Abstract
Milk production is highly dependent on the extensive development of the mammary epithelium, which occurs during puberty. It is therefore essential to distinguish the epithelial cells committed to development from the related epithelial hierarchy. Using cell phenotyping and sorting, we highlighted four cell sub-populations within the bovine mammary gland at puberty. The CD49fhighCD24neg cells expressing CD10, KRT14, vimentin and PROCR corresponded to cells committed to the basal lineage. The CD49flow sub-population contained two cell subsets (CD49flowCD24neg and CD49flowCD24pos). Both subsets expressed hormone receptors including ER, PR and PRLR, as well as ALDH1 activity but only the CD49flowCD24pos subset expressed ELF5. These data indicated that the CD49flow sub-population is mainly composed of cells displaying a luminal phenotype and that this population comprises two luminal cell subsets, namely the CD24neg and CD24pos cells, likely committed to ductal and alveolar lineage, respectively. The putative mammary stem cell (MaSC) fraction was recovered in the CD49fhighCD24pos sub-population which were shown to form mammospheres in vitro. These cells differentially expressed CD10, KRT14 and KRT7, suggesting the existence of several putative MaSC sub-fractions. In-depth characterization of these epithelial sub-populations provides new insights into the bovine mammary epithelial cell lineage and suggests a common developmental lineage in mammals.
Similar content being viewed by others
Introduction
The mammary gland undergoes dynamic morphological changes over the lifetime of female mammals. At birth, bovine mammary parenchyma consists of a rudimentary duct network connected to a small cisternal cavity. At the onset of puberty, the mammary rudiment develops and starts to expand into the stroma upon stimulation by the ovarian steroid hormones, including estradiol and progesterone, and by growth factors1. Ductal elongation occurs through the growth, development, and subsequent extension of terminal ductal lobular units (TDLU) in a process referred to as branching morphogenesis. Bovine mammary TDLUs initially consist of solid cords of epithelial cells that penetrate into the stroma. As these cords extend into the mammary fat pad, lateral outgrowths emerge. This parenchymal development continues through puberty, until the mammary fat pad becomes filled. In addition, during gestation, the tissue continues its differentiation with the formation of lobulo-alveolar structures and the maturation of TDLUs in response to circulating hormones, notably prolactin. At the end of its development, the mammary epithelium has the appearance of an elaborate tree of ducts and alveoli. After parturition, the alveolar epithelium starts to be fully functional, with mammary epithelial cells secreting milk proteins into the lumen of the alveoli for lactation2.
The ability of the mammary gland to undergo many cycles of lactation, with their stages of tissue proliferation and involution, suggests that the epithelial compartment contains resident cells capable of generating the entire epithelial architecture. Evidence for the existence of mammary stem cells (MaSCs) has been primarily derived from transplantation studies with murine mammary tissues. These studies revealed that the ductal architecture could be regenerated in vivo when isolated parenchymal explants were transplanted into cleared mammary fat pads3,4,5. More recent assays showed that an entire and functional mammary gland can be reconstituted from the transplantation of the progeny of a single “stem-like” cell6,7. Since these pioneering demonstrations, many studies in murine and human species have focused on identifying and isolating MaSC populations in order to establish the hierarchical cell organization and the molecular players in the regulation of the epithelium8,9. The epithelial hierarchy can be described as a pyramidal setup of the epithelial cell populations with stem cells at the apex and differentiated mature cells at the base of the pyramid. Between these two cell populations are the multiple progenitors that originate from the division and activation of stem cells and that progressively differentiate into mature cell lineages. Of note, the mammary structures are described as being composed of two major lineages: the luminal and basal cells, the latter including the myoepithelial cells. Luminal and basal cells can be distinguished by either their location in the epithelial structure or their protein expression profiles. Cells of these two lineages are considered immature during development as compared to the differentiated (mature) cells that constitute the functional secretory tissue.
In contrast, in bovines, only a few groups have attempted to elucidate the epithelial hierarchy via the identification of progenitor/stem cell populations10,11. We recently participated in this research effort by providing original data on the mammary epithelial hierarchy committed to lactation during a lactation cycle in bovines12. In this study, we used flow cytometry analysis and fluorescence activated cell sorting based on the expression of classic markers previously identified in the murine, human and bovine species. These markers are cell surface proteins, including the cluster of differentiation (CD) 24 (heat-stable antigen), CD29 (ß1-integrin) or CD49f (α6-integrin), and CD1013,14. These approaches led us to isolate putative populations of MaSCs, a prerequisite for further study of these target cell populations.
Research on MaSC biology in dairy mammals is important and relates to their potential use to improve animal robustness through the enhancement of lactation efficiency and infection resistance. A better understanding of the epithelial hierarchy at each developmental stage is therefore a prerequisite for the optimization of lactation in cows. Until now, literature describing the epithelial cell populations at key developmental stages (after puberty) and the regulators governing the bovine epithelial hierarchy has been scant. In this context, our study aims to further characterize the cells that make up the epithelial lineage at the branching morphogenesis stage in order to provide new insights into the epithelial hierarchy.
Results
Discrimination between cell sub-populations within the mammary epithelium of pubertal cows using the cell surface markers CD49f, CD24 and CD10
Since puberty is a key period of mammary gland development during which the different epithelial lineages, basal/myoepithelial and luminal cells, are committed to the process of branching morphogenesis and are identifiable, we used mammary gland samples from pubertal cows for our study.
