Abstract
Active microbes likely have larger impact on gut health status compared to inactive or dormant microbes. We investigate the composition of active and total mucosal microbiota of treatment-naïve ulcerative colitis (UC) patients to determine the microbial picture at the start-up phase of disease, using both a 16S rRNA transcript and gene amplicon sequencing. DNA and RNA were isolated from the same mucosal colonic biopsies. Our aim was to identify active microbial members of the microbiota in early stages of disease and reveal which members are present, but do not act as major players. We demonstrated differences in active and total microbiota of UC patients when comparing inflamed to non-inflamed tissue. Several taxa, among them the Proteobacteria phyla and families therein, revealed lower transcriptional activity despite a high presence. The Bifidobacteriaceae family of the Actinobacteria phylum showed lower abundance in the active microbiota, although no difference in presence was detected. The most abundant microbiota members of the inflamed tissue in UC patients were not the most active. Knowledge of active members of microbiota in UC patients could enhance our understanding of disease etiology. The active microbial community composition did not deviate from the total when comparing UC patients to non-IBD controls.
Similar content being viewed by others
Introduction
Ulcerative colitis (UC) and Crohn’s disease represent two of the most common types of inflammatory bowel disease (IBD). These are complex immune-mediated disorders characterized by chronic intestinal inflammation, where genetics, the environment and the gut microbiota are all factors related to disease initiation and progression1,2,3. The role of the gut microbiota in the pathogenesis of IBD is still poorly understood, despite extensive research efforts during the last several decades. Several studies have revealed a pattern of reduced alpha diversity and abnormal microbial community composition with a decrease in a number of taxa within the phylum Firmicutes and an increase in the phyla Bacteriodetes and Proteobacteria4,5. At the species level, some candidates have been reported to have an altered abundance in diseased versus healthy subjects, but a microbial signature of disease has not been found3,6,7. The normal, healthy human gut microbiota, seen at the phylogenetic level, exhibits a generally high variability between and within individuals over time4,8,9. However, functional redundancy can reduce the effect of variations in microbial community structure, and the healthy gut microbiota is relatively stable at a functional level8,10.
16S rRNA gene sequencing is widely used for the determination of the total microbial community composition. A potential drawback in using 16S rRNA gene sequencing is that its results comprise a total mixture of active, dormant and dead bacteria. There are methods available to exclude detection of dead microbes11,12,13. Recent studies have utilized propidium monoazid (PMA) treatment prior to metagenomics sequencing (PMAseq) to exclude dead bacterial cell DNA from further analysis14,15. Our study focus on the mucosal microbiota and the PMAseq method cannot be used on stabilized tissue biopsies due to the stabilization process itself and the biopsy sample consistency16.
We, together with other researchers, have used the transcribed 16S rRNA counts, as opposed to the gene counts, to indicate the potential bacterial metabolic activity of a sample as rRNA levels correlate with the protein synthesis potential of the microorganisms and hence can be used to estimate their activity17,18,19. The potential active microbial community composition differs from the total microbial community composition in healthy and diseased individuals. The observation that active microbes seem to have a larger impact on gut health status than inactive or dormant microbes highlights the importance of studying metabolically active microbiota in both diseased and healthy individuals20,21,22.
In the present study, we investigated the composition of both the active and total microbiota of newly diagnosed, treatment-naïve UC patients to uncover the microbial picture at the time of disease diagnosis.
The functional capacity of the microbiota was further elucidated using Piphillin23. Our aim was to understand which microbial community members are active at diagnosis and which members are present, but may not be major players, in the early stages of the disease.
Results
Forty-six UC patients and 39 symptomatic non-IBD controls were included in the study, and 114 and 39 biopsy samples were collected, respectively. All the biopsies yielded sufficient DNA and RNA quantities and qualities for further analyses. The DNA and RNA samples were subjected to microbial profiling using 16S rRNA gene and gene transcript sequencing on the Illumina MiSeq platform with 250 bp overlapping paired-end reads. The samples with fewer than 3 000 and 10 000 raw reads, set from the DNA and RNA blanks, respectively, were excluded from further analysis. This cut-off left 137 of the 153 DNA samples and 129 of the 153 RNA samples from 44 of 46 UC patients and 35 of 39 symptomatic non-IBD controls to be further analysed (Tables 1, S1 and S2).
After quality filtering and assembling of overlapping paired-end reads in mothur, more than 7.9 million sequences were retained in both the DNA and the RNA datasets (with a mean of 67 071 per sample).
The sequences were clustered into 2043 OTUs at 97% identity. The OTUs spanned 122 microbial families and 20 phyla. Aggregating the OTU counts into families and phyla for abundance analyses resulted in 26 and 24 families remaining in the DNA and RNA datasets, respectively.
