culture experiments
Nodularia spumigena (KAC64) cultures were grown on artificial seawater medium (BG-11) adjusted to a salinity of 7 and without the addition of fixed N. Growth conditions were set to room temperature (~20 °C) and a 12:12 h light/dark cycle. To determine single-cell CO2 and DIP-uptake rates cultures were transferred to fresh medium (150 nmol DIP L−1) 48 hours prior to the incubation experiment. For the incubation experiments, medium was enriched with NaH13CO3 (≥98 atom%, Sigma-Aldrich), resulting in a final enrichment of 13.6 atom% 13C in the DIC pool, and transferred carefully to 6-mL Exetainers. To each of these Exetainers a single hand-picked Nodularia colony was added. Afterwards, ~3 MBq 33PO43−, corresponding to 1.57 pmol 33P, were added to each Exetainer. This resulted in an isotope enrichment of 0.16 atom% 33P (at 150 nmol DIP L−1) with no significant natural background of 33P. Incubations lasted 24 hours and were terminated and prepared for further analysis as described below.
Field experiments
In June 2015, field sampling took place at the Swedish National Monitoring site B1 (5848′18″N, 17°37′52″E) close to the south-eastern coast of Sweden using a small motor-driven boat. Bulk water and Aphanizomenon colonies (the only abundant cyanobacterium) were collected with 30-L high-density polyethylene (HDPE) containers (Nalgene) and a plankton net (mesh size 20 μm), respectively, on 13 June 2015. Samples were transported back to the Askö Laboratory (Baltic Sea Center) within 30 minutes after sampling. Bulk seawater was subsampled for nutrient concentrations while Aphanizomenon colonies were incubated with H13CO3−, 15N2 and 33PO43− to measure single-cell CO2, N2 and DIP assimilation rates, respectively, using the developed method (see ‘Single-cell CO2, N2 and DIP assimilation’ below). Incubations were carried out in a temperature- and light-controlled room that was set for in situ conditions measured on the sampling day at station B1 at a 1–2 m water depth (11.7 °C; 280 µmol photons m−2 s−1 light irradiance; 18 h:6 h light:dark cycle).
In August 2015, field sampling was carried out as described above on three different occasions (06, 08 and 12 August 2015) at the same site. Samples were transported back to the Askö Laboratory (Baltic Sea Center) within 30 minutes after sampling.
In the first experiment, surface water was subsampled for nutrient (NH4+, NO2−/NO3− and PO43−) concentrations and the abundance of filamentous cyanobacteria, while the colonies were used to determine colony-based DIP uptake rates using the radiotracer 33P. The second experiment was carried out as described above with the addition of incubations for single-cell rates of CO2 fixation, N2 fixation and DIP uptake using a triple isotope approach with H13CO3−, 15N2 and 33PO43−, respectively. A third experiment was carried out as described for the first experiment with additional colony incubations under artificially increased (+1 µmol L−1) DIP concentrations and with the addition of stable isotopes H13CO3− and 15N2. All incubations were carried out in a temperature- and light-controlled room that was set for in situ conditions measured on the sampling day at station B1 at a 1–2 m water depth (17.4–18.0 °C; ~280 µmol photons m−2 s−1 light irradiance; 18 h:6 h light:dark cycle).
Nutrient concentrations
Samples for ammonium (NH4+) concentration were taken immediately upon arrival at the laboratory and subsequently measured fluorometrically63. Subsamples for phosphate (PO43−; dissolved inorganic phosphorus, DIP) and nitrite/nitrate (NO2−/NO3−) concentrations were pre-filtered (0.45 µm pore size Millex® syringe filter; Millipore) and frozen at −20 °C until analysis while samples for total phosphorus (TP) were frozen untreated at −20 °C. DIP concentration was measured with an auto-analyzer (TRAACS 800 Bran Luebbe) equipped with a 1-m long capillary system64 (World Precision Instruments) at the Max-Planck-Institute for Marine Microbiology within three months of sampling. Both nitrite/nitrate (NO2−/NO3−) and total phosphorus (TP) concentrations were determined by the Stockholm University Baltic Sea Monitoring Group using a standard flow injection analysis (FIA, QuikChem 8000 method: 31-107-06-1-A Lachat Instruments), and oxidation of phosphorus by pyrosulfate coupled to a segmented flow system7 (ALPKEM Flow Solution IV). The total organic phosphorus concentration (TOP) was then calculated by subtracting DIP from the TP concentrations. Nutrient concentrations at station B1 before and after our sampling campaign were retrieved from the Swedish Meteorological and Hydrological Institute (SMHI) database at www.smhi.se, open access data under the Creative Commons Attribution 4.0.