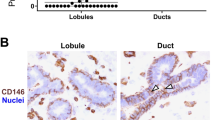
In agreement with this, tissue staining with hematoxylin and eosin showed numerous neo-formed ductal and alveolar structures constituting an epithelium that largely formed the mammary parenchyma (Fig. S1). To identify the cell sub-populations of the epithelial lineages acting in the building of this parenchyma in the most exhaustive way possible, we focused our analysis on three cell surface markers that are well known to be specific for mammary epithelial cells: CD49f, CD24 and CD10. To validate our approach, we first analyzed the in situ localization of the cells expressing these markers by immunofluorescence. As shown in Fig. 1, cells of the ductal trees at the origin of future TDLUs were clearly stained by anti-CD49f antibodies (Fig. 1, left panels). The outer cells of these epithelial structures formed a monolayer and were strongly stained at their basal side, whereas the inner cells were weakly stained. In contrast, CD24 was expressed apically by epithelial cells located in the lumen of ductal structures in development (Fig. 1, middle panels). As to CD10, which has been described as a cell surface marker of basal cells, it was clearly expressed by cells surrounding the developing duct structures (Fig. 1, right panels). In this case, stained cells were exclusively localized to the outer epithelium layer, or sometimes appeared in small clusters (see the little structure at the top right of the image; Fig. 1, right panels). These immuno-histological results having confirmed the relevance of using these markers, we decided to evaluate the proportion of each cell sub-population of the mammary tissue expressing them by flow cytometry.
The cell surface markers CD49f, CD24 and CD10 are located in the luminal and basal cells within the ductal mammary epithelium of cows at puberty. Cryo- (CD49f and CD24) and paraffin sections (CD10) from the mammary tissue of pubertal cows were processed for immunofluorescence for the indicated antigens. Nuclei were counterstained with Hoechst 33342. Note that the CD24 images were obtained with an Apotome microscope. The basal membrane of the outer cell layer of the epithelium was highly stained for CD49f whereas luminal cells were weakly stained (left panels, green). CD24-positive cells were located within mammary epithelial ductal structures (middle panels, green). Antibodies against CD10 nicely stained the outer cells of the developing ductal structures (right panels, red). Images are representative of three cows. Scale bars, 100 µm.
As shown in the cytometric profile of CD49f expression (Fig. S2a, upper plot), 62% (±1.8%) of total single cells prepared from the mammary tissue of pubertal cows were CD49fpos cells. Moreover, it was possible to distinguish two distinct sub-populations within these cells: the CD49flow (38%) and CD49fhigh (24%) sub-populations. To further identify the cell types that compose the mammary gland tissue of the pubertal cow, total single cells were sorted based on CD49f expression. A set of proteins known to be specifically expressed in the epithelial lineage was then quantified in both negative and positive cell sub-populations by Western blotting (Fig. S3). What was highly noticeable was the higher expression level of all epithelial lineage protein markers in the CD49fpos cells as compared to the CD49fneg cells (Fig. S3a,b). Only the CD49fpos cells expressed the epithelial cadherin (CDH1, Fig. S3a, left graph), the basal marker keratin (KRT) 14 and the myoepithelial marker alpha-smooth muscle actin (αSMA), as well as the luminal KRT7, KRT19 and KRT18 (see also Fig. S4 for the in situ lineage-specific localization of KRT). Finally, these cells significantly overexpressed the basal marker CD10 (Fig. S3b). Altogether, these data strongly suggested that CD49f cell sorting allowed the recovery of epithelial cells of both basal and luminal origins.
When cells were analyzed for CD24 expression, a unique heterogeneous population of CD24pos cells was observed (Fig. S2a, middle plot). It accounted for 32% (±9.8%) of total single mammary cells. Western blotting showed that the epithelial marker CDH1 was expressed in both CD24neg and CD24pos cells (Fig. S3b) but was much more abundant in the latter cells. As a whole, the CD24neg cells preferentially expressed the basal markers, i.e., CD10, αSMA and KRT14, whereas the luminal markers were more highly expressed in the CD24pos cells. Indeed, both CD24neg and CD24pos cells expressed KRT7, KRT18 and KRT19, but all the luminal keratins were expressed at significantly higher levels in the CD24pos population (Fig. S3a, middle graph and Fig. S3b). We concluded that CD24 is a marker that allows the distinction of epithelial sub-populations within the basal and the luminal lineage.
Finally, we identified two cell sub-populations expressing CD10 (CD10low and CD10high), the sum of which accounted for 41% (±7.7%) of total mammary cells (Fig. S2a, bottom plot). KRT14 was only present in the CD10pos cells (Fig. S3a, right graph and Fig. S3b). In addition, αSMA was almost 6-fold more abundant in the CD10pos population than in the CD10neg population (15.2% ± 4% vs 3.1% ± 0.5%). Interestingly, the luminal KRT19, KRT18 and KRT7 were expressed in both the CD10neg and CD10pos cell sub-populations with no significant difference, except for KRT7 which was expressed at 6-fold higher level in the CD10pos cells (3.79% ± 0.97% vs. 0,55% ± 0.18%). In summary, our data confirm that CD10 expression is characteristic of basal cells, making it a pertinent marker to discriminate the basal lineage from the luminal lineage.
Determination of the cell sub-populations involved in mammary gland development at puberty
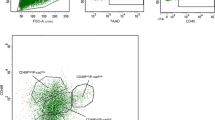
To further delineate the different cell sub-populations involved in the development of the mammary gland in pubertal cows, we analyzed all combinations of cell co-staining with CD49f, CD24 and CD10 by flow cytometry. Co-staining for CD49f and CD24 revealed four distinct positive cell sub-populations in addition to the double-negative population (Fig. 2a). The majority of the cells was CD49fposCD24neg (42% ± 0.8% of total cells) and equally distributed into CD49flow and CD49fhigh cells (see Table 1 for the relative proportion of each sub-populations). The CD49fposCD24pos sub-populations represented 20% (±3.7%) of total single cells with a large proportion of CD49flowCD24pos cells (Fig. 2a and Table 1). Interestingly, each of these sub-populations (CD49flowCD24pos, CD49flowCD24neg and CD49fhighCD24neg) approximately accounted for one third of the total CD49fpos cells (see Fig. S5). Finally, we found that only 2% (±0.1) of total single cells were CD49fnegCD24pos. Co-staining for CD49f and CD10 revealed five distinct sub-populations (Fig. S2b, middle plot, Table 1). Finally, co-staining for CD10 and CD24 revealed heterogeneous sub-populations (Fig. S2B, bottom plot, Table 1). Altogether, these data highlighted the multiple cell sub-populations present within the mammary tissue during pubertal development.
