Abstract
Systemic sclerosis (SSc) is a prototypic systemic fibrotic disease with unclearly characterized genetic basis. We have discovered that mutations in family with sequence similarity 111, member B (FAM111B) gene cause hereditary fibrosing poikiloderma with tendon contractures, myopathy, and pulmonary fibrosis, a multisystem fibrotic condition with clinical similarities to SSc. This observation has established FAM111B as a candidate gene for SSc. Patients and Methods: Demographic and clinical characteristics of consenting adults with definite SSc were recorded. Blood DNA analysis was performed using the High-Resolution Melt technique, and samples with abnormal electropherograms were selected for Sanger sequencing to identify mutations. Ethnically-matched controls from the general South African population were used to verify the frequency of variants in FAM111B. Public databases such as 1000 Genomes and ExAC were also used to verify the frequency of variants in FAM111B. Results: Of 131 patients, 118 (90.1%) were female, and 78 (59.5%) were black Africans. Genetic analysis revealed two FAM111B genetic variants. The c.917 A > G variant (rs200497516) was found in one SSc patients, and one control, and was classified as a missense variant of unknown significance. The c.988 C > T variant (rs35732637) occurred in three SSc patients and 42/243 (17.3%) of healthy controls, and is a known polymorphism. Conclusion: One rare variant was found in a patient with SSc but has no functional or structural impact on the FAM111B gene. In this cohort, FAM111B gene mutations are not associated with SSc.
Similar content being viewed by others
Introduction
Systemic sclerosis (SSc) is a rare autoimmune disease characterized by widespread fibrosis, micro-vascular alterations and autoantibody production1. The disease is incurable, with a 5-year mortality of up to 50%, with respiratory failure accounting for over a third of deaths2,3. The pathogenesis is a poorly understood complex interaction between vascular dysfunction, dysregulation of the innate and adaptive immune systems, and excess activation of fibroblasts and related cells4.
Although SSc is not inherited in a Mendelian fashion, and heritability of the disease remains low, a family history of SSc is the strongest risk factor for developing the condition, and siblings of affected individuals have a 15-fold increased risk of SSc5. Frech et al. studied Utah Americans and demonstrated that the presence of SSc in first degree relatives increases susceptibility to SSc, Raynaud’s phenomenon, fibrotic lung disease and other autoimmune diseases6.
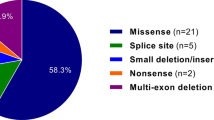
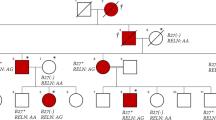
Recently, family with sequence similarity 111, member B (FAM111B) mutations were shown to cause a rare autosomal dominant disease, hereditary fibrosing poikiloderma7. This condition has clinical similarities to SSc, and is characterized by childhood-onset poikiloderma with tendon contractures, myopathy, and pulmonary fibrosis (POIKTMP). Whole exome sequencing revealed three single-nucleotide mutations on the FAM111B gene, presumably affecting the function of the gene and resulting in multisystem fibrosis.
We undertook this study on a cohort of South African SSc patients to determine the presence of FAM111B mutations.
Patients and Methods
This cross-sectional study was conducted at rheumatology outpatient departments of two tertiary hospitals between June 2013 and December 2015. All patients met the 2013 American College of Rheumatology/European League of Arthritis and Rheumatism criteria for SSc and signed informed consent before participating8. Approval for the study was obtained from the University of Cape Town Human Research Ethics Committee and the University of the Witwatersrand Committee for Research on Human Subjects. All methods were performed in accordance with the relevant guidelines and regulations.
Demographic particulars and self-reported ethnic background, clinical details, the presence of serum autoantibodies (antinuclear factor (ANA), anti-topoisomerase 1 and anti-centromere antibodies), and investigations including chest x-ray (CXR), lung function tests, high resolution computed tomography (HRCT), barium studies, gastroscopy, and echocardiograms, were documented. Physical examination included the modified Rodnan skin score (mRSS). Eosophageal involvement was considered when a patient had a clinical complaint of dysphagia or heartburn; and/or barium swallow revealed esophageal dysmotility or reflux disease on gastroscopy. Pulmonary fibrosis was diagnosed when a patient presented with infiltrates or honeycombing on chest X ray and/or on high resolution computed tomography (HRCT) and had abnormal pulmonary function test (reduced forced vital and diffusion capacity). Pulmonary arterial hypertension (PAH) was defined as an elevated right ventricular systolic pressure (>45 mmHg) on echocardiography.