Abundance of filamentous cyanobacteria
Aphanizomenon sp., Dolichospermum sp. and Nodularia spumigena, were identified and counted in (acid) Lugol-preserved seawater as described in the HELCOM Guidelines65.
Colony-based DIP-uptake rates
Cyanobacterial colonies were identified using a stereomicroscope, hand-picked with a needle, washed in filtered seawater (0.2 μm pore size) and transferred to 6 mL gas-tight Exetainer® filled with filtered seawater. The colony-based DIP uptake rates were measured using radiolabeled phosphate (150 kBq 33PO43− in 6 mL of filtered (0.2 µm) water, specific activity 3000 Ci mmol−1, t1/2 25.4 d; Hartmann Analytic) in two different sets of incubations: for the first set, each Exetainer® contained ten colonies for rate measurements using liquid scintillation counting (LSC), for the second set each Exetainer® contained a single colony for rate measurements using digital autoradiography (DAR) analysis. Each of these two sets contained triplicate Exetainer® incubations per time point (0, 6, 12, 18 and 24 h). At each time point, the triplicate sets were filtered through polycarbonate filters (GTTP, 0.2 µm pore size, 25 mm diameter, Millipore) and washed twice with 5 mL of filtered-seawater for LSC The filters were then transferred to a 6 mL scintillation vial containing 5 mL scintillation fluid (Irga-safe plus, PerkinElmer). For DAR analysis, the samples were preserved with formaldehyde (1% w/v final concentration) for 30 minutes at room temperature prior to filtering. DAR samples were mounted onto microscope glass slides, using a small drop of 0.1% low-melting-point agarose, dried and stored at room temperature before analysis using a real-time radio imaging system (Micro Imager, BioSpace Lab) as described in Goldhammer et al.66.
Total radioactivity was determined from every incubation bottle using 100 µL of water that were directly transferred into a scintillation vial. All samples from one experiment were measured on the same day using a liquid scintillation counter (type: 425-034, Hidex). Control incubations were carried out with heat-killed (80 °C, 30 minutes) or no colonies. Both controls were used to check for any unspecific binding of the radiotracer to the bottle, filter or organic material. Colony-based DIP uptake rates were calculated using the linear regression for the radiotracer incorporation into biomass between time points, the total amount of radiotracer added and the DIP concentration in the water. Uptake rates were corrected for radiotracer decay, and unspecific radiotracer binding using the controls. Uptake rates were normalized to the number of colonies per incubation.
Single-cell CO2, N2 and DIP assimilation
Single-cell CO2, N2 and DIP-uptake rates were determined using a triple isotope approach followed by nanometer-scale secondary ion mass spectrometry (nanoSIMS). In detail, a 2.5-L polycarbonate bottle with filtered seawater (0.2 μm pore size) was enriched with NaH13CO3 (≥98 atom%, Sigma-Aldrich) and 15N2 (≥99 atom%; lot # 1-17299, Cambridge Isotope Laboratories; modified bubble method as in Klawonn et al.67). The prepared water was carefully transferred to 6-mL Exetainers to which a single hand-picked cyanobacterial colony was added. Subsequently, ~10 MBq 33PO43−, corresponding to 4.95 pmol 33P, were added to each Exetainer. This resulted in an isotope enrichment of 0.28 atom% 33P (294 nmol DIP L−1) in June and 2.43 atom% in August (34 nmol DIP L−1), with no significant natural background of 33P. These 33P enrichments led to detection limits of ~0.107 and 0.012 of relative P incorporation for June and August, respectively (Supplementary Fig. S1). After the addition of 33PO43−, the Exetainers were closed headspace-free and incubated as described above. Per cyanobacterial genera three samples were terminated after 24 hours (t24) for June and August and a single sample was terminated at time zero (t0) (i.e. immediately after the last tracer was added) for August only. One of the three sampled colonies was later used for SEM-EDS analysis while the other colonies were used for nanoSIMS measurements. Total radioactivity was determined from each Exetainer as described above except that the 100 µL of incubation water were first diluted ten-fold with nanopure water and 100 µL from this dilution were used for the measurement. The remaining sample was fixed with formaldehyde (1% w/v final concentration) for 30 minutes at room temperature and then filtered through a gold-sputtered polycarbonate filter (0.22 μm pore size; 25 mm diameter; Millipore). The filters were washed with nanopure water, dried at room temperature and mounted onto microscope glass slides using a small drop of 0.1% low-melting-point agarose. Microscope slides were stored in the dark at room temperature before single-cell analyses.