Sub-populations of epithelial cells exhibit distinct lineage types in the developing bovine mammary gland. (a) Cells dissociated from pubertal bovine mammary tissue were co-stained with anti-CD49f -FITC (CD49f) and anti-CD24-APC (CD24) antibodies and analyzed by flow cytometry. The gating for each control immunoglobulin isotype is indicated by the solid lines. (b) Cells expressing low or high levels of CD49f and/or CD24 were subjected to flow cytometry analysis for either CD10 expression or ALDH1 activity. The mean percentages of cells (±SEM) expressing CD10 or showing ALDH1 activity in each sub-population from three independent experiments (3 cows) are summarized in a table. Abbreviations: ALDH1, Aldehyde dehydrogenase 1.
Characterization of the cell sub-populations composing the mammary epithelial hierarchy
Since mammary stem cells and progenitors were reported to belong to a subset of CD49fposCD24pos cells, we decided to further investigate the phenotype of this cell sub-population. We found that the CD49flowCD24neg cells were predominantly negative for CD10 whereas almost all CD49fhigh cells expressed CD10 at high level (Fig. 2b). Within the CD49flowCD24pos sub-population, 75% of the cells were positive for CD10. To confirm the double lineage of this sub-population, we analyzed sorted cells by immunofluorescence and found that the CD49flowCD24pos cells co-expressed CD10 and KRT8, a luminal marker protein (Fig. 3a). Similarly, we evaluated the activity of aldehyde dehydrogenase 1 (ALDH1) in the aforementioned CD49fpos sub-populations. Indeed, ALDH1 activity has been previously identified as a marker of luminal cells and it has been shown to distinguish progenitor from mature mammary luminal cells in different species15. We found that 78 to 87% of the CD49flow cells, namely the CD49flowCD24neg and the CD49flowCD24pos cells, exhibited ALDH1 activity, as well as 68% of the CD49fhigh CD24pos cells (Fig. 2b). It is therefore reasonable to assume that these three sub-populations belong or are related to the luminal lineage.
Analysis of luminal and basal marker protein expression highlights heterogeneity of the MaSC fraction. Cells dissociated from pubertal bovine mammary tissue were sorted according to the level of expression of both CD49f and CD24 and analyzed by immunofluorescence. (a) CD49flow CD24pos cells were co-stained with anti-CD10 and -KRT8 antibodies. (b) CD49fhigh CD24pos cells were co-stained with anti-keratin (KRT) 14 and KRT7 antibodies. Nuclei were counterstained with Hoechst 33342. Scale bars, 100 µm.
We next investigated the expression of target genes by RT-qPCR (Fig. 4). Hormone receptivity of the pubertal mammary tissue was also assessed beforehand by immunofluorescence (Fig. S6a). These analysis revealed the expression of the progesterone (PR) and estradiol (ERα) receptors by the epithelial cells (22% ± 2.4% and 11% ± 1% of total cells for the PR and ERα-stained cells, respectively) (Fig. S6b). We found that the genes known to be expressed by stromal cells, namely vimentin, ALDH1 and the Protein C receptor (PROCR) were significantly more expressed in the CD49fnegCD24neg sub-population. In contrast, this sub-population under-expressed genes of the KRT family and the differentiation/receptivity markers. On the other hand, the two CD49flow sub-populations expressed higher levels of KRT19, KRT18 and KRT7 as compared to the CD49fneg sub-population, confirming their luminal origin. However, the CD49flowCD24neg and CD49flowCD24pos sub-populations presented differences in KRT expression (2 and 2.6-fold more abundant for KRT19 and KRT18, respectively, in the CD49flowCD24neg sub-population than in the CD49flowCD24pos sub-population) and in their hormonal receptivity (2.5-fold more abundant for PR and prolactin receptor (PRLR) in the CD49flowCD24neg sub-population than in the CD49flowCD24pos sub-population). The CD49flowCD24neg sub-population was characterized by expression of the three luminal keratins and of both PR and PRLR. The CD49flowCD24pos sub-population especially expressed the luminal KRT7, the stemness markers ALDH1 and the receptivity markers PR and E74-like factor 5 (ELF5). As for the CD49fhigh sub-populations, they significantly expressed KRT14, confirming their basal origin. Finally, the CD49fhighCD24neg sub-population was characterized by a moderate abundance of the vimentin and PROCR genes whereas the CD49fhighCD24pos sub-population expressed the KRT7, ALDH1 and ELF5 genes. Cells of this latest sub-population were also found to express the basal markers CD10 and KRT14, inferring some concerns about its homogeneity. In line with this, immunofluorescence analysis of sorted cells confirmed that the CD49fhighCD24pos sub-population contained both cells expressing only KRT14, and cells co-expressing KRT14 and KRT7 (Fig. 3b). In conclusion, each CD49f CD24 sub-population exhibited a unique phenotype and molecular signature which allowed them to be catalogued in a lineage type.