Genetic analysis
Genomic DNA was extracted from peripheral leucocytes and mutational screening of FAM111B was performed using a High Resolution Melt (HRM) technique. An HRM reaction with a total volume of 25 ul/sample was prepared using 0.5 U GoTaq® Flexi DNA Polymerase (Promega, Madison, WI, USA), 1X Colorless GoTaq® Flexi Buffer (Promega), 3 mMMgCl2 (Promega), 0.8 μM dNTPs (Bioline, London, United Kingdom), 0.4x EvaGreen dye (Biotium, Hayward, CA, USA), 0.4, μM of each primer (forward and reverse) and 50 ng/ul DNA. HRM reactions were conducted using the RotorGene 6000 (Corbett Life Sciences – Qiagen, Venlo, Limber, Netherlands) and the cycling conditions were set at 95 °C for 10 minutes; 50 cycles of 95 °C for 5 seconds, 55 °C for 10 seconds and 72 °C for10 seconds; and a high resolution melt from 72 °C to 95 °C with 0.1 °C increases in temperature. Samples with abnormal electropherograms were selected for Sanger sequencing to identify mutations.
Samples were purified using Exonuclease I (New England Biolabs, Ipswich, MA, USA) and FastAPTM Thermosensitive Alkaline Phosphatase (Promega) using a Mastercycler® pro thermal cycler (Eppendorf, Hamburg, Germany); conditions were 37 °C for 1 hour and 75 °C for 15 minutes. Sequencing reactions were prepared using the BigDye® Direct Cycle Sequencing Kit (Applied Biosystems – Life Technologies, Carlsbad, CA, USA). Reaction conditions were 96 °C for 5 minutes and 25 cycles of 96 °C for 30 seconds, 50 °C for 15 seconds and 60 °C for 4 minutes. Sequencing products were analyzed using capillary electrophoresis with the ABI PRISM® 3130 × l Genetic Analyser (Applied Biosystems)9.
Normal Controls
All variants that were identified in the SSc cohort were similarly screened in a cohort of 243 normal South African population controls made up of 107 (44.0%) Caucasians, 64 (26.3%) black Africans, and 72 (29.6%) individuals of mixed ancestry.
Bioinformatic analysis
Public databases were also used to verify the frequency of variants in FAM111B. These included: 1000 Genomes (http://www.internationalgenome.org/) and ExAC (http://exac.broadinstitute.org/).
The missense variants were investigated using the following pathogenicity prediction tools: PolyPhen2 (http://genetics.bwh.harvard.edu/pph2/), SIFT http://sift.jcvi.org/), MutationTaster (http://www.mutationtaster.org/), MutationAssessor (http://mutationassessor.org/r3/), PROVEAN (http://provean.jcvi.org/index.php), AlignGVGD (http://agvgd.iarc.fr/), fathmm (http://fathmm.biocompute.org.uk/) and M-CAP (http://bejerano.stanford.edu/mcap/).
Results
131 patients were genotyped, 13 men and 118 women, and the mean (SD) age of diagnosis was 47.2 (13.4) years (Table 1). The majority of patients were black Africans (59.5%). More than half (55.7%) had the diffuse cutaneous subtype (dcSSc). The vast majority had Raynaud’s phenomenon (90.1%), and were ANA positive (90.4%). As expected, skin scores in dcSSc patients were significantly higher compared to patients with the limited cutaneous subtype (lcSSc), and musculoskeletal manifestations (tendon friction rubs, joint contractures, muscle weakness and elevated creatinine kinase) were significantly more common in dcSSc than lcSSc patients. Over a quarter (26.7%) of patients had pulmonary fibrosis, and 12.2% had PAH.
Genetic analysis of the SSc patients revealed two genetic variants of FAM111B in only four patients: c.917 A > G in one patient and c.988 C > T in three patients (Table 2). The c.917 A > G variant (rs200497516) was detected in one of the 243 healthy controls, and had a reported frequency of <0.01 in the 1000 Genomes and ExAC databases. This missense variant was thus classified as a genetic variant of unknown significance (GVUS). Although the amino acid was conserved across species, all bioinformatic pathogenicity prediction tools showed the variant to be benign and thus unlikely to have a structural or functional impact on the FAM111B (Table 3). The second variant, c.988 C > T variant (rs35732637) is a known polymorphism, having a reported frequency of >0.01 in the 1000 Genomes and ExAC databases, and was detected in 42/243 (17.3%) of the normal control population.
Discussion
To the best of our knowledge, this is the first study to investigate FAM111B mutations in SSc. Our findings of a missense variant and a known polymorphism with no functional or structural impact on FAM111B suggest that, in this cohort, FAM111B gene mutations are not associated with SSc. Thus, despite some clinical similarities, POIKTMP and SSc do not share genetic basis and are likely to have different pathogenetic pathways. The different pathogenetic pathways of these disorders are underscored by the absence of auto-antibodies in POIKTMP compared to SSc where there is definite evidence of auto-immunity.
The search for SSc susceptibility genes by candidate gene studies, and more recently by genome wide association studies (GWAS), have yielded numerous susceptibility genes, both in the major histocompatibility complex- human leucocyte antigen (MHC-HLA) region and outside this region10. Candidate genes for SSc fall into three groups that reflect the three main areas of disease expression and pathogenesis: immune responses, vasculopathy and fibrosis11. To date, the majority of gene mutations described are implicated in auto-immune dysregulation, with relatively few genetic variants pertaining to fibrosis or vasculopathy. It is possible that several polymorphisms occurring in an individual contribute to the risk, or that epigenetic alterations in gene expression underlie this complex disease.