Single-cell elemental ratios
Single-cell C:N, C:P and N:P elemental ratios were obtained in August samples using a scanning electron microscope (SEM; Quanta FEG 250, FEI) equipped with an energy-dispersive X-ray spectroscopy (EDS) double detector system (QUANTAX EDS, Bruker Nano GmbH) at the Max Planck Institute for Marine Microbiology. The detectors (XFlash 6/30) had a detector area of 30 mm2 and an energy resolution at Mn K α line of <123 eV, which allows for quantification of light elements. The NBS SRM 1155 ANSI 316 stainless steel standard was used to check the performance of the EDS system. Prior to EDS measurements, we tested the EDS spectra with an accelerating voltage of 10 kV and 20 kV. The same elements up to Ca could be detected with both voltages, sufficient to obtain ratios for the light elements. In order to reduce the penetration depth, and thereby reduce potential signals from the filter, the 10 kV accelerating voltage was used for further measurements. Monte-Carlo simulations showed that a 10 kV accelerating voltage led to a penetration depth of ~2 µm indicating that the obtained elemental ratios were representative for the cellular composition due to minimal interference from the filter. The analysis of the cellular elemental content was done using the P/B-ZAF method (Quantax 400 software, version 1.9; Bruker) in which the elements C, N, O, Na, Mg, Al, P, S, Cl, K and Ca were included. This analysis relies on the ratio of the X-ray intensities relative to the intensities of the Bremsstrahlung, which is simultaneously recorded by the EDS detector. The P/B-ZAF method is a standardless evaluation without the need for normalization and is particularly useful for the analysis of rough surfaces and particles. Analyses were done from samples of the beginning and the end of the incubation for all three different cyanobacteria in August. Significant differences in the elemental ratios were tested using a Kruskal-Wallis test (for non-normal distribution) (Fig. 7 and Supplementary Table S1).
nanoSIMS analysis
Single-cell CO2, N2 and DIP uptake rates were determined based on the incorporation of DI13C, 15N2 and 33PO43− into the cyanobacteria cell biomass. The radiotracer 33P decays to the stable 33S with a half-life of only 25.4 days and becomes part of the cellular sulfur pool. Therefore, DIP-uptake rates were measured based on the labelling excess of 33S/32S (natural abundance ratio of 0.0078) in the biomass using a nanoSIMS 50 L (CAMECA) at the Max Planck Institute for Marine Microbiology in Bremen, Germany. Measurements were done after at least four half-lives had passed to ensure that the majority of the 33P had decayed.
Aphanizomenon, Dolichospermum or Nodularia filaments were identified based on auto-fluorescence and morphology and were marked using a laser micro-dissection (LMD) microscope (6000B, Leica). After loading the sample into the nanoSIMS, marked filaments were pre-sputtered with the Cs+ primary ion beam of 300 pA to remove surface contamination, implant Cs+ ions into the sample and to achieve an approximately stable ion emission rate. A primary Cs+ ion beam with a beam current between 1 and 1.2 pA and a beam diameter between 50 and 100 nm was rastered across the cells for analysis. For individual cells, secondary ion images of 12C−, 13C−, 12C14N−, 12C15N−, 31P−, 32S− and 33S− were simultaneously recorded using seven electron multipliers. The analysis areas were between 20 × 20 and 50 × 50 µm in size and an image size of 256 × 256 pixels with a dwell time of 1 ms per pixel and a minimum of 50 planes. To minimize interferences for 12C15N− and 33S− the instrument was tuned for high mass resolution (>7000 MRP). All nanoSIMS measurements were analyzed using the Matlab-based software package Look@NanoSims68 (available at http://nanosims.geo.uu.nl/nanosims-wiki/doku.php/nanosims:lans). For every measurement, secondary ion images were drift-corrected and accumulated. Regions of interest (ROIs) were drawn around cells using the secondary ion images, and 13C/12C, 15N/14N (inferred from 12C15N/12C14N), 33S/32S and 31P/12C ratios were calculated.
Accuracy and precision of the 33S/32S measurements were determined with an in-house sulfate standard (BaSO4) which had natural abundance values. Cells were regarded as significantly enriched when their 33S/32S ratio was above the natural abundance +3 × standard deviation of the mean of the sulfate standard, i.e. a 33S/32S ratio of 0.0084 was the detection limit with a natural abundance value of 0.0078.