Small systematic differences in diversity
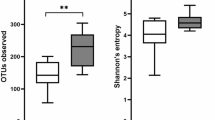
There was no significant difference in the alpha diversity between the UC patients (inflamed biopsies and non-inflamed biopsies) and the symptomatic controls, when comparing either the DNA or RNA samples (Fig. 1a and b). A non-significant lower diversity of the symptomatic non-IBD control group was observed in the RNA dataset (Fig. 1a). The Bland-Altman plot of the paired RNA and DNA samples showed some heterogeneity in the alpha diversity across the samples but few systematic differences between groups (Fig. 1c). Most of the samples showed a difference within + − 0.5. These differences cannot be explained by the disease status or the inflammation status.
Alpha diversity in inflamed and non-inflamed tissue. Distribution of alpha diversity in the inflamed and non-inflamed samples from UC patients and control patients, showing (a) the DNA and (b) the RNA datasets, and (c) a scatterplot of the alpha diversity in the paired RNA and DNA samples, with the average alpha diversity on the x-axis and the difference on the y-axis.
We performed a PCoA analysis with UniFrac distances to investigate whether the sample location could differentiate the samples and thus interfere with further analyses. The results show that alpha diversity is a main factor differentiating the samples and not the sampling location, inflammation status or disease status (Figures S1, S2 and S3).
The PCoA plot of UniFrac distances in the DNA and RNA datasets shows that the biopsy samples from the same patient are similar to each other and that there is no separation between the samples from the UC patients and the symptomatic non-IBD controls (Fig. 2a and b). Investigating the RNA and DNA data from the same samples, these are found near each other in the PCoA plot, indicating an overall pattern of agreement between the two datasets but not complete correspondence (Fig. 2c).
UniFrac distances between different diagnosis groups and datasets. PCoA plot for (a) the DNA dataset and (b) the RNA dataset. Each sample is coloured according to the disease status and inflammation status. The black lines connect the samples from the same patient. c) PCoA plot of a combined analysis from both the DNA and RNA dataset. Paired RNA-DNA samples are connected with black lines.
Taxonomic differences in total and active microbiota between inflamed and non-inflamed mucosa of UC patients
When comparing differences in the total (rRNA gene) microbial community composition between the inflamed and non-inflamed mucosa of UC patients, a significantly higher abundance of Enterobacteriaceae (phylum Proteobacteria) and a higher but not significant abundance in the Firmicutes family XI Incertae Sedis and in the Actinobacteria family Actinomycetaceae were observed in the inflamed mucosa (Tables 2, S3 and S4). However, none of these differences were observed when active (rRNA) microbial community compositions were compared. For the active microbiota, a significantly lower transcriptional activity in the Actinobacteria family Bifidobacteriaceae and a slightly lower transcriptional activity in the Proteobacteria family Alcaligenaceae was found in the inflamed mucosa compared to the non-inflamed mucosa. In addition, although not significant, the Prevotellaceae showed a higher transcriptional activity (Table S4).
Interestingly, when performing the analysis at the phylum level, a significantly higher abundance of Proteobacteria was found in the total microbiota in the inflamed tissues, while a significantly lower transcriptional activity was found in the active microbiota. Also a significantly lower transcriptional activity of Actinobacteria was found, and a similar but not significant difference in the abundance in the DNA dataset.
Taxonomic differences in total and active microbiota of non-inflamed mucosa between UC and symptomatic non-IBD controls
At the phylum level, a significantly lower abundance of Bacteroidetes was found in the total microbiota of the UC patients compared to the symptomatic non-IBD controls (Tables 3 and S5). A significantly higher abundance and transcriptional activity was found in Proteobacteria in UC patients compared to the symptomatic non-IBD controls in both the total and active microbiota, respectively.
Several bacterial families were found to have an increased or decreased fold change (FC) in both the total and active microbiota, but this was not significant at the 0.05 level (Table S6).
Paired comparisons of total and active microbiota
Our nucleic acid extraction method allowed for the direct comparison of total and active microbial community compositions in each biopsy. The transcriptional activity and the total abundances at the family level generally agreed and were highly positively correlated (Fig. 3). A paired Wilcoxon test of family-level abundances showed significant differences for most families, but the only family where the differences were of any meaningful magnitude was Ruminococcaceae, with a mean abundance of approximately 10 percentage points higher in the active microbiota (Table S7). These associations hold across disease status and inflammation status.
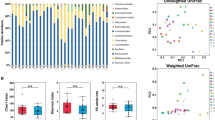
Comparing the microbial composition on the taxonomic family level in the active and total microbiota of each biopsy, collected from inflamed and non-inflamed tissues from UC patients and symptomatic non-IBD controls, revealed compositional similarities (Fig. 4). The same three families Ruminococcaceae, Lachnospiraceae and Bacteroidaceae dominated in both datasets. Some overall differences were found, such as a larger proportion of active Ruminococcaceae compared to total Ruminococcaceae in the symptomatic non-IBD controls, and a larger proportion of Verrucomicrobiaceae was seen in the total microbiota compared to the active microbiota.