Gene expression levels in epithelial cell sub-populations. Cells dissociated from pubertal bovine mammary tissue were co-stained with anti-CD49f -FITC (CD49f) and anti-CD24-APC (CD24) antibodies and five sub-populations were sorted. The level of expression of the indicated genes was measured in each sub-population by RT-qPCR. Data are expressed as the mean of Delta Ct calculation ± SEM from three independent experiments (3 cows). Different letters (a–d) indicate significant differences. Abbreviations: ALDH1, Aldehyde dehydrogenase 1; ELF5, E74-like factor 5; ERα, Estradiol receptor alpha; KRT, keratin; NOTCH1, Notch homolog 1, translocation-associated; PR, Progesterone receptor; PRLR, prolactin receptor; PROCR, Protein C receptor.
It was shown that stem cells survive, proliferate and form spheres of cells (called mammospheres) when grown in three-D culture in the presence of Matrigel. In fact, the formation of mammospheres now accounts for a functional assay of stem cells. We therefore tested the ability of the sorted epithelial cell sub-populations to form spheres (Fig. 5). After 7 days of culture in Matrigel, both the CD49flowCD24neg and CD49flowCD24pos sub-populations failed to form mammospheres. In striking contrast, the CD49fhighCD24pos and CD49fhighCD24neg cells formed well-shaped mammospheres of 186 µm (±12 µm) and 173 µm (±8 µm) diameters, respectively.
CD49fhigh sub-populations displayed functional properties to form mammospheres. Cells dissociated from pubertal bovine mammary tissue were sorted according to the level of expression of both CD49f and CD24. The resulting epithelial cell sub-populations were cultured in Matrigel during 7 days to assess the formation of mammospheres. Images are representative of three independent experiments (3 cows). Scale bars, 200 µm.
Discussion
After puberty, each estrous cycle is accompanied by periods of enhanced cell proliferation and differentiation in the mammary gland until the fat pad is filled with parenchymal tissue. Therefore, this post-pubertal stage is a key period to search for and identify the most epithelial cell categories, including progenitor cells. Based on the literature, the analysis of the expression of the specific cell surface markers CD49f, CD24 and CD10 in bovine mammary epithelial cells at puberty by flow cytometry allowed for the identification and isolation of prospective key cell sub-populations. Of course, it was of the utmost interest to further analyze the molecular signatures of these sub-populations to improve our knowledge of the mammary epithelial cell lineage.
We first found that the CD49fhighCD24neg sub-population expressed KRT14, a well-known marker of the basal lineage classically associated with basal/myoepithelial cells16,17. This sub-population also substantially expressed CD10, another marker of basal cells17. Finally, immunohistological observations revealed that the cells of the outer epithelium layer were strongly stained at their basal side by anti-CD49f antibodies. In summary, these data indicate that the CD49fhighCD24neg sub-population is from the basal lineage. This is in agreement with a previous bovine study on the characterization of the epithelial cells present in the mammary gland a few months after birth10. More recently, this group reported that, at early developmental stages, the basal cells were CD49fposCD24neg, and specified that their phenotype was CD49fhighCD24neg18. In the present study, we further characterized this basal CD49fhighCD24neg sub-population by, notably, studying the expression of the vimentin and PROCR genes. Indeed, vimentin filaments are expressed, inter alia, in the basal epithelial cell population of the mammary gland19 and it has recently been demonstrated that vimentin deficiency in vimentin KO mice affects mammary ductal development by altering progenitor cell activity19. This suggested a regulatory role of vimentin in the basal MaSC/progenitor cell population. The fact that the bovine CD49fhighCD24neg cell analysed here expressed vimentin therefore suggest that they might be progenitor cells. This is also supported by the observation that these cells also expressed high levels of PROCR. Indeed, this gene which was found to be relatively abundant in the basal cells of murine mammary epithelium was suggested to be a marker of mammary stem cells20. This possibility was previously envisioned in a model of human breast cancers in which the receptor was one of the molecular markers for stem/progenitor-like populations21. However, our current knowledge of the bovine model does not allow us to firmly conclude whether the CD49fhighCD24neg cells account for the basal progenitor cells, but only that they are most likely committed to basal/myoepithelial lineage.
We observed by immunofluorescence that the cells localized into the inner epithelium layer expressed low levels of CD49f. Also, our cytometric profiles showed that two mammary epithelial cell sub-populations expressed low levels of CD49f, namely the CD49flowCD24neg and CD49flowCD24pos cell sub-populations. This is in agreement with the aforementioned study in bovines and with studies in mouse, in which the luminal population was reported as being CD49flow 18,22,23. In addition to these data, we showed by Western blotting that KRT19, KRT18 and KRT7 were expressed by the CD49fpos cells, including the CD49flow cells. The abundance of these luminal KRTs was also demonstrated at the mRNA level in the two CD49flow sub-populations. These data confirm that the CD49flow sub-populations belong to the luminal lineage.
Many studies have reported that mammary development at puberty is triggered by the main steroid hormones estradiol and progesterone (for review see2). Indeed, experiments with PR-deficient mice demonstrated that, at puberty, progesterone is not essential for ductal elongation but is critical in inducing side-branching24. This observation suggests that progesterone, independent from estradiol, could intervene late in branching morphogenesis to promote side branching and later in the formation of lobulo-alveolar structures25. Moreover, it has been found that a large number of luminal cells are PR-positive in adult virgin mice at an advanced stage of puberty26. We also observed that PR staining is restricted to luminal cells. As to the key role of prolactin, it has been found that deletion of the PRLR in mice resulted in defects in side branching and further alveolar formation27. Conversely, overexpression of prolactin in mice has been shown to increase lateral ductal budding and to increase epithelial progenitor sub-populations23. Here, we found that both the CD49flowCD24neg and CD49flowCD24pos sub-populations expressed the hormone receptors (ER, PR and PRLR). Additionally, we observed that the CD49flowCD24pos sub-population expressed ELF5 in contrast to the CD49flowCD24neg sub-population. Interestingly, the transcription factor ELF5 is known to orient the fate of luminal cells during alveogenesis28. In the murine model, the CD49flowCD24neg cell sub-population was described to represent the ductal cell population29. Altogether, our data indicated that the CD49flow population is mainly composed of cells displaying a luminal phenotype and that this population comprises two major luminal subsets, namely CD24neg and CD24pos, likely committed to ductal and alveolar lineage, respectively.