Single-cell CO2, N2 and DIP uptake rates
Individual cell biovolumes were calculated based on cell-diameter measurements, obtained via nanoSIMS analysis, in combination with a geometrical shape model. For Aphanizomenon and Dolichospermum we used a cylinder +2 half spheres-shape and for Nodularia a cylinder-shape69. Cellular C content was determined based on the biovolume and a volume-to-carbon conversion factor of 17.5 fmol C μm−3 (ref.70). Cellular N and P content was calculated using the molar C:N and C:P ratios obtained via SEM-EDS described above.
CO2 and N2 fixation rates were calculated using the cellular C and N content in combination with the single-cell 15N2 and DI13C incorporation, analogous to the calculations described in Musat et al.71. Single-cell DIP-uptake rates were calculated using the cellular P-content (Pcell) obtained via the SEM-EDS, DI33P incorporated in the cell measured as 33S excess (33Sexcess), the DI33P labelling yield (33Pexcess-inc) and incubation times (t) as follows:
Growth rates
Growth rates were determined based on the incorporation of DI13C (Cexcess) into the biomass assuming an even distribution of the isotope in the biomass during cell division as follows:
with Cexcess-inc representing atom%-labelling of DIC in the bottle and t the incubation time. A Kruskal-Wallis test was used to determine significant differences in C-based growth rates between species and incubation conditions (in situ or added DIP).
Contribution of DIP and organic P to cellular growth
To determine the contribution of DIP to cellular growth, we compared the experimental CO2:DIP uptake ratio, obtained from the nanoSIMS analysis, with the predicted ratio of CO2:DIP (CO2:P). The latter was determined from the mass-balanced cellular growth between the beginning and the end of the incubation as follows:
which is then solved for Ruptake, i.e. the predicted ratio
with M = median biomass at the start (Mstart; obtained from cell size and volume conversion) or end (Mfinal; calculated by growth rates of start biomass) of the incubation and the newly formed biomass (Mnew; difference between start and end biomass) during the incubation and with R = median elemental ratio of the biomass at the start (Rstart) and end (Rfinal) of the incubation (both obtained from the SEM-EDS) and the median uptake ratio of CO2:DIP (Ruptake) during the incubation.
The contribution of DIP and organic P uptake to cellular growth was then calculated by dividing the ratio of uptake rates obtained by nanoSIMS (CO2:DIP) by the predicted ratio (CO2:P, i.e. Ruptake). To obtain the fraction of DIP supporting cellular growth the resulting number (P/DIP) was inverted (DIP/P). It follows that the cellular growth supported by organic P = (1 − DIP/P) with DIP/P + organicP/P = 1.
References
Stal, L. J. et al. BASIC: Baltic Sea cyanobacteria. An investigation of the structure and dynamics of water blooms of cyanobacteria in the Baltic Sea - responses to a changing environment. Cont. Shelf Res. 23, 1695–1714, https://doi.org/10.1016/j.csr.2003.06.001 (2003).
Michalak, A. M. et al. Record-setting algal bloom in Lake Erie caused by agricultural and meteorological trends consistent with expected future conditions. Proc. Natl. Acad. Sci. USA 110, 6448–6452, https://doi.org/10.1073/pnas.1216006110 (2013).
Elser, J. J. et al. Global analysis of nitrogen and phosphorus limitation of primary producers in freshwater, marine and terrestrial ecosystems. Ecol. Lett. 10, 1135–1142, https://doi.org/10.1111/j.1461-0248.2007.01113.x (2007).
Sañudo-Wilhelmy, S. A. et al. Phosphorus limitation of nitrogen fixation by Trichodesmium in the central Atlantic Ocean. Nature 411, 66, https://doi.org/10.1038/35075041.
Kuypers, M. M. M., Marchant, H. K. & Kartal, B. The microbial nitrogen-cycling network. Nat. Rev. Microbiol. 16, 263, https://doi.org/10.1038/nrmicro.2018.9 (2018).
Van Mooy, B. A. S. et al. Phytoplankton in the ocean use non-phosphorus lipids in response to phosphorus scarcity. Nature 458, 69–72, https://doi.org/10.1038/nature07659 (2009).
Walve, J. & Larsson, U. Blooms of Baltic Sea Aphanizomenon sp (cyanobacteria) collapse after internal phosphorus depletion. Aquat. Microb. Ecol. 49, 57–69, https://doi.org/10.3354/ame01130 (2007).
Degerholm, J., Gundersen, K., Bergman, B. & Söderbäck, E. Phosphorus-limited growth dynamics in two Baltic Sea cyanobacteria, Nodularia sp and Aphanizomenon sp. FEMS Microbiol. Ecol. 58, 323–332, https://doi.org/10.1111/j.1574-6941.2006.00180.x (2006).