The characteristics of patients in our cohort are comparable to those in the EULAR Scleroderma Trials and Research (EUSTAR) group database12, with some major differences. The predominance of the dcSSc subset in this cohort was also described in an earlier study of SSc in black South Africans where 65% of the patients had dcSSc compared to 37% of the EUSTAR cohort13. Less pulmonary fibrosis and PAH was diagnosed our cohort, probably because lung function tests, HRCT and echocardiograms were not routinely performed in all patients12. In addition, of the patients with antibody results, significantly fewer were anti-topoisomerase 1 and anti-centromere positive than described in the EUSTAR cohort. The absence of anti-centromere antibody was a feature of the other South African SSc study.
The majority of patients in our study were black Africans, and it is possible that mutations in FAM111B are present in other population groups, although this is unlikely based on the autoimmune basis of SSc compared to a hereditary origin of POIKTMP. Future studies should examine the role of the FAM111B mutation in the pathogenesis of related idiopathic conditions such as pulmonary fibrosis.
In conclusion, our study found one SSc patient with a missense variant of FAM111B, suggesting that, in this population, FAM111B is not associated with SSc.
Data Availability
The datasets generated during and analysed during the current study are available from the corresponding author on reasonable request.
References
Akter, T., Silver, R. M. & Bogatkevich, G. S. Recent advances in understanding the pathogenesis of scleroderma-interstitial lung disease. Curr Rheumatol Rep 16, 411, https://doi.org/10.1007/s11926-014-0411-1 (2014).
Solomon, J. J. et al. Scleroderma lung disease. Eur Respir Rev 22, 6–19, https://doi.org/10.1183/09059180.00005512 (2013).
Manetti, M. & Matucci-Cerinic, M. The new frontier in systemic sclerosis: from epigenetics to new treatments. Rheumatology (Oxford) 54, 1757–1758, https://doi.org/10.1093/rheumatology/kev264 (2015).
Abraham, D. J., Krieg, T., Distler, J. & Distler, O. Overview of pathogenesis of systemic sclerosis. Rheumatology (Oxford) 48(Suppl 3), iii3–7, https://doi.org/10.1093/rheumatology/ken481 (2009).
Arnett, F. C. et al. Familial occurrence frequencies and relative risks for systemic sclerosis (scleroderma) in three United States cohorts. Arthritis Rheum 44, 1359–1362, https://doi.org/10.1002/1529-0131(200106)44:6<1359::AID-ART228>3.0.CO;2-S (2001).
Frech, T. et al. Heritability of vasculopathy, autoimmune disease, and fibrosis in systemic sclerosis: a population-based study. Arthritis Rheum 62, 2109–2116, https://doi.org/10.1002/art.27469 (2010).
Mercier, S. et al. CUGC for hereditary fibrosing poikiloderma with tendon contractures, myopathy, and pulmonary fibrosis (POIKTMP). Eur J Hum Genet 24, https://doi.org/10.1038/ejhg.2015.205 (2016).
van den Hoogen, F. et al. Classification criteria for systemic sclerosis: an American College of Rheumatology/European League against Rheumatism collaborative initiative. Arthritis Rheum 65, 2737–2747, https://doi.org/10.1002/art.38098 (2013).
Fish, M. et al. Mutation analysis of the phospholamban gene in 315 South Africans with dilated, hypertrophic, peripartum and arrhythmogenic right ventricular cardiomyopathies. Scientific reports 6, 22235, https://doi.org/10.1038/srep22235 (2016).
Murdaca, G., Contatore, M., Gulli, R., Mandich, P. & Puppo, F. Genetic factors and systemic sclerosis. Autoimmun Rev 15, 427–432, https://doi.org/10.1016/j.autrev.2016.01.016 (2016).
Romano, E. et al. The genetics of systemic sclerosis: an update. Clin Exp Rheumatol 29, S75–86 (2011).
Walker, U. A. et al. Clinical risk assessment of organ manifestations in systemic sclerosis: a report from the EULAR Scleroderma Trials And Research group database. Ann Rheum Dis 66, 754–763, https://doi.org/10.1136/ard.2006.062901 (2007).
Tager, R. E. & Tikly, M. Clinical and laboratory manifestations of systemic sclerosis (scleroderma) in Black South Africans. Rheumatology (Oxford) 38, 397–400 (1999).
Author information
Authors and Affiliations
Contributions
A.G. and G.D. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gcelu, A., Deshpande, G., Shaboodien, G. et al. Mutations of FAM111B gene are not associated with Systemic Sclerosis. Sci Rep 8, 15988 (2018). https://doi.org/10.1038/s41598-018-34341-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-34341-7
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.