Furthermore, we found that both CD49flowCD24neg and CD49flowCD24pos sub-populations exhibited ALDH1 activity, a feature that characterizes the differentiation status of the luminal cells. Indeed, a previous study in human mammary gland demonstrated that ALDH1 activity was upregulated at the transition of progenitor cells into the luminal lineage15. ALDH1 activity has also been used in the bovine model to define luminal-restricted progenitors30 and in the mouse model to distinguish the relatively undifferentiated luminal progenitors from the differentiated ones31,32. From these reports and our data, we could hypothesise that the two CD49flow sub-populations are luminal progenitors. However, further investigations are required to confirm the progenitor commitment of these sub-populations.
In many species, whether human, murine or bovine, the MaSC population has been described as being CD49fhighCD24pos9,33,34. Here, this cell sub-population represented 3.4% of total mammary cells. This relatively small percentage was consistent with what is usually reported for the MaSC-enriched fraction in the literature (5% of total mammary cells in mice and 2.43% in post-pubertal bovines35). Recently, we showed that the proportion of CD49fhighCD24pos cells in the bovine lactating mammary gland range from 0.7% to 3.3%12. In the present study, we found that this sub-population also expressed the two basal markers CD10 and KRT14. This was consistent with the observation that MaSCs appeared localized to the basal compartment, sharing characteristics with the surrounding basal cells36,37,38. As observed previously8 and confirmed here, the CD49fhighCD24pos sub-population formed mammospheres when cultured in the presence of Matrigel. The above considerations suggest that the CD49fhighCD24pos sub-population is the putative MaSC fraction. Since this fraction exhibited heterogeneous phenotypes with variable ALDH1 activity, KRT7 and KRT14 expression, one can conclude that the CD49fhighCD24pos sub-population contains several MaSC sub-fractions. This notion has recently been raised in an elegant study of the murine MaSCs39 in which the dynamics of branching morphogenesis were monitored by highlighting the behaviour of the different lineage-committed MaSCs using a “confetti” cell strategy. Indeed, it was concluded that a pool of MaSCs is engaged in the development of the tissue whereas another stays quiescent. By analogy, we can hypothesize that the putative MaSC sub-populations exhibiting ALDH1 activity and KRT14/KRT7 expression represent the lineage-restricted “activated” MaSC engaged in epithelial development.
Even if it could be confusing to compare mammary epithelial cell lineages between species, notably because investigators regularly use different cell markers40, the mammary epithelial cell lineages we described here shares many common characteristics with that proposed for the murine model9. In agreement with this, our data lead us to hypothesize that the putative bovine MaSC fraction is most likely the CD49fhighCD24pos sub-population, as proposed to date in several reports34,40,41. Refining our knowledge on MaSC in dairy animals is of utmost importance in order to properly manipulate MaSC differentiation potential and/or expansion. Indeed, it would provide means to increase the robustness traits of dairy cows by increasing the number of fully functional mammary secretory cells and by improving the capacity of replacement of senescent or damaged cells during lactation.
Materials and Methods
All the animal procedures were discussed and approved by the CNREEA No. 07 (Local Ethics Committee in Animal Experiment of Rennes) in compliance with French regulations (Decree No. 2013–118, February 07, 2013).
Animals
The Holstein cows (bos taurus) used in this study were housed at the experimental farm of Méjusseaume INRA-Rennes (France). The pubertal cows were sacrificed at 17 months of age at the slaughterhouse of Gallais Viande (Montauban-de-Bretagne, France) following standard commercial practices. The mammary glands were collected at the time of slaughter and immediately transported on ice to the laboratory to be sampled for further analyses.
Mammary tissue sampling
Total parenchyma of the mammary gland was dissected and sampled. Samples destined for tissue dissociation were manually cut into small explants (≈1 mm3), suspended in 90% fetal bovine serum (10270-106; Gibco Invitrogen Saint Aubin, France)/ 10% dimethyl sulfoxide (DMSO, D2650, Sigma-Aldrich, Saint-Quentin Fallavier, France), and stored at −150 °C. For immunohistological analysis, tissue pieces (≈5 mm3) were fixed in 4% paraformaldehyde (FOR007OAF59001, VWR, Fontenay-sous-Bois, France) and were either mounted in OCT embedding compound (00411243, Labonord, Templemars, France) and frozen at −80 °C, or dehydrated in ethanol and embedded in paraffin.
Flow cytometry and cell sorting
Mammary tissue fragments were thawed and enzymatically dissociated as previously described12 to obtain a single cell suspension. Dissociated cells were incubated with the relevant antibodies for 20 min at 4 °C, washed and re-suspended in MACS buffer (130-091-222, Miltenyi Biotec, Paris, France) with 2% bovine serum albumin (130-091-376; Miltenyi Biotec) for flow cytometry analysis or cell sorting.
Flow cytometry was performed using a MACSQuant Analyzer 10 cytometer (Miltenyi Biotec). The controls and gating strategy used in the present study have been previously detailed12. Note that isotype control antibodies were used as negative controls in the flow cytometry experiment. Data were analyzed using MACSQuantify analysis software (Miltenyi Biotec) and results expressed in percentage of cells out of 20,000 events.