Vahtera, E., Laamanen, M. & Rintala, J. M. Use of different phosphorus sources by the bloom-forming cyanobacteria Aphanizomenon flos-aquae and Nodularia spumigena. Aquat. Microb. Ecol. 46, 225–237, https://doi.org/10.3354/ame046225 (2007).
Paerl, H. W. & Pinckney, J. L. A mini-review of microbial consortia: Their roles in aquatic production and biogeochemical cycling. Microb. Ecol. 31, 225–247 (1996).
Frischkorn, K. R., Rouco, M., Van Mooy, B. A. S. & Dyhrman, S. T. Epibionts dominate metabolic functional potential of Trichodesmium colonies from the oligotrophic ocean. ISME J. 11, 2090–2101, https://doi.org/10.1038/ismej.2017.74 (2017).
Zubkov, M. V. et al. Microbial control of phosphate in the nutrient-depleted North Atlantic subtropical gyre. Environ. Microbiol. 9, 2079–2089, https://doi.org/10.1111/j.1462-2920.2007.01324.x (2007).
Orchard, E. D., Ammerman, J. W., Lomas, M. W. & Dyhrman, S. T. Dissolved inorganic and organic phosphorus uptake in Trichodesmium and the microbial community: The importance of phosphorus ester in the Sargasso Sea. Limnol. Oceanogr. 55, 1390–1399, https://doi.org/10.4319/lo.2010.55.3.1390 (2010).
Cottrell, M. T. & Kirchman, D. L. Contribution of major bacterial groups to bacterial biomass production (thymidine and leucine incorporation) in the Delaware estuary. Limnol. Oceanogr. 48, 168–178, https://doi.org/10.4319/lo.2003.48.1.0168 (2003).
Longnecker, K., Lomas, M. W. & Van Mooy, B. A. S. Abundance and diversity of heterotrophic bacterial cells assimilating phosphate in the subtropical North Atlantic Ocean. Environ. Microbiol. 12, 2773–2782, https://doi.org/10.1111/j.1462-2920.2010.02247.x (2010).
Nielsen, J. L. & Nielsen, P. H. In Handbook of Hydrocarbon and Lipid Microbiology (ed. Kenneth N. Timmis) 4093–4102 (Springer Berlin Heidelberg, 2010).
Sebastián, M., Pitta, P., González, J. M., Thingstad, T. F. & Gasol, J. M. Bacterioplankton groups involved in the uptake of phosphate and dissolved organic phosphorus in a mesocosm experiment with P-starved Mediterranean waters. Environ. Microbiol. 14, 2334–2347, https://doi.org/10.1111/j.1462-2920.2012.02772.x (2012).
Sintes, E. & Herndl, G. J. Quantifying substrate uptake by individual cells of marine bacterioplankton by catalyzed reporter deposition fluorescence in situ hybridization combined with microautoradiography. Appl. Environ. Microb. 72, 7022–7028, https://doi.org/10.1128/aem.00763-06 (2006).
Casey, J. R. et al. Phytoplankton taxon-specific orthophosphate (Pi) and ATP utilization in the western subtropical North Atlantic. Aquat. Microb. Ecol. 58, 31–44, https://doi.org/10.3354/ame01348 (2009).
Michelou, V. K., Lomas, M. W. & Kirchman, D. L. Phosphate and adenosine-5′-triphosphate uptake by cyanobacteria and heterotrophic bacteria in the Sargasso Sea. Limnol. Oceanogr. 56, 323–332, https://doi.org/10.4319/lo.2011.56.1.0323 (2011).
Benitez-Nelson, C. R. & Buesseler, K. O. Variability of inorganic and organic phosphorus turnover rates in the coastal ocean. Nature 398, 502–505, https://doi.org/10.1038/19061 (1999).
Azam, F. & Hodson, R. E. Dissolved ATP in the sea and its utilization by marine bacteria. Nature 267, 696–698, https://doi.org/10.1038/267696a0 (1977).
Björkman, K. M. & Karl, D. M. Presence of dissolved nucleotides in the North Pacific Subtropical Gyre and their role in cycling of dissolved organic phosphorus. Aquat. Microb. Ecol. 39, 193–203, https://doi.org/10.3354/ame039193 (2005).
Lomas, M. W. et al. Sargasso Sea phosphorus biogeochemistry: an important role for dissolved organic phosphorus (DOP). Biogeosciences 7, 695–710, https://doi.org/10.5194/bg-7-695-2010 (2010).
