Abstract
Changes in gene expression can rapidly influence adaptive traits in the early stages of lineage diversification. Venom is an adaptive trait comprised of numerous toxins used for prey capture and defense. Snake venoms can vary widely between conspecific populations, but the influence of lineage diversification on such compositional differences are unknown. To explore venom differentiation in the early stages of lineage diversification, we used RNA-seq and mass spectrometry to characterize Sidewinder Rattlesnake (Crotalus cerastes) venom. We generated the first venom-gland transcriptomes and complementary venom proteomes for eight individuals collected across the United States and tested for expression differences across life history traits and between subspecific, mitochondrial, and phylotranscriptomic hypotheses. Sidewinder venom was comprised primarily of hemorrhagic toxins, with few cases of differential expression attributable to life history or lineage hypotheses. However, phylotranscriptomic lineage comparisons more than doubled instances of significant expression differences compared to all other factors. Nevertheless, only 6.4% of toxins were differentially expressed overall, suggesting that shallow divergence has not led to major changes in Sidewinder venom composition. Our results demonstrate the need for consensus venom-gland transcriptomes based on multiple individuals and highlight the potential for discrepancies in differential expression between different phylogenetic hypotheses.
Similar content being viewed by others
Introduction
Lineage diversification allows for evolutionary processes such as selection to act independently upon a trait across different clades. For traits tightly linked to an organism’s survival, adaptation within a lineage can occur rapidly through the evolution of gene-expression patterns (i.e. regulatory evolution), where few changes in the regulatory machinery yield large, cascading changes in the expressed phenotype1,2. Regulatory evolution contrasts with the evolution of coding sequences in focal genes, which is reliant on the comparatively slow process of mutation in coding gene exons3. Notably, variation in gene-expression patterns has contributed to adaptation in many species and may be a key source of variability among populations in the early stages of lineage diversification1. However, as many ecologically-important traits are polygenic and involve complex pathways, they can be difficult to track genetically. Traits with a tractable genetic basis and high ecological importance can therefore permit investigations of the mode, tempo, and correlates of adaptive regulatory evolution in young lineages2.
Venomous animals provide exceptional systems to investigate the role of regulatory evolution on adaptive traits in the early stages of lineage diversification4. Venom is a polygenic trait that has evolved multiple times across the tree of life, where it serves in prey capture and predator defense4,5. Unlike many polygenic traits2,6, venom is the result of a relatively direct pathway from the transcription of toxin genes to the translation of toxin proteins, which are then stored as a protein mixture prior to use7,8. Therefore, by combining venom-gland transcriptomics and venom proteomics, we can accurately map the progression from genotype to phenotype in this adaptive trait4.
Prior proteomic, enzymatic, and transcriptomic studies have found significant variation in the composition and expression of snake venom across all levels of biological organization, including between populations of the same species9. This extensive variation is both biologically significant and medically important: for example, venomous snake families (i.e. Viperidae, Elapidae) often share similar toxins based on phylogenetic relatedness10. However, within rattlesnakes (Crotalus and Sistrurus), little evidence exists for phylogenetic signal in patterns of venom expression between species, indicating rapid local adaptation in allopatry10,11,12,13. Most studies of venom variation in rattlesnakes focus on populations of single species, where geographic variation is commonly detected and associated with variable diets14,15, local environments16, or interactions with coevolving prey16,17. These patterns are largely attributed to differential gene expression17,18,19,20. However, few intraspecific studies have explored the role of differential gene expression in driving venom variation where there is phylogenetic structure within a species across its range. At this level, if we find patterns of differential toxin expression, then it could be due to general accumulation of change as the lineages diversify in addition to local adaptation to prey17,19,21.
Here, we attempt to understand the effect of shallow lineage divergence on venom through quantification of the transcriptomes and proteomes of distinct lineages of Sidewinder Rattlesnakes (Crotalus cerastes). Sidewinders are small rattlesnakes native to the warm deserts of the southwestern United States and northwestern Mexico, easily recognized by their conspicuous, raised supraoculars and specialized side-winding locomotion22,23. Previous studies have noted some differences in venom enzymatic activity between populations, but the proteomic basis for this activity is unknown24,25. Three subspecies are currently recognized within Crotalus cerastes26: C. c. cerastes found in the Mojave Desert, C. c. cercobombus in the Sonoran Desert, and C. c. laterorepens in the Colorado Desert (Fig. 1). However, these subspecific assignments are not consistent with phylogenetic lineages. Mitochondrial phylogenetic analyses of C. cerastes by Pece27 and Douglas et al.28 recovered four shallow lineages in the United States, corresponding to the North and South Mojave, Colorado, and Sonoran Deserts; Douglas et al.28 recovered a fifth lineage from the southern portion of the Sonoran Desert in Mexico. It is well-documented that phylogenies built solely on mitochondrial data may not accurately represent the true evolutionary history or relationships of the study taxa, as the mitochondrial genome is rapidly evolving and matrilineal29,30,31,32,33,34. Therefore, it is important to verify these lineage assignments with larger genomic datasets. Sidewinder subspecies and mitochondrial lineages are suitable as initial hypotheses to test for differential venom expression, but the considerable number of nontoxin genes captured by venom-gland transcriptomics allows for a more robust investigation of the evolutionary relationships of these populations.
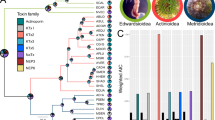
Sampling and approximate distribution of Crotalus cerastes by subspecies. Phylogeny in upper panel based on 1,508 nontoxin loci and 1,401,828 bp, rooted at the midpoint. Phylogeny in the lower panel based on the mtDNA loci ATPase 8 and 6, using samples from GenBank (Douglas et al.28) and transcriptomes generated herein. Sample colors correspond to recovered clade, not assignment based on proposed subspecies ranges. Note the difference in the position of CAS 259903 between trees. Pie charts correspond to toxin gene expression in each sample, showing the five most highly expressed toxin families. Maps were created using ArcGIS 10.5.1 (ESRI Inc., USA. http://www.esri.com) using freely available spatial data (www.diva-gis.org/Data), then annotated in Inkscape 0.92 (https://inkscape.org); species ranges were approximated based on83.
Based on previous studies suggesting variable enzymatic activity within Crotalus cerastes venom24,25,35 and the well-documented intraspecific variation in venom from other co-distributed Crotaline taxa36,37,38, we hypothesize that lineage diversification has led to venom divergence within the species and predict this divergence will correspond to lineage, subspecies, and/or life history. To test this hypothesis, we describe the first venom-gland transcriptomes and corresponding venom proteomes of eight Crotalus cerastes, including samples assignable to all subspecies and mitochondrial lineages known from the southwestern United States. We test whether life history traits (sex or age) or lineage assignments (subspecies, mitochondrial or phylotranscriptomic lineages) best explain differences in toxin expression, and subsequently explore the influence of lineage diversification on differential toxin expression.
Methods
Specimen, venom, and venom gland collection
We collected representative samples from each of the recognized subspecies of C. cerastes. In total, eight snakes (six males, two females) were collected between October 2012 and July 2014 in California and Arizona (Fig. 1; Table 1). Snout-to-vent lengths (SVL) ranged from 30.9–48.4 cm (43.12 ± 7.12 cm) for males and 37.7–40.8 cm (39.2 ± 1.55 cm) for females. Snakes with SVL less than 34 cm (males) or 38 cm (females) were consider juveniles39,40. We collected venom from the snakes, then four days later collected venom glands from each snake to maximize transcription of venom genes41. Venom was collected by either allowing the snakes to bite onto a parafilm-sealed sterile collection cup or by electrostimulation42, then dehydrated and stored at −80 °C until analyzed. Four days after venom extraction, snakes were euthanized with a single-step sodium pentobarbital (100 mg/kg) injection following standard approved AVMA guidelines43, and both the left and right venom glands of each snake were excised. The excised venom glands were immediately transferred to RNAlater following dissection and stored briefly at 4 °C prior to long term storage at −80 °C. Snakes were handled and collected under the following permits and Animal Care and Use Protocols: State of Arizona Game and Fish Department (SP673390), California Natural Resources Agency Department of Fish and Wildlife (SC-12985), and University of Central Florida IACUC (13–17 W).