ALDH1 activity was measured in 500.000 cells with the Aldefluor kit (01700, Stem cell technologies, Grenoble, France) according to the manufacturer’s recommendations. Cells were then centrifuged at 250 G, re-suspended in Aldefluor assay buffer and labeled with antibodies against CD49f and CD24.
For cell sorting, cells were incubated with the relevant antibodies for 20 min at 4 °C in the dark. Single live cells were gated by DAPI exclusion and sorted on a BD FACS ARIA II flow cytometer (BIOSIT CytomeTRI technical Platform – Villejean Campus, Rennes, France). Sorted cells were centrifuged at 300 G for 5 min at 4 °C and stored at −80 °C. The antibodies used are described in the Supplementary Table S1.
Mammosphere assays
Three-dimensional cell culture was performed on 48-well plates coated with Matrigel (356230, Life Sciences, Amsterdam, Netherlands). Wells received 100 µL of Matrigel and plates were left for 30 min at 37 °C for Matrigel gelation. Suspensions of sorted cells were centrifuged at 300 G during 5 min (4 °C) and the cell pellets were resuspended in Epicult medium (Epicult Basal medium, 05602, Stem cell technologies) supplemented with Epicult B supplement (05602, Stem cell technologies), L-Glutamine (2 mM final concentration), hydrocortisone (0.48 µg/mL final concentration) and 2% of Matrigel. Cells were plated at a concentration of 100.000 cells/well and cultured in Epicult medium for 7 days. Cells were observed at 10 × magnification using phase contrast microscopy (Zeiss Axio; Zeiss France, Fougères, France) and images were digitalized. Mammosphere mean diameters were determined from at least 12 frames per conditions using the Zen analysis software (Zeiss, France).
Protein extraction and Western Blotting
Proteins were extracted from sorted cell populations, quantified using the BCA assay kit (23227, Thermo Fisher, Illkirch, France) and analyzed by Western blotting as previously described42, except that the amount of loaded protein was reduced to 2.5 µg. ECL signal was digitalized using the ImageQuant LAS4000 Imager digital system (GE Healthcare, Velizy-Villacoublay, France) and quantified with the ImageQuant TL software (GE Healthcare). Transferred proteins on PVDF membrane were stained using Coomassie brilliant blue R-250 (161-0436, Biorad, Marnes-la-Coquette, France) to normalize data. Coomassie blue stained PVDF membrane were digitized and total protein in each track was quantified as described above for the ECL signal. ECL signals were expressed as the percentage of total transferred proteins. The antibodies used are described in Supplementary Table S1.
mRNA extraction and quantitative PCR
RNA extraction was performed using the Nucleospin RNA XS kit (740902, Macherey-Nagel, Hoerdt, France) according to the manufacter’s instructions. Reverse transcription and quantitative PCR were performed as previously described12. Raw cycle threshold (Ct) values obtained from StepOne Software version 2.3 (Applied Biosystems) were transformed into quantities using the delta Ct method. This calculation of Delta Ct (Ct of target gene/Ct of reference gene) was chosen to highlight the level of expression for each sub-population considered as independent sub-populations. The endogenous control gene, the Ribosomal Protein Large P0 (RPLP0), was selected as the most stable gene within a panel of 3 genes (18S rRNA, Ribosomal Protein S5 and RPLP0) using the Normfinder algorithm. The primers used in this study are described in Supplementary Table S2.
Histological and immunohistochemical staining
Hematoxylin and eosin staining were performed on paraffin sections (8 µm) after rehydration as previously described12. CD49f and CD24 immunostaining (see below) were performed on frozen sections (5 µm) mounted on Superfrost Plus slides (4951PLUS4, Thermo Fisher). CD10 immunostaining was done on paraffin sections (8 µm) as previously detailed12 with the following modifications. After deparaffinization, slides were first incubated with 50 mM ammonium chloride (A0171, Sigma-Aldrich) for 10 min and then with 0.1% Sudan black B (S2380, Sigma-Aldrich) in 70% ethanol for 20 min to quench the autofluorescence of immune cells. Slides were then rinsed with Tris-buffered saline (TBS) with 0.02% Tween-20 (P1379, Sigma-Aldrich). Tissue sections were then subjected to heat-induced epitope retrieval in 1 mM ethylenediaminetetraacetic acid (EDTA, E9884, Sigma-Aldrich), pH8, using a microwave at 800 watts for 2 × 5 min. For immunofluorescence analysis of sorted cells, cells were deposited on poly-lysine coated slides using cytospin at 800 rpm for 3 min and fixed for 30 min with 4% paraformaldehyde at room temperature. Sorted cells and sections from both frozen and paraffin-embedded tissue were then permeabilized with 0.25% Triton X-100 (T9284, Sigma-Aldrich). Nonspecific-antibody binding was blocked with 2% bovine serum albumin (A2153, Sigma-Aldrich) in TBS. Tissue slices were then sequentially incubated with primary and secondary antibodies (Table S1) at 37 °C for 1h30 and 45 min, respectively. After washing, nuclei were counterstained with Hoechst 33342 (14533, VWR) at 1 µg/mL for 2 min. Slides were mounted using Vectashield mounting medium (H-1000; Vector Laboratories, Burlingame, CA). Images were obtained with either an E400 Nikon microscope (Nikon France, Le Pallet, France) for the CD49f and CD10 staining using the NIS-Elements BR4.20.00 software (Nikon) or an Apotome and Zen software (Zeiss france) for CD24 (Fig. 1), KRT7, KRT14, KRT8 and CD10 staining of sorted cells.