Perry, M. J. Alkaline phosphatase activity in subtropical central North Pacific waters using a sensitive fluorometric method. Mar. Biol. 15, 113–119, https://doi.org/10.1007/bf00353639 (1972).
Huang, B. et al. Alkaline phosphatase activity of phytoplankton in East China Sea coastal waters with frequent harmful algal bloom occurrences. Aquat. Microb. Ecol. 49, 195–206, https://doi.org/10.3354/ame01135 (2007).
Løvdal, T., Tanaka, T. & Thingstad, T. F. Algal—bacterial competition for phosphorus from dissolved DNA, ATP, and orthophosphate in a mesocosm experiment. Limnol. Oceanogr. 52, 1407–1419, https://doi.org/10.4319/lo.2007.52.4.1407 (2007).
Sivonen, K. et al. Occurrence of the hepatotoxic cyanobacterium Nodularia spumigena in the Baltic Sea and structure of the toxin. Appl. Environ. Microb. 55, 1990–1995 (1989).
Halinen, K., Jokela, J., Fewer, D. P., Wahlsten, M. & Sivonen, K. Direct evidence for production of microcystins by Anabaena strains from the Baltic Sea. Appl. Environ. Microb. 73, 6543–6550, https://doi.org/10.1128/AEM.01377-07 (2007).
Sellner, K. G., Doucette, G. J. & Kirkpatrick, G. J. Harmful algal blooms: causes, impacts and detection. J. Ind. Microbiol. Biot. 30, 383–406, https://doi.org/10.1007/s10295-003-0074-9 (2003).
Kahru, M. & Elmgren, R. Multidecadal time series of satellite-detected accumulations of cyanobacteria in the Baltic Sea. Biogeosciences 11, 3619–3633, https://doi.org/10.5194/bg-11-3619-2014 (2014).
Larsson, U., Hajdu, S., Walve, J. & Elmgren, R. Baltic Sea nitrogen fixation estimated from the summer increase in upper mixed layer total nitrogen. Limnol. Oceanogr. 46, 811–820, https://doi.org/10.4319/lo.2001.46.4.0811 (2001).
Nausch, M., Nausch, G., Wasmund, N. & Nagel, K. Phosphorus pool variations and their relation to cyanobacteria development in the Baltic Sea: A three-year study. J. Marine Syst. 71, 99–111, https://doi.org/10.1016/j.jmarsys.2007.06.004 (2008).
Wilson, R. G., Stevie, F. A. & Magee, C. W. Secondary ion mass spectrometry: a practical handbook for depth profiling and bulk impurity analysis (1989).
Mohr, W., Vagner, T., Kuypers, M. M. M., Ackermann, M. & LaRoche, J. Resolution of conflicting signals at the single-cell level in the regulation of cyanobacterial photosynthesis and nitrogen fixation. PLoS ONE 8, https://doi.org/10.1371/journal.pone.0066060 (2013).
Schreiber, F. et al. Phenotypic heterogeneity driven by nutrient limitation promotes growth in fluctuating environments. Nat. Microbiol. 1, https://doi.org/10.1038/nmicrobiol.2016.55 (2016).
Nikolic, N. et al. Cell-to-cell variation and specialization in sugar metabolism in clonal bacterial populations. PloS Genet. 13, https://doi.org/10.1371/journal.pgen.1007122 (2017).
Ploug, H. et al. Carbon and nitrogen fluxes associated with the cyanobacterium Aphanizomenon sp. in the Baltic Sea. ISME J. 4, 1215–1223, https://doi.org/10.1038/ismej.2010.53 (2010).
Foster, R. A., Sztejrenszus, S. & Kuypers, M. M. M. Measuring carbon and N2 fixation in field populations of colonial and free-living unicellular cyanobacteria using nanometer-scale secondary ion mass spectrometry. J. Phycol. 49, 502–516, https://doi.org/10.1111/jpy.12057 (2013).
Martínez-Pérez, C. et al. The small unicellular diazotrophic symbiont, UCYN-A, is a key player in the marine nitrogen cycle. Nat. Microbiol. 1, https://doi.org/10.1038/nmicrobiol.2016.163 (2016).
Wasmund, N. Occurrence of cyanobacterial blooms in the Baltic Sea in relation to environmental conditions. Int. Rev. Ges. Hydrobio. Hydrogr. 82, 169–184, https://doi.org/10.1002/iroh.19970820205 (1997).