In most families, there was a proportional relationship between the total and active microbiota (Table S7). However, some important exceptions were detected (Fig. 5a–d). Ruminococcaceae showed increased activity (Fig. 5a), whereas Bacteroidaceae and Verrucomicrobiaceae were more genetically abundant (Fig. 5b and d). The Enterobacteriaceae members, usually reported as important contributors to IBD-associated dysbiosis, were found to be less transcribed, indicating a partially dead or dormant population (Fig. 5c).
The functional dysbiosis in the mucosal tissue
The Piphillin tool returned a total of 295 KEGG pathway abundances (Tables S8–S11). The functional predictions based on DNA sequencing revealed the following results. Comparing the non-inflamed mucosal samples from UC patients with symptomatic non-IBD patients revealed significant differences among the metabolic pathways of chemical carcinogenesis (ko05201), drug metabolism – cytochrome P450 (ko00982), metabolism of xenobiotics by cytochrome P450 (ko00980) and carotenoid biosynthesis (ko00906), all of which were more present in total microbiota in the UC patients.
The functional analysis based on RNA (cDNA) sequencing did not show any significant differences (Tables S9 and S11).
Discussion
Our results support the use of RNA as a study tool in microbiota studies. The present study shows that the most abundant bacterial taxa, found in the inflamed and non-inflamed human mucosal tissue microbiota in UC patients, differed from the most active taxa. Using 16S rRNA [11] to reveal the potential active microbial taxa, we found a significant decrease in members of the phylum Proteobacteria, while the 16S rRNA gene revealed a significant increase in the total microbiota. Comparing UC patients to symptomatic non-IBD controls revealed more corresponding results between the 16S rRNA and 16S rRNA gene amplicon counts at the phylum level.
In a recent study by Heinsen and colleagues17, it was discovered that OTU abundance and potential activity in a bacterial community do not necessarily conform to each other. As such, we aimed to study the active and total microbial members of the colonic mucosal microbiota in treatment-naïve UC patients in search of bacterial taxa that may have a larger pathological role defined by their potential activity. In UC, the colonic mucosal tissue is inflamed, resulting in microbes with direct access to the epithelial layer. We believe that microbes thriving and having a potentially high metabolic activity in this environment will have a more important role in the disease than the total bacterial community occupying the same niche. The mucosal biopsy samples in the present study were subjected to an extensive nucleic acid extraction protocol optimized to extract and preserve DNA and RNA from both hard-to-lyse and more-prone-to-lyse bacterial taxa. This is an important step to obtain a balanced nucleotide purification and thus get a better representation of the microbes present in the samples. The DNA and RNA of each sample were isolated from the same single mucosal colonic biopsy, making it possible to perform a direct comparison between the active and total microbiota24,25. Other groups have purified DNA and RNA from single biopsies in their work on microbiota, but the purification methods have not been optimized for microbial nucleic acids17,26
Comparing the paired RNA and DNA samples revealed a modest heterogeneity in the alpha diversity (Fig. 1c). The alpha diversity analyses showed no significant differences when comparing the UC patients, the inflamed and non-inflamed tissue, and the symptomatic non-IBD control group in either dataset (Fig. 1a and b). An explanation for the lack of differences in alpha diversity and the lack of large differences in the active and total microbial community compositions when analysing both within UC patients and between UC patients and symptomatic non-IBD controls could be the result of mixing disease severity in our study group (Table S1). Walujkar27 and colleagues found in their study different shifts in the mucosal microbiota following UC severity stages. In the present study, we included patients suffering from different degrees of severity. Possible shifts in the diseased microbiota could be masked due to this mixing, hence shielding differences within the patient group. The control group of symptomatic non-IBD patients included patients suffering from IBD-like symptoms but not experiencing tissue inflammation. This patient group will most likely have a disturbed microbiota but probably to a lesser extent than the IBD patients. Thus, the analyses between the UC patients and the symptomatic non-IBD control group could also be affected by the mixing of disease severity in the former group.
Overall composition analyses with UniFrac showed that between-patient differences were more important than other potential sources of differentiation, such as diagnosis status and inflammation status (Fig. 2a and b). Pairwise differences between the active and total microbiota were also small (Fig. 2c).