Statistical analysis
Data were expressed as means ± SEM. PCR and cytometry results were subjected to an analysis of variance (ANOVA) using the R Studio software. Post-hoc Tukey pairwise comparisons were used. Different letters in Fig. 3 and Table 1 indicate significant differences (p < 0.05). For statistical analysis of Western blot results (Fig. S3) we used the non-parametric Mann-Whitney U test. Significant differences were considered at p < 0.05 and trends at p < 0.10.
References
Yart, L., Lollivier, V., Marnet, P. G. & Dessauge, F. Role of ovarian secretions in mammary gland development and function in ruminants. Animal 8, 72–85, https://doi.org/10.1017/S1751731113001638 (2014).
McBryan, J. & Howlin, J. Pubertal Mammary Gland Development: Elucidation of In Vivo Morphogenesis Using Murine Models. Methods Mol Biol 1501, 77–114, https://doi.org/10.1007/978-1-4939-6475-8_3 (2017).
Ormerod, E. J. & Rudland, P. S. Regeneration of mammary glands in vivo from isolated mammary ducts. J Embryol Exp Morphol 96, 229–243 (1986).
Deome, K. B., Faulkin, L. J. Jr., Bern, H. A. & Blair, P. B. Development of mammary tumors from hyperplastic alveolar nodules transplanted into gland-free mammary fat pads of female C3H mice. Cancer Res 19, 515–520 (1959).
Smith, G. H. & Medina, D. A morphologically distinct candidate for an epithelial stem cell in mouse mammary gland. J Cell Sci 90(Pt 1), 173–183 (1988).
Shackleton, M. et al. Generation of a functional mammary gland from a single stem cell. Nature 439, 84–88, https://doi.org/10.1038/nature04372 (2006).
Stingl, J. et al. Purification and unique properties of mammary epithelial stem cells. Nature 439, 993–997, https://doi.org/10.1038/nature04496 (2006).
Dontu, G. & Ince, T. A. Of mice and women: a comparative tissue biology perspective of breast stem cells and differentiation. J Mammary Gland Biol Neoplasia 20, 51–62, https://doi.org/10.1007/s10911-015-9341-4 (2015).
Visvader, J. E. & Stingl, J. Mammary stem cells and the differentiation hierarchy: current status and perspectives. Genes Dev 28, 1143–1158, https://doi.org/10.1101/gad.242511.114 (2014).
Rauner, G. & Barash, I. Cell hierarchy and lineage commitment in the bovine mammary gland. PLoS One 7, e30113, https://doi.org/10.1371/journal.pone.0030113 (2012).
Martignani, E., Eirew, P., Eaves, C. & Baratta, M. Functional identification of bovine mammary epithelial stem/progenitor cells. Vet Res Commun 33(Suppl 1), 101–103, https://doi.org/10.1007/s11259-009-9254-z (2009).
Perruchot, M. H. et al. Mammary Epithelial Cell Hierarchy in the Dairy Cow Throughout Lactation. Stem Cells Dev 25, 1407–1418, https://doi.org/10.1089/scd.2016.0098 (2016).
Sleeman, K. E., Kendrick, H., Ashworth, A., Isacke, C. M. & Smalley, M. J. CD24 staining of mouse mammary gland cells defines luminal epithelial, myoepithelial/basal and non-epithelial cells. Breast Cancer Res 8, R7, https://doi.org/10.1186/bcr1371 (2006).
Inman, J. L., Robertson, C., Mott, J. D. & Bissell, M. J. Mammary gland development: cell fate specification, stem cells and the microenvironment. Development 142, 1028–1042, https://doi.org/10.1242/dev.087643 (2015).
Eirew, P. et al. Aldehyde dehydrogenase activity is a biomarker of primitive normal human mammary luminal cells. Stem Cells 30, 344–348, https://doi.org/10.1002/stem.1001 (2012).
Dairkee, S. H., Puett, L. & Hackett, A. J. Expression of basal and luminal epithelium-specific keratins in normal, benign, and malignant breast tissue. J Natl Cancer Inst 80, 691–695 (1988).
Safayi, S. et al. Myoepithelial cell differentiation markers in prepubertal bovine mammary gland: effect of ovariectomy. J Dairy Sci 95, 2965–2976, https://doi.org/10.3168/jds.2011-4690 (2012).
Rauner, G. & Barash, I. Enrichment for Repopulating Cells and Identification of Differentiation Markers in the Bovine Mammary Gland. J Mammary Gland Biol Neoplasia 21, 41–49, https://doi.org/10.1007/s10911-015-9348-x (2016).
Peuhu, E., Virtakoivu, R., Mai, A., Warri, A. & Ivaska, J. Epithelial vimentin plays a functional role in mammary gland development. Development, https://doi.org/10.1242/dev.154229 (2017).
Wang, D. et al. Identification of multipotent mammary stem cells by protein C receptor expression. Nature 517, 81–84, https://doi.org/10.1038/nature13851 (2015).
Shipitsin, M. et al. Molecular definition of breast tumor heterogeneity. Cancer Cell 11, 259–273, https://doi.org/10.1016/j.ccr.2007.01.013 (2007).
Asselin-Labat, M. L. et al. Steroid hormone receptor status of mouse mammary stem cells. J Natl Cancer Inst 98, 1011–1014, https://doi.org/10.1093/jnci/djj267 (2006).
O’Leary, K. A., Shea, M. P., Salituro, S., Blohm, C. E. & Schuler, L. A. Prolactin Alters the Mammary Epithelial Hierarchy, Increasing Progenitors and Facilitating Ovarian Steroid Action. Stem Cell Reports 9, 1167–1179, https://doi.org/10.1016/j.stemcr.2017.08.011 (2017).
Atwood, C. S. et al. Progesterone induces side-branching of the ductal epithelium in the mammary glands of peripubertal mice. J Endocrinol 167, 39–52 (2000).
Brisken, C. & Ataca, D. Endocrine hormones and local signals during the development of the mouse mammary gland. Wiley Interdiscip Rev Dev Biol 4, 181–195, https://doi.org/10.1002/wdev.172 (2015).
Seagroves, T. N., Lydon, J. P., Hovey, R. C., Vonderhaar, B. K. & Rosen, J. M. C/EBPbeta (CCAAT/enhancer binding protein) controls cell fate determination during mammary gland development. Mol Endocrinol 14, 359–368, https://doi.org/10.1210/mend.14.3.0434 (2000).
Ormandy, C. J. et al. Investigation of the transcriptional changes underlying functional defects in the mammary glands of prolactin receptor knockout mice. Recent Prog Horm Res 58, 297–323 (2003).
Oakes, S. R. et al. The Ets transcription factor Elf5 specifies mammary alveolar cell fate. Genes Dev 22, 581–586, https://doi.org/10.1101/gad.1614608 (2008).
Van Keymeulen, A. et al. Lineage-Restricted Mammary Stem Cells Sustain the Development, Homeostasis, and Regeneration of the Estrogen Receptor Positive Lineage. Cell Rep 20, 1525–1532, https://doi.org/10.1016/j.celrep.2017.07.066 (2017).
Martignani, E., Eirew, P., Accornero, P., Eaves, C. J. & Baratta, M. Human milk protein production in xenografts of genetically engineered bovine mammary epithelial stem cells. PLoS One 5, e13372, https://doi.org/10.1371/journal.pone.0013372 (2010).
Shehata, M. et al. Phenotypic and functional characterisation of the luminal cell hierarchy of the mammary gland. Breast Cancer Res 14, R134, https://doi.org/10.1186/bcr3334 (2012).
Bach, K. et al. Differentiation dynamics of mammary epithelial cells revealed by single-cell RNA sequencing. Nat Commun 8, 2128, https://doi.org/10.1038/s41467-017-02001-5 (2017).
Borena, B. M., Bussche, L., Burvenich, C., Duchateau, L. & Van de Walle, G. R. Mammary stem cell research in veterinary science: an update. Stem Cells Dev 22, 1743–1751, https://doi.org/10.1089/scd.2012.0677 (2013).
Rauner, G., Ledet, M. M. & Van de Walle, G. R. Conserved and variable: Understanding mammary stem cells across species. Cytometry A, https://doi.org/10.1002/cyto.a.23190 (2017).
Osinska, E. et al. Comparison of stem/progenitor cell number and transcriptomic profile in the mammary tissue of dairy and beef breed heifers. J Appl Genet 55, 383–395, https://doi.org/10.1007/s13353-014-0213-1 (2014).
Bachelard-Cascales, E. et al. The CD10 enzyme is a key player to identify and regulate human mammary stem cells. Stem Cells 28, 1081–1088, https://doi.org/10.1002/stem.435 (2010).
Prater, M. D. et al. Mammary stem cells have myoepithelial cell properties. Nat Cell Biol 16(942–950), 941–947, https://doi.org/10.1038/ncb3025 (2014).
Van Keymeulen, A. et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature 479, 189–193, https://doi.org/10.1038/nature10573 (2011).
Scheele, C. L. et al. Identity and dynamics of mammary stem cells during branching morphogenesis. Nature 542, 313–317, https://doi.org/10.1038/nature21046 (2017).
Stingl, J. Detection and analysis of mammary gland stem cells. J Pathol 217, 229–241, https://doi.org/10.1002/path.2457 (2009).
Asselin-Labat, M. L. et al. Delineating the epithelial hierarchy in the mouse mammary gland. Cold Spring Harb Symp Quant Biol 73, 469–478, https://doi.org/10.1101/sqb.2008.73.020 (2008).
Arevalo Turrubiarte, M., Perruchot, M. H., Finot, L., Mayeur, F. & Dessauge, F. Phenotypic and functional characterization of two bovine mammary epithelial cell lines in 2D and 3D models. Am J Physiol Cell Physiol 310, C348–356, https://doi.org/10.1152/ajpcell.00261.2015 (2016).
Acknowledgements
We thank Laurent Deleurme and Gersende Lacombe from the BIOSIT CytomeTRI platform of Rennes (France) for technical assistance. Acknowledgements are also extended to the staff at the INRA dairy farm of Méjusseaume (UMR1348 PEGASE, Le Rheu, France) and Frédérique Mayeur-Nickel for laboratory analyses. This work was supported by the Animal Physiology & Livestock System Department of the French National Institute for Agricultural Research (INRA).
Author information
Authors and Affiliations
Contributions
L.F. performed experiments, data interpretation, statistics and manuscript preparation. F.D. supervised the project conception, and contributed to the design of experiments and to the writing of the manuscript. E.C. contributed to data interpretation and to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Finot, L., Chanat, E. & Dessauge, F. Molecular signature of the putative stem/progenitor cells committed to the development of the bovine mammary gland at puberty. Sci Rep 8, 16194 (2018). https://doi.org/10.1038/s41598-018-34691-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-34691-2
Keywords
This article is cited by
-
Expression of NR5A2, NUP153, HNF4A, USP15 and FNDC3B is consistent with their use as novel biomarkers for bovine mammary stem/progenitor cells
Journal of Molecular Histology (2021)
-
Colostrogenesis: Role and Mechanism of the Bovine Fc Receptor of the Neonate (FcRn)
Journal of Mammary Gland Biology and Neoplasia (2021)
-
Mammary Epithelial Cell Lineage Changes During Cow’s Life
Journal of Mammary Gland Biology and Neoplasia (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.