Talarmin, A. et al. Improved methodology to measure taxon-specific phosphate uptake in live and unfiltered samples. Limnol. Oceanogr.-Meth. 9, 443–453, https://doi.org/10.4319/lom.2011.9.443 (2011).
Musat, N. et al. The effect of FISH and CARD-FISH on the isotopic composition of 13C- and 15N-labeled Pseudomonas putida cells measured by nanoSIMS. Syst. Appl. Microbiol. 37, 267–276, https://doi.org/10.1016/j.syapm.2014.02.002 (2014).
Stevens, S. E., Paone, D. A. M. & Balkwill, D. L. Accumulation of cyanophycin granules as a result of phosphate limitation in Agmenellum quadruplicatum. Plant Physiol. 67, 716–719, https://doi.org/10.1104/pp.67.4.716 (1981).
Kromkamp, J. Formation and functional significance of storage products in cyanobacteria. New Zeal. J. Mar. Fresh. 21, 457–465, https://doi.org/10.1080/00288330.1987.9516241 (1987).
Dyhrman, S. T. et al. The transcriptome and proteome of the diatom Thalassiosira pseudonana reveal a diverse phosphorus stress response. PLoS ONE 7, e33768, https://doi.org/10.1371/journal.pone.0033768 (2012).
Martin, P., Dyhrman, S. T., Lomas, M. W., Poulton, N. J. & Van Mooy, B. A. S. Accumulation and enhanced cycling of polyphosphate by Sargasso Sea plankton in response to low phosphorus. Proc. Natl. Acad. Sci. USA 111, 8089–8094, https://doi.org/10.1073/pnas.1321719111 (2014).
Wang, H. et al. Genome-derived insights into the biology of the hepatotoxic bloom-forming cyanobacterium Anabaena sp. strain 90. BMC Genomics 13, 613, https://doi.org/10.1186/1471-2164-13-613 (2012).
Voß, B. et al. Insights into the physiology and ecology of the brackish-water-adapted cyanobacterium Nodularia spumigena CCY9414 based on a genome-transcriptome analysis. PLoS ONE 8, e60224, https://doi.org/10.1371/journal.pone.0060224 (2013).
Šulčius, S. et al. Draft genome sequence of the cyanobacterium Aphanizomenon flos-aquae strain 2012/KM1/D3, isolated from the Curonian Lagoon (Baltic Sea). Genome Announc. 3, e01392–01314, https://doi.org/10.1128/genomeA.01392-14 (2015).
Teikari, J. E. et al. Strains of the toxic and bloom-forming Nodularia spumigena (cyanobacteria) can degrade methylphosphonate and release methane. ISME J. 12, 1619–1630, https://doi.org/10.1038/s41396-018-0056-6 (2018).
Seymour, J. R., Amin, S. A., Raina, J.-B. & Stocker, R. Zooming in on the phycosphere: the ecological interface for phytoplankton–bacteria relationships. Nat. Microbiol. 2, 17065, https://doi.org/10.1038/nmicrobiol.2017.65.
Bar-Yosef, Y., Sukenik, A., Hadas, O., Viner-Mozzini, Y. & Kaplan, A. Enslavement in the water body by toxic Aphanizomenon ovalisporum, inducing alkaline phosphatase in phytoplanktons. Curr. Biol. 20, 1557–1561, https://doi.org/10.1016/j.cub.2010.07.032 (2010).
Luo, H., Benner, R., Long, R. A. & Hu, J. Subcellular localization of marine bacterial alkaline phosphatases. Proc. Natl. Acad. Sci. USA 106, 21219–21223, https://doi.org/10.1073/pnas.0907586106 (2009).
Hoppe, H. G. Blue-Green algae agglomeration in surface water: a microbiotope of high bacterial activity. Kiel. Meeresforsch. (Sonderh.) 5, 291–303 (1981).
Klawonn, I. et al. Cell-specific nitrogen- and carbon-fixation of cyanobacteria in a temperate marine system (Baltic Sea). Environ. Microbiol. 18, 4596–4609, https://doi.org/10.1111/1462-2920.13557 (2016).
Laanemets, J. et al. A fuzzy logic model to describe the cyanobacteria Nodularia spumigena blooms in the Gulf of Finland, Baltic Sea. Hydrobiologia 554, 31–45, https://doi.org/10.1007/s10750-005-1004-x (2006).
Wasmund, N., Nausch, G. & Voss, M. Upwelling events may cause cyanobacteria blooms in the Baltic Sea. J. Marine Syst. 90, 67–76, https://doi.org/10.1016/j.jmarsys.2011.09.001 (2012).