To compare the mucosal microbiota between UC patients and symptomatic non-IBD controls, non-inflamed tissue biopsies were collected in the terminal ileum or the ascending colon if the terminal ileum could not be reached. This sampling area was chosen in an attempt to use data from all the UC patients, including the total colitis patients. As the sampling area of the non-inflamed tissue biopsies varied between the two parts of the intestine, analyses were performed to investigate whether the sampling could affect the further microbiota analysis. The analyses did not reveal any significant difference in either the 16S rRNA gene or the transcribed 16S rRNA counts between the non-inflamed tissues sampled from different gut locations within the patients (Fig. S1). Furthermore, our analysis revealed that even inflamed and non-inflamed tissue samples within the same patient were more similar with regard to both total and active microbiota than microbiota from the same location and with the same inflammation status in other patients (Fig. 2). These findings support other studies, indicating that there is greater variability between subjects, regardless of diagnosis, location in the gut and inflammation status, than the internal variability1.
Analyses of the inflamed and non-inflamed mucosal microbiota in UC patients revealed a greater abundance of Proteobacteria in the total microbial community composition of the inflamed tissues, with an FC of 1.331 (Table 2). This finding was contrasted by the active microbial community composition, which showed a lower transcriptional activity in the inflamed tissues, with an FC of 0.846. The Proteobacteria family level revealed a 2.455-fold higher abundance in Enterobacteriaceae of the total microbiota. The active microbiota was not found to have a significantly higher transcriptional activity, indicating that Enterobacteriaceae might not have an increased function or a more pathophysiological role in the inflamed tissue compared to the non-inflamed tissues of UC patients. The pairwise comparison within biopsies supported this finding, with less transcriptional activity of the bacterial family in the RNA dataset and higher abundance in the DNA dataset (Fig. 5c). In the active microbiota, a 0.808-fold lower transcriptional activity was found in the Proteobacteria family Alcaligenaceae. This result was not found in the total microbiota. The contrasting difference in abundance and activity in tissues with different inflammatory statuses may indicate that Proteobacteria have important pathophysiological functions in UC patients, being more abundant and active compared to symptomatic non-IBD controls, but do not have any increased function in the area of the diseased tissue. The discrepancies observed between the active and total microbiota could possibly be explained by the increased activity of human immune system in inflamed tissue.
The active microbiota showed a lower transcriptional activity in the phylum Actinobacteria, with a 0.563-fold decrease in the family Bifidobacteriaceae, suggesting a less prominent functional role of this bacterial family in the inflamed mucosa of UC patients. The total microbiota did not show this finding. The lower transcriptional activity could be related to the observed higher transcriptional activity of the aerotolerant anaerobic, saccharolytic, H2-producing, mucus-associated Prevotellaceae family, for which Prevotellaceae may possibly occupy the niche of Bifidobacteriaceae.
The paired biopsy analyses showed that at the family level, the active and total abundances generally agreed, but Ruminococcaceae had a larger transcriptional activity than its abundance for both the UC patients, regardless of inflammation status of the tissue samples, and the symptomatic non-IBD controls (Figs 4 and 5a).
Our results contrast with the study of Forbes and colleagues28, which found no significant variations in any phyla when comparing inflamed and non-inflamed mucosa within a group of UC patients. Reasons for this discrepancy could be differences in the protocol of DNA purification, including different enzymes used in the lysis step, as well as the use of different regions of the variable 16S rRNA gene for next generation sequencing (V5 versus V4, in our study).
The controls in our study were symptomatic non-IBD controls who had predefined symptoms of IBD but did not meet the diagnostic criteria. Elucidation of this group was not performed in the IBD-Character study, but the group probably consists of several patients who were classified as IBS-like. Proteobacteria has been found to have a higher abundance in UC patients compared to healthy individuals29, but the relation to IBS patients has not been convincing30. In the present study, the Proteobacteria phylum in the UC patient group, for both the total and active microbiota, revealed a considerably higher abundance and transcriptional activity compared to the symptomatic non-IBD control group (Table 3). Both the total and active microbiota revealed corresponding patterns with regard to significant differences in the fourth- to fifth-fold greater presence of the Proteobacteria family Enterobacteriaceae and the Firmicutes family Peptostreptococcaceae, indicating an important contribution of these two families to the diseased microbiota. Both bacterial families have been associated with disease and UC1,31,32,33, and they seem to have a greater function in the mucosal tissue of UC patients compared to the diseased non-IBD patients. Both bacterial families have been found to participate in xenobiotic/drug metabolism34, which is also a functional pathway found to be more present in UC patients in the present study. Enterobacteriaceae is a family of facultative anaerobes with an increased ability to adhere to the intestinal mucosa and have diverse energy sources. Peptostreptococcaceae, on the other hand, is a family of strict anaerobes with a fermentative metabolism. The latter family contains saccharolytic members and has been linked to colon cancer, but its function in the gut microbiota is still largely unclear33. The results of the functional analyses revealed a significantly greater abundance in the chemical carcinogenesis pathway in UC patients. UC is a well-known risk factor for colon cancer, and the results hint at a possible role for bacteria in modulating cancer risk35.
The Bacteroidetes family Prevotellaceae was found to be significantly less present and to have a lower transcriptional activity in the UC patients36. This is a dysbiosis pattern, seen when comparing UC patients to healthy controls, supporting the view of a less dysbiotic microflora in possible IBS patients30.
The pairwise analyses from non-IBD patients revealed, like the analyses of the UC patients, a high presence of the Clostridiales family Ruminococcaceae in the active microbiota and a low presence in the total microbiota. Schirmer and colleagues22 found that R. gnavus showed a large increase in transcriptional activity in Crohn’s disease and UC patients compared to non-IBD controls. In the present study, changes at the species level could not be detected, thus leaving this possible variation undetected.
This study has limitations. Strict ethical regulations in Norway hinder the collection of ileocolonic biopsies from healthy individuals. For this reason, our control group was comprised of symptomatic non-IBD controls who may have also suffered from a disturbed microbiota, although likely to a lesser extent than UC patients.
In conclusion, we find support for the use of RNA as a study tool in microbiota studies. Knowledge of the active members of microbiota in UC patients could aid in the understanding of the disease ethology. With the mucosal microbiota being in direct contact with the host, the active members could be of particular importance. In the present study, the most abundant microbiota members of the inflamed tissue in UC patients were not the most active. The active microbial community composition did not deviate from the total microbial community composition when comparing UC patients to non-IBD controls; thus, the choice of which microbiota to study depends on the questions raised.
Material and Methods
Study population and sample collection
Treatment-naïve UC patients were included at the time of disease diagnosis. The clinical data were collected at the time of inclusion (Table S1). Correspondingly, the patients referred for colonoscopy because of suspected IBD, but who had a normal endoscopic investigation and no elevated faecal calprotectin (Bühlmann Laboratories AG, Basel, Switzerland), were included as symptomatic controls. Celiac disease was excluded by negative serology. Suspected IBD was defined by the presence of predefined symptoms, including abdominal pain, diarrhoea and/or blood in the stool for more than 10 days37, with no evidence of enteric infection (Table S1). In the present study, we selected all the available symptomatic controls and an excess of UC patients, who were gender and age matched to the controls. All the patients were recruited from the Akershus University Hospital in Lørenskog, Norway and the study population was part of the EU study IBD-Character (EU ref no 305676) (http://www.ibdcharacter.eu/).
Prior to colonoscopy, the subjects went through bowel cleansing with Picoprep (Ferring Legemidler AS, Oslo, Norway), according to the manufacturer’s instructions. Colonic mucosal biopsies of 2–3 mm in size were collected from patients during colonoscopy at the time of inclusion. For the UC patients, biopsies from macroscopically inflamed tissue and non-inflamed tissue were collected from the same segment, if possible, in addition to a control sample from the ascending colon or the terminal ileum, if accessible. From the non-IBD control group, biopsies were collected from the ascending colon or the terminal ileum, when possible. All the biopsies were placed in Allprotect Tissue Reagent (Qiagen, Hilden, Germany) and stored according to the manufacturer’s instructions.
Ethics approval and consent to participate
The study, including data collection and analyses, was approved by the Regional Committee for Medical Research Ethics, South-Eastern Norway, reference no. REK sør-øst 2009/2015 and the representative of privacy protection at Akershus University Hospital, reference no. 13–126. All experiments were performed in accordance with and following the Declaration of Helsinki Principles. All methods were performed in accordance with the relevant guideline and regulations. All patients gave their written informed consent prior to colonoscopy and their inclusion in the IBD-Character study.
Nucleic acid purification
The microbial RNA and DNA were purified using the AllPrep DNA/RNA Mini kit (Qiagen) following a modified protocol published by our group previously24. In short, the biopsies were subjected to a thorough lysis and homogenization procedure, involving both enzymatic and mechanical lysis steps. This procedure ensured the protection of RNA by removing the lysate after each homogenization step. The DNA and RNA were eluted using 40 µl RNase-free water and were stored at -20 °C and -80 °C, respectively. The concentrations of the DNA and RNA samples were assessed using a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and using the OD260 for calculation. For the DNA, the quality was obtained using the OD260/OD280 and OD260/OD230 ratios for purity assessment of the samples. The RNA quality, indicated by the RNA integrity number (RIN), was assessed with an Agilent 2100 Bioanalyser, Agilent 2100 Expert software and Agilent RNA 6000 Nano Kit (Agilent Technologies Inc., Santa Clara, CA, USA).
cDNA synthesis
The cDNA was synthesised from 1 µg RNA from each sample using the AccuScript High Fidelity 1st Strand cDNA Synthesis Kit (Agilent Technologies Inc.) and random hexamers, according to the manufacturer’s instructions. Two random RNA samples were run in the absence of reverse transcriptase to assess the degree of contaminating genomic DNA.
16S rRNA and 16S rRNA gene amplicon sequencing
Two 16S amplicon libraries, consisting of a 16S rRNA gene and 16S rRNA transcript, were made according to a protocol published by Kozich and colleagues38,39. In short, the genomic DNA and cDNA were subjected to 16S PCR amplifications of the v4 gene region using a 500 ng and 250 ng template, respectively. The AccuPrime TM Pfx Supermix (Agilent Technologies Inc.) concentration, PCR primer combinations and thermal cycling conditions described in the MiSeq Wet Lab SOP39 were followed, with the exception of the annealing temperature and extension step, which were set to 50 and 68 °C, respectively. The two amplicon libraries were semi quantified using 1% agarose gel quantification and pooled according to band intensity into three pools of high, medium and low concentrations. From each library pool, 200 µl was purified by a 2% agarose gel using QIAquick Gel Extraction Kit (Qiagen) to reduce non-specific amplification products and ensure the removal of primer dimers. The purified products were quantified using the Kapa Library Quantification Kit (Universal) (Kapa Biosystems Inc., Wilmington, MA, USA) and finally pooled according to the concentrations and number of samples. The sequencing was performed using the Illumina MiSeq platform (Illumina Inc., San Diego, CA, USA) and the MiSeq reagent kit v/2 (500 cycles) according to the manufacturer’s instructions, with the addition of custom sequencing primers and index and 8% PhiX, as described in the MiSeq Wet Lab SOP39.
Data analysis
Blank samples were run through the wet lab procedure, from the nucleic acid purification step through the sequencing process, to detect possible reagent contamination. The minimum numbers of reads from each sample was set from the number of reads on the DNA and RNA blanks. MiSeq Reporter software (Illumina Inc.) was used for demultiplexing the reads and for fastQ file generation.
Sequence processing
The total sequence data were processed with mothur v.1.36.1 according to the MiSeq SOP38. The processed sequences were clustered into operational taxonomic units (OTUs) at 97% identity, and the SILVA reference database release 119 provided taxonomic information. The most abundant sequence in each OTU was picked as the representative sequence for the purpose of downstream sequence-based analyses, including UniFrac distances and functional analyses.
Diversity and abundance analysis
OTU-based analyses were done in R40. The phyloseq package was used for further data handling, diversity, and ordination analyses41. UniFrac distances and Principal Coordinates Analysis (PcoA) were used in the ordination analyses.
A differential abundance analysis was completed at both the family and phylum taxonomic levels. Lower taxonomic levels were not used due to known limitations in performing species-level identification from 16S rRNA sequencing42. For each analysis, the OTUs that shared taxonomic rank at each level were merged into one. The number of reads was modelled using a Negative Binomial regression model43. When we compared the inflamed and non-inflamed samples in UC patients, we included a random intercept to account for the paired samples. If several non-inflamed samples from the same patient were available, we compared the biopsies taken closest to the inflamed biopsy only. In the comparison between the symptomatic controls and the UC patients, we used the non-inflamed biopsy taken closest to the ileum.
OTUs with an average abundance of less than 0.01% reads and patients who had used antibiotics in the last 13 months were excluded from these analyses. The regression models also included a term for the sample gut location and a scaling factor to account for the total number of reads in the sample. The lme4 R package was used to fit the random effects models, while the MASS R package was used to fit the model without the random effects.
Differences in the diversity indices were investigated with linear regression models, with the same predictor variables as in the abundance analysis.
Paired analyses of the total and active microbial community composition from the same sample was completed using paired Wilcoxon tests on all the families.
Functional analyses
To investigate the functional capabilities of the total and active microbiota, we used Piphillin to infer KEGG (version 78.1, May 2016) pathway abundances23. Differential abundance analyses were performed in the same manner as in the OTU-based analyses.
Availability of Data and Material
The fastq files generated during the current study are available from the corresponding author on reasonable request.
References
Gevers, D. et al. The treatment-naïve microbiome in new-onset Crohn’s disease. Cell host & microbe 15, 382–392 (2014).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Manichanh, C., Borruel, N., Casellas, F. & Guarner, F. The gut microbiota in IBD. Nat. Rev. Gastroenterol. Hepatol 9, 599–608 (2012).
Kostic, A. D., Xavier, R. J. & Gevers, D. The Microbiome in Inflammatory Bowel Diseases: Current Status and the Future Ahead. Gastroenterology, https://doi.org/10.1053/j.gastro.2014.02.009 (2014).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13 (2012).
Martinez-Medina, M. & Garcia-Gil, L. J. Escherichia coli in chronic inflammatory bowel diseases: An update on adherent invasive Escherichia coli pathogenicity. World J. Gastrointest. Pathophysiol. 5, 213–227, https://doi.org/10.4291/wjgp.v5.i3.213 (2014).
Lopez-Siles, M. et al. Mucosa-Associated Faecalibacterium prausnitzii Phylotype Richness Is Reduced in Patients with Inflammatory Bowel Disease. Appl. Environ. Microbiol. 81, 7582–7592, https://doi.org/10.1128/aem.02006-15 (2015).
HMP-Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214, http://www.nature.com/nature/journal/v486/n7402/abs/nature11234.html#supplementary-information (2012).
Costello, E. K. et al. Bacterial Community Variation in Human Body Habitats Across Space and Time. Science 326, 1694–1697, https://doi.org/10.1126/science.1177486 (2009).
Moya, A. & Ferrer, M. Functional Redundancy-Induced Stability of Gut Microbiota Subjected to Disturbance. Trends Microbiol. 24, 402–413, https://doi.org/10.1016/j.tim.2016.02.002 (2016).
Nocker, A., Cheung, C. Y. & Camper, A. K. Comparison of propidium monoazide with ethidium monoazide for differentiation of live vs. dead bacteria by selective removal of DNA from dead cells. J. Microbiol. Methods 67, 310–320, https://doi.org/10.1016/j.mimet.2006.04.015 (2006).
Sanchez, M. C. et al. Quantitative real-time PCR combined with propidium monoazide for the selective quantification of viable periodontal pathogens in an in vitro subgingival biofilm model. J. Periodontal Res. 49, 20–28, https://doi.org/10.1111/jre.12073 (2014).
Emerson, J. B. et al. Schrodinger’s microbes: Tools for distinguishing the living from the dead in microbial ecosystems. Microbiome 5, 86, https://doi.org/10.1186/s40168-017-0285-3 (2017).
Chu, N. D., Smith, M. B., Perrotta, A. R., Kassam, Z. & Alm, E. J. Profiling Living Bacteria Informs Preparation of Fecal Microbiota Transplantations. PLoS One 12, e0170922, https://doi.org/10.1371/journal.pone.0170922 (2017).
Nocker, A., Richter-Heitmann, T., Montijn, R., Schuren, F. & Kort, R. Discrimination between live and dead cellsin bacterial communities from environmental water samples analyzed by 454 pyrosequencing. Int. Microbiol. 13, 59–65, https://doi.org/10.2436/20.1501.01.111 (2010).
Li, R. et al. Comparison of DNA-, PMA-, and RNA-based 16S rRNA Illumina sequencing for detection of live bacteria in water. Sci. Rep. 7, 5752, https://doi.org/10.1038/s41598-017-02516-3 (2017).
Heinsen, F.-A. et al. Dynamic changes of the luminal and mucosa-associated gut microbiota during and after antibiotic therapy with paromomycin. Gut Microbes 6, 243–254, https://doi.org/10.1080/19490976.2015.1062959 (2015).
Gosalbes, M. J. et al. Metatranscriptomic approach to analyze the functional human gut microbiota. PLoS One 6, e17447, https://doi.org/10.1371/journal.pone.0017447 (2011).
Blazewicz, S. J., Barnard, R. L., Daly, R. A. & Firestone, M. K. Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J 7, 2061–2068, https://doi.org/10.1038/ismej.2013.102 (2013).
Rehman, A. et al. Transcriptional activity of the dominant gut mucosal microbiota in chronic inflammatory bowel disease patients. J. Med. Microbiol. 59, 1114–1122, https://doi.org/10.1099/jmm.0.021170-0 (2010).
Peris-Bondia, F., Latorre, A., Artacho, A., Moya, A. & D’Auria, G. The active human gut microbiota differs from the total microbiota. PLoS One 6, e22448, https://doi.org/10.1371/journal.pone.0022448 (2011).
Schirmer, M. et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat Microbiol, https://doi.org/10.1038/s41564-017-0089-z (2018).
Iwai, S. et al. Piphillin: Improved Prediction of Metagenomic Content by Direct Inference from Human Microbiomes. PLoS One 11, e0166104, https://doi.org/10.1371/journal.pone.0166104 (2016).
Moen, A. E. et al. Simultaneous purification of DNA and RNA from microbiota in a single colonic mucosal biopsy. BMC Res. Notes 9, 328, https://doi.org/10.1186/s13104-016-2110-7 (2016).
Valles-Colomer, M. et al. Meta-omics in Inflammatory Bowel Disease Research: Applications, Challenges, and Guidelines. J Crohns Colitis 10, 735–746, https://doi.org/10.1093/ecco-jcc/jjw024 (2016).
Rehman, A. et al. Geographical patterns of the standing and active human gut microbiome in health and IBD. Gut 65, 238–248, https://doi.org/10.1136/gutjnl-2014-308341 (2016).
Walujkar, S. A. et al. Characterization of bacterial community shift in human Ulcerative Colitis patients revealed by Illumina based 16S rRNA gene amplicon sequencing. Gut Pathog. 6, 22–22, https://doi.org/10.1186/1757-4749-6-22 (2014).
Forbes, J. D., Van Domselaar, G. & Bernstein, C. N. Microbiome Survey of the Inflamed and Noninflamed Gut at Different Compartments Within the Gastrointestinal Tract of Inflammatory Bowel Disease Patients. Inflamm. Bowel Dis. 22, 817–825, https://doi.org/10.1097/MIB.0000000000000684 (2016).
Mukhopadhya, I., Hansen, R., El-Omar, E. M. & Hold, G. L. IBD—what role do Proteobacteria play? Nature Reviews Gastroenterology &Amp; Hepatology 9, 219, https://doi.org/10.1038/nrgastro.2012.14 (2012).
Casen, C. et al. Deviations in human gut microbiota: a novel diagnostic test for determining dysbiosis in patients with IBS or IBD. Aliment. Pharmacol. Ther. 42, 71–83, https://doi.org/10.1111/apt.13236 (2015).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 104, 13780–13785, https://doi.org/10.1073/pnas.0706625104 (2007).
Lavelle, A. et al. Spatial variation of the colonic microbiota in patients with ulcerative colitis and control volunteers. Gut 64, 1553–1561, https://doi.org/10.1136/gutjnl-2014-307873 (2015).
Rajilic-Stojanovic, M. & de Vos, W. M. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol. Rev. 38, 996–1047, https://doi.org/10.1111/1574-6976.12075 (2014).
Das, A., Srinivasan, M., Ghosh, T. S. & Mande, S. S. Xenobiotic Metabolism and Gut Microbiomes. PLoS One 11, e0163099, https://doi.org/10.1371/journal.pone.0163099 (2016).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766, https://doi.org/10.15252/msb.20145645 (2014).
Lepage, P. et al. Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology 141, 227–236, https://doi.org/10.1053/j.gastro.2011.04.011 (2011).
Lennard-Jones, J. E. Classification of inflammatory bowel disease. Scand. J. Gastroenterol. Suppl. 170, 2–6; discussion 16–19 (1989).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Appl. Environ. Microbiol. 79, 5112–5120, https://doi.org/10.1128/AEM.01043-1-20 (2013).
Kozich, J., Schloss, P., Baxter, N., Jenior, M. & Koumpouras, C. 16S rRNA Sequencing with the Illumina MiSeq: Library Generation, QC, & Sequencing, https://github.com/SchlossLab/MiSeq_WetLab_SOP/blob/master/MiSeq_WetLab_SOP_v4.md (2013).
R: A language and environment for statistical computing. R Foundation for Statistical Computing (Vienna, Austria, 2016).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8, e61217, https://doi.org/10.1371/journal.pone.0061217 (2013).
Schloss, P. D. The Effects of Alignment Quality, Distance Calculation Method, Sequence Filtering, and Region on the Analysis of 16S rRNA Gene-Based Studies. PLoS Comput. Biol 6, e1000844, https://doi.org/10.1371/journal.pcbi.1000844 (2010).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol 10, e1003531, https://doi.org/10.1371/journal.pcbi.1003531 (2014).
Acknowledgements
The project was funded by the EU study IBD-Character (EU ref no 305676). We want to thank Engineer Sumana Kalyanasundaram, Institute of Informatics, University of Oslo, Norway for the bioinformatical help during the startup phase of the project. European IBD Character Consortium, FP7, Grant Agreement Number: 305676.
Author information
Authors and Affiliations
Author notes
A comprehensive list of consortium members appears at the end of the paper
Consortia
Contributions
A.E.F.M. and T.M.T. conceived of and designed the study and performed the laboratory work. S.V. recruited subjects and collected the biopsy samples. J.C.M. and T.M.T. performed the bioinformatical and biostatistical parts of the project, and together with A.E.F.M., M.V., P.R. and J.J., interpreted the results. A.E.F.M., T.M.T. and J.C.L. drafted the article. J.J., M.V. and P.R. performed a critical revision of the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moen, A.E.F., Lindstrøm, J.C., Tannæs, T.M. et al. The prevalence and transcriptional activity of the mucosal microbiota of ulcerative colitis patients. Sci Rep 8, 17278 (2018). https://doi.org/10.1038/s41598-018-35243-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35243-4
Keywords
This article is cited by
-
Constituents of stable commensal microbiota imply diverse colonic epithelial cell reactivity in patients with ulcerative colitis
Gut Pathogens (2024)
-
The interplay between alterations in esophageal microbiota associated with Th17 immune response and impaired LC20 phosphorylation in achalasia
Journal of Gastroenterology (2024)
-
Mucosa-associated gut microbiota reflects clinical course of ulcerative colitis
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.