RNA extraction and sequencing
We isolated RNA via a standard TRIzol extraction following Rokyta et al.20,44,45. Briefly, venom glands were finely diced and placed in TRIzol solution (Invitrogen). The mixture was homogenized and transferred to a phase lock heavy gel tube (5Prime). Once cells were lysed, total RNA was isolated using chloroform and purified via isopropyl alcohol and ethanol precipitation. RNA quantification was performed using a Qubit RNA BroadRange kit. RNA quality was checked using a Bioanalyzer 2100 with an RNA 6000 Pico Kit (Agilent Technologies) or Agilent TapeStation 2200 with a RNA ScreenTape to ensure sufficient quantity and quality RNA for library preparation and sequencing.
We produced cDNA libraries from isolated mRNA using magnetic bead isolation of mRNA followed by cDNA synthesis and PCR amplification. First, we isolated mRNA using the NEBNext Poly(A) mRNA Magnetic Isolation Module (NEB #E7490S) with equal amounts of mRNA from each of the left and right venom glands. Following bead isolation and cleanup, cDNA libraries were prepared from isolated mRNA using a NEB Next Ultra RNA Library Prep Kit for Illumina (NEB #E7530) following manufacturer’s recommendations. We used a fragmentation time of 13 minutes, 30 seconds to achieve a target mean fragment size of 400 bp, and 14 PCR cycles for amplification of double stranded cDNA libraries. Library yield and quality was quantified on a Bioanalyzer 2100. The total amplifiable concentration of cDNA in each library was then determined using KAPA qPCR at the Florida State University Molecular Cloning Facility. Equal concentrations of samples were then pooled in groups of 12 samples for sequence. The final concentration and quality of the pooled libraries samples was then assessed on the Bioanalyzer and via KAPA qPCR. Pooled libraries were sequenced with 150 base pair (bp) paired-end reads on an Illumina HiSeq2500 platform at the Florida State University College of Medicine Translational Science Laboratory (Tallahassee, FL, USA).
Transcriptome assembly and annotation
We cleaned and assembled the raw reads using a combination of custom python scripts and established software. The raw 150 bp paired-end reads were checked for potential cross-contamination, caused by mis-allocation of reads during demultiplexing, using custom python scripts. The scripts first obtain counts of all 57-mers in the raw reads using Jellyfish v. 2.2.646. The scripts then obtain all pairwise comparisons of these k-mer counts for samples in a given lane, and removes reads that were 25% constructed of these k-mers. Cleaned reads were then trimmed to keep base calls with a phred score of 5 or greater using Trim Galore! v. 0.4.4 and merged using PEAR v. 0.9.1047. The trimmed, merged reads were de novo assembled using two different methods: Extender44 and SeqMan NGen (using the Lasergene DNAStar software package; Madison, WI: https://www.dnastar.com/t-nextgen-seqman-ngen.aspx). Extender assemblies used 1000 merged reads as seeds, extending the seeds based on exact overlaps of 120 base pairs. This method has efficiently assembled toxin sequences in a variety of snake taxa, including other Crotalus species19,20,44,45,48,49,50. De novo NGen assemblies were performed with default settings in NGen v. 14.
Assembled contigs from both Extender and NGen were annotated via blastx searches against the UniProt animal venom proteins and toxins database (http://www.uniprot.org/program/Toxins) with a minimum e-value of 10−4. Both toxins and nontoxins (e.g. housekeeping genes) were annotated by clustering sequences using cd-hit-est51 to a known database of previously annotated snake toxins and C. horridus nontoxins48. Sequences and associated signal peptides with a match percentage of 80 were automatically annotated. The remaining toxin contigs were manually annotated by comparing the sequences to the blastx results. We then combined annotated sequences from both assembly methods, removed duplicates, and screened for chimeric sequences by aligning merged reads to the annotated toxins using BWA-MEM52 and removing reads with any mismatches via gaps or nucleotide differences. Transcripts with zero coverage at any base were automatically removed, and transcripts with more than 10-fold coverage differentials across their length were removed if they showed signs of chimeric transcripts (multimodal coverage distributions). The remaining transcripts were clustered with a sequence identity threshold of 98% using cd-hit to reduce redundancy of repeat transcripts and cluster allelic variation at single loci51. Finally, the de novo-assembled transcriptomes of each individual were combined and clustered using cd-hit with a sequence identity threshold of 98% to produce a species consensus transcriptome.
Phylogenetic analysis
We assigned our samples to known phylogeographic lineages first using previously published sequence data as well as using the annotated nontoxins recovered from the transcriptomes. First, we obtained partial sequences (665 bp) of the mitochondrial loci ATPase 8 and 6 from our samples by mapping each individual’s merged reads to a reference sequence obtained from GenBank using Bowtie 2 (DQ493803, see below)53. These sequences were aligned with eleven sequences obtained from GenBank corresponding to the five mitochondrial lineages of Crotalus cerastes recovered by Douglas et al.28 (DQ493803–DQ493813), the most complete molecular sampling of the species available to date (Fig. 1). An additional sequence assigned to C. helleri (AF462375; taxonomy following Davis et al.54) was included as an outgroup. Our goal was not to explore evolutionary relationships with only a small dataset of mtDNA, rather to cluster our samples with the appropriate phylogeographic clades corresponding to the mitochondrial lineages recovered by Douglas et al.28.
We also used a subset of the automatically annotated nontoxins for a more robust phylotranscriptomic analysis. First, merged reads for each individual were mapped to the consensus transcriptome using BWA-MEM52 and reads with more than three mismatches—via gaps or nucleotide differences—were removed. Picard (http://broadinstitute.github.io/picard/) was used to sort and index the aligned reads prior to using bedtools to calculate the coverage of each site within each transcript. Using a custom R script, any transcript which had zero coverage for any site in a transcript was marked as absent. Nontoxins which were present across all eight of our individuals were then pulled and used to re-map the merged reads with BWA-MEM again. Alleles were phased using a combination of samtools, Picard, and GATK (following55) and concatenated using Sequence Matrix56. The final dataset consisted of 1,508 nontoxin loci, and 1,401,828 bp.
We used both maximum likelihood and Bayesian phylogenetic approaches to recover lineage assignments for our samples. We carried out maximum likelihood (ML) analyses in RAxML v. 8.2.457 with 1000 bootstrap pseudoreplicates under the default GTR + G substitution model. Bayesian inference (BI) was performed using MrBayes 3.5.258,59. For the Bayesian analysis, ATPase 8 and 6 were partitioned by codon, and we used PartitionFinder v. 1.1.160 to determine best fit models of nucleotide substitution, with a greedy search algorithm and the model search restricted to those implementable in MrBayes. The GTR + G model was chosen as the best model for all codons except ATPase 6 codons 2 (GTR + I) and 3 (HKY). For the nontoxin dataset, we used the GTR + G model partitioned by locus. The analyses consisted of two parallel runs of four Markov chains (three heated, one cold) run for 100 million generations, sampling every 10,000, with a random starting tree and the first 20% discarded as burnin.
Venom proteomics
To generate a genotype-phenotype map and verify toxin expression proteomically, we performed quantitative mass spectrometry (qMS) on whole venom samples following Rokyta & Ward61 and Ward et al.62. We first quantified venom protein samples using the Qubit Protein Assay kit with a Qubit 1.0 Fluorometer (Thermo Fisher Scientific). For each sample, we then digested approximately 5 μg of whole venom using the Calbiochem ProteoExtract All-in-One Trypsin Digestion Kit (Merck, Darmstadt, Germany) according to the manufacturer’s instructions and using LC/MS grade solvents, leaving an overall yield of approximately 4.3 μg of digested venom protein after digestion. Samples were dried using a SpeedVac at 25 °C for 1 hour and stored at −20 °C until use. To initiate the mass spectrometry run, the resulting dried and digested tryptic peptides were resuspended in 0.1% formic acid at a final concentration of 250 ng/μL. Three digested Escherichia coli proteins-purchased from Abcam at known concentrations and mixed in the specified proportions (1000×) prior to digestion-were used as internal standards: 25 fmol of P00811 (Beta-lactamase ampC), 250 fmol of P31658 (Protein deglycase 1), and 2500 fmol of P31697 (Chaperone protein FimC) per injection. Final sample concentrations were achieved by infusing the internal standard peptide mix into samples. For the LCMS/MS run, a 2 μL aliquot was analyzed using an externally calibrated Thermo Q Exactive HF (high-resolution electrospray tandem mass spectrometer) in conjunction with Dionex UltiMate3000 RSLCnano System. A 2 μL sample was aspirated into a 50 μL loop and loaded onto the trap column (Thermo μ-Precolumn 5 mm, with nanoViper tubing 30 μm i.d. ×10 cm). For separation on the analytical column (Acclaim pepmap RSLC 75 μMx 15 cm nanoviper), the flow rate was set to 300 nl/min. Mobile phase A was composed of 99.9% H2O (EMD Omni Solvent) and 0.1% formic acid, and mobile phase B was composed of 99.9% ACN and 0.1% formic acid. We performed a 60 min linear gradient from 3% to 45% B. The LC eluent was directly nanosprayed into the Q Exactive HF mass spectrometer (Thermo Scientific), and during the chromatographic separation, the Q Exactive HF was operated in a data-dependent mode and under direct control of the Thermo Excalibur 3.1.66 (Thermo Scientific). Resulting MS data were acquired using a data-dependent top-20 method for the Q Exactive HF platform, dynamically choosing the most abundant not-yet-sequenced precursor ions from the survey scans (350–1700).
Sequencing was performed via higher energy collisional dissociation fragmentation with a target value of 105 ions determined with predictive automatic gain control. Full scans (350–1700 m/z) were performed at 60,000 resolution in profile mode. MS2 were acquired in centroid mode at 15,000 resolution. We excluded ions with a single charge, charges more than seven, or unassigned charge. A 15-s dynamic exclusion window was used. All measurements were performed at room temperature, and done with three technical replicates to account for machine-related variability and to facilitate label-free quantification. We searched the resulting raw files with Proteome Discoverer 1.4 using SequestHT as the search engine with custom-generated FASTA databases and percolator as peptide validator. The SequestHT search parameters used were: enzyme name = Trypsin, maximum missed cleavage = 2, minimum peptide length = 6, maximum peptide length = 144, maximum delta Cn = 0.05, precursor mass tolerance = 10 ppm, fragment mass tolerance = 0.2 Da, dynamic modifications, carbamidomethyl + 57.021 Da(C) and oxidation + 15.995 Da(M). Protein identities were validated using Scaffold (version 4.3.4, Proteome Software Inc., Portland, OR, USA) software. We accepted protein identities based on a 1.0% false discovery rate (FDR) using the Scaffold Local FDR algorithm and a minimum of one recognized peptide. We considered a transcript proteomically detected if it was found in at least one of the three replicates per individual. Any protein not detected in all three replicates was excluded from further analyses. From all individuals we also excluded Bradykinin-potentiation peptides due to the potential for extensive proteolytic cleavage49, as well as myotoxins due to low quality assignments. Finally, individual toxins that appeared to be ambiguously assigned in all three replicates of a sample were omitted.
To estimate proteomic abundances, we followed Rokyta & Ward61 and Ward et al.62 and calculated separate conversion factors for each of three replicates based on the known concentrations of the Escherichia coli control proteins and their observed quantitative values (normalized spectral counts) determined by Scaffold. These conversion factors were calculated by finding the slope of the best fit line of the known control concentrations and the observed normalized spectral counts, with an intercept at the origin. The conversions factors were then used to convert the normalized spectral counts for each venom protein in each replicate to concentrations; final concentrations for each sample were then averaged across each individual’s corresponding replicates. The centered log-ratio (clr) transformation63 was applied to all our transcriptome and proteome abundances for these analyses; this transformation preserves rank and is equivalent to a log transformation for linear relationships49.
The methods used here are designed to provide information on relative protein abundances, primarily to validate transcriptomic findings. In snakes, the internal standard approach has previously been shown to provide a strong agreement between venom proteins and transcripts across multiple species49. Using this approach to determine absolute protein abundances has limitations, however, due to protein-specific properties which can affect downstream abundance calculations. For this reason, all analyses are done in a compositional framework. Prior work has shown that different approaches to quantification do not affect correlation between mRNA and protein levels using the internal standard approach49. Although additional proteomic and/or analytical methods for identification and quantification may afford greater detectability of specific toxins (e.g. BPPs, myotoxins), these methods introduce other potential biases to quantification. Rather than potentially biasing our results by using different parameters for different proteins, we chose instead to treat all proteins equally using the same parameters across all samples.
Comparative transcriptome analyses
We measured relative expression of each venom gene by calculating the number of transcripts per million reads (TPM) and compared our individuals with hierarchical clustering. Using RSEM64 with default Bowtie 2 alignment settings53, merged reads from each individual were mapped to the consensus transcriptome to calculate the transcripts per million reads (TPM). To avoid zero values in the dataset and allow transformations, we used the ‘cmultRepl’ function in the R package zCompositions65. We used the R package ‘pheatmap’66 to generate a heatmap hierarchically-clustered by expression similarity (natural log-transformed TPM) between individuals in order to visualize differences in the expression of individual toxin transcripts.
To determine if consensus transcripts were potentially absent in any individuals, we aligned the merged reads of each individual to the consensus transcriptome following Rokyta et al.20 and calculated coverage of each site. Briefly, alignments were carried out using BWA MEM52 and reads with more than three mismatches—via gaps or nucleotide differences—were removed. Picard (http://broadinstitute.github.io/picard/) was used to sort and index the aligned reads prior to using bedtools to calculate the coverage of each site within each transcript. Using a custom R script, transcripts with less than 5x coverage for more than 10% of the coding sequence were considered absent from that individual’s transcriptome20. This cutoff was not applied to differential expression analyses; all transcripts were considered present in subsequent analyses.
We tested for differential venom gene expression between life history traits, phylogenetic structure, and subspecific assignment of each individual snake (Table 1). We used DESeq267 and edgeR68 to test for differential expression—based on RSEM TPM counts for each venom transcript—across five factors: body size (SVL, corresponding to life stage), sex, mitochondrial lineage, nontoxin (i.e. phylotranscriptomic) lineage, and historical subspecies assignment. Mitochondrial and nontoxin lineages largely corresponded to the deserts in which the snakes were found, and are therefore named as in Table 1. Pairwise contrasts were made between the discrete factors: i.e. different clades (e.g. South Mojave vs. North Mojave), subspecies (e.g. C. c. cerastes vs. C. c. cercobombus), and sex. Seventeen pairwise comparisons were made in total (Supplementary Tables S1 and S2). For analysis with DESeq2, we performed Wald significance tests with a local fit of dispersions for all comparisons except nontoxin lineage comparisons which was able to use a parametric fit of dispersions. Independent filtering was applied at P = 0.05 and P-values were corrected for false-discovery rate (FDR). For analysis with edgeR, negative binomial generalized linear models were fit for each gene and significance tested with likelihood ratio tests. Similar to DESeq2, P-values were corrected for FDR. We used α = 0.05 for detecting differentially expressed genes following FDR-correction in both DESeq2 and edgeR analyses. Genes were considered significantly differentially expressed if they were found to be significant in both DESeq2 and edgeR analyses. We also quantified the phylogenetic signal of venom gene expression by calculating Blomberg’s K69 in the R package ‘picante’70 using the nontoxin phylogeny rooted at the midpoint. This method assumes evolution under Brownian motion, and provides a measure of whether the evolution of a trait matches what is expected (K = 1), is less similar between closely related taxa than expected (K < 1), or is more similar between closely related taxa than expected (K > 1)69.
Lastly, we sought to detect toxin class-specific variation in gene expression. To this end, we summed TPM counts by toxin class–across paralogous toxins from the same gene family–to account for potential differences in venom not captured by individual transcript analysis. The new counts of toxin class abundance were similarly used to generate hierarchically clustered heatmaps and test for differential expression as described above.
Accession codes
SRA: SRR6768682–SRR6768689; TSA: GGMJ00000000
Results
Phylogenetic analysis
Our phylogenetic analyses recovered incongruent topologies between the mtDNA-only dataset (two loci, 665 bp) and the nontoxin phylotranscriptomic dataset (1,508 loci, ~1.4 million bp). The mtDNA topologies were poorly supported at several nodes (Fig. 1), but clustered our samples with four of the five known mtDNA lineages. The remaining mitochondrial lineage is found only in South Sonora, Mexico, which we were unable to sample as part of this study. The transcriptomically-derived nontoxin topologies were all strongly supported (PP = 1, BS = 100 for all nodes), largely clustering our samples into clades corresponding to the mtDNA assignments. However, one sample, CAS 259903, which was recovered with the South Mojave lineage in the mtDNA analyes, was recovered most closely related to the North Mojave lineage (CAS 259901), distinct from the other two samples assignable to the South Mojave group (CAS 259905, CAS 259917). This was geographically located between the individuals consistently recovered from the North Mojave and South Mojave lineages, suggesting gene flow may be occurring between these lineages.
The Sidewinder venom-gland transcriptome
De novo assembly and annotation of the venom-gland transcriptome from our eight C. cerastes individuals recovered an average of 1,167 unique nontoxin transcripts (range: 909–1,409) and 38 unique toxin transcripts (range: 33–44) per individual. We merged the eight independently assembled transcriptomes to create a consensus venom-gland transcriptome for this species. Our consensus transcriptome consisted of 1,977 unique nontoxin transcripts and 62 unique toxin transcripts (Fig. 2).
Average expression of the venom-gland transcriptome of Crotalus cerastes. (A) Expression of each recovered toxin transcript. An asterisk (*) above a toxin indicates it was verified proteomically in at least one individual. (B) Proportion of toxin gene expression by class. (C) Toxin and nontoxin gene expression in the venom gland. Inset photo: C. cerastes by Travis Fisher.
Like many other rattlesnakes, C. cerastes venom was dominated by snake venom metalloproteinases (SVMP), phospholipase A2 (PLA2), C-type lectins (CTL), snake venom serine proteinases (SVSP), and a bradykinin-protentiating peptide (BPP). Overall, toxins accounted for 63.8% of the transcriptome’s expression based on RSEM-mapped reads across the eight individuals. Five toxin classes made up the majority of Crotalus cerastes venom (Fig. 2). SVMPs were the most diverse and abundant toxin class, with a total of 16 unique transcripts (6 SVMPIIs, 10 SVMPIIIs) accounting for 25.8% of toxin-specific expression (Fig. 2). The most highly expressed toxin, a single PLA2, accounted for 23.4% of toxin expression (Fig. 2). The 11 recovered CTLs and 15 SVSPs accounted for 18.3% and 15.1% of the toxin expression, respectively (Fig. 2). Lastly, the second most highly expressed toxin, a single BPP, accounted for 6.4% of toxin expression.
Many less-abundant toxins were present in the venom-gland transcriptome. These included a L-amino-acid oxidase (LAAO), two vascular endothelial growth factors (VEGF), two myotoxins (MYO), a nerve growth factor (NGF), a cysteine-rich secretory protein (CRISP), two 3-fingered toxins (3FTx), a vespryn, a nucleotidase (NUC), and a phosphodiesterase (PDE). The least abundant toxin genes in the venom-gland transcriptome were those that may only have indirect toxic function or no toxic function at all71. These include a hyaluronidase (HYAL), a Kunitz-type proteinase inhibitor (KUN), a waprin, two ficolin, and a fused toxin.
Comparative transcriptome analysis
In contrast to our original hypothesis, similar toxin gene composition was found across all eight sampled individuals; however, differences in the presence or absence of toxins genes were recovered, providing some support for apparent variation in Sidewinder venoms as previously reported. While most toxins displayed similar expression across all individuals, 19 of the 62 toxin transcripts (31%) were absent in at least one individual, and none of the eight individuals contained all 62 transcripts (Fig. 3). Individual transcriptomes were missing between three and eleven toxin transcripts (4.8–17.7%), with an average of 7.4 (12%) missing. The majority of missing toxins were in lowly-expressed genes, though some highly-expressed toxins, such as CTL-3, were missing from one or more individuals (Fig. 3).
Hierarchical clustering of Crotalus cerastes expression similarity based on individual natural log-transformed toxin transcript TPM. Heatmap of toxins is ordered from greatest to least average ln expression with warmer colors indicating higher expression. Toxins with more than 10% of the coding sequence containing less than 5x coverage are marked as absent in that individual’s transcriptome. Detected significance for each transcript, based on consensus DESeq and edgeR analysis, is indicated with asterisks. Significant comparisons displayed in Fig. 4. SVL is reported in centimeters.
Unexpectedly, few toxin transcripts were differentially expressed across the majority of tested factor comparisons, contradicting our earlier prediction that Sidewinder venom variation would correspond to life history or lineage assignment. Overall, only four of 62 toxin transcripts (10 of 1,054 comparisons) and three of 19 toxin classes (7 of 323 comparisons) were recovered as differentially expressed across any tested factors (Fig. 3; Supplementary Table S1). Only three cases of differential expression (consisting of three toxin transcripts) were recovered across 682 comparisons of life stage (Figs 3 and 4A), sex (no difference), subspecies (Figs 3 and 4B), and mitochondrial lineage assignment (Figs 3 and 4C). Similarly, expression of each toxin class lacked major patterns of differential expression across these factors (Fig. 5; Supplementary Table S2), as only two of 19 toxin classes (3 of 209 comparisons) were recovered as differentially expressed across life stage (Figs 5 and 6A), sex (no differences), subspecies (Figs 5 and 6B), and mitochondrial lineage comparisons (Figs 5 and 6C; Supplementary Table S1).
Significant relationships recovered by the consensus of DESeq2 and edgeR seen in Fig. 3. (A) SVMPIII-10 expression changes with size. Dotted-line and standard error represent the average toxin trend. (B) Expression of VEGF-1 in C. c. cercobombus relative to C. c. laterorepens. (C) MYO-1 expression in the mitochondrial Colorado lineage relative to the Sonoran lineage. (D) MYO-1 expression in the nontoxin Colorado lineage relative to all three of the remaining lineages; additionally, the Sonoran lineage maintained significantly higher levels of MYO-1 compared to the South Mojave lineage. (E) BPP-1 expression in the nontoxin North Mojave lineage relative to both the South Mojave and Sonoran lineages; given only one BPP was recovered, this relationship was retained in toxin class expression analyses in Fig. 5. (F) VEGF-1 expression in the Sonoran lineage relative to the South Mojave lineage.
Hierarchical clustering of Crotalus cerastes expression similarity based on summed and natural log-transformed toxin classes. Heatmap of toxins is ordered from greatest to least average ln expression with warmer colors indicating higher expression. Detected significance for each transcript, based on consensus DESeq and edgeR analysis, is indicated with asterisks. Significant comparisons displayed in Fig. 6. SVL is reported in centimeters.
Significant relationships recovered by the consensus of DESeq2 and edgeR seen in Fig. 5. (A) Expression of combined myotoxins (n = 2) with size. Dotted-line and standard error represent the average toxin trend. (B) Expression of combined VEGFs (n = 2) in C. c. cercobombus relative to C. c. laterorepens. (C) Expression of combined myotoxins (n = 2) in the mitochondrial South Mojave lineage relative to the Sonoran lineage. (D) Expression of combined VEGFs (n = 2) in the nontoxin Sonoran lineage relative to both the South Mojave lineage and Colorado lineage.
Notably, despite few instances of differential expression overall, more instances of expression differences were recovered between comparisons of nontoxin (phylotranscriptomic) lineages than all other comparisons combined. While only one individual changed lineage assignment using the more robust dataset, the number of significant results more than doubled. Seven of the remaining 372 comparisons for individual transcripts–consisting of a total of three toxin transcripts–were differentially expressed (Figs 3 and 4D–F); Supplementary Table S1). The number of significant toxin class comparisons also increased: VEGFs and BPPs were recovered as differentially expressed–four of the remaining 114 comparisons for toxin classes (Figs 5 and 6D; Supplementary Table S2). Given that only one BPP is present in the venom of Crotalus cerastes, this result reflected those of independent transcript analysis (Fig. 4E). This pattern of increased differential expression was mirrored with nontoxin transcripts as only 32 of 21,747 comparisons were found to be significant across all comparisons of life stage, sex, subspecies, and mitochodrial lineage assignments (Supplementary Table S1). However, across all comparisons of nontoxin lineage assignments, 55 of 11,862 comparisons were found to be significantly differentially expressed (Supplementary Table S1). In addition to the lack of significant differential expression, we detected significant phylogenetic signal in only one transcript, CTL-5 (K = 1.35; P = 0.02; Supplementary Table S3). As K was greater than one, it suggests the similarity in the expression of this toxin between closely related individuals was higher than expected under Brownian motion. This may have been driven by the absence of CTL-5 in the two most closely related Sonoran lineage samples (ASU 36167 and KW1744; Fig. 3), as the toxin was moderately to highly expressed in all other individuals.
Venom proteomics and transcript versus protein abundance estimates
The majority of toxin transcripts were verified proteomically, and the unverified toxins were nearly all the lowest expressed across all individuals. With qMS, 49 of the 62 (79%) toxin transcripts were found in at least one individual; of these, 26 (53%) were conserved across all individuals, and nine additional transcripts were missing from two or fewer individuals. Toxins that were not verified in any individual were largely those most lowly expressed, and included all 3FTx, Ficolin, FusedToxin, KUN, and Waprin transcripts, as well as VEGF-2 and several CTLs (9, 10, and 11) (Fig. 2). Only CTL-4 was highly expressed but not verified proteomically at a 1% FDR; this transcript was recovered in four individuals at less stringent FDRs (2% in one individual, 10% in three others). Seven of the individual proteomes each contained 40–45 confirmed toxins; a juvenile—CAS 259893—had only 34; this was the smallest individual and similarly contained the highest number of suspected absent toxins in the transcriptome. Two of the most common toxin families, SVMPs and SVSPs, were conserved across nearly all individuals, as 24 of the 29 confirmed in those two families (83%) were found in all or all but one of the individuals. CTLs, on the other hand, were more variable: only two of seven confirmed (29%) were found across all individuals. Comparisons of transcriptomic and proteomic abundances were positively correlated, though the strength of the correlation was weaker in some individuals compared to others (Spearman’s rank correlation coefficients: 0.48 ≤ ρ ≤ 0.77; Pearson’s correlation coefficients: 0.53 ≤ R ≤ 0.76; coefficients of determination: 0.28 ≤ R2 ≤ 0.58; Fig. 7). As protein identity was based on the species-consensus transcriptome rather than individual transcriptomes, such differences in correlation were expected. Rokyta et al.49 found that venom-gland transcriptomes and venom proteomes were highly correlated in snakes (0.47 ≤ ρ ≤ 0.89; 0.58 ≤ R ≤ 0.92; 0.34 ≤ R2 ≤ 0.85), and the majority of our relationships were similarly correlated.
Correlation of venom transcript and protein abundances for eight Crotalus cerastes, showing positive correlation in all individuals. Data were centered log-ratio transformed (clr). Abbreviations: n - number of transcripts; ρ (Spearman’s rank correlation coefficient; R - Pearson’s correlation coefficient; R2 - coefficient of determination.
Discussion
In this study, we first characterized venom-gland transcriptomes together with complementary venom proteomes of Sidewinder Rattlesnakes (Crotalus cerastes) for the first time. We also tested for differential expression across hypothesized lineages, subspecies, and life history traits to understand the effect of shallow lineage divergence on venom. Similar to other rattlesnakes, we found that C. cerastes venom largely consisted of SVMPs, SVSPs, a PLA2, CTLs, and a BPP. The most highly expressed toxins in C. cerastes venom are primarily responsible for hemorrhagic activity and necrosis (SVMPs), as well as myotoxic activity and the inhibition of platelet aggregation (PLA2s)10,72. The abundance of SVMPs and presence of PLA2 supports previous reports of moderate hemorrhagic activity and low-to-moderate PLA2 activity in C. cerastes relative to other Crotalus35.
Variation in Sidewinder venom-gland transcriptomes was largely not attributable to hypothesized lineages, subspecies, or life history traits, as we had originally predicted. Similarly, we recovered only one instance of significant phylogenetic signal in venom gene expression across our phylogeny, which appeared to be driven by the absence of that toxin in two individual transcriptomes. In fact, most toxin transcripts were not differentially expressed in the majority of the comparisons tested. One exception were myotoxins, supporting previous work by Bober et al.24 that detected myotoxins in the venom of samples of C. c. cerastes from southwestern Utah, but not from pooled commercial samples of unknown origin. We found MYO-1 to be expressed significantly less in the nontoxin Colorado lineage than any other lineage. This difference appeared to be based on transcript presence/absence: the gene encoding MYO-1 may be absent from the genome of the Colorado lineage, or not expressed. It also may be the result of an ontogenetic change rather than lineage-specific adaptations, as the Colorado individual was the smallest juvenile, and myotoxin class expression increased with size. One additional transcript, SVMPIII-10, decreased expression with size. As Crotalus cerastes undergo a known ontogenetic dietary shift40, these shifts in venom expression suggest a possible trade-off between these toxins at different life stages.
The most notable factor influencing differential expression in our dataset was lineage assignment based on analysis of the large nontoxin dataset. This analysis only re-assigned CAS 259903 from the South Mojave to the North Mojave lineage, but this single change more than doubled the number of significantly differentially expressed comparisons of toxins and a similar increase was found when comparing nontoxin transcripts. Discordance among morphological (subspecies), mitochondrial, and genomic datasets is common in systematics and phylogeography, and can be caused by differences in rates of evolution, inheritance patterns, introgression, or sex-biased dispersal29,30,31,32,33,34. Here, we demonstrate that improper assignment of lineages can misinform conclusions regarding the differential expression of transcripts across a phylogeny. Therefore, robust phylogenomic datasets, such as the thousands of nontoxins already sequenced as “bycatch” during venom gland transcriptomic studies, should be considered when testing for differential expression between clades.
Unlike some rattlesnake species, including taxa co-distributed through much of their range, Sidewinder venom did not form multiple discrete phenotypes, or follow a geographic pattern of variation. Mojave Rattlesnakes (C. scutulatus), a species sympatric with C. cerastes in much of the southwestern United States, exhibits dramatic cases of venom differentiation with hemorrhagic and neurotoxic phenotypes in short geographic distance, which seems to be driven by the presence or absence of neurotoxic PLA2 genes in their genomes38,73,74,75,76,77,78. Similar cases of dramatic venom differentiation at short geographic distance can be observed in Timber Rattlesnakes (C. horridus) and Southern Pacific Rattlesnakes (C. helleri)36,37,45. The lack of differential expression in Sidewinder venom is especially notable in light of a recent study examining intraspecific venom variation in two populations of the Common Lancehead (Viperidae: Bothrops atrox)21. Using similar methods to our own, 124 of 152 toxins (81.5%) were found to be differentially expressed between two genetically-distinct, allopatric populations21. In contrast, only four of the 62 (6.4%) toxins found in the Sidewinder venom-gland transcriptome were differentially expressed across four lineages. This lack of expression variation in Sidewinder venom between lineages and life history may be due to a variety of evolutionary processes, such as migration patterns or relaxed selection pressures.
While geographic structure of venom expression is well-documented in many species, examples of species lacking expression differences are known. For example, Micrurus fulvius populations across Florida were found to have a conserved venom phenotype19, which was hypothesized to be the result of either a species-wide selective sweep or a recent range expansion. Within Sidewinders, the lack of differential expression does not necessarily signify a conserved venom phenotype. While the expression of toxin classes, such as SVMPs, CTLs, and SVSPs, was largely similar across all individuals, the expression of individual toxin transcripts themselves was variable. The minimal differential expression in Sidewinder venom composition despite individual variation suggests that gene flow is likely occurring between the shallow lineages. Gene flow across the range of the species, as evidenced by the incongruence between our mitochondrial and nontoxin phylogenies, may have precluded local adaptation shaping one or more discrete venom phenotypes by swamping local adaptation to local prey abundance79,80. The individual variability could also be associated with a flat adaptive landscape or relaxed selective constraints on some more lowly-expressed toxins, allowing variation to arise through non-deleterious mutations over time81.
Further exploration of the drivers of venom variation in Crotalus cerastes should focus on sequence evolution rather than differential expression. While sequence evolution generally occurs more slowly than regulatory evolution, if Sidewinders are under relaxed selection or maintain high gene flow between lineages, these patterns would be apparent in the sequence data and could be explicitly tested. Most importantly, further studies involving venom variation of C. cerastes requires more thorough sampling, including the remaining south Sonoran mitochondrial lineage, and pairs of adults and juveniles within each lineage. As our sampling of juveniles was sparse, our conclusions regarding both ontogenetic shifts and pairwise lineage comparisons were limited. Similarly, our sample size was on the low end of what is required for detectable significance in tests of phylogenetic signal69; more comprehensive sampling would greatly increase the statistical power of such analyses.
In many snake venom-gland transcriptome studies, a single individual’s transcriptome is presented as characteristic of the entire species44,82. The differences in toxin gene composition across eight Crotalus cerastes transcriptomes reinforces the importance of using multiple samples to fully characterize the toxin arsenal of a species20; while a single sample will provide the majority of toxins for a species, it may lack some toxins that are present, even highly expressed, in others. In one case, the second most abundant toxin gene (BPP-1) was absent from the transcriptome of one individual (CAS 259903), and others were missing specific copies of other gene families (e.g. CTL-3 and CTL-5 missing from four and two individuals, respectively). This is especially true when de novo assembling and annotating transcriptomes, as 19 of 62 transcripts were missing from at least one individual. Without sampling the transcriptomes of multiple individuals and across the distribution, entire transcripts may be missed, and potential variation overlooked.
In conclusion, we provide the first venom-gland transcriptomes and complementary venom proteomes of Crotalus cerastes from individuals across their southwestern United States range. Our original hypothesis—that Sidewinder venom composition and gene expression is variable as a result of lineage diversification and traceable to specific life history or lineage assignments—was only partially supported, as individual variation in composition did not translate to extensive differential expression in the factors we tested. The relative lack of differential expression may be the result of high gene flow, relaxed selective pressures, individual stochasticity in venom overwhelming any potential signal from the factors tested, or local adaptation. Importantly, we found differences in the number of differentially expressed toxins between subspecies, mitochondrial lineage, and phylotranscriptomic lineage comparisons. Indeed, incorporating robust phylogenetic hypotheses, especially for taxa with poorly-explored evolutionary histories, is necessary for accurately detecting differential expression in transcriptomic studies relying on lineage assignments as hypotheses. Finally, studies looking to generate a species-consensus venom-gland transcriptome must sample multiple individuals in order to capture individual variation and the full venom repertoire of a species.
References
Fraser, D. J., Weir, L. K., Bernatchez, L., Hansen, M. M. & Taylor, E. B. Extent and scale of local adaptation in salmonid fishes: review and meta-analysis. Heredity 106, 404–420, https://doi.org/10.1038/hdy.2010.167 (2011).
Romero, I. G., Ruvinsky, I. & Gilad, Y. Comparative studies of gene expression and the evolution of gene regulation. Nature Reviews Genetics 13, 505–516, https://doi.org/10.1038/nrg3229 (2012).
Lamichhaney, S. et al. A beak size locus in Darwin’s finches facilitated character displacement during a drought. Science 352, 470–474, https://doi.org/10.1126/science.aad8786 (2016).
Casewell, N. R., Wüster, W., Vonk, F. J., Harrison, R. A. & Fry, B. G. Complex cocktails: the evolutionary novelty of venoms. Trends in Ecology & Evolution 28, 219–229, https://doi.org/10.1016/j.tree.2012.10.020 (2013).
Margres, M. J. et al. Functional characterizations of venom phenotypes in the eastern diamondback rattlesnake (Crotalus adamanteus) and evidence for expression-driven divergence in toxic activities among populations. Toxicon 119, 28–38, https://doi.org/10.1016/j.toxicon.2016.05.005 (2016).
Shapiro, M. D. et al. Genetic and developmental basis of evolutionary pelvic reduction in threespine sticklebacks. Nature 428, 717–723, https://doi.org/10.1038/nature02415 (2004).
Casewell, N. R. et al. Medically important differences in snake venom composition are dictated by distinct postgenomic mechanisms. Proceedings of the National Academy of Sciences 111, 9205–9210, https://doi.org/10.1073/pnas.1405484111 (2014).
Margres, M. J. et al. Linking the transcriptome and proteome to characterize the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). Journal of Proteomics 96, 145–158, https://doi.org/10.1016/j.jprot.2013.11.001 (2014).
Chippaux, J.-P., Williams, V. & White, J. Snake venom variability: methods of study, results and interpretation. Toxicon 29, 1279–1303, https://doi.org/10.1016/0041-0101(91)90116-9 (1991).
Mackessy, S. P. Handbook of Venoms and Toxins of Reptiles (CRC Press, Boca Raton, F. L. USA, 2010).
Mackessy, S. P. Venom composition in rattlesnakes: trends and biological significance. In Hayes, W., Beaman, K., Cardwell, M. & Bush, S. (eds) The Biology of Rattlesnakes, 495–510 (Loma Linda University Press, Loma Linda, CA USA, 2008).
Gibbs, H. L., Sanz, L., Sovic, M. G. & Calvete, J. J. Phylogeny-based comparative analysis of venom proteome variation in a clade of rattlesnakes (Sistrurus sp.). PLoS ONE 8, e67220, https://doi.org/10.1371/journal.pone.0067220 (2013).
Lomonte, B. et al. Snake venomics and toxicological profiling of the arboreal pitviper Bothriechis supraciliaris from Costa Rica. Toxicon 59, 592–599, https://doi.org/10.1016/j.toxicon.2012.01.005 (2012).
Daltry, J. C., Wüster, W. & Thorpe, R. S. Diet and snake venom evolution. Nature 379, 537–540, https://doi.org/10.1038/379537a0 (1996).
Creer, S. et al. Genetic and ecological correlates of intraspecific variation in pitviper venom composition detected using matrix-assisted laser desorption time-of-flight mass spectrometry (MALDI-TOF-MS) and isoelectric focusing. Journal of Molecular Evolution 56, 317–329, https://doi.org/10.1007/s00239-002-2403-4 (2003).
Holding, M. L., Biardi, J. E. & Gibbs, H. L. Coevolution of venom function and venom resistance in a rattlesnake predator and its squirrel prey. Proceedings of the Royal Society B: Biological Sciences 283, 20152841, https://doi.org/10.1098/rspb.2015.2841 (2016).
Margres, M. J. et al. Quantity, not quality: rapid adaptation in a polygenic trait proceeded exclusively through expression differentiation. Molecular Biology and Evolution 34, 3099–3110, https://doi.org/10.1093/molbev/msx231 (2017).
Casewell, N. R., Huttley, G. A. & Wüster, W. Dynamic evolution of venom proteins in squamate reptiles. Nature Communications 3, 1066, https://doi.org/10.1038/ncomms2065 (2012).
Margres, M. J. et al. Contrasting modes and tempos of venom expression evolution in two snake species. Genetics 199, 165–176, https://doi.org/10.1534/genetics.114.172437 (2015).
Rokyta, D. R., Margres, M. J., Ward, M. J. & Sanchez, E. E. The genetics of venom ontogeny in the eastern diamondback rattlesnake (Crotalus adamanteus). PeerJ 5, e3249, https://doi.org/10.7717/peerj.3249 (2017).
Amazonas, D. R. et al. Molecular mechanisms underlying intraspecific variation in snake venom. Journal of Proteomics, https://doi.org/10.1016/j.jprot.2018.03.032 (2018).
Klauber, L. M. Rattlesnakes: their habits, life histories, and influence on mankind. Vol. 1., 2 edn.(University of California Press, Berkeley and Los Angeles, CA USA, 1972).
Ernst, C. H. & Ernst, E. M. Venomous Reptiles of the United States, Canada, and Northern Mexico. Vol. 2: Crotalus. (The John Hopkins University Press, Baltimore, MD USA, 2012).
Bober, M. A., Glenn, J. L., Straight, R. C. & Ownby, C. L. Detection of myotoxin a-like proteins in various snake venoms. Toxicon 26, 665–673, https://doi.org/10.1016/0041-0101(88)90248-6 (1988).
Mackessy, S. P. Venom ontogeny in the Pacific Rattlesnakes Crotalus viridis helleri and C. v. oreganus. Copeia 1988, 92, https://doi.org/10.2307/1445927 (1988).
Reiserer, R. Sidewinder, Crotalus cerastes (Hallowell 1854). In Schuett, G., Feldner, M., Smith, C. & Reiserer, R. (eds) Rattlesnakes of Arizona, Vol. 1, 395–432 (Eco Publishing, Rodeo, NM USA, 2016).
Pece, A. J. Phylogeography of the Sidewinder (Crotalus cerastes), with implications for the historical biogeography of Southwestern North American Deserts. Ph.D. thesis, San Diego State University (2004).
Douglas, M. E., Douglas, M. R., Schuett, G. W. & Porras, L. W. Evolution of rattlesnakes (Viperidae; Crotalus) in the warm deserts of western North America shaped by Neogene vicariance and Quaternary climate change. Molecular ecology 15, 3353–74, https://doi.org/10.1111/j.1365-294X.2006.03007.x (2006).
Shaw, K. L. Conflict between nuclear and mitochondrial DNA phylogenies of a recent species radiation: What mtDNA reveals and conceals about modes of speciation in Hawaiian crickets. Proceedings of the National Academy of Sciences 99, 16122–16127, https://doi.org/10.1073/pnas.242585899 (2002).
Leaché, A. D. & McGuire, J. A. Phylogenetic relationships of horned lizards (Phrynosoma) based on nuclear and mitochondrial data: Evidence for a misleading mitochondrial gene tree. Molecular Phylogenetics and Evolution 39, 628–644, https://doi.org/10.1016/j.ympev.2005.12.016 (2006).
Leaché, A. D. Species trees for spiny lizards (Genus Sceloporus): Identifying points of concordance and conflict between nuclear and mitochondrial data. Molecular Phylogenetics and Evolution 54, 162–171, https://doi.org/10.1016/j.ympev.2009.09.006 (2010).
Wiens, J. J., Kuczynski, C. A. & Stephens, P. R. Discordant mitochondrial and nuclear gene phylogenies in emydid turtles: Implications for speciation and conservation. Biological Journal of the Linnean Society 99, 445–461, https://doi.org/10.1111/j.1095-8312.2009.01342.x (2010).
Mulcahy, D. G. et al. Estimating divergence dates and evaluating dating methods using phylogenomic and mitochondrial data in squamate reptiles. Molecular Phylogenetics and Evolution 65, 974–991, https://doi.org/10.1016/j.ympev.2012.08.018 (2012).
Papakostas, S. et al. Integrative taxonomy recognizes evolutionary units despite widespread mitonuclear discordance: Evidence from a Rotifer cryptic species complex. Systematic Biology 65, 508–524, https://doi.org/10.1093/sysbio/syw016 (2016).
Tan, N.-H. & Ponnudurai, G. A comparative study of the biological activities of rattlesnake (genera Crotalus and Sistrurus) venoms. Comparative Biochemistry and Physiology Part C: Comparative. Pharmacology 98, 455–461, https://doi.org/10.1016/0742-8413(91)90233-J (1991).
French, W. J. et al. Mojave toxin in venom of Crotalus helleri (Southern Pacific Rattlesnake): molecular and geographic characterization. Toxicon 44, 781–791, https://doi.org/10.1016/j.toxicon.2004.08.008 (2004).
Sunagar, K. et al. Intraspecific venom variation in the medically significant Southern Pacific Rattlesnake (Crotalus oreganus helleri): biodiscovery, clinical and evolutionary implications. Journal of Proteomics 99, 68–83, https://doi.org/10.1016/j.jprot.2014.01.013 (2014).
Strickland, J. L., Mason, A. J., Rokyta, D. R. & Parkinson, C. L. Phenotypic variation in Mojave Rattlesnake (Crotalus scutulatus) venom is driven by four toxin families. Toxins 10, 1–23, https://doi.org/10.3390/toxins10040135 (2018).
Goldberg, S. Reproductive cycle of the sidewinder, Crotalus cerastes (Serpentes: Viperidae), from California. The Texas Journal of Science 56, 55–62 (2004).
Webber, M. M., Jezkova, T. & Rodríguez-Robles, J. A. Feeding ecology of Sidewinder Rattlesnakes, Crotalus cerastes (Viperidae). Herpetologica 72, 324–330, https://doi.org/10.1655/Herpetologica-D-15-00031.1 (2016).
Rotenberg, D., Bamberger, E. S. & Kochva, E. Studies on ribonucleic acid synthesis in the venom glands of Vipera palaestinae (Ophidia, Reptilia). The Biochemical Journal 121, 609–612 (1971).
McCleary, R. J. & Heard, D. J. Venom extraction from anesthetized Florida cottonmouths, Agkistrodon piscivorus conanti, using a portable nerve stimulator. Toxicon 55, 250–255, https://doi.org/10.1016/j.toxicon.2009.07.030 (2010).
Leary, S. et al. AVMA Guidelines for the Euthanasia of Animals: 2013 Edition. Tech. Rep., American Veterinary Medical Association (2013).
Rokyta, D. R., Lemmon, A. R., Margres, M. J. & Aronow, K. The venom-gland transcriptome of the eastern diamondback rattlesnake (Crotalus adamanteus). BMC Genomics 13, 312, https://doi.org/10.1186/1471-2164-13-312 (2012).
Rokyta, D. R., Wray, K. P., McGivern, J. J. & Margres, M. J. The transcriptomic and proteomic basis for the evolution of a novel venom phenotype within the Timber Rattlesnake (Crotalus horridus). Toxicon 98, 34–48, https://doi.org/10.1016/j.toxicon.2015.02.015 (2015).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620, https://doi.org/10.1093/bioinformatics/btt593 (2014).
Rokyta, D. R., Wray, K. P. & Margres, M. J. The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics. BMC Genomics 14, 394, https://doi.org/10.1186/1471-2164-14-394 (2013).
Rokyta, D. R., Margres, M. J. & Calvin, K. Post-transcriptional mechanisms contribute little to phenotypic variation in snake venoms. G3: Genes|Genomes|Genetics 5, 2375–2382, https://doi.org/10.1534/g3.115.020578 (2015).
Holding, M., Margres, M., Mason, A., Parkinson, C. & Rokyta, D. Evaluating the performance of de novo assembly methods for venom-gland transcriptomics. Toxins 10, 249, https://doi.org/10.3390/toxins10060249 (2018).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152, https://doi.org/10.1093/bioinformatics/bts565 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv:1303.3997 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Davis, M. A., Douglas, M. R., Collyer, M. L. & Douglas, M. E. Deconstructing a species-complex: geometric morphometric and molecular analyses define species in the Western Rattlesnake (Crotalus viridis). PLOS ONE 11, e0146166, https://doi.org/10.1371/journal.pone.0146166 (2016).
Alexander, A. M. Phase_hybrid_from_next_gen, https://github.com/laninsky/Phase_hybrid_from_next_gen. (2015).
Vaidya, G., Lohman, D. J. & Meier, R. SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27, 171–180, https://doi.org/10.1111/j.1096-0031.2010.00329.x (2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313, https://doi.org/10.1093/bioinformatics/btu033 (2014).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755, https://doi.org/10.1093/bioinformatics/17.8.754 (2001).
Ronquist, F. et al. MrBayes 3.2: efficient bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61, 539–542, https://doi.org/10.1093/sysbio/sys029 (2012).
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molecular Biology and Evolution 29, 1695–1701, https://doi.org/10.1093/molbev/mss020 (2012).
Rokyta, D. R. & Ward, M. J. Venom-gland transcriptomics and venom proteomics of the Black-back Scorpion (Hadrurus spadix) reveal detectability challenges and an unexplored realm of animal toxin diversity. Toxicon 128, 23–37, https://doi.org/10.1016/j.toxicon.2017.01.014 (2017).
Ward, M. J., Ellsworth, S. A. & Rokyta, D. R. Venom-gland transcriptomics and venom proteomics of the Hentz Striped Scorpion (Centruroides hentzi; Buthidae) reveal high toxin diversity in a harmless member of a lethal family. Toxicon 142, 14–29, https://doi.org/10.1016/j.toxicon.2017.12.042 (2018).
Aitchison, J. The statistical analysis of compositional data. Journal of the Royal Statistical Society. Series B. Methodological 44, 139–177, https://doi.org/10.2307/2345821 (1982).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323, https://doi.org/10.1186/1471-2105-12-323 (2011).
Palarea-Albaladejo, J. & Martin-Fernandez, J. zCompositions – R package for multivariate imputation of left-censored data under a compositional approach. Chemometrics and Intelligent Laboratory Systems 143, 85–96 (2015).
Kolde, R. pheatmap: Pretty Heatmaps. R package version 1.0.8, https://cran.r-project.org/package=pheatmap (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 15, 550, https://doi.org/10.1186/s13059-014-0550-8 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
Blomberg, S. P., Garland, T. & Ives, A. R. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57, 717–745, https://doi.org/10.1111/j.0014-3820.2003.tb00285.x (2003).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464, https://doi.org/10.1093/bioinformatics/btq166 (2010).
Hargreaves, A. D., Swain, M. T., Hegarty, M. J., Logan, D. W. & Mulley, J. F. Restriction and recruitment—gene duplication and the origin and evolution of snake venom toxins. Genome Biology and Evolution 6, 2088–2095, https://doi.org/10.1093/gbe/evu166 (2014).
Manjunatha Kini, R. Excitement ahead: structure, function and mechanism of snake venom phospholipase A2 enzymes. Toxicon 42, 827–840, https://doi.org/10.1016/j.toxicon.2003.11.002 (2003).
Dowell, N. L. et al. The deep origin and recent loss of venom toxin genes in rattlesnakes. Current Biology 26, 2434–2445, https://doi.org/10.1016/j.cub.2016.07.038 (2016).
Massey, D. J. et al. Venom variability and envenoming severity outcomes of the Crotalus scutulatus scutulatus (Mojave rattlesnake) from Southern Arizona. Journal of Proteomics 75, 2576–2587, https://doi.org/10.1016/j.jprot.2012.02.035 (2012).
Glenn, J. & Straight, R. Mojave rattlesnake Crotalus scutulatus scutulatus venom: variation in toxicity with geographical origin. Toxicon 16, 81–84, https://doi.org/10.1016/0041-0101(78)90065-X (1978).
Glenn, J. L., Straight, R. C., Wolfe, M. C. & Hardy, D. L. Geographical variation in Crotalus scutulatus scutulatus (Mojave rattlesnake) venom properties. Toxicon 21, 119–130, https://doi.org/10.1016/0041-0101(83)90055-7 (1983).
Borja, M. et al. Biological and proteolytic variation in the venom of Crotalus scutulatus scutulatus from Mexico. Toxins 10, 35, https://doi.org/10.3390/toxins10010035 (2018).
Whittington, A. C., Mason, A. J. & Rokyta, D. R. A single mutation unlocks cascading exaptations in the origin of a potent pitviper neurotoxin. Molecular Biology and Evolution 35, 887–898, https://doi.org/10.1093/molbev/msx334 (2018).
Wiens, J. J. et al. Niche conservatism as an emerging principle in ecology and conservation biology. Ecology Letters 13, 1310–1324, https://doi.org/10.1111/j.1461-0248.2010.01515.x (2010).
Crisp, M. D. & Cook, L. G. Phylogenetic niche conservatism: what are the underlying evolutionary and ecological causes? New Phytologist 196, 681–694, https://doi.org/10.1111/j.1469-8137.2012.04298.x (2012).
Aird, S. D. et al. Population genomic analysis of a pitviper reveals microevolutionary forces underlying venom chemistry. Genome Biology and Evolution 9, 2640–2649, https://doi.org/10.1093/gbe/evx199 (2017).
Margres, M. J., Aronow, K., Loyacano, J. & Rokyta, D. R. The venom-gland transcriptome of the eastern coral snake (Micrurus fulvius) reveals high venom complexity in the intragenomic evolution of venoms. BMC Genomics 14, https://doi.org/10.1186/1471-2164-14-531. (2013).
Crotalus cerastes. The IUCN Red List of Threatened Species 2007, https://doi.org/10.2305/IUCN.UK.2007.RLTS.T64315A12764960.en (2007).
Acknowledgements
We thank T. Burkhardt for donating a sample from his private collection for use in this study, along with K.P. Wray and the late R. Govreau for assistance collecting additional samples used herein. E.J. Ely (California Academy of Science) and C. Johnson (Arizona State University) provided specimen measurements. M.J. Margres provided comments on an early draft that greatly improved the quality of this manuscript. Funding was provided by the National Science Foundation to D.R.R. (DEB 1145987 and DEB 1638902) and C.L.P. (DUE 1161228, DEB 1822417, and DEB 1638879), Clemson University (C.L.P.), PrairieBiotic Research Inc. (J.L.S.), Sigma Xi Grants-in-aid-of-research (J.L.S.), the SnakeDays Research Grant (J.L.S.), the Southwestern Association of Naturalists Howard McCarley Research Grant (J.L.S.), and the Theodore Roosevelt Memorial Fund through the American Museum of Natural History (J.L.S. and A.J.M.). Clemson University generously provided computational resources on their Palmetto Cluster. The Clemson University Department of Biological Sciences and the Clemson University Libraries Open Access Publishing Fund generously provided funding in support of publication costs. Two anonymous reviewers provided helpful comments and suggestions that improved the quality of this work.
Author information
Authors and Affiliations
Contributions
E.P.H., R.M.R. and C.L.P. designed the study. J.L.S. collected samples. E.P.H., R.M.R., J.L.S., M.P.H., A.J.M. and D.R.R. generated data. E.P.H., R.M.R., M.L.H., M.P.H., A.J.M and D.R.R. analyzed the data. D.R.R. and C.L.P. provided materials, reagents, computational resources, and lab space. E.P.H., R.M.R. and M.L.H. wrote the manuscript. All authors edited, reviewed, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hofmann, E.P., Rautsaw, R.M., Strickland, J.L. et al. Comparative venom-gland transcriptomics and venom proteomics of four Sidewinder Rattlesnake (Crotalus cerastes) lineages reveal little differential expression despite individual variation. Sci Rep 8, 15534 (2018). https://doi.org/10.1038/s41598-018-33943-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33943-5
Keywords
This article is cited by
-
Sequence Divergence in Venom Genes Within and Between Montane Pitviper (Viperidae: Crotalinae: Cerrophidion) Species is Driven by Mutation–Drift Equilibrium
Journal of Molecular Evolution (2023)
-
Venom-gland transcriptomic, venomic, and antivenomic profiles of the spine-bellied sea snake (Hydrophis curtus) from the South China Sea
BMC Genomics (2021)
-
Trait differentiation and modular toxin expression in palm-pitvipers
BMC Genomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.