Meier, H. E. M. et al. Modeling the combined impact of changing climate and changing nutrient loads on the Baltic Sea environment in an ensemble of transient simulations for 1961–2099. Clim. Dynam. 39, 2421–2441, https://doi.org/10.1007/s00382-012-1339-7 (2012).
Conley, D. J. et al. Hypoxia-Related Processes in the Baltic Sea. Environ. Sci. Technol. 43, 3412–3420 (2009).
Paerl, H. W. & Huisman, J. Blooms Like It Hot. Science 320, 57–58, https://doi.org/10.1126/science.1155398 (2008).
Anderson, D., Glibert, P. & Burkholder, J. Harmful algal blooms and eutrophication: nutrient sources, compositions, and consequences. Estuaries 25, 704–726, https://doi.org/10.1016/j.hal.2008.08.017 (2002).
Holmes, R. M., Aminot, A., Kerouel, R., Hooker, B. A. & Peterson, B. J. A simple and precise method for measuring ammonium in marine and freshwater ecosystems. Can. J. Fish. Aquat. Sci. 56, 1801–1808, https://doi.org/10.1139/cjfas-56-10-1801 (1999).
Patey, M. D. et al. Determination of nitrate and phosphate in seawater at nanomolar concentrations. TrAC-Trend Anal. Chem. 27, 169–182 (2008).
HELCOM. Guidelines for the Baltic Monitoring Programme for the Third Stage: Biological determinands (1988).
Goldhammer, T., Brüchert, V., Ferdelman, T. G. & Zabel, M. Microbial sequestration of phosphorus in anoxic upwelling sediments. Nat. Geosci. 3, 557–561, https://doi.org/10.1038/ngeo913 (2010).
Klawonn, I. et al. Simple approach for the preparation of 15–15N2-enriched water for nitrogen fixation assessments: Evaluation, application and recommendations. Front. Microbiol. 6, https://doi.org/10.3389/fmicb.2015.00769 (2015).
Polerecky, L. et al. Look@NanoSIMS – a tool for the analysis of nanoSIMS data in environmental microbiology. Environ. Microbiol. 14, 1009–1023, https://doi.org/10.1111/j.1462-2920.2011.02681.x (2012).
Sun, J. & Liu, D. Geometric models for calculating cell biovolume and surface area for phytoplankton. J. Plankton Res. 25, 1331–1346, https://doi.org/10.1093/plankt/fbg096 (2003).
Svedén, J. B. et al. High cell-specific rates of nitrogen and carbon fixation by the cyanobacterium Aphanizomenon sp. at low temperatures in the Baltic Sea. FEMS Microbiol. Ecol. 91, https://doi.org/10.1093/femsec/fiv131 (2015).
Musat, N. et al. A single-cell view on the ecophysiology of anaerobic phototrophic bacteria. Proc. Natl. Acad. Sci. USA 105, 17861–17866, https://doi.org/10.1073/pnas.0809329105 (2008).
Öberg, J. Cyanobacteria blooms in the Baltic Sea. HELCOM Baltic Sea Environmental Fact Sheet 2015. Online (2015).
Acknowledgements
We thank Daniela Tienken, Katharina Kitzinger, Clara Martínez-Pérez, Osmund Darril Rebello, Gabriele Klockgether and Gaute Lavik from the MPI Bremen for their help with sampling support and (laboratory) analyses. Jacob Walve (Stockholm University) generously provided total phosphorus and nitrate/nitrite data, and Jörgen Öberg (SMHI) provided the satellite data of the cyanobacterial bloom extent. Igor Nemeth and Max Patzschke from Bruker Nano GmbH are thanked for the Monte-Carlo simulations and discussions on single-cell EDS data processing and quality control. Financial support was provided by the Max Planck Society as well as the Askö Laboratory (Baltic Sea Center) through travel grants and provision of accommodation, lab space and sampling equipment.
Author information
Authors and Affiliations
Contributions
N.J.S., W.M., T.G.F., M.V.Z., H.P. and M.M.M.K. designed the study and method development. N.J.S., J.D., T.G.F. carried out experiments. N.J.S., W.M., S.L. and M.M.M.K. analyzed data. N.J.S., W.M. and M.M.M.K. wrote the manuscript with the contribution of all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schoffelen, N.J., Mohr, W., Ferdelman, T.G. et al. Single-cell imaging of phosphorus uptake shows that key harmful algae rely on different phosphorus sources for growth. Sci Rep 8, 17182 (2018). https://doi.org/10.1038/s41598-018-35310-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35310-